Development and Characterization of 15 Novel Genomic SSRs for Viburnum farreri
Abstract
1. Introduction
2. Results and Discussion
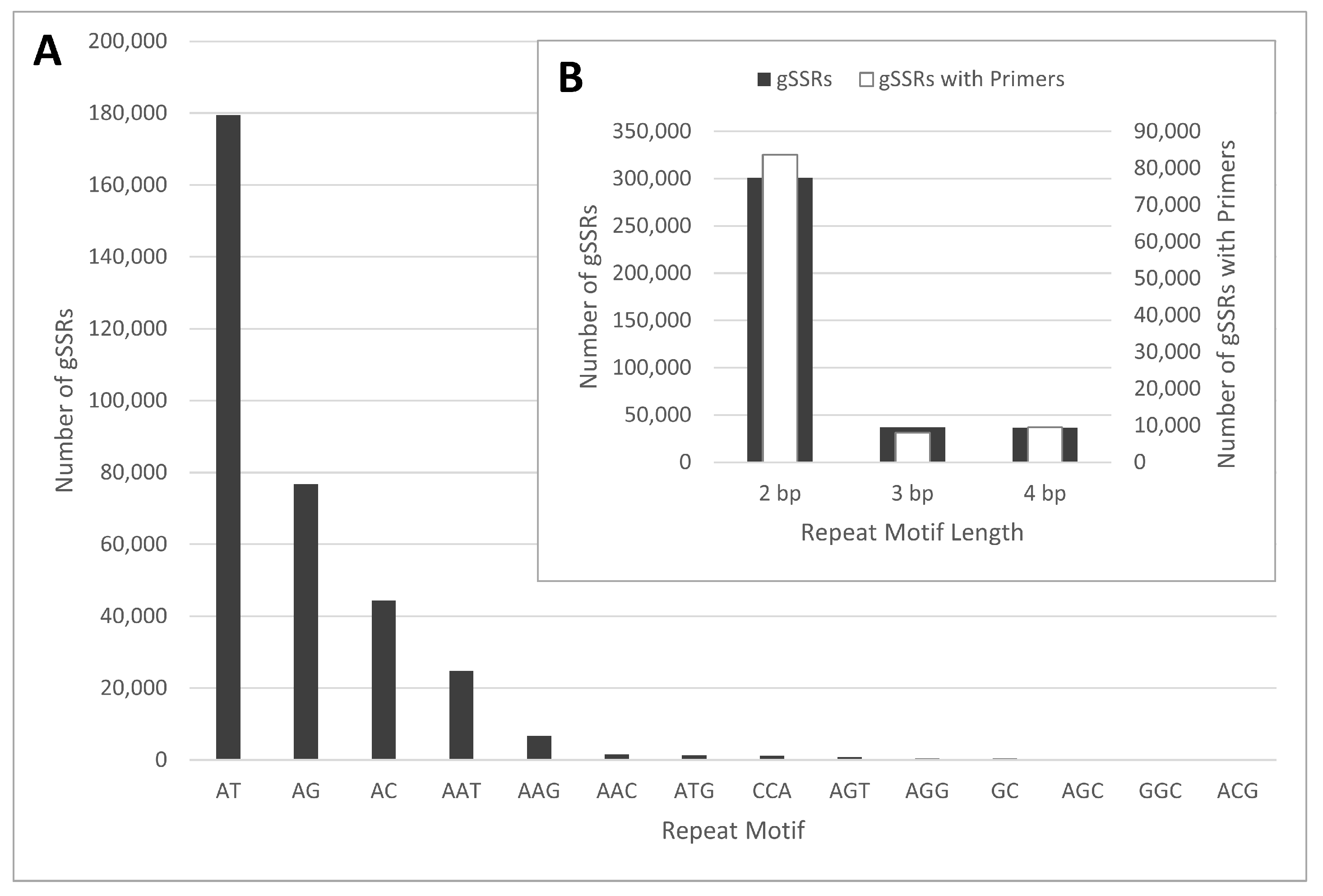
2.1. gSSR Development
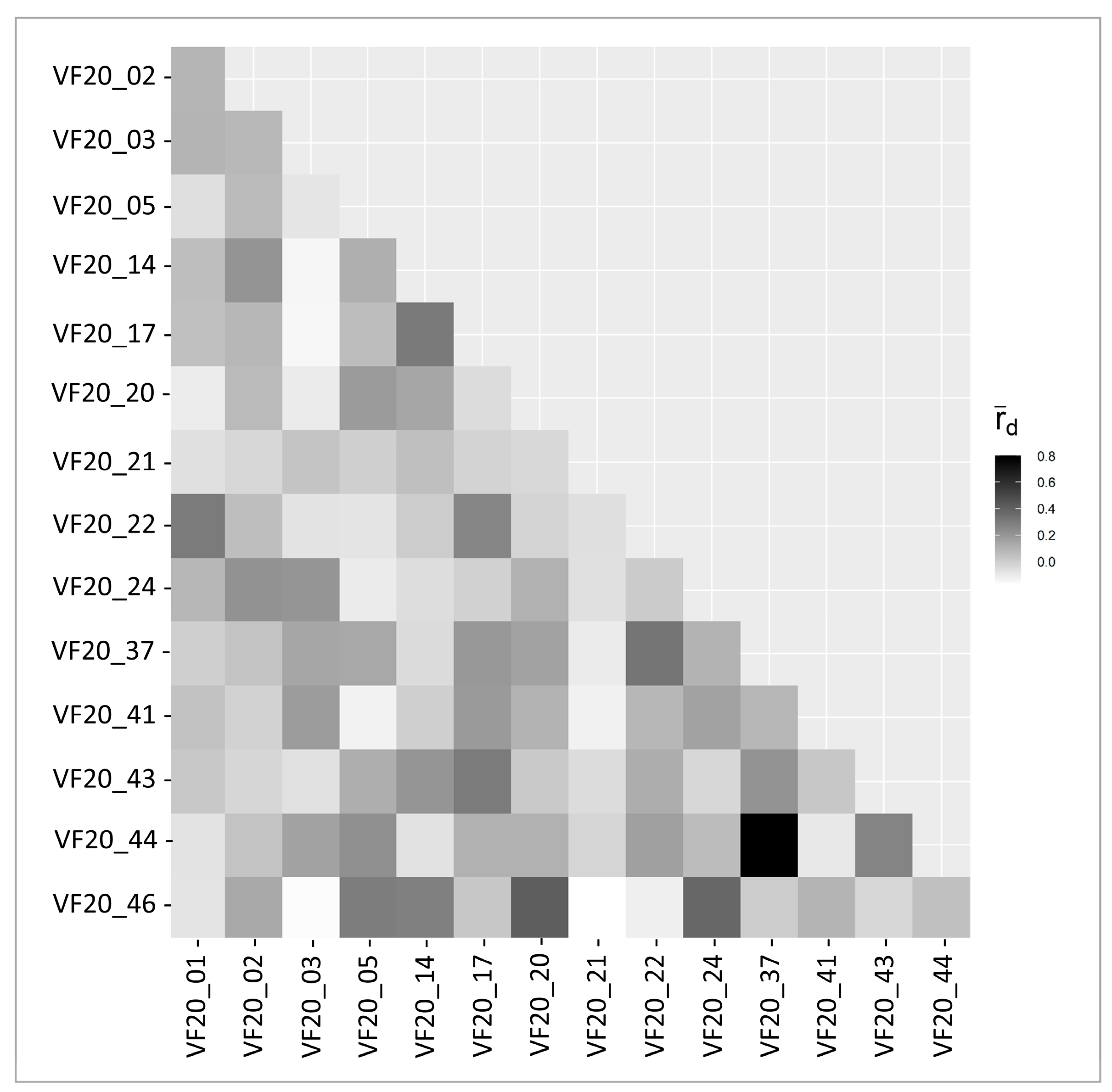
2.2. gSSR Characteristics and Diversity Indices
3. Materials and Methods
3.1. Plant Materials and gDNA Extraction
3.2. gSSR Development and Screening
3.3. Estimation of Diversity Indices
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Li, A. An update of the angiosperm phylogeny group classification for the orders and families of flowering plants: APG II. Bot. J. Linn. Soc. 2003, 141, 399–436. [Google Scholar]
- Landis, M.J.; Eaton, D.A.R.; Clement, W.L.; Park, B.; Spriggs, E.L.; Sweeney, P.W.; Edwards, E.J.; Donoghue, M.J. Joint phylogenetic estimation of geographic movements and biome shifts during the global diversification of Viburnum. Syst. Biol. 2021, 70, 67–85. [Google Scholar] [CrossRef]
- Hoch, W.; Zeldin, E.; Nienhuis, J.; McCown, B. Generation and identification of new Viburnum hybrids. J. Environ. Hortic. 1995, 13, 193–195. [Google Scholar] [CrossRef]
- Winkworth, R.C.; Donoghue, M.J. Viburnum phylogeny based on combined molecular data: Implications for taxonomy and biogeography. Am. J. Bot. 2005, 92, 653–666. [Google Scholar] [CrossRef]
- Morton, C.V. The Mexican and Central American Species of Viburnum. In Systematic Plant Studies: Mainly Tropical America; Government Printing Office: Washington, DC, USA, 1933; pp. 339–366. [Google Scholar]
- Rehder, A. The Viburnums of Eastern Asia. In Trees and Shrubs; Sargent, C.S., Ed.; Houghton Mifflin: Boston, MA, USA, 1908; Volume 2, pp. 105–116. [Google Scholar]
- Donoghue, M.J. A Preliminary analysis of phylogenetic relationships in Viburnum (Caprifoliaceae s.1.). Syst. Bot. 1983, 8, 45. [Google Scholar] [CrossRef]
- Nicolson, D.H.; Hara, H. A revision of Caprifoliaceae of Japan with Reference to Allied Plants in Other Districts and the Adoxaceae; Academia Scientific Book: Tokyo, Japan, 1983. [Google Scholar]
- Dirr, M. Viburnums: Flowering Shrubs for Every Season; Timber Press: Portland, OR, USA, 2007. [Google Scholar]
- USDA-NASS. Census of Horticultural Specialties: Table Deciduous Shrubs; USDA-NASS: Washington, DC, USA, 2020.
- Edwards, E.J.; Chatelet, D.S.; Chen, B.-C.; Ong, J.Y.; Tagane, S.; Kanemitsu, H.; Tagawa, K.; Teramoto, K.; Park, B.; Chung, K.-F.; et al. Convergence, consilience, and the evolution of temperate deciduous forests. Am. Nat. 2017, 190, S87–S104. [Google Scholar] [CrossRef] [PubMed]
- Lens, F.; Vos, R.A.; Charrier, G.; Van Der Niet, T.; Merckx, V.; Baas, P.; Gutierrez, J.A.; Jacobs, B.; Dória, L.C.; Smets, E.; et al. Scalariform-to-simple transition in vessel perforation plates triggered by differences in climate during the evolution of Adoxaceae. Ann. Bot. 2016, 118, 1043–1056. [Google Scholar] [CrossRef]
- Spriggs, E.L.; Clement, W.L.; Sweeney, P.W.; Madriñán, S.; Edwards, E.J.; Donoghue, M.J. Temperate radiations and dying embers of a tropical past: The diversification of Viburnum. N. Phytol. 2014, 207, 340–354. [Google Scholar] [CrossRef] [PubMed]
- Clement, W.L.; Donoghue, M.J. Dissolution of Viburnum section megalotinus (Adoxaceae) of Southeast Asia and its implications for morphological evolution and biogeography. Int. J. Plant. Sci. 2011, 172, 559–573. [Google Scholar] [CrossRef]
- Choi, Y.G.; Youm, J.W.; Lim, C.E.; Oh, S.-H. Phylogenetic analysis of Viburnum (Adoxaceae) in Korea using DNA sequences. Korean J. Plant. Taxon. 2018, 48, 206–217. [Google Scholar] [CrossRef][Green Version]
- Eaton, D.A.R.; Spriggs, E.L.; Park, B.; Donoghue, M.J. Misconceptions on missing data in RAD-seq phylogenetics with a deep-scale example from flowering plants. Syst. Biol. 2016, 66, 399–412. [Google Scholar] [CrossRef] [PubMed]
- Clement, W.L.; Donoghue, M.J. Barcoding success as a function of phylogenetic relatedness in Viburnum, a clade of woody angiosperms. BMC Evol. Biol. 2012, 12, 73. [Google Scholar] [CrossRef] [PubMed]
- Donoghue, M.J.; Baldwin, B.G.; Li, J.; Winkworth, R.C. Viburnum phylogeny based on chloroplast trnK intron and nuclear ribosomal ITS DNA sequences. Syst. Bot. 2004, 29, 188–198. [Google Scholar] [CrossRef]
- Winkworth, R.C.; Donoghue, M.J. Viburnum phylogeny: Evidence from the duplicated nuclear gene GBSSI. Mol. Phylogenetics Evol. 2004, 33, 109–126. [Google Scholar] [CrossRef]
- Donoghue, M.J.; Olmstead, R.G.; Smith, J.F.; Palmer, J.D. Phylogenetic relationships of dipsacales based on rbcL sequences. Ann. Mo. Bot. Gard. 1992, 79, 333. [Google Scholar] [CrossRef]
- Clement, W.L.; Arakaki, M.; Sweeney, P.W.; Edwards, E.J.; Donoghue, M.J. A chloroplast tree for Viburnum (Adoxaceae) and its implications for phylogenetic classification and character evolution. Am. J. Bot. 2014, 101, 1029–1049. [Google Scholar] [CrossRef] [PubMed]
- Donoghue, M. Flowering times in Viburnum. Arnoldia 1980, 40, 2–22. [Google Scholar]
- Stearn, W.T. Viburnum farreri, a new name for V. fragrans Bunge. Taxon 1966, 15, 22–23. [Google Scholar]
- Spriggs, E.L.; Eaton, D.A.R.; Sweeney, P.W.; Schlutius, C.; Edwards, E.J.; Donoghue, M.J. Restriction-site-associated DNA sequencing reveals a cryptic Viburnum species on the North American coastal plain. Syst. Biol. 2018, 68, 187–203. [Google Scholar] [CrossRef] [PubMed]
- Powell, W.; Machray, G.C.; Provan, J. Polymorphism revealed by simple sequence repeats. Trends Plant Sci. 1996, 1, 215–222. [Google Scholar] [CrossRef]
- Ellegren, H. Microsatellites: Simple sequences with complex evolution. Nat. Rev. Genet. 2004, 5, 435–445. [Google Scholar] [CrossRef]
- Gupta, P.; Varshney, R. The development and use of microsatellite markers for genetic analysis and plant breeding with emphasis on bread wheat. Euphytica 2000, 113, 163–185. [Google Scholar] [CrossRef]
- Edwards, T.P.; Trigiano, R.N.; Ownley, B.H.; Windham, A.S.; Wyman, C.R.; Wadl, P.A.; Hadziabdic, D. Genetic diversity and conservation status of Helianthus verticillatus, an endangered sunflower of the Southern United States. Front. Genet. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Ony, M.A.; Nowicki, M.; Boggess, S.L.; Klingeman, W.E.; Zobel, J.M.; Trigiano, R.N.; Hadziabdic, D. Habitat fragmentation influences genetic diversity and differentiation: Fine-scale population structure of Cercis canadensis (Eastern redbud). Ecol. Evol. 2020, 10, 3655–3670. [Google Scholar] [CrossRef] [PubMed]
- Nowicki, M.; Schilling, E.E.; Boggess, S.L.; Houston, L.C.; Huff, M.L.; Staton, M.E.; Lampley, J.A.; Trigiano, R.N. Development and characterization of genic microsatellites for the ornamental plant green and gold (Chrysogonum virginianum). HortScience 2019, 54, 395–400. [Google Scholar] [CrossRef]
- Luber, J.; Christ, J.A.; Ferreira, M.F.D.S.; Carrijo, T.T. Species delimitation within Campomanesia (Myrtaceae) using morphology and amplification profiles of microsatellite markers. Braz. J. Bot. 2020, 43, 131–137. [Google Scholar] [CrossRef]
- Tabbasam, N.; Zafar, Y.; Rahman, M.-U. Pros and cons of using genomic SSRs and EST-SSRs for resolving phylogeny of the genus Gossypium. Plant. Syst. Evol. 2013, 300, 559–575. [Google Scholar] [CrossRef][Green Version]
- Nowicki, M.; Houston, L.C.; Boggess, S.L.; Aiello, A.S.; Payá-Milans, M.; Staton, M.E.; Hayashida, M.; Yamanaka, M.; Eda, S.; Trigiano, R.N.; et al. Species diversity and phylogeography of Cornus kousa (Asian dogwood) captured by genomic and genic microsatellites. Ecol. Evol. 2020, 10, 8299–8312. [Google Scholar] [CrossRef] [PubMed]
- Peakall, R.; Gilmore, S.; Keys, W.; Morgante, M.; Rafalski, A. Cross-species amplification of soybean (Glycine max) simple sequence repeats (SSRs) within the genus and other legume genera: Implications for the transferability of SSRs in plants. Mol. Biol. Evol. 1998, 15, 1275–1287. [Google Scholar] [CrossRef] [PubMed]
- Ellis, J.R.; Burke, J.M. EST-SSRs as a resource for population genetic analyses. Heredity 2007, 99, 125–132. [Google Scholar] [CrossRef]
- Ouyang, P.; Kang, D.; Mo, X.; Tian, E.; Hu, Y.; Huang, R. Development and characterization of high-throughput EST-based SSR markers for Pogostemon cablin using transcriptome sequencing. Molecules 2018, 23, 2014. [Google Scholar] [CrossRef] [PubMed]
- Arnold, C.; Rossetto, M.; McNally, J.; Henry, R.J. The application of SSRs characterized for grape (Vitis vinifera) to conservation studies in Vitaceae. Am. J. Bot. 2002, 89, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Wadl, P.A.; Trigiano, R.N.; Werner, D.J.; Pooler, M.R.; Rinehart, T.A. Simple sequence repeat markers from Cercis canadensis show wide cross-species transfer and use in genetic studies. J. Am. Soc. Hortic. Sci. 2012, 137, 189–201. [Google Scholar] [CrossRef]
- Wadl, P.A.; Wang, X.; Moulton, J.K.; Hokanson, S.C.; Skinner, J.A.; Rinehart, T.A.; Reed, S.M.; Pantalone, V.R.; Trigiano, R.N. Transfer of Cornus florida and C. kousa simple sequence repeats to selected Cornus (Cornaceae) species. J. Am. Soc. Hortic. Sci. 2010, 135, 279–288. [Google Scholar] [CrossRef]
- Hatmaker, E.A.; Wadl, P.A.; Mantooth, K.; Scheffler, B.E.; Ownley, B.H.; Trigiano, R.N. Development of microsatellites from Fothergilla × intermedia (Hamamelidaceae) and cross transfer to four other genera within Hamamelidaceae. Appl. Plant. Sci. 2015, 3, 1400123. [Google Scholar] [CrossRef]
- Dean, D.; Wadl, P.A.; Wang, X.; Klingeman, W.E.; Ownley, B.H.; Rinehart, T.A.; Scheffler, B.E.; Trigiano, R.N. Screening and characterization of 11 novel microsatellite markers from Viburnum dilatatum. HortScience 2011, 46, 1456–1459. [Google Scholar] [CrossRef]
- Dean, D.; Wadl, P.A.; Hadziabdic, D.; Klingeman, W.E.; Ownley, B.H.; Rinehart, T.A.; Dattilo, A.J.; Scheffler, B.; Trigiano, R.N. Analysis of genetic diversity and population structure for the native tree Viburnum rufidulum occurring in Kentucky and Tennessee. J. Am. Soc. Hortic. Sci. 2015, 140, 523–531. [Google Scholar] [CrossRef]
- Barish, S.; Arakaki, M.; Edwards, E.J.; Donoghue, M.J.; Clement, W.L. Characterization of 16 microsatellite markers for the Oreinotinus cade of Viburnum (Adoxaceae). Appl. Plant. Sci. 2016, 4, 1600103. [Google Scholar] [CrossRef]
- Dean, D. Assessing the Genetic Diversity of the Genus Viburnum Using Simple Sequence Repeats. Ph.D. Thesis, University of Tennessee, Knoxville, TN, USA, 2014. [Google Scholar]
- Lin, E.; Zhuang, H.; Yu, J.; Liu, X.; Huang, H.; Zhu, M.; Tong, Z. Genome survey of Chinese fir (Cunninghamia lanceolata): Identification of genomic SSRs and demonstration of their utility in genetic diversity analysis. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Nowicki, M.; Zhao, Y.; Boggess, S.L.; Fluess, H.; Payá-Milans, M.; Staton, M.E.; Houston, L.C.; Hadziabdic, D.; Trigiano, R.N. Taraxacum kok-saghyz (Rubber dandelion) genomic microsatellite loci reveal modest genetic diversity and cross-amplify broadly to related species. Sci. Rep. 2019, 9, 1–17. [Google Scholar] [CrossRef]
- Agapow, P.-M.; Burt, A. Indices of multilocus linkage disequilibrium. Mol. Ecol. Notes 2001, 1, 101–102. [Google Scholar] [CrossRef]
- Ambreen, H.; Kumar, S.; Variath, M.T.; Joshi, G.; Bali, S.; Agarwal, M.; Kumar, A.; Jagannath, A.; Goel, S. Development of genomic microsatellite markers in Carthamus tinctorius L. (Safflower) using next generation sequencing and assessment of their cross-species transferability and utility for diversity analysis. PLoS ONE 2015, 10, e0135443. [Google Scholar] [CrossRef]
- Diversity Arrays Technology. Plant DNA Extraction Protocol for DArT. Available online: https://www.diversityarrays.com/orderinstructions/plant-dna-extraction-protocol-for-dart/ (accessed on 20 May 2020).
- Kubisiak, T.L.; Nelson, C.D.; Staton, M.E.; Zhebentyayeva, T.; Smith, C.; Olukolu, B.A.; Fang, G.-C.; Hebard, F.V.; Anagnostakis, S.; Wheeler, N.; et al. A transcriptome-based genetic map of Chinese chestnut (Castanea mollissima) and identification of regions of segmental homology with peach (Prunus persica). Tree Genet. Genomes 2012, 9, 557–571. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 8 July 2020).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Simpson, J.T.; Wong, K.; Jackman, S.D.; Schein, J.E.; Jones, S.J.; Birol, I. ABySS: A parallel assembler for short read sequence data. Genome Res. 2009, 19, 1117–1123. [Google Scholar] [CrossRef] [PubMed]
- Morgulis, A.; Gertz, E.M.; Schäffer, A.A.; Agarwala, R. A fast and symmetric DUST implementation to mask low-complexity DNA sequences. J. Comput. Biol. 2006, 13, 1028–1040. [Google Scholar] [CrossRef] [PubMed]
- Staton, M.E.; Ficklin, S. Finding SSRs-Findssrs_altered.pl. Available online: https://github.com/statonlab/Finding-SSRs/blob/master/findSSRs_altered.pl (accessed on 8 July 2020).
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef]
- Don, R.H.; Cox, P.T.; Wainwright, B.J.; Baker, K.; Mattick, J.S. “Touchdown” PCR to circumvent spurious priming during gene amplification. Nucleic Acids Res. 1991, 19, 4008. [Google Scholar] [CrossRef]
- Amos, W.; Hoffman, J.I.; Frodsham, A.; Zhang, L.; Best, S.; Hill, A.V.S. Automated binning of microsatellite alleles: Problems and solutions. Mol. Ecol. Notes 2006, 7, 10–14. [Google Scholar] [CrossRef]
- Team, R.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. Peer J. 2014, 2, e281. [Google Scholar] [CrossRef]
- Goudet, J. hierfstat, a package for r to compute and test hierarchical F-statistics. Mol. Ecol. Notes 2005, 5, 184–186. [Google Scholar] [CrossRef]
| Locus | GenBank # | Primer Sequences | Repeat Motif | Allele Size Range (bp) | N | Missing (%) | H’ | Ho | He |
|---|---|---|---|---|---|---|---|---|---|
| VF20_01 | MW326735 | F: ACGATAAATGTGTATGCTCGC | [AT]6 | 203–205 | 2 | 0.00 | 0.66 | 0.00 | 0.48 |
| R: AACCCGGGAAGAAAGGTTACC | |||||||||
| VF20_02 | MW326736 | F: GAACCCTTTGAACACATGGCC | [AC]13 | 280–300 | 6 | 0.00 | 1.61 | 0.16 | 0.80 |
| R: CCAAGAAGCTTCGAAACTAGTTCC | |||||||||
| VF20_03 | MW326737 | F: AGCAATGTTCTAGGTCAGGGC | [GA]6 | 177–197 | 7 | 0.00 | 1.24 | 0.26 | 0.62 |
| R: CGATTTGCCCTAATCTTAGCGC | |||||||||
| VF20_05 | MW326738 | F: TGAAATGCAGACTGAAACGC | [AT]7 | 290–315 | 6 | 0.00 | 1.43 | 0.00 | 0.72 |
| R: GTTTGGTTCACGTCTGGTTGG | |||||||||
| VF20_14 | MW326739 | F: GGTTCACTGTTCATATGAATGATGC | [TC]7 | 218–245 | 6 | 5.26 | 1.51 | 0.11 | 0.75 |
| R: ATAAAGAAGTGCCACCCGTCC | |||||||||
| VF20_17 | MW326740 | F: GATGGTGCCAACTGATGAAGC | [AT]12 | 366-385 | 8 | 5.26 | 1.62 | 0.83 | 0.74 |
| R: GACTTCTAGGAGGTTGGTGCC | |||||||||
| VF20_20 | MW326741 | F: AATGCTCAAATTGCTTACGC | [TA]9 | 116–130 | 5 | 0.00 | 1.47 | 0.42 | 0.76 |
| R: TCTTAGAGCCTTGGATACTCCG | |||||||||
| VF20_21 | MW326742 | F: TAGATGCCTTGTTGTTGTTGC | [TAT]7 | 176–196 | 6 | 0.00 | 1.33 | 0.37 | 0.67 |
| R: CAAACGTGATTGCTGGATGGG | |||||||||
| VF20_22 | MW326743 | F: TCAATCAGAGCCTTGTTTGTGC | [GTA]6 | 117–119 | 2 | 0.00 | 0.58 | 0.00 | 0.40 |
| R: ATTGTTTGTTGCAGCTTTGGC | |||||||||
| VF20_24 | MW326744 | F: GGAGGAGATATGAGTGGGTTGG | [TAT]6 | 358–392 | 6 | 15.79 | 1.20 | 0.12 | 0.59 |
| R: AGATGATGATGATGAGTGTACC | |||||||||
| VF20_37 | MW326745 | F: GTTGACAGCGTTATGAAATTGG | [AAAT]4 | 390–395 | 2 | 5.26 | 0.69 | 0.11 | 0.51 |
| R: CCATAACCTAGGATCCTTGAGC | |||||||||
| VF20_41 | MW326746 | F: TCAGGTTGGCTCATGATACCG | [TCCC]4 | 391–394 | 2 | 21.05 | 0.69 | 0.00 | 0.51 |
| R: ATGGAACCACTACAACCAACC | |||||||||
| VF20_43 | MW326747 | F: TTCACGGTGAGTCAAGGAACC | [TTTA]5 | 284–314 | 3 | 5.26 | 0.85 | 0.11 | 0.55 |
| R: ATTGAAATGCAAGGGTCGACC | |||||||||
| VF20_44 | MW326748 | F: ATTTGACAACAACCCTACGCG | [TCTT]4 | 363–376 | 4 | 0.00 | 1.37 | 0.84 | 0.76 |
| R: GGCATGAGTAGGATGAAATGTTGG | |||||||||
| VF20_46 | MW326749 | F: ACATGCTTTGCACATGAAGGG | [TTTA]4 | 150–182 | 2 | 0.00 | 0.21 | 0.11 | 0.10 |
| R: AACAACCCGAACCTGACTTGC | |||||||||
| Mean | 4.47 | 3.86 | 1.10 | 0.23 | 0.60 |
| Species/Cultivars Analyzed | Specimen Origin/Accession Number a | Provenance | Year Collected |
|---|---|---|---|
| V. farreri | MtA 785 | no record | no record |
| V. farreri | UWBG 1190-49 | Garden Origin | no record |
| V. farrerib | NA 0111167 | Garden Origin | 1938 |
| V. farrerib | NA 0111164 | Garden Origin | 1941 |
| V. farreri | MA 314-49*2 | Garden Origin | 1949 |
| V. farreri | NA 0111168 | Garden Origin | 1966 |
| V. farreri | NA 0111169 | Garden Origin | 1966 |
| V. farreri | MA 533-70*2 | Garden Origin | 1970 |
| V. farreri | MA 398-83*1 | Garden Origin | 1983 |
| V. farreri | NA 0111163 | Garden Origin | 1985 |
| V. farreri | USNA 59728-H | Garden Origin | 1987 |
| V. farreri | USNA 59728-J | Garden Origin | 1987 |
| V. farreri | NA 0111166 | Garden Origin | 1987 |
| V. farreri | MA 915-2005*2 | Known wild origin | 2005 |
| V. farreri | NA 0052257 | Garden Origin | 2007 |
| V. farreri ‘Album’ | MA 1036-40*1 | Garden Origin | 1940 |
| V. farreri ‘Candidissimum’ | UWBG 1052-52 | Garden Origin | no record |
| V. farreri ‘Candidissimum’ | MtA 200704033 5664 | no record | 2007 |
| V. farreri ‘KLMW’ | MA 120-2012*1 | Garden Origin | 2012 |
| V. farreri ‘Nanum’ | MA 252-2002*1 | Garden Origin | 2002 |
| V. farreri ‘Nanum’ | AA 293-2003*C | Garden Origin | 2003 |
| V. farreri ‘Nanum’ c | NCSU 2020-063 | Garden Origin | 2020 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hamm, T.P.; Nowicki, M.; Boggess, S.L.; Klingeman, W.E.; Hadziabdic, D.; Huff, M.L.; Staton, M.E.; Trigiano, R.N. Development and Characterization of 15 Novel Genomic SSRs for Viburnum farreri. Plants 2021, 10, 487. https://doi.org/10.3390/plants10030487
Hamm TP, Nowicki M, Boggess SL, Klingeman WE, Hadziabdic D, Huff ML, Staton ME, Trigiano RN. Development and Characterization of 15 Novel Genomic SSRs for Viburnum farreri. Plants. 2021; 10(3):487. https://doi.org/10.3390/plants10030487
Chicago/Turabian StyleHamm, Trinity P., Marcin Nowicki, Sarah L. Boggess, William E. Klingeman, Denita Hadziabdic, Matthew L. Huff, Margaret E. Staton, and Robert N. Trigiano. 2021. "Development and Characterization of 15 Novel Genomic SSRs for Viburnum farreri" Plants 10, no. 3: 487. https://doi.org/10.3390/plants10030487
APA StyleHamm, T. P., Nowicki, M., Boggess, S. L., Klingeman, W. E., Hadziabdic, D., Huff, M. L., Staton, M. E., & Trigiano, R. N. (2021). Development and Characterization of 15 Novel Genomic SSRs for Viburnum farreri. Plants, 10(3), 487. https://doi.org/10.3390/plants10030487







