Genome-Wide Identification and Characterization of the CsFHY3/FAR1 Gene Family and Expression Analysis under Biotic and Abiotic Stresses in Tea Plants (Camellia sinensis)
Abstract
1. Introduction
2. Results
2.1. Identification and Analysis of CsFHY3/FAR1 Genes
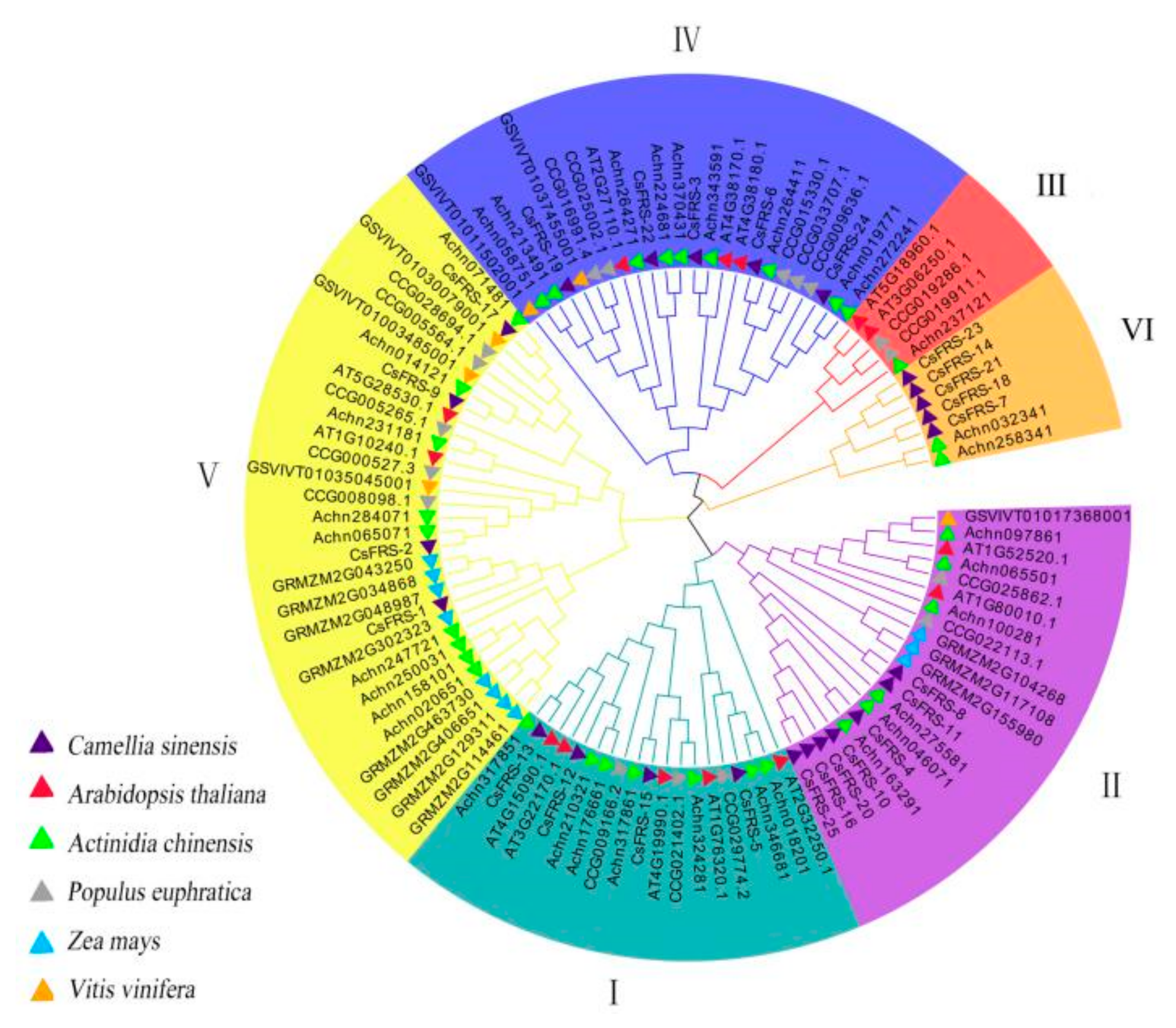
2.2. Phylogenetic Analysis of CsFHY3/FAR1s
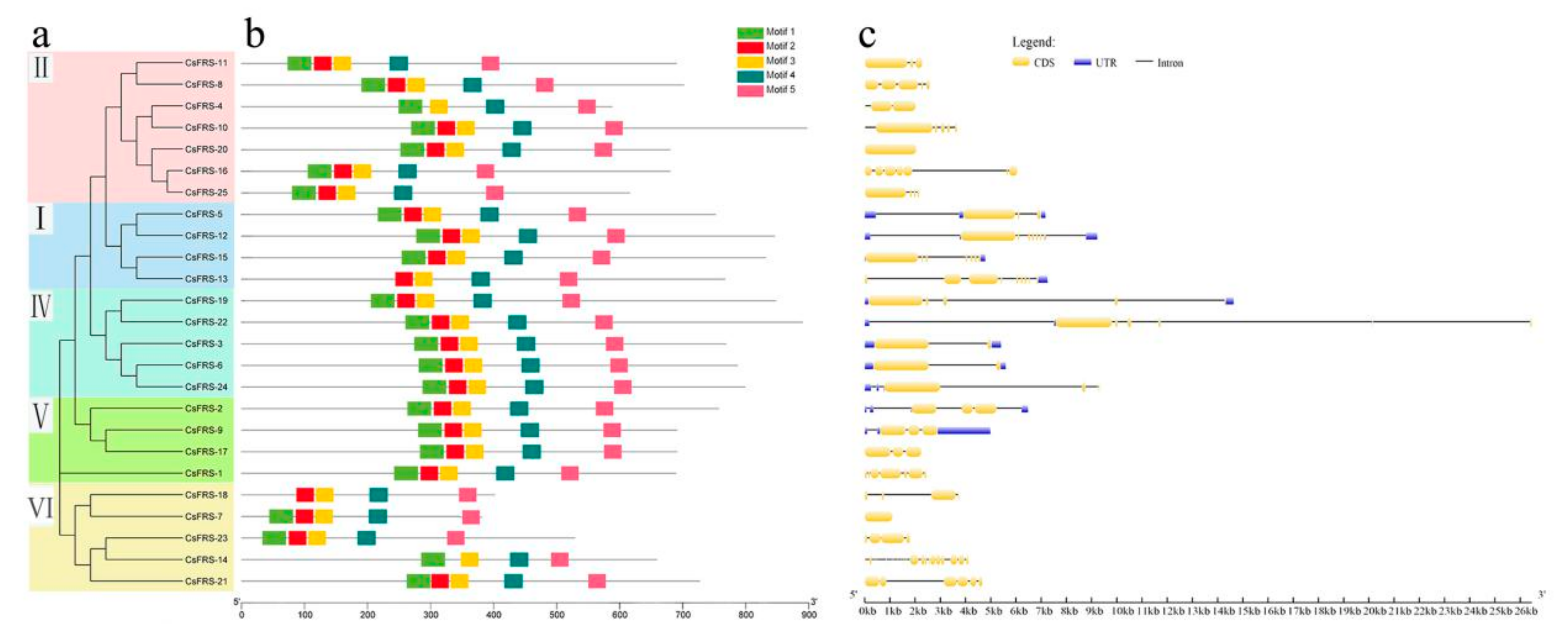
2.3. Gene Structure and Motif Analysis
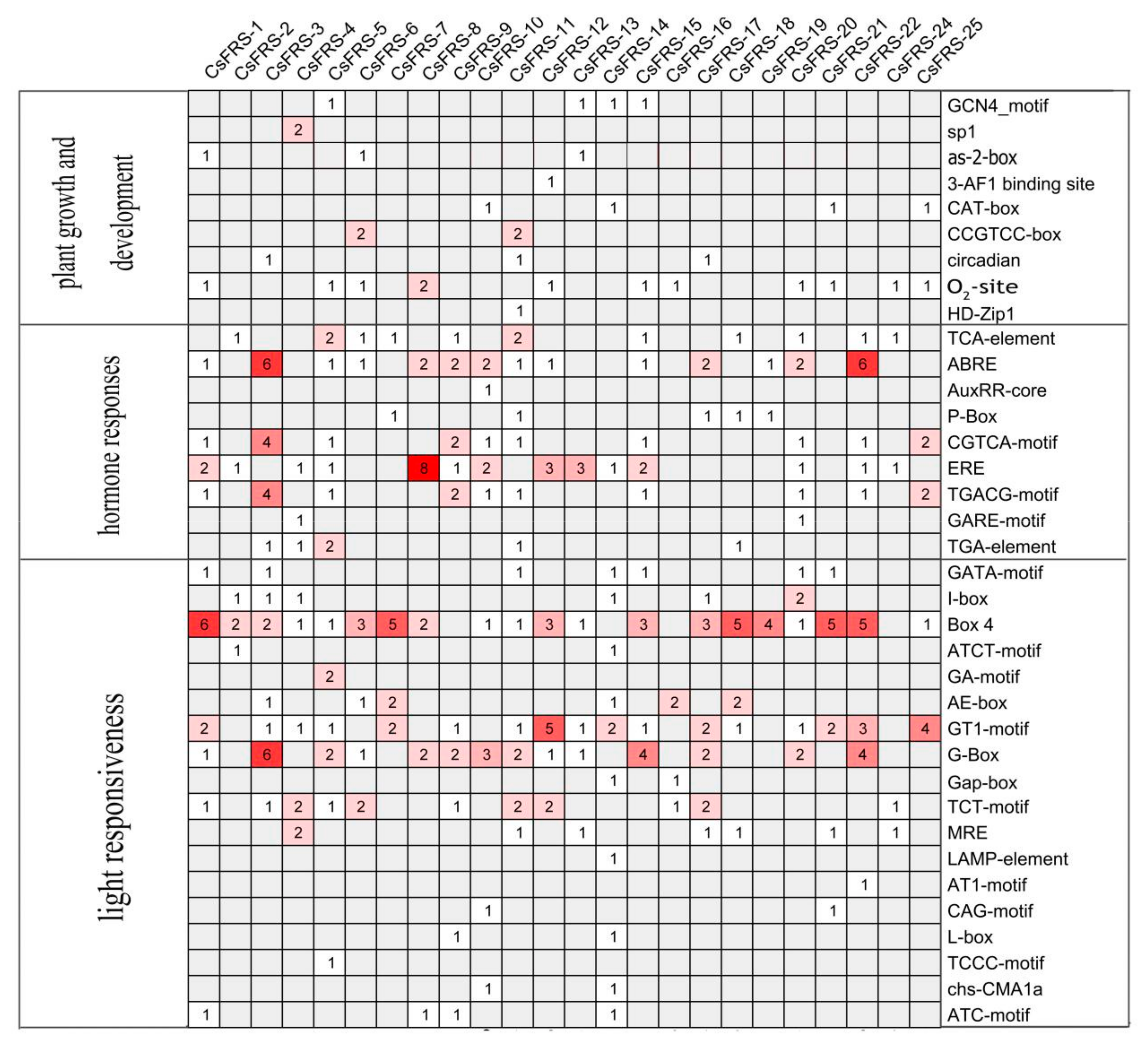
2.4. Analysis of Cis-Acting Elements in the Promoters of CsFHY3/FAR1
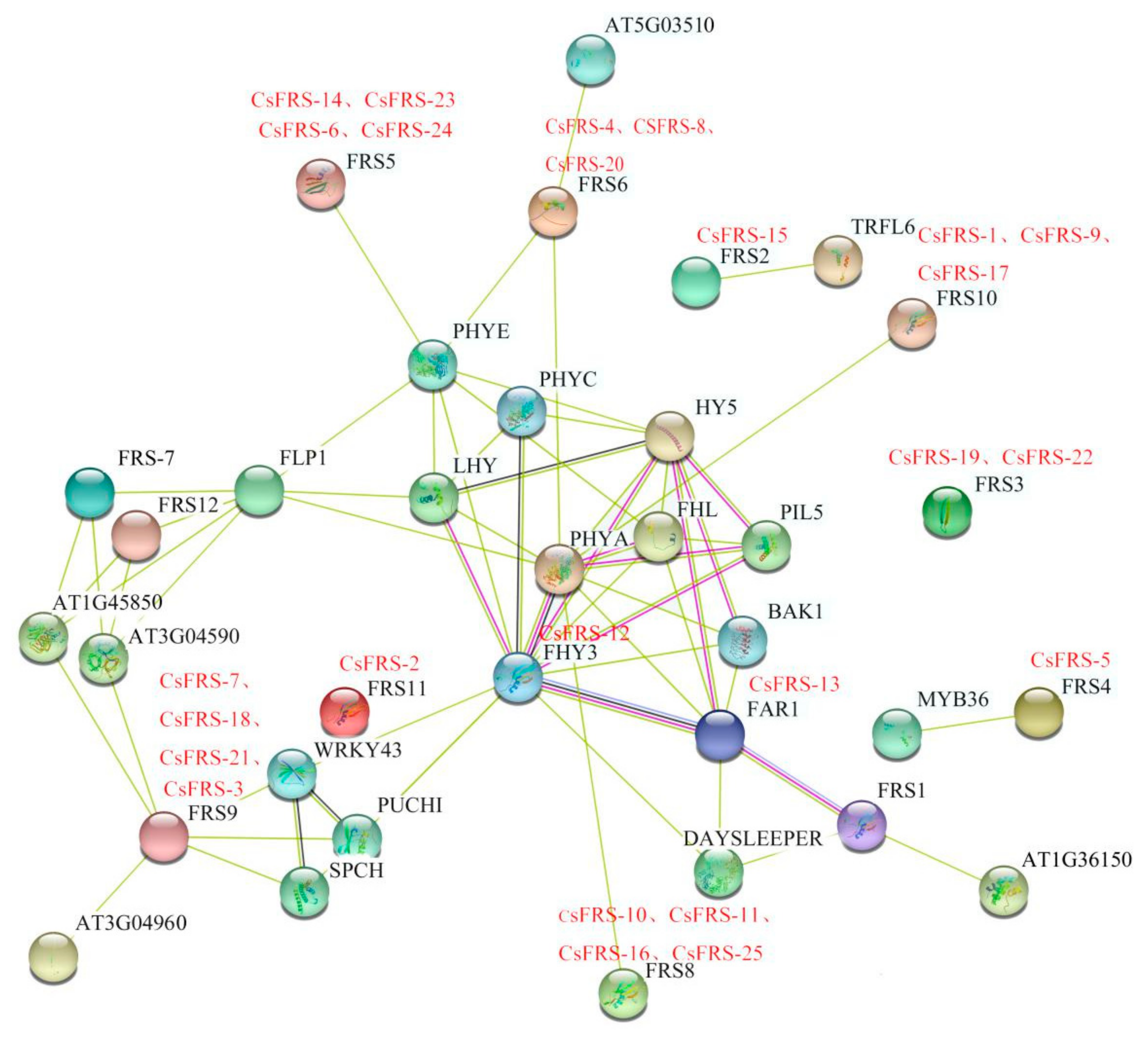
2.5. Protein Interaction Network of CsFHY3/FAR1 Proteins
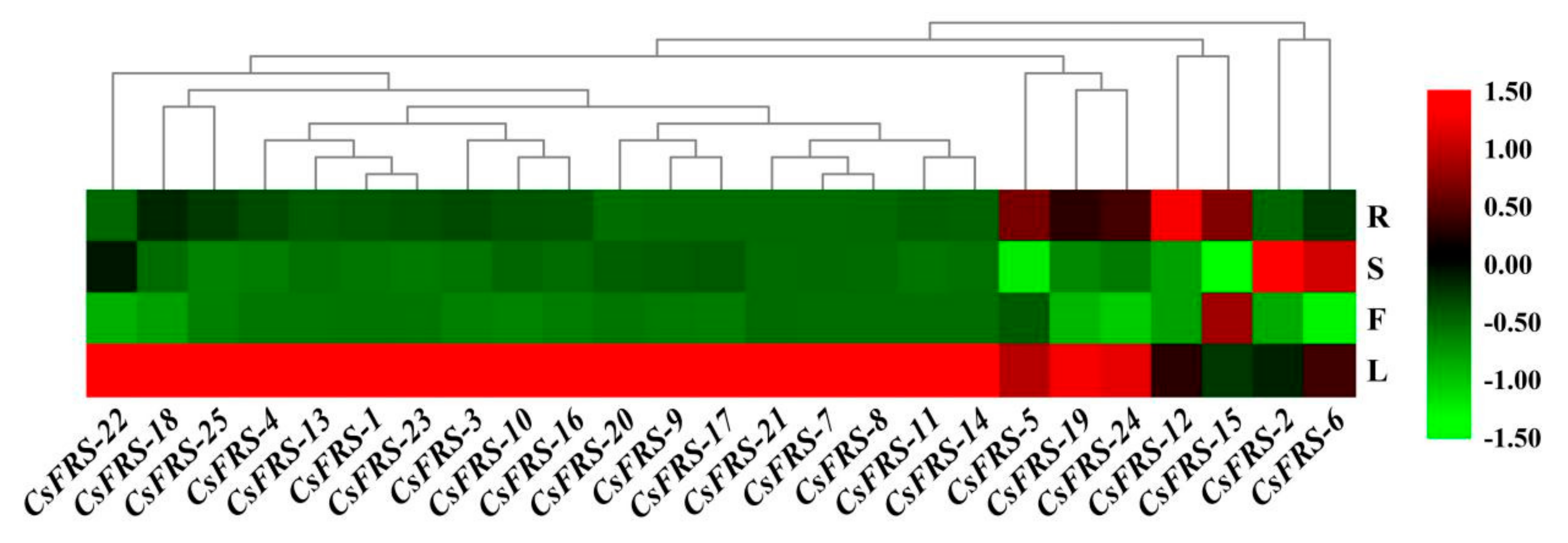
2.6. Expression Analysis in the Different Tissues of Tea Plants
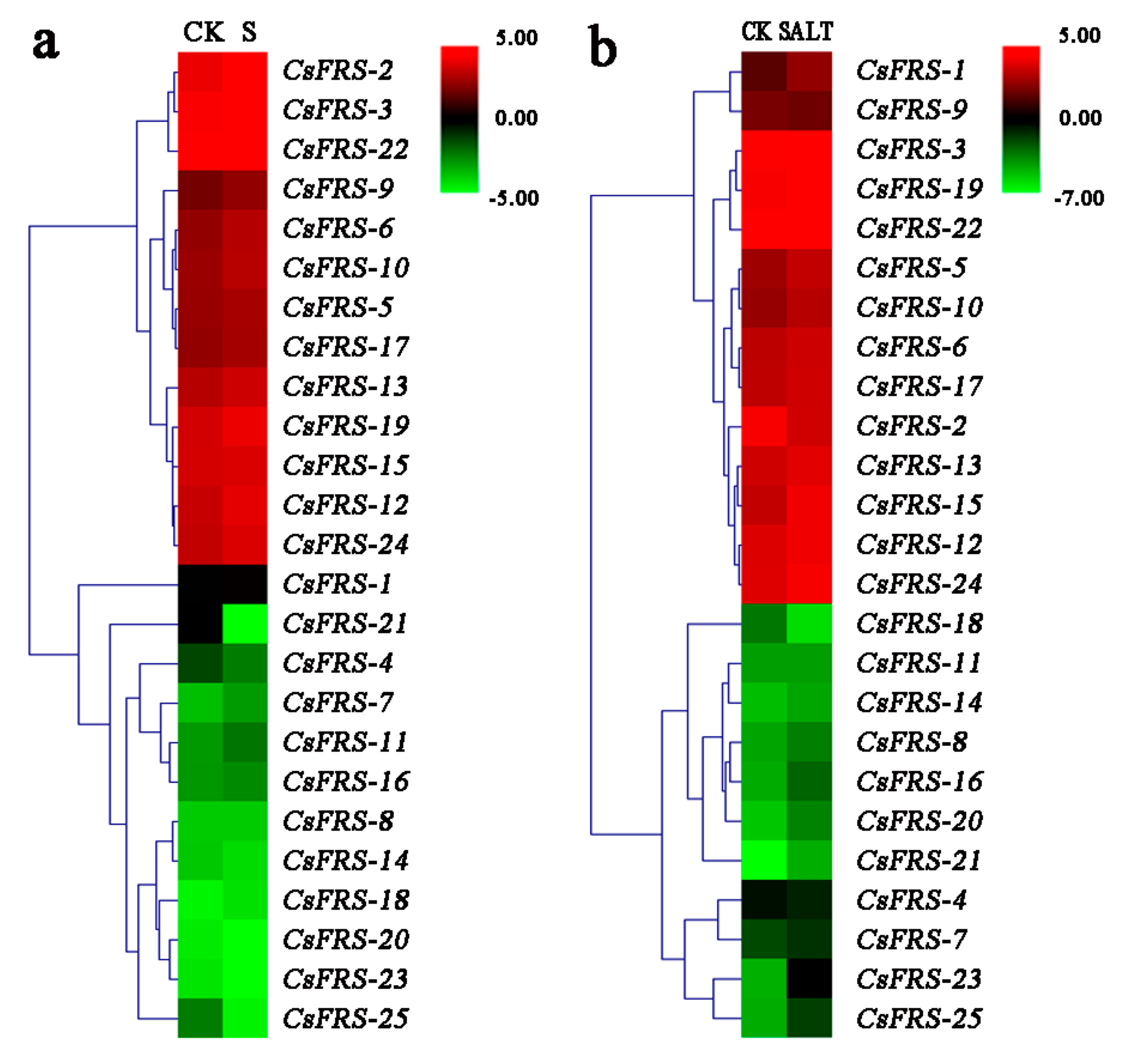
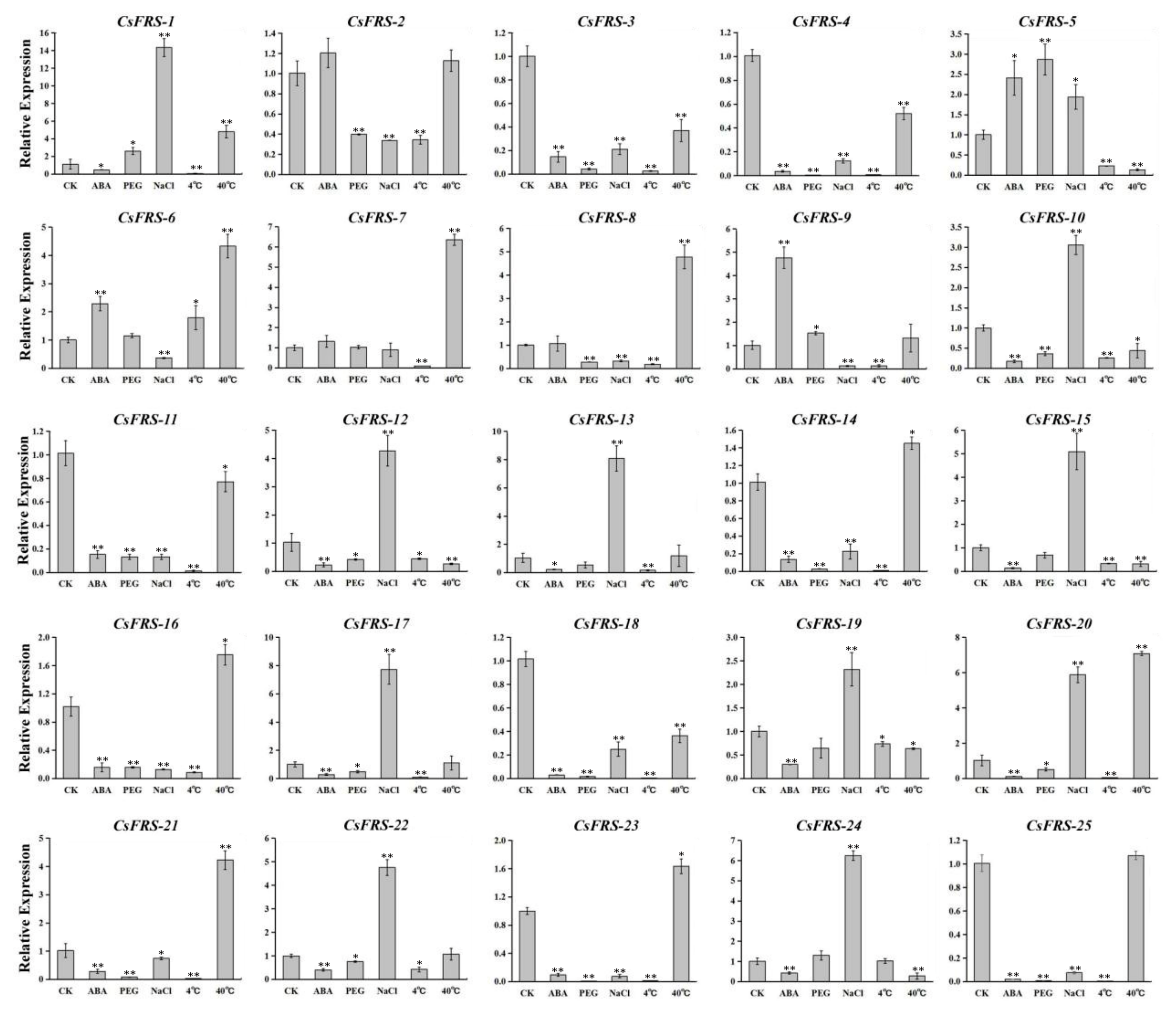
2.7. Expression Analysis of CsFHY3/FAR1s under Different Stresses
3. Discussion
4. Conclusions
5. Materials and Methods
5.1. Plant Materials and Stress Treatments
5.2. Identification of the CsFHY3/FAR1 Gene Family in Tea Plants
5.3. Sequence Analysis and Phylogenetic Tree Construction
5.4. Analysis of Gene Structure and Conserved Motifs
5.5. Prediction of the Cis-Acting Elements and Protein Interaction Network
5.6. Transcriptome Analysis
5.7. Quantitative Real-Time PCR (qRT-PCR) Analysis
5.8. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huai, J.; Zhang, X.; Li, J.; Ma, T.; Zha, P.; Jing, Y.; Lin, R. SEUSS and PIF4 coordinately regulate light and temperature signaling pathways to control plant growth. Mol. Plant 2018, 11, 928–942. [Google Scholar] [CrossRef] [PubMed]
- Krzeszowiec, W.; Novokreshchenova, M.; Gabryś, H. Chloroplasts in C3 grasses move in response to blue-light. Plant Cell Rep. 2020, 39, 1331–1343. [Google Scholar] [CrossRef]
- Casal, J.J. Phytochromes, cryptochromes, phototropin: Photoreceptor interactions in plants. Photochem. Photobiol. 2000, 71, 1. [Google Scholar] [CrossRef]
- Heijde, M.; Ulm, R. UV-B photoreceptor-mediated signalling in plants. Trends Plant Sci. 2012, 17, 230–237. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Ouyang, X.; Yang, P.; Lau, O.S.; Li, G.; Li, J.; Chen, H.; Deng, X.W. Arabidopsis FHY3 and HY5 positively mediate induction of COP1 transcription in response to photomorphogenic UV-B light. Plant Cell 2012, 24, 4590–4607. [Google Scholar] [CrossRef]
- Franklin, K.A.; Davis, S.J.; Stoddart, W.M.; Vierstra, R.D.; Whitelam, G.C. Mutant analyses define multiple roles for phytochrome C in Arabidopsis photomorphogenesis. Plant Cell 2003, 15, 1981–1989. [Google Scholar] [CrossRef]
- Jung, J.H.; Domijan, M.; Klose, C.; Biswas, S.; Ezer, D.; Gao, M.; Khattak, A.K.; Box, M.S.; Charoensawan, V.; Cortijo, S.; et al. Phytochromes function as thermosensors in Arabidopsis. Science 2016, 354, 886–889. [Google Scholar] [CrossRef]
- Oide, M.; Hikima, T.; Oroguchi, T.; Kato, T.; Yamaguchi, Y.; Yoshihara, S.; Yamamoto, M.; Nakasako, M.; Okajima, K. Molecular shape under far-red light and red light-induced association of Arabidopsis phytochrome B. FEBS J. 2020, 287, 1612–1625. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.; Ding, L.; Casola, C. Transposase-derived transcription factors regulate light signaling in Arabidopsis. Science 2007, 318, 1302–1305. [Google Scholar] [CrossRef]
- Wang, H.; Deng, X. Arabidopsis FHY3 defines a key phytochrome A signaling component directly interacting with its homologous partner FAR1. EMBO J. 2002, 21, 1339–1349. [Google Scholar] [CrossRef]
- Wang, H.; Wang, H. Multifaceted roles of FHY3 and FAR1 in light signaling and beyond. Trends Plant Sci. 2015, 20, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, G.; Gao, S.; Martinez, C.; He, G.; Zhou, Z.; Huang, X.; Lee, J.H.; Zhang, H.; Shen, Y.; et al. Arabidopsis transcription factor ELONGATED HYPOCOTYL5 plays a role in the feedback regulation of Phytochrome a signaling. Plant Cell 2010, 22, 3634–3649. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Siddiqui, H.; Teng, Y.; Lin, R.; Wan, X.; Li, J.; Lau, O.S.; Ouyang, X.; Dai, M.; Wan, J.; et al. Coordinated transcriptional regulation underlying the circadian clock in Arabidopsis. Nat. Cell Biol. 2011, 13, 616–622. [Google Scholar] [CrossRef]
- Hudson, M.E.; Lisch, D.R.; Quail, P.H. The FHY3 and FAR1 genes encode transposase-related proteins involved in regulation of gene expression by the phytochrome A-signaling pathway. Plant J. 2003, 34, 453–471. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.; Wang, H. Arabidopsis FHY3/FAR1 gene family and distinct roles of its members in light control of Arabidopsis development. Plant Physiol. 2004, 136, 4010–4022. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Li, G. FAR1-RELATED SEQUENCE (FRS) AND FRS-RELATED FACTOR (FRF) family proteins in Arabidopsis growth and development. Front. Plant Sci. 2018, 9, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Yan, X.; Maurais, S.; Fu, H.; O′Brien, D.G.; Mottinger, J.; Dooner, H.K. Jittery, a mutator distant relative with a paradoxical mobile behavior: Excision without reinsertion. Plant Cell 2004, 16, 1105–1114. [Google Scholar] [CrossRef]
- Lin, R.; Teng, Y.; Park, H.J.; Ding, L.; Black, C.; Fang, P.; Wang, H. Discrete and essential roles of the multiple domains of Arabidopsis FHY3 in mediating phytochrome A signal transduction. Plant Physiol. 2008, 148, 981–992. [Google Scholar] [CrossRef]
- Li, D.; Fu, X.; Guo, L.; Huang, Z.; Li, Y.; Liu, Y.; He, Z.; Cao, X.; Ma, X.; Zhao, M.; et al. FAR-RED ELONGATED HYPOCOTYL3 activates SEPALLATA2 but inhibits CLAVATA3 to regulate meristem determinacy and maintenance in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, 9375–9380. [Google Scholar] [CrossRef] [PubMed]
- O′Malley, R.C.; Huang, S.S.C.; Song, L.; Lewsey, M.G.; Bartlett, A.; Nery, J.R.; Galli, M.; Gallavotti, A.; Ecker, J.R. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 2016, 165, 1280–1292. [Google Scholar] [CrossRef]
- Ouyang, X.; Li, J.; Li, G.; Li, B.; Chen, B.; Shen, H.; Huang, X.; Mo, X.; Wan, X.; Lin, R.; et al. Genome-wide binding site analysis of FAR-RED ELONGATED HYPOCOTYL3 reveals its novel function in Arabidopsis development. Plant Cell 2011, 23, 2514–2535. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Liu, H.; An, C.; Shi, Y.; Liu, X.; Yuan, W.; Zhang, B.; Yang, J.; Yu, C.; Gao, H. Arabidopsis FRS4/CPD25 and FHY3/CPD45 work cooperatively to promote the expression of the chloroplast division gene ARC5 and chloroplast division. Plant J. 2013, 75, 795–807. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ma, M.; Li, G.; Yuan, L.; Xie, Y.; Wei, H.; Ma, X.; Li, Q.; Devlin, P.F.; Xu, X.; et al. Transcription factors FHY3 and FAR1 regulate light-induced circadian clock AsSOCIATED1 gene expression in Arabidopsis. Plant Cell 2020, 32, 1464–1478. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Liu, Y.; Ma, M.; Zhou, Q.; Zhao, Y.; Zhao, B.; Wang, B.; Wei, H.; Wang, H. Arabidopsis FHY3 and FAR1 integrate light and strigolactone signaling to regulate branching. Nat. Commun. 2020, 11, 1–13. [Google Scholar] [CrossRef]
- Xie, Y.; Zhou, Q.; Zhao, Y.; Li, Q.; Liu, Y.; Ma, M.; Wang, B.; Shen, R.; Zheng, Z.; Wang, H. FHY3 and FAR1 integrate light signals with the miR156-SPL module-mediated aging pathway to regulate Arabidopsis flowering. Mol. Plant 2020, 13, 483–498. [Google Scholar] [CrossRef]
- Tang, W.; Wang, W.; Chen, D.; Ji, Q.; Jing, Y.; Wang, H.; Lin, R. Transposase-derived proteins FHY3/FAR1 interact with PHYTOCHROME-INTERACTING FACTOR1 to regulate chlorophyll biosynthesis by modulating HEMB1 during deetiolation in Arabidopsis. Plant Cell 2012, 24, 1984–2000. [Google Scholar] [CrossRef]
- Ma, L.; Xue, N.; Fu, X.; Zhang, H.; Li, G. Arabidopsis thaliana FAR-RED ELONGATED HYPOCOTYLS3 (FHY3) and FAR-RED-IMPAIRED RESPONSE1 (FAR1) modulate starch synthesis in response to light and sugar. New Phytol. 2017, 213, 1682–1696. [Google Scholar] [CrossRef]
- Tang, W.; Ji, Q.; Huang, Y.; Jiang, Z.; Bao, M.; Wang, H.; Lin, R. FAR-RED ELONGATED HYPOCOTYl3 and FAR-RED IMPAIRED RESPONSE1 transcription factors integrate light and abscisic acid signaling in Arabidopsis. Plant Physiol. 2013, 163, 857–866. [Google Scholar] [CrossRef]
- Ma, L.; Tian, T.; Lin, R.; Deng, X.W.; Wang, H.; Li, G. Arabidopsis FHY3 and FAR1 regulate light-induced myo-inositol biosynthesis and oxidative stress responses by transcriptional activation of MIPS1. Mol. Plant 2016, 9, 541–557. [Google Scholar] [CrossRef]
- Wang, W.; Tang, W.; Ma, T.; Niu, D.; Jin, J.B.; Wang, H.; Lin, R. A pair of light signaling factors FHY3 and FAR1 regulates plant immunity by modulating chlorophyll biosynthesis. J. Integr. Plant Biol. 2016, 58, 91–103. [Google Scholar] [CrossRef]
- Liu, Y.; Xie, Y.; Wang, H.; Ma, X.; Yao, W.; Wang, H. Light and ethylene coordinately regulate the phosphate starvation response through transcriptional regulation of PHOSPHATE STARVATION RESPONSE1. Plant Cell 2017, 29, 2269–2284. [Google Scholar] [CrossRef]
- Liu, Y.; Wei, H.; Ma, M.; Li, Q.; Kong, D.; Sun, J.; Ma, X.; Wang, B.; Chen, C.; Xie, Y.; et al. Arabidopsis FHY3 and FAR1 regulate the balance between growth and defense responses under shade conditions. Plant Cell 2019, 31, 2089–2106. [Google Scholar] [CrossRef]
- Stirnberg, P.; Zhao, S.; Williamson, L.; Ward, S.; Leyser, O. FHY3 promotes shoot branching and stress tolerance in Arabidopsis in an AXR1-dependent manner. Plant J. 2012, 71, 907–920. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, F.; Kimura, K.; Saba, T.; Ogino, A.; Yamaguchi, S.; Tanaka, J. Worldwide core collections of tea (Camellia sinensis) based on SSR markers. Tree Genet. Genomes 2014, 10, 1555–1565. [Google Scholar] [CrossRef]
- Zhao, L.; Li, Z.; Feng, G.; Ji, X.; Tan, Y.; Li, H.; Gunter, M.J.; Xiang, Y. Tea drinking and risk of cancer incidence: A meta-analysis of prospective cohort studies and evidence evaluation. Adv. Nutr. 2020, 1–11. [Google Scholar] [CrossRef]
- Zhou, L.; Xu, H.; Zhu, X.; Chen, X.; Wang, Y.; Fang, W.; Li, X. Effect of abscisic acid on physiological characteristics of tea plant under drought stress. J. Tea Sci. 2014, 473–480. [Google Scholar] [CrossRef]
- Wei, C.; Yang, H.; Wang, S.; Zhao, J.; Liu, C.; Gao, L.; Xia, E.; Lu, Y.; Tai, Y.; She, G.; et al. Draft genome sequence of Camellia sinensis var. sinensis provides insights into the evolution of the tea genome and tea quality. Proc. Natl. Acad. Sci. USA 2018, 115, E4151–E4158. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Han, Y.; Meng, D.; Li, D.; Jin, Q.; Lin, Y.; Cai, Y. Structural, evolutionary, and functional analysis of the class III peroxidase gene family in Chinese pear (pyrus bretschneideri). Front. Plant Sci. 2016, 7, 1–12. [Google Scholar] [CrossRef]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon-intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van De Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Ma, Q.; Zou, Z.; Sun, K.; Yao, Y.; Tao, J.; Kaleri, N.A.; Li, X. Molecular link between leaf coloration and gene expression of flavonoid and carotenoid biosynthesis in Camellia sinensis cultivar ‘Huangjinya’. Front. Plant Sci. 2017, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Wan, S.; Wang, W.; Zhou, T.; Zhang, Y.; Chen, J.; Xiao, B.; Yang, Y.; Yu, Y. Transcriptomic analysis reveals the molecular mechanisms of Camellia sinensis in response to salt stress. Plant Growth Regul. 2018, 84, 481–492. [Google Scholar] [CrossRef]
- Jeffares, D.C.; Penkett, C.J.; Bähler, J. Rapidly regulated genes are intron poor. Trends Genet. 2008, 24, 375–378. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Liu, J.; Lu, J.; Zhai, Y.; Wang, H.; Gong, Z.; Wang, S.; Lu, M. Genome-wide analysis of the CaHsp20 gene family in pepper: Comprehensive sequence and expression profile analysis under heat stress. Front. Plant Sci. 2015, 6, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Yuan, N.; Wang, T.; Liu, T.; Yang, Y.; Du, J. Genome-wide analysis of the FAR1/FHY3 gene family in Cotton. Cott. Sci. 2018, 14, 1–11. [Google Scholar]
- Yao, L.; Hao, X.; Cao, H.; Ding, C.; Yang, Y.; Wang, L.; Wang, X. ABA-dependent bZIP transcription factor, CsbZIP18, from Camellia sinensis negatively regulates freezing tolerance in Arabidopsis. Plant Cell Rep. 2020, 39, 553–565. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.H.; Wan, S.Q.; Wang, W.D.; Chen, J.F.; Huang, L.L.; Duan, M.S.; Yu, Y.B. Genome-wide identification and characterization of the CsSnRK2 family in Camellia sinensis. Plant Physiol. Biochem. 2018, 132, 287–296. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef]
- Xia, E.; Zhang, H.; Sheng, J.; Li, K.; Zhang, Q.; Kim, C.; Zhang, Y.; Liu, Y.; Zhu, T.; Li, W.; et al. The tea tree genome provides insights into tea flavor and independent evolution of caffeine biosynthesis. Mol. Plant 2017, 10, 866–877. [Google Scholar] [CrossRef]
- Li, Y.; Liang, L.; Fan, F.; Shi, Y. Gene cloning and expression analysis of GAGP in tea plant. Bioinformatics 2015, 64–72. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
| Gene Name | Gene ID | Amino Acids | MW (kDa) | pI | GRAVY | Instability Index | Aliphatic Index | Subcellular Location |
|---|---|---|---|---|---|---|---|---|
| CsFRS-1 | CSS002622.1 | 689 | 80.13 | 9.10 | −0.434 | 46.00 | 77.21 | chlo:4, vacu:3, cyto:2, nucl:1.5, cysk_nucl:1.5, mito:1, plas:1, extr:1 |
| CsFRS-2 | CSS016876.1 | 757 | 86.03 | 6.20 | −0.318 | 47.61 | 77.64 | nucl:13, vacu:1 |
| CsFRS-3 | CSS003256.1 | 769 | 87.81 | 5.79 | −0.509 | 53.55 | 72.65 | nucl:13, cyto:1 |
| CsFRS-4 | CSS001271.1 | 588 | 68.71 | 8.91 | −0.560 | 48.43 | 68.59 | nucl:7, pero:2, cysk:1.5, cysk_plas:1.5, cyto:1, mito:1, vacu:1 |
| CsFRS-5 | CSS001125.1 | 752 | 86.58 | 6.47 | −0.539 | 50.51 | 76.77 | nucl:13, vacu:1 |
| CsFRS-6 | CSS013735.1 | 787 | 90.41 | 6.26 | −0.565 | 52.42 | 71.60 | nucl:14 |
| CsFRS-7 | CSS042110.1 | 368 | 43.19 | 8.42 | −0.428 | 39.24 | 65.76 | cyto:7, cysk:5, nucl:1, golg:1 |
| CsFRS-8 | CSS022313.1 | 702 | 80.33 | 9.28 | −0.385 | 41.50 | 80.24 | nucl:11, cyto:2, cysk:1 |
| CsFRS-9 | CSS018104.1 | 691 | 80.13 | 6.04 | −0.351 | 42.06 | 77.54 | nucl:7, chlo:4, mito:2, cysk:1 |
| CsFRS-10 | CSS039560.1 | 897 | 103.05 | 7.36 | −0.568 | 49.04 | 70.51 | nucl:13, plas:1 |
| CsFRS-11 | CSS007312.1 | 690 | 79.47 | 9.06 | −0.436 | 39.05 | 70.04 | nucl:12, cyto:1, cysk:1 |
| CsFRS-12 | CSS040860.1 | 846 | 97.07 | 6.98 | −0.687 | 52.72 | 63.43 | nucl:14 |
| CsFRS-13 | CSS040864.1 | 767 | 88.40 | 7.38 | −0.692 | 57.85 | 66.75 | nucl:6, pero:4, chlo:2, cyto:1, mito_plas:1 |
| CsFRS-14 | CSS035662.1 | 659 | 75.35 | 9.11 | −0.428 | 39.71 | 68.21 | chlo:9, nucl:4, vacu:1 |
| CsFRS-15 | CSS052639.1 | 832 | 95.68 | 7.49 | −0.501 | 54.13 | 74.13 | nucl:13, vacu:1 |
| CsFRS-16 | CSS022572.1 | 680 | 79.60 | 8.97 | −0.558 | 47.81 | 77.25 | cyto:8, nucl:4, mito:1, plas:1 |
| CsFRS-17 | CSS003022.1 | 691 | 79.70 | 6.03 | −0.357 | 44.10 | 79.58 | nucl:6, mito:3.5, cyto_mito:2.5, chlo:2, cysk:2 |
| CsFRS-18 | CSS008927.1 | 402 | 47.13 | 8.91 | −0.491 | 48.24 | 67.69 | cyto:8, nucl:2, plas:2, mito:1, pero:1 |
| CsFRS-19 | CSS040724.1 | 848 | 96.73 | 8.70 | −0.525 | 45.91 | 70.06 | nucl:13, cyto:1 |
| CsFRS-20 | CSS025022.1 | 680 | 79.13 | 6.56 | −0.594 | 55.50 | 68.50 | nucl:13, vacu:1 |
| CsFRS-21 | CSS052868.1 | 727 | 83.96 | 8.33 | −0.583 | 35.12 | 64.68 | nucl:12, plas:1, vacu:1 |
| CsFRS-22 | CSS032048.1 | 890 | 101.02 | 6.70 | −0.532 | 44.93 | 67.07 | nucl:12, cyto:2 |
| CsFRS-23 | CSS025195.1 | 529 | 61.07 | 8.92 | −0.460 | 41.99 | 65.80 | chlo:11, nucl:2, mito:1 |
| CsFRS-24 | CSS043153.1 | 799 | 90.80 | 6.04 | −0.467 | 47.21 | 72.60 | nucl:13, chlo:1 |
| CsFRS-25 | CSS050875.1 | 616 | 71.73 | 6.65 | −0.501 | 47.28 | 76.25 | nucl:9, cyto:4, plas:1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Z.; An, C.; Zhao, Y.; Xiao, Y.; Bao, L.; Gong, C.; Gao, Y. Genome-Wide Identification and Characterization of the CsFHY3/FAR1 Gene Family and Expression Analysis under Biotic and Abiotic Stresses in Tea Plants (Camellia sinensis). Plants 2021, 10, 570. https://doi.org/10.3390/plants10030570
Liu Z, An C, Zhao Y, Xiao Y, Bao L, Gong C, Gao Y. Genome-Wide Identification and Characterization of the CsFHY3/FAR1 Gene Family and Expression Analysis under Biotic and Abiotic Stresses in Tea Plants (Camellia sinensis). Plants. 2021; 10(3):570. https://doi.org/10.3390/plants10030570
Chicago/Turabian StyleLiu, Zhengjun, Chuanjing An, Yiqing Zhao, Yao Xiao, Lu Bao, Chunmei Gong, and Yuefang Gao. 2021. "Genome-Wide Identification and Characterization of the CsFHY3/FAR1 Gene Family and Expression Analysis under Biotic and Abiotic Stresses in Tea Plants (Camellia sinensis)" Plants 10, no. 3: 570. https://doi.org/10.3390/plants10030570
APA StyleLiu, Z., An, C., Zhao, Y., Xiao, Y., Bao, L., Gong, C., & Gao, Y. (2021). Genome-Wide Identification and Characterization of the CsFHY3/FAR1 Gene Family and Expression Analysis under Biotic and Abiotic Stresses in Tea Plants (Camellia sinensis). Plants, 10(3), 570. https://doi.org/10.3390/plants10030570







