Nematicidal Activity of the Endophyte Serratia ureilytica against Nacobbus aberrans in Chili Plants (Capsicum annuum L.) and Identification of Genes Related to Biological Control
Abstract
:1. Introduction
2. Results
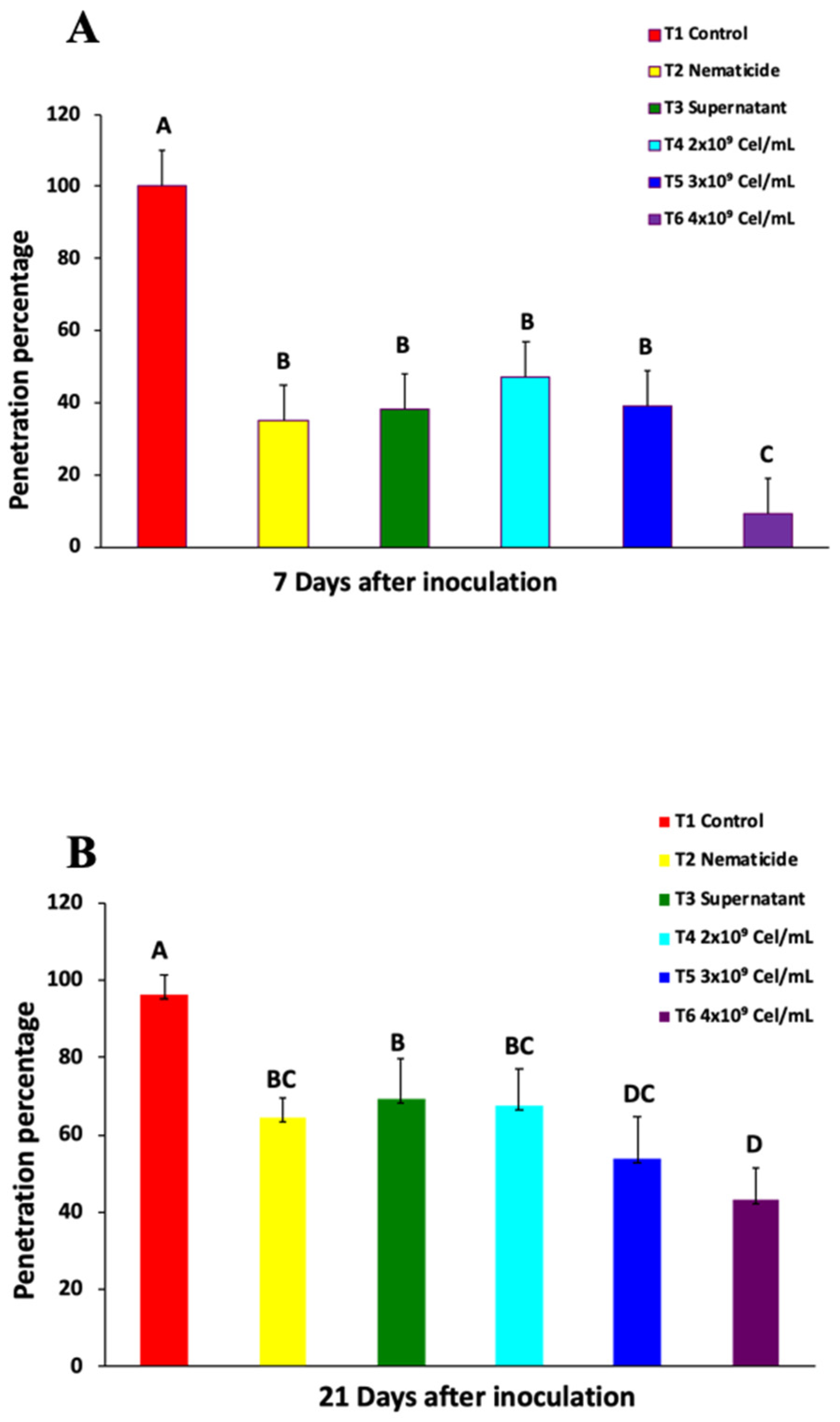
2.1. Nematode Penetration
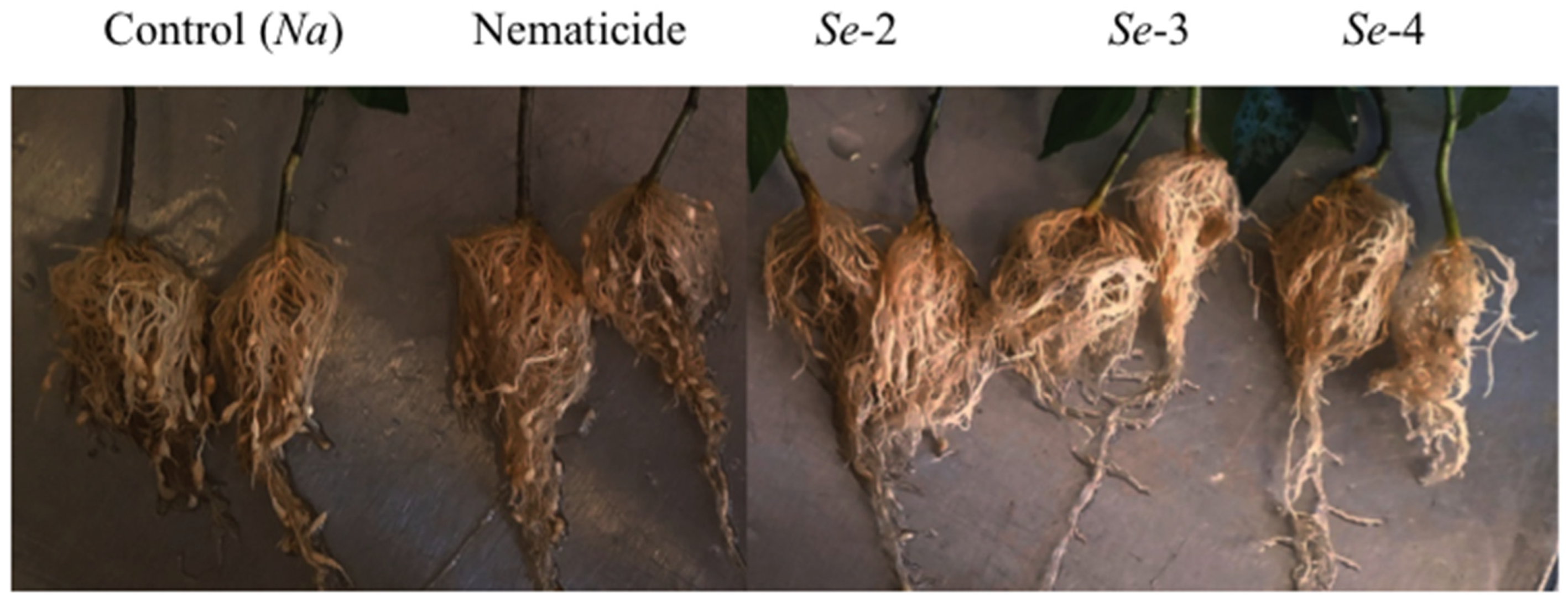
2.2. Galls, Egg Masses, Eggs, and Reproduction Factor
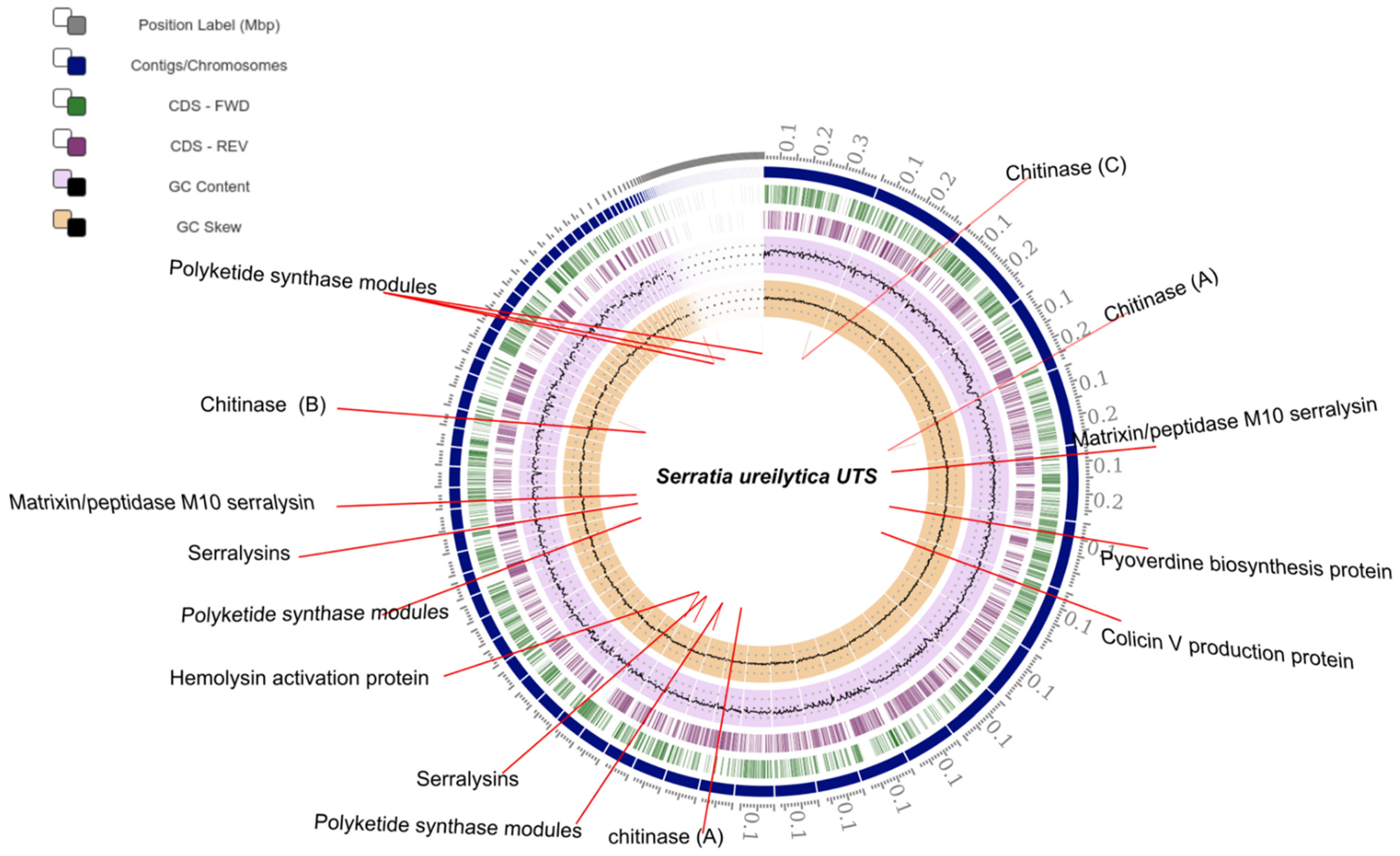
2.3. Genome Assembly and Annotation
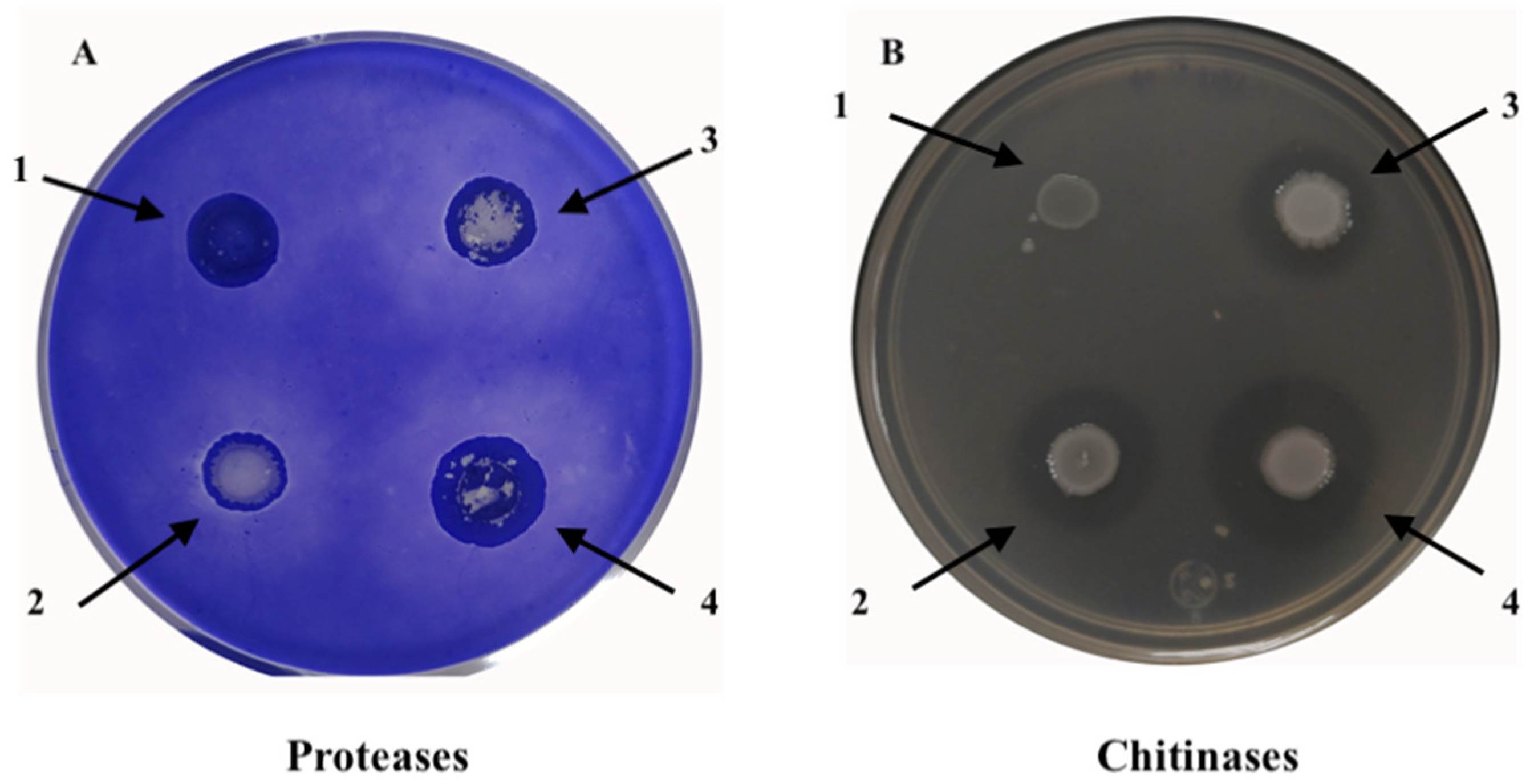
2.4. Protease and Chitinase Degradation Assays
2.5. Figures and Tables
3. Discussion
4. Materials and Methods
4.1. Plant Material
4.2. Obtaining and Inoculation of Serratia sp. (Se)
4.3. Obtaining Inoculum and Inoculation with N. aberrans (Na)
4.4. Experiment Establishment and Evaluation
4.5. Statistical Analysis
4.6. DNA Extraction, Library Preparation, and Sequencing
4.7. Genome Assembly and Annotation and Taxonomic Classification
4.8. Homologous Identification
4.9. Accession of the Genome Sequences
4.10. Protease and Chitinase Degradation Assays
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jones, J.T.; Haegeman, A.; Danchin, E.G.J.; Gaur, H.S.; Jones, M.G.K.; Kikuchi, T.; Manzanilla-López, R.; Palomares-Rius, J.E.; Wesemael, W.M.L.; Perry, R.N. Top 10 plant-parasitic nematodes in molecular plant pathology. Mol. Plant Pathol. 2013, 14, 946–961. [Google Scholar] [CrossRef]
- Cid del Prado-Vera, I.; Franco-Navarro, F.; Godinez-Vidal, D. Plant parasitic nematodes and management strategies of major crops in Mexico. In Plant Parasitic Nematodes in Sustainable Agriculture of North America, 1st ed.; Subbotin, S.A., Chitambar, J.J., Eds.; Springer: Sacramento, CA, USA, 2018; pp. 31–68. [Google Scholar] [CrossRef]
- Nicol, J.M.; Turner, S.J.; Coyne, D.L.; den Nijs, L.; Hockland, S.; Maafi, Z.T. Key nematodes threatening major agricultural crops of importance worldwide. In Genomics and Molecular Genetics of Plant-Nematode Interactions; Jones, J.T., Gheysen, G., Fenoll, C., Eds.; Springer: Dordrecht, Germany, 2011; pp. 21–44. [Google Scholar] [CrossRef]
- Castillo, P.; Stanley, J.; Inserra, R.N.; Manzanilla, L.R.H. Pratylenchidae—the lesión nematodes. In Practical Plant Nematology; Manzanilla-López, R.H., Marban-Mendoza, N., Eds.; Biblioteca Básica de Agricultura: Ciudad de Mexico, Mexico, 2012; pp. 411–478. [Google Scholar]
- Manzanilla, L.R.H.; Costilla, M.A.; Doucet, M.; Franco, J.; Inserra, R.N.; Lehman, P.S.; Cid del Prado-Vera, I.; Souza, R.M.; Evans, K. The genus Nacobbus Thorne & Allen, 1944 (Nematoda: Pratylenchidae): Systematics, distribution, biology and management. Nematropica 2002, 32, 149–227. [Google Scholar]
- Chen, S.; Dickson, D.W. Biological control of plant-parasitic nematodes. In Practical Plant Nematology; Manzanilla-López, R.H., Marbán-Mendoza, N., Eds.; Biblioteca Básica de Agricultura: Ciudad de Mexico, México, 2012; pp. 761–768. [Google Scholar]
- Stirling, G.R. Ecosystem services and the concept of ‘Integrated soil biology management’. In Biological Control of Plant-Parasitic Nematodes: Soil Ecosystem Management in Sustainable Agriculture; Stirling, G.R., Ed.; CABI: Wallingford, UK, 2014; pp. 3–11. [Google Scholar]
- Almaghrabi, O.A.; Massoud, S.I.; Abdelmoneim, T.S. Influence of inoculation with plant growth promoting rhizobacteria (PGPR) on tomato plant growth and nematode reproduction under greenhouse conditions. Saudi J. Biol. Sci. 2013, 20, 57–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Castaneda, A.C.; Aballay, E. Rhizobacteria with nematicide aptitude: Enzymes and compounds associated. World J. Microbiol. Biotechnol. 2016, 32, 2–7. [Google Scholar] [CrossRef]
- Mercer, C.F.; Greenwood, D.R.; Grant, J.L. Effect of Plant and Microbial Chitinases On the Eggs and Juveniles of Meloidogyne hapla Chitwood (Nematoda: Tylenchida). Nematologica 1992, 38, 227–236. [Google Scholar] [CrossRef]
- Zhao, D.; Zhao, H.; Zhao, D.; Zhu, X.; Wang, Y.; Duan, Y.; Xuan, Y.; Chen, L. Isolation and identification of bacteria from rhizosphere soil and their effect on plant growth promotion and root-knot nematode disease. Biol. Control 2018, 119, 12–19. [Google Scholar] [CrossRef]
- Paiva, G.; Proença, D.N.; Francisco, R.; Verissimo, P.; Santos, S.S.; Fonseca, L.; Abrantes, I.M.; Morais, P.V. Nematicidal bacteria associated to pinewood nematode produce extracellular proteases. PLoS ONE 2013, 7, e79705. [Google Scholar] [CrossRef] [Green Version]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef]
- Abebe-Akele, F.; Tisa, L.S.; Cooper, V.S.; Hatcher, P.J.; Abebe, E.; Thomas, W.K. Genome sequence and comparative analysis of a putative entomopathogenic Serratia isolated from Caenorhabditis briggsae. BMC Genom. 2015, 8, 531. [Google Scholar] [CrossRef] [Green Version]
- Su, C.; Liu, Y.; Sun, Y.; Li, Z. Complete genome sequence of Serratia sp. YD25 (KCTC 42987) presenting strong antagonistic activities to various pathogenic fungi and bacteria. J. Biotechnol. 2017, 45, 9–13. [Google Scholar] [CrossRef]
- Méndez-Santiago, E.W.; Sánchez-Cruz, R.; Folch Mallol, J.L.; Aguilar-Marcelino, L.; Hernández-Velázquez, V.M.; Gómez-Rodríguez, O.; Villar-Luna, E.; Wong-Villarreal, A. Serratia sp., an endophyte of Mimosa pudica nodules with nematicidal, antifungal activity and growth promoting characteristics. Arch. Microbiol. 2020, 203, 549–559. [Google Scholar] [CrossRef]
- Kim, M.; Oh, H.S.; Park, S.C.; Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 2014, 64, 346–351. [Google Scholar] [CrossRef]
- Richter, M.; Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA 2009, 106, 19126–19131. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suzuki, K.; Sugawara, N.; Suzuki, M.; Uchiyama, T.; Katouno, F.; Nikaidou, N.; Watanabe, T. Chitinases A, B, and C1 of Serratia marcescens 2170 produced by recombinant Escherichia coli: Enzymatic properties and synergism on chitin degradation. Biosci. Biotechnol. Biochem. 2002, 66, 1075–1083. [Google Scholar] [CrossRef]
- Page, A.P.; Stepek, G.; Winter, A.D.; Pertab, D. Enzymology of the nematode cuticle: A potential drug target? Int. J. Parasitol. Drugs Drug Resist. 2014, 4, 133–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Page, A.P.; Johnstone, I.L. The cuticle. In The C. Elegans Research Community; WormBook: Pasadena, CA, USA, 2007. [Google Scholar] [CrossRef] [Green Version]
- Veronico, P.; Gray, L.J.; Jones, J.T.; Bazzicalupo, P.; Arbucci, S.; Cortese, M.R.; Di Vito, M.; De Giorgi, C. Nematode chitin synthases: Gene structure, expression and function in Caenorhabditis elegans and the plant parasitic nematode Meloidogyne artiellia. Mol. Genet. Genom. 2001, 266, 28–34. [Google Scholar] [CrossRef]
- Millew, P.M.; Sands, D.C. Effects of hydrolytic enzymes on plant parasitic nematodes. J. Nematol. 1977, 9, 192–197. [Google Scholar]
- Sánchez, P.J.F. Efecto de quitina y quitosano sobre huevos y juveniles de nematodos formadores de nodulos radicales, Nacobbus aberrans y Meloidogyne incognita, bajo condiciones in vitro e in vivo. Master’s Thesis, Colegio de Postgraduados, Texcoco, México, 2010. [Google Scholar]
- Hegazy, M.I.; Salama, A.S.A.; El-Ashry, R.M.; Othman, A.E.I.O. Serratia marcescens and Pseudomonas aeruginosa are promising candidates as biocontrol agents against root-knot nematodes (Meloidogyne spp.). Middle East J. Agricul. Res. 2019, 8, 828–838. [Google Scholar]
- Zeinat, K.M.; El-Sayed, S.A.; Radwan, T.E.E.; El-Wahab, G.S.A. Potency evaluation of Serratia marcescens and Pseudomonas fluorescens as biocontrol agents for root-knot nematodes in Egypt. J. Appl. Sci. Res. 2009, 4, 93–102. [Google Scholar]
- Eves-van den Akker, S.; Lilley, C.J.; Danchin, E.G.J.; Rancurel, C.; Cock, P.J.A.; Urwin, P.E.; Jones, J.T. The transcriptome of Nacobbus aberrans reveals insights into the evolution of sedentary endoparasitism in plan-parasitic nematodes. Genome Biol. Evol. 2014, 6, 2181–2194. [Google Scholar] [CrossRef] [Green Version]
- Perry, R.N.; Moens, M. Introduction to plant-parasitic nematodes; modes of parasitism. In Genomics and Molecular Genetics of Plant-Nematode Interactions; Jones, J., Gheysen, G., Fenoll, C., Eds.; Springer: Dordrecht, Germany, 2011. [Google Scholar] [CrossRef]
- Cristóbal, A.J.; Cid del Prado, V.I.; Marbán-Mendoza, N.; Sánchez, G.P.; Mora-Aguilera, G.; Manzanilla, L.R.H. Survival of life stages of Nacobbus aberrans under field conditions. Nematropica 2001, 31, 229–235. [Google Scholar]
- Suryawanshi, R.; Chandrashekhar, P.; Hemant, B.; Chandrakant, N.; Lax-mikantSatish, S.P. Nematicidal activity of microbialpigment from Serratia marcescens. Nat. Prod. Res. 2014, 28, 1399–1404. [Google Scholar] [CrossRef]
- Marques-Pereira, C.; Proença, D.N.; Morais, P.V. Genome Sequences of Serratia Strains Revealed Common Genes in Both Serratomolides Gene Clusters. Biology 2020, 20, 482. [Google Scholar] [CrossRef]
- Wang, S.L.; Lin, C.L.; Liang, T.W.; Liu, K.C.; Kuo, Y.H. Conversion of squid pen by Serratia ureilytica for the production of enzymes and antioxidants. Bioresour. Technol. 2000, 100, 316–323. [Google Scholar] [CrossRef] [PubMed]
- Vrain, T.C. A technique for the collection of larvae of Meloidogyne spp. and a comparison of eggs and larvae as inocula. J. Nematol. 1977, 9, 249–251. [Google Scholar] [CrossRef]
- Teillet, A.; Dybal, K.; Kerry, B.R.; Miller, A.J.; Curtis, R.H.C.; Hedden, P. Transcriptional changes of the root-knot nematode Meloidogyne incognita in response to Arabidopsis thaliana root signals. PLoS ONE 2013, 8, e61259. [Google Scholar] [CrossRef] [Green Version]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef] [Green Version]
- Wattam, A.R.; Abraham, D.; Dalay, O.; Disz, T.L.; Driscoll, T.; Gabbard, J.L.; Gillespie, J.J.; Gough, R.; Hix, D.; Kenyon, R.; et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic Acids Res. 2014, 42, D581–D591. [Google Scholar] [CrossRef] [Green Version]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST Server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaumeil, P.A.; Mussig, A.J.; Hugenholtz, P.; Parks, D.H. GTDB-Tk: A toolkit to classify genomes with the Genome Taxonomy Database. Bioinformatics 2019, 36, 1925–1927. [Google Scholar] [CrossRef]
- Kejela, T.; Thakkar, V.R.; Thakor, P. Bacillus species (BT42) isolated from Coffea arabica L. rhizosphere antagonizes Colletotrichum gloeosporioides and Fusarium oxysporum and also exhibits multiple plant growth promoting activity. BMC Microbiol. 2016, 18, 277. [Google Scholar] [CrossRef] [Green Version]

| Treatments | Galls | Egg Masses | Eggs | RF (%) |
|---|---|---|---|---|
| T1 | 41.0 ± 1.4 a | 16 ± 1.6 a | 584 ± 60 a | 3.31 ± 0.45 a |
| T2 | 22.5 ± 1.1 b | 9 ± 1.3 ab | 145 ± 12.6 cd | 2.28 ± 0.64 ab |
| T3 | 29.0 ± 0.9 ab | 9 ± 1.1 ab | 168 ± 18.8 b | 2.29 ± 0.56 ab |
| T4 | 25.5 ± 1.1 ab | 7 ± 1.2 b | 151 ± 16.8 bcd | 1.75 ± 0.57 b |
| T5 | 27.1 ± 0.8 ab | 8 ± 0.5 ab | 161 ± 19.6 bc | 1.87 ±0.38 ab |
| T6 | 21.6 ± 1.0 b | 6 ± 0.8 b | 51 ± 7.9 d | 1.26 ± 0.41 b |
| CV (%) | 16.7 | 25.1 | 26.8 | 25.1 |
| Features | Chromosome |
|---|---|
| Contigs | 134 |
| Genome size | 5,401,403 bp |
| GC content (%) | 59.32561 |
| Coding gene | 5303 |
| tRNA | 87 |
| rRNA | 6 |
| Hypothetical proteins | 954 |
| Proteins with functional assignments | 4349 |
| Genomic Location | Serratia ureilytica UTS Protein |
|---|---|
| 1–6213 87709–93531 3–2393 1–408 3–206 | Putative peptide synthetase, containing non-ribosomal peptide synthetase |
| 117538–118815 | Chitinase (A) |
| 76512–78203 | Chitinase (A) |
| 7930–9444 | Serine 3-dehydrogenase |
| 56070–57587 | Colicin V production protein |
| 64025–64888 | Pyoverdine biosynthesis protein |
| 38269–39687 | Serine 3-dehydrogenase |
| 36474–38090 | Protease C |
| 50375–51874 | Chitinase (B) |
| 49385–50767 | Matrixin/peptidase M10 serralysin |
| 58625–59740 | Chitinase (C) |
| 23187–28013 | Hemolysin |
| 28059–29738 | Hemolysin activation protein |
| 7930–9444 | Serralysin |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wong-Villarreal, A.; Méndez-Santiago, E.W.; Gómez-Rodríguez, O.; Aguilar-Marcelino, L.; García, D.C.; García-Maldonado, J.Q.; Hernández-Velázquez, V.M.; Yañez-Ocampo, G.; Espinosa-Zaragoza, S.; I. Ramírez-González, S.; et al. Nematicidal Activity of the Endophyte Serratia ureilytica against Nacobbus aberrans in Chili Plants (Capsicum annuum L.) and Identification of Genes Related to Biological Control. Plants 2021, 10, 2655. https://doi.org/10.3390/plants10122655
Wong-Villarreal A, Méndez-Santiago EW, Gómez-Rodríguez O, Aguilar-Marcelino L, García DC, García-Maldonado JQ, Hernández-Velázquez VM, Yañez-Ocampo G, Espinosa-Zaragoza S, I. Ramírez-González S, et al. Nematicidal Activity of the Endophyte Serratia ureilytica against Nacobbus aberrans in Chili Plants (Capsicum annuum L.) and Identification of Genes Related to Biological Control. Plants. 2021; 10(12):2655. https://doi.org/10.3390/plants10122655
Chicago/Turabian StyleWong-Villarreal, Arnoldo, Erick Williams Méndez-Santiago, Olga Gómez-Rodríguez, Liliana Aguilar-Marcelino, Daniel Cerqueda García, José Q. García-Maldonado, Victor M. Hernández-Velázquez, Gustavo Yañez-Ocampo, Saúl Espinosa-Zaragoza, Sandra I. Ramírez-González, and et al. 2021. "Nematicidal Activity of the Endophyte Serratia ureilytica against Nacobbus aberrans in Chili Plants (Capsicum annuum L.) and Identification of Genes Related to Biological Control" Plants 10, no. 12: 2655. https://doi.org/10.3390/plants10122655
APA StyleWong-Villarreal, A., Méndez-Santiago, E. W., Gómez-Rodríguez, O., Aguilar-Marcelino, L., García, D. C., García-Maldonado, J. Q., Hernández-Velázquez, V. M., Yañez-Ocampo, G., Espinosa-Zaragoza, S., I. Ramírez-González, S., & Sanzón-Gómez, D. (2021). Nematicidal Activity of the Endophyte Serratia ureilytica against Nacobbus aberrans in Chili Plants (Capsicum annuum L.) and Identification of Genes Related to Biological Control. Plants, 10(12), 2655. https://doi.org/10.3390/plants10122655









