Functional and Comparative Genomics of Hoxa2 Gene cis-Regulatory Elements: Evidence for Evolutionary Modification of Ancestral Core Element Activity
Abstract
1. Introduction
2. Materials and Methods
2.1. Tol2 Plasmid Construction
2.2. Medaka Genomic DNA Extraction
2.3. Amplification of Medaka hoxa2a and ψhoxa2b UER(K20-RE5)s
2.4. PCR-Mediated Deletion Mutagenesis of the Medaka hoxa2a and ψhoxa2b UER(K20-RE5)s
2.5. Microinjection of Medaka Embryos
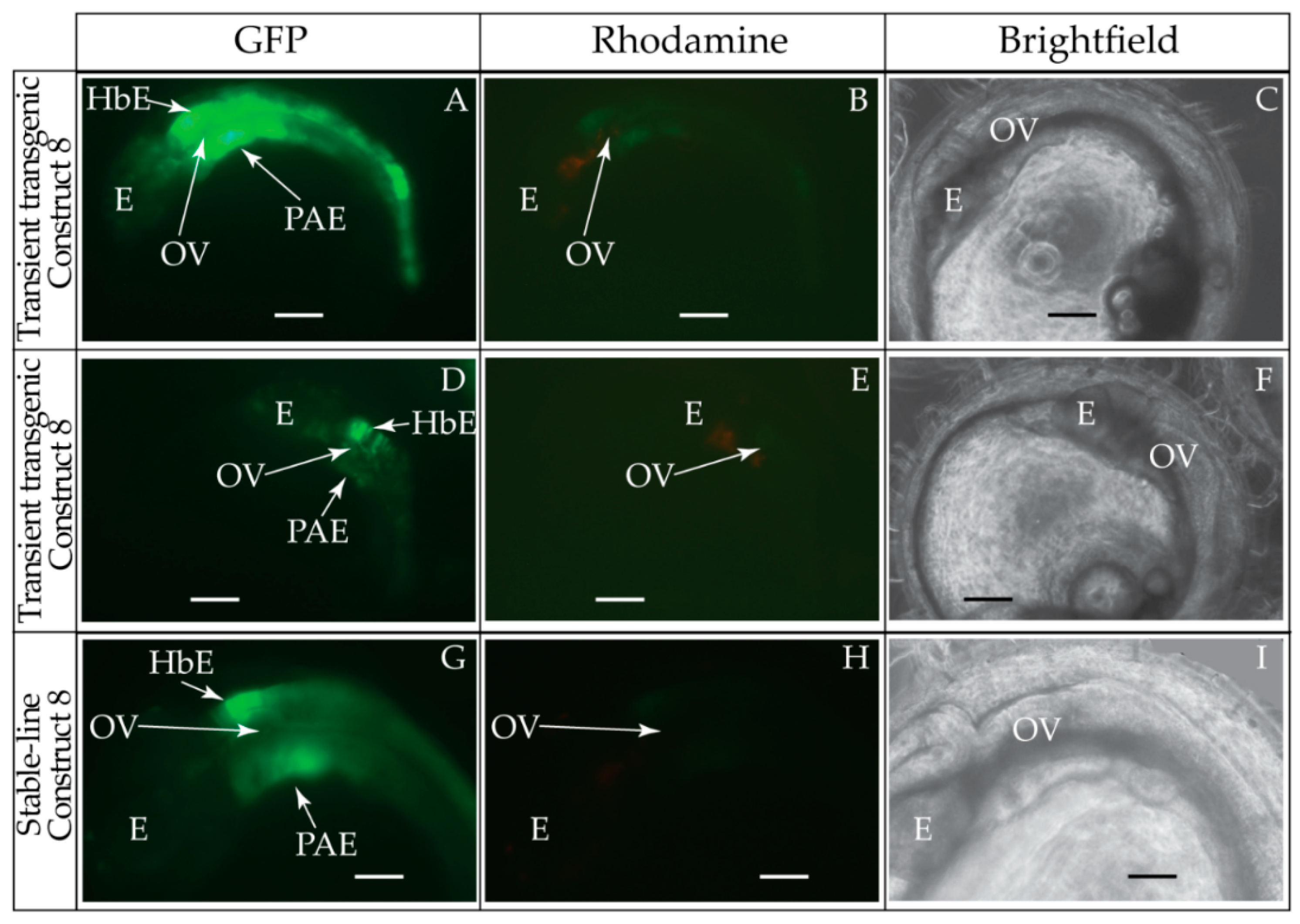
2.6. Generation and Visualization of Transient and Stable-Line Transgenic Medaka Embryos
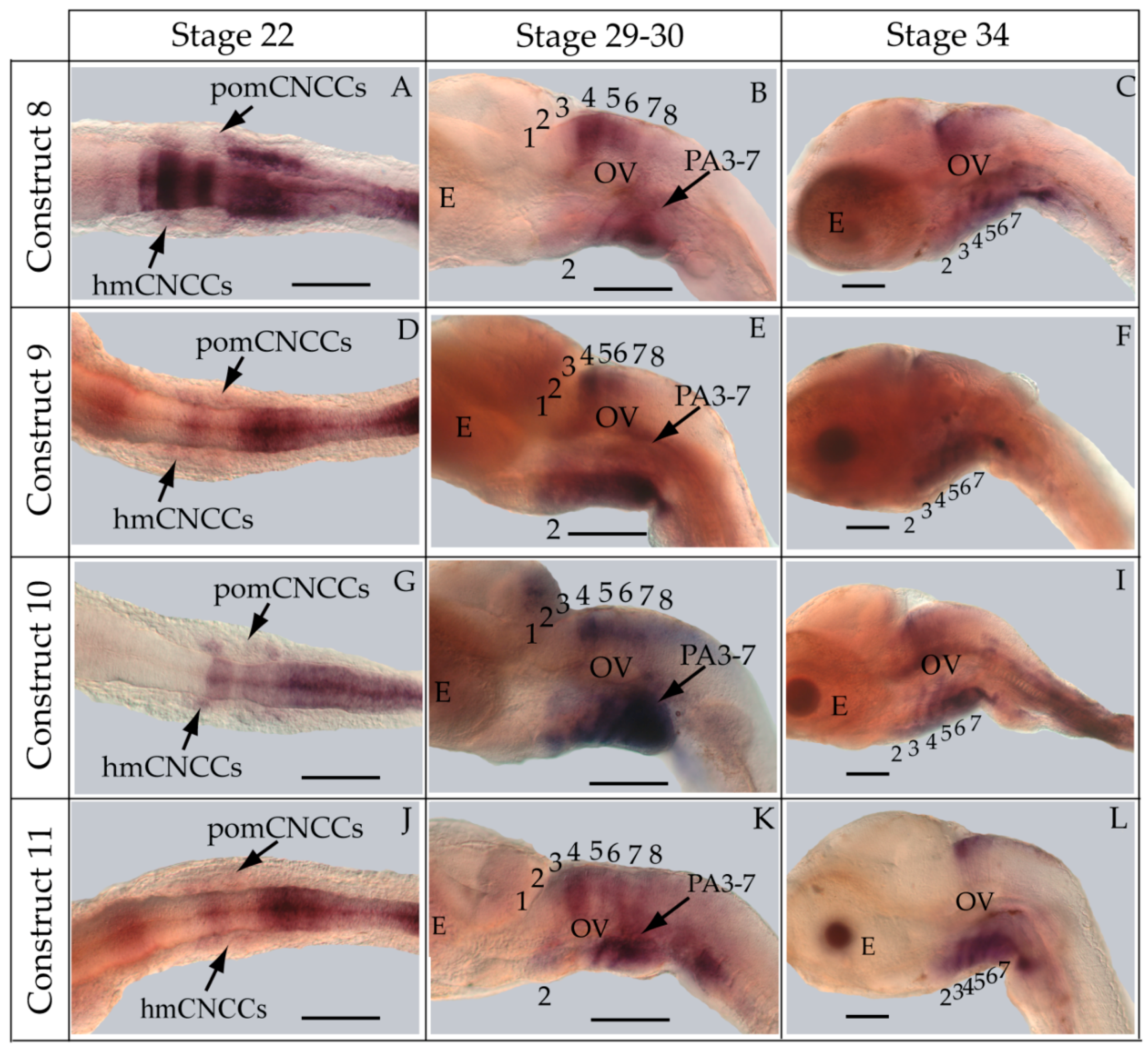
2.7. Whole-Mount in Situ Hybridization
2.8. Comparative Genomic Sequence Analysis
3. Results
3.1. Validation of the Tol2 Transposon System for Medaka Embryos
3.2. Functional Genomic Analysis of the Medaka Hoxa2a UER(K20-RE5)
3.3. Functional Genomic Analysis of the Medaka ψHoxa2b UER(K20-RE5)
4. Discussion
4.1. The Use of Medaka in Reporter Gene Expression Analyses
4.2. Medaka Hoxa2a-Directed Gene Expression in the Hindbrain
4.3. Hoxa2a-Directed Gene Expression in the Cranial Neural Crest Cells
4.4. Functional Nature of the Medaka ψHoxa2b r3/5 Enhancer Region
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| A-P | anterior-posterior |
| bp | base pair |
| CNCC | cranial neural crest cell |
| CNE | conserved noncoding element |
| CRE | cis-regulatory element |
| eGFP | enhanced green fluorescent protein |
| ERM | embryo rearing medium |
| E. coli | Escherichia coli |
| GFP | green fluorescent protein |
| NC | neural crest |
| PA | pharyngeal arch |
| PG | paralog group |
| r | rhombomere |
| RE | rhombomeric element |
| UER | upstream enhancer region |
| UER(K20-RE5) | upstream enhancer region spanning 5′ from Krox20 binding element to the 3′ RE5 element |
References
- McGinnis, W.; Krumlauf, R. Homeobox genes and axial patterning. Cell 1992, 68, 283–302. [Google Scholar] [CrossRef]
- Ferrier, D.E.; Minguillon, C.; Holland, P.W.; Garcia-Fernandez, J. The amphioxus Hox cluster: Deuterostome posterior flexibility and Hox14. Evolut. Dev. 2000, 2, 284–293. [Google Scholar] [CrossRef]
- Holland, P.W.; Garcia-Fernandez, J. Hox genes and chordate evolution. Dev. Biol. 1996, 173, 382–395. [Google Scholar] [CrossRef] [PubMed]
- Powers, T.P.; Amemiya, C.T. Evidence for a Hox14 paralog group in vertebrates. Curr. Biol. 2004, 14, R183–R184. [Google Scholar] [CrossRef] [PubMed]
- Amores, A.; Force, A.; Yan, Y.L.; Joly, L.; Amemiya, C.; Fritz, A.; Ho, R.K.; Langeland, J.; Prince, V.; Wang, Y.L.; et al. Zebrafish hox clusters and vertebrate genome evolution. Science 1998, 282, 1711–1714. [Google Scholar] [CrossRef] [PubMed]
- Amores, A.; Suzuki, T.; Yan, Y.L.; Pomeroy, J.; Singer, A.; Amemiya, C.; Postlethwait, J.H. Developmental roles of pufferfish Hox clusters and genome evolution in ray-fin fish. Genome Res. 2004, 14, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Hoegg, S.; Boore, J.L.; Kuehl, J.V.; Meyer, A. Comparative phylogenomic analyses of teleost fish Hox gene clusters: Lessons from the cichlid fish Astatotilapia burtoni. BMC Genom. 2007, 8, 317. [Google Scholar] [CrossRef] [PubMed]
- Moghadam, H.; Ferguson, M.; Danzmann, R. Evolution of Hox Clusters in Salmonidae: A Comparative Analysis Between Atlantic Salmon (Salmo salar) and Rainbow Trout (Oncorhynchus mykiss). J. Mol. Evol. 2005, 61, 636–649. [Google Scholar] [CrossRef] [PubMed]
- Mungpakdee, S.; Seo, H.; Angotzi, A.R.; Dong, X.; Akalin, A.; Chourrout, D. Differential evolution of the 13 Atlantic salmon hox clusters. Mol. Biol. Evol. 2008, 25, 1333–1343. [Google Scholar] [CrossRef] [PubMed]
- Prince, V. The Hox paradox: More complex(es) than imagined. Dev. Biol. 2002, 249, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Stellwag, E.J. Hox gene duplication in fish. Semin. Cell Dev. Biol. 1999, 10, 531–540. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.; Scemama, J.L.; Stellwag, E.J. Japanese medaka Hox paralog group 2: Insights into the evolution of Hox PG2 gene composition and expression in the Osteichthyes. J. Exp. Zool. B Mol. Dev. Evol. 2008, 310, 623–641. [Google Scholar] [CrossRef] [PubMed]
- Davis, A.; Stellwag, E.J. Spatio-temporal patterns of Hox paralog group 3–6 gene expression during Japanese medaka (Oryzias latipes) embryonic development. Gene Expr. Patterns 2010, 10, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Le Pabic, P.; Stellwag, E.J.; Brothers, S.N.; Scemama, J.L. Comparative analysis of Hox paralog group 2 gene expression during Nile tilapia (Oreochromis niloticus) embryonic development. Dev. Genes Evol. 2007, 217, 749–758. [Google Scholar] [CrossRef] [PubMed]
- Lyon, R.S.; Davis, A.; Scemama, J.L. Spatio-temporal expression patterns of anterior Hox genes during Nile tilapia (Oreochromis niloticus) embryonic development. Gene Expr. Patterns 2013, 13, 104–108. [Google Scholar] [CrossRef] [PubMed]
- Hurley, I.A.; Scemama, J.; Prince, V.E. Consequences of Hoxb1 duplication in teleost fish. Evol. Dev. 2007, 9, 540–554. [Google Scholar] [CrossRef] [PubMed]
- Le Pabic, P.; Stellwag, E.J.; Scemama, J. Embryonic development and skeletogenesis of the pharyngeal jaw apparatus in the cichlid Nile tilapia (Oreochromis niloticus). Anat. Rec. 2009, 292, 1780–1800. [Google Scholar] [CrossRef] [PubMed]
- Mungpakdee, S.; Seo, H.; Chourrout, D. Spatio-temporal expression patterns of anterior Hox genes in Atlantic salmon (Salmo salar). Gene Expr. Patterns 2008, 8, 508–514. [Google Scholar] [CrossRef] [PubMed]
- Scemama, J.L.; Vernon, J.L.; Stellwag, E. Differential expression of Hoxa2a and Hoxa2b genes during striped bass embryonic development. Gene Expr. Patterns 2006, 6, 843–848. [Google Scholar] [CrossRef] [PubMed]
- Soshnikova, N.; Dewaele, R.; Janvier, P.; Krumlauf, R.; Duboule, D. Duplications of hox gene clusters and the emergence of vertebrates. Dev. Biol. 2013, 378, 194–199. [Google Scholar] [CrossRef] [PubMed]
- Tümpel, S.; Cambronero, F.; Wiedemann, L.M.; Krumlauf, R. Evolution of cis elements in the differential expression of two Hoxa2 coparalogous genes in pufferfish (Takifugu rubripes). Proc. Natl. Acad. Sci. USA 2006, 103, 5419–5424. [Google Scholar] [CrossRef] [PubMed]
- Gendron-Maguire, M.; Mallo, M.; Zhang, M.; Gridley, T. Hoxa-2 mutant mice exhibit homeotic transformation of skeletal elements derived from cranial neural crest. Cell 1993, 75, 1317–1331. [Google Scholar] [CrossRef]
- Rijli, F.M.; Mark, M.; Lakkaraju, S.; Dierich, A.; Dolle, P.; Chambon, P. A homeotic transformation is generated in the rostral branchial region of the head by disruption of Hoxa-2, which acts as a selector gene. Cell 1993, 75, 1333–1349. [Google Scholar] [CrossRef]
- Amin, S.; Donaldson, I.J.; Zannino, D.A.; Hensman, J.; Rattray, M.; Losa, M.; Spitz, F.; Ladam, F.; Sagestrom, C.; Bobola, N. Hoxa2 selectively enhances Meis binding to change a branchial arch ground state. Dev. Cell 2015, 32, 265–277. [Google Scholar] [CrossRef] [PubMed]
- Frasch, M.; Chen, X.; Lufkin, T. Evolutionary-conserved enhancers direct region-specific expression of the murine Hoxa-1 and Hoxa-2 loci in both mice and Drosophila. Development 1995, 121, 957–974. [Google Scholar] [PubMed]
- Lampe, X.; Samad, O.A.; Guiguen, A.; Matis, C.; Remacle, S.; Picard, J.J.; Rijli, F.M.; Rezsohazy, R. An ultraconserved Hox-Pbx responsive element resides in the coding sequence of Hoxa2 and is active in rhombomere 4. Nucleic Acids Res. 2008, 36, 3214–3225. [Google Scholar] [CrossRef] [PubMed]
- Maconochie, M.K.; Krishnamurthy, R.; Nonchev, S.; Meier, P.; Manzanares, M.; Mitchell, P.J.; Krumlauf, R. Regulation of Hoxa2 in cranial neural crest cells involves members of the AP-2 family. Development 1999, 126, 1483–1494. [Google Scholar] [PubMed]
- Maconochie, M.K.; Nonchev, S.; Manzanares, M.; Marshall, H.; Krumlauf, R. Differences in Krox20-dependent regulation of Hoxa2 and Hoxb2 during hindbrain development. Dev. Biol. 2001, 233, 468–481. [Google Scholar] [CrossRef] [PubMed][Green Version]
- McEllin, J.A.; Alexander, T.B.; Tümpel, S.; Wiedemann, L.M.; Krumlauf, R. Analyses of fugu hoxa2 genes provide evidence for subfunctionalization of neural crest cell and rhombomere cis-regulatory modules during vertebrate evolution. Dev. Biol. 2016, 409, 530–542. [Google Scholar] [CrossRef] [PubMed]
- Nonchev, S.; Maconochie, M.; Vesque, C.; Aparicio, S.; Ariza-McNaughton, L.; Manzanares, M.; Maruthainar, K.; Kuroiwa, A.; Brenner, S.; Charnay, P.; et al. The conserved role of Krox-20 in directing Hox gene expression during vertebrate hindbrain segmentation. Proc. Natl. Acad. Sci. USA 1996, 93, 9339–9345. [Google Scholar] [CrossRef] [PubMed]
- Nonchev, S.; Vesque, C.; Maconochie, M.; Seitanidou, T.; Ariza-McNaughton, L.; Frain, M.; Marshall, H.; Sham, M.H.; Krumlauf, R.; Charnay, P. Segmental expression of Hoxa-2 in the hindbrain is directly regulated by Krox-20. Development 1996, 122, 543–554. [Google Scholar] [PubMed]
- Tümpel, S.; Maconochie, M.; Wiedemann, L.M.; Krumlauf, R. Conservation and diversity in the cis-regulatory networks that integrate information controlling expression of Hoxa2 in hindbrain and cranial neural crest cells in vertebrates. Dev. Biol. 2002, 246, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Tümpel, S.; Cambronero, F.; Ferretti, E.; Blasi, F.; Wiedemann, L.M.; Krumlauf, R. Expression of Hoxa2 in rhombomere 4 is regulated by a conserved cross-regulatory mechanism dependent upon Hoxb1. Dev. Biol. 2007, 302, 646–660. [Google Scholar] [CrossRef] [PubMed]
- Tümpel, S.; Cambronero, F.; Sims, C.; Krumlauf, R.; Wiedemann, L.M. A regulatory module embedded in the coding region of Hoxa2 controls expression in rhombomere 2. Proc. Natl. Acad. Sci. USA 2008, 105, 20077–20082. [Google Scholar] [CrossRef] [PubMed]
- Parker, H.A.; Bronner, M.E.; Krumlauf, R. A Hox regulatory network of hindbrain segmentation is conserved to the base of vertebrates. Nature 2014, 514, 490–493. [Google Scholar] [CrossRef] [PubMed]
- Sham, M.H.; Vesque, C.; Nonchev, S.; Marshall, H.; Frain, M.; Gupta, R.D.; Whiting, J.; Wilkinson, D.; Charnay, P.; Krumlauf, R. The zinc finger gene Krox20 regulates HoxB2 (Hox2.8) during hindbrain segmentation. Cell 1993, 72, 183–196. [Google Scholar] [CrossRef]
- Ferretti, E.; Marshall, H.; Popperl, H.; Maconochie, M.; Krumlauf, R.; Blasi, F. Segmental expression of Hoxb2 in r4 requires two separate sites that integrate cooperative interactions between Prep1, Pbx and Hox proteins. Development 2000, 127, 155–166. [Google Scholar] [PubMed]
- Vesque, C.; Maconochie, M.; Nonchev, S.; Ariza-McNaughton, L.; Kuroiwa, A.; Charnay, P.; Krumlauf, R. Hoxb-2 transcriptional activation in rhombomeres 3 and 5 requires an evolutionarily conserved cis-acting element in addition to the Krox-20 binding site. EMBO J. 1996, 15, 5383–5396. [Google Scholar] [PubMed]
- Barrow, J.R.; Capecchi, M.R. Targeted disruption of the Hoxb-2 locus in mice interferes with expression of Hoxb-1 and Hoxb-4. Development 1996, 122, 3817–3828. [Google Scholar] [PubMed]
- Barrow, J.R.; Stadler, H.S.; Capecchi, M.R. Roles of Hoxa1 and Hoxa2 in patterning the early hindbrain of the mouse. Development 2000, 127, 933–944. [Google Scholar] [PubMed]
- Davenne, M.; Maconochie, M.K.; Neun, R.; Pattyn, A.; Chambon, P.; Krumlauf, R.; Rijli, F.M. Hoxa2 and Hoxb2 control dorsoventral patterns of neuronal development in the rostral hindbrain. Neuron 1999, 22, 677–691. [Google Scholar] [CrossRef]
- Gavalas, A.; Davenne, M.; Lumsden, A.; Chambon, P.; Rijli, F.M. Role of Hoxa-2 in axon pathfinding and rostral hindbrain patterning. Development 1997, 124, 3693–3702. [Google Scholar] [PubMed]
- Gavalas, A.; Ruhrberg, C.; Livet, J.; Henderson, C.E.; Krumlauf, R. Neuronal defects in the hindbrain of Hoxa1, Hoxb1 and Hoxb2 mutants reflect regulatory interactions among these Hox genes. Development 2003, 130, 5663–5679. [Google Scholar] [CrossRef] [PubMed]
- Oury, F.; Murakami, Y.; Renaud, J.S.; Pasqualetti, M.; Charnay, P.; Ren, S.Y.; Rijli, F.M. Hoxa2- and rhombomere-dependent development of the mouse facial somatosensory map. Science 2006, 313, 1408–1413. [Google Scholar] [CrossRef] [PubMed]
- Baltzinger, M.; Ori, M.; Pasqualetti, M.; Nardi, I.; Rijli, F.M. Hoxa2 knockdown in Xenopus results in hyoid to mandibular homeosis. Dev. Dyn. 2005, 234, 858–867. [Google Scholar] [CrossRef] [PubMed]
- Grammatopoulos, G.A.; Bell, E.; Toole, L.; Lumsden, A.; Tucker, A.S. Homeotic transformation of branchial arch identity after Hoxa2 overexpression. Development 2000, 127, 2355–5365. [Google Scholar]
- Kitazawa, T.; Fujisawa, K.; Narboux-Nême, N.; Arima, Y.; Kawamura, Y.; Inoue, T.; Wada, Y.; Kohro, T.; Aburatani, H.; Kodama, T.; et al. Distinct effects of Hoxa2 overexpression in cranial neural crest populations reveal that the mammalian hyomandibular-ceratohyal boundary maps within the styloid process. Dev. Biol. 2015, 402, 162–174. [Google Scholar] [CrossRef] [PubMed]
- Minoux, M.; Antonarakis, G.S.; Kmita, M.; Duboule, D.; Rijli, F.M. Rostral and caudal pharyngeal arches share a common neural crest ground pattern. Development 2009, 136, 637–645. [Google Scholar] [CrossRef] [PubMed]
- Pasqualetti, M.; Ori, M.; Nardi, I.; Rijli, F.M. Ectopic Hoxa2 induction after neural crest migration results in homeosis of jaw elements in Xenopus. Development 2000, 127, 5367–5378. [Google Scholar] [PubMed]
- Hurley, I.A.; Mueller, R.L.; Dunn, K.A.; Schmidt, K.J.; Friedman, M.; Ho, R.K.; Prince, V.E.; Yang, Z.; Thomas, M.G.; Coates, M.I. A new time-scale for ray-finned fish evolution. Proc. R. Soc. B 2007, 274, 489–498. [Google Scholar] [CrossRef] [PubMed]
- Meyer, A.; van de Peer, Y. From 2R to 3R: Evidence for a fish-specific genome duplication (FSGD). Bioessays 2005, 27, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Postlethwait, J.; Amores, A.; Cresko, W.; Singer, A.; Yan, Y.L. Subfunction partitioning, the teleost radiation and the annotation of the human genome. Trends Genet. 2004, 20, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Santini, F.; Harmon, L.J.; Carnevale, G.; Alfaro, M.E. Did genome duplication drive the origin of teleosts? A comparative study of diversification in ray-finned fishes. BMC Evol. Biol. 2009, 9, 194. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.S.; van de Peer, Y.; Meyer, A. Revisiting recent challenges to the ancient fish-specific genome duplication hypothesis. Curr. Biol. 2001, 11, R1005–R1008. [Google Scholar] [CrossRef]
- Moens, C.B.; Selleri, L. Hox cofactors in vertebrate development. Dev. Biol. 2006, 291, 193–206. [Google Scholar] [CrossRef] [PubMed]
- Hunter, M.; Prince, V.E. Zebrafish Hox paralogue group 2 genes function redundantly as selector genes to pattern the second pharyngeal arch. Dev. Biol. 2002, 247, 367–389. [Google Scholar] [CrossRef] [PubMed]
- Prince, V.; Lumsden, A. Hoxa-2 expression in normal and transposed rhombomeres: Independent regulation in the neural tube and neural crest. Development 1994, 120, 911–923. [Google Scholar] [PubMed]
- Steinke, D.; Salzburger, W.; Meyer, A. Novel relationships among ten fish model species based on a phylogenomic analysis using ESTs. J. Mol. Evol. 2006, 62, 772–784. [Google Scholar] [CrossRef] [PubMed]
- Tümpel, S.; Wiedemann, L.M.; Krumlauf, R. Hox genes and segmentation of the vertebrate hindbrain. Curr. Top. Dev. Biol. 2009, 88, 103–137. [Google Scholar] [PubMed]
- Davidson, E.H.; Erwin, D.H. Gene regulatory networks and the evolution of animal body plans. Science 2006, 311, 796–800. [Google Scholar] [CrossRef] [PubMed]
- Barrallo-Gimeno, A.; Holzschuh, J.; Driever, W.; Knapik, E.W. Neural crest survival and differentiation in zebrafish depends on mont blanc/tfap2a gene function. Development 2004, 131, 1463–1477. [Google Scholar] [CrossRef] [PubMed]
- Crump, J.G.; Swartz, M.E.; Eberhart, J.K.; Kimmel, C.B. Moz-dependent Hox expression controls segment-specific fate maps of skeletal precursors in the face. Development 2006, 133, 2661–2669. [Google Scholar] [CrossRef] [PubMed]
- Deflorian, G.; Tiso, N.; Ferretti, E.; Meyer, D.; Blasi, F.; Bortolussi, M.; Argenton, F. Prep1.1 has essential genetic functions in hindbrain development and cranial neural crest cell differentiation. Development 2004, 131, 613–627. [Google Scholar] [CrossRef] [PubMed]
- Le Pabic, P.; Scemama, J.; Stellwag, E.J. Role of Hox PG2 genes in Nile tilapia pharyngeal arch specification: Implications for gnathostome pharyngeal arch evolution. Evol. Dev. 2010, 12, 45–60. [Google Scholar] [CrossRef] [PubMed]
- Melvin, V.S.; Feng, W.; Hernandez-Lagunas, L.; Artinger, K.B.; Williams, T. A morpholino-based screen to identify novel genes involved in craniofacial morphogenesis. Dev. Dyn. 2013, 242, 817–831. [Google Scholar] [CrossRef] [PubMed]
- Selleri, L.; Depew, M.J.; Jacobs, Y.; Chanda, S.K.; Tsang, K.Y.; Cheah, K.S.; Rubenstein, J.L.; O’Gorman, S.; Cleary, M.L. Requirement for Pbx1 in skeletal patterning and programming chondrocyte proliferation and differentiation. Development 2001, 128, 3543–3557. [Google Scholar] [PubMed]
- Ferretti, E.; Cambronero, F.; Tümpel, S.; Longobardi, E.; Wiedemann, L.M.; Blasi, F.; Krumlauf, R. The Hoxb1 enhancer and control of rhombomere 4 expression: Complex interplay between PREP1-PBX1-HOXB1 binding sites. Mol. Cell. Biol. 2005, 25, 8541–8552. [Google Scholar] [CrossRef] [PubMed]
- Pöpperl, H.; Bienz, M.; Studer, M.; Chan, S.; Aparicio, S.; Brenner, S.; Mann, R.; Krumlauf, R. Segmental expression of Hoxb1 is controlled by a highly conserved autoregulatory loop dependent on exd/Pbx. Cell 1995, 81, 1031–1042. [Google Scholar] [CrossRef]
- Urasaki, A.; Morvan, G.; Kawakami, K. Functional dissection of the Tol2 transposable element identified the minimal cis-sequences and a highly repetitive sequence in the subterminal region essential for transposition. Genetics 2006, 174, 639–649. [Google Scholar] [CrossRef] [PubMed]
- Kawakami, K. Tol2: A versatile gene transfer vector in vertebrates. Genome Biol. 2007, 8, S7. [Google Scholar] [CrossRef] [PubMed]
- Kurosawa, G.; Takamatsu, N.; Takahashi, M.; Sumitomo, M.; Sanaka, E.; Yamada, K.; Nishii, K.; Matsuda, M.; Asakawa, S.; Ishiguro, H.; et al. Organization and structure of hox gene loci in medaka genome and comparison with those of pufferfish and zebrafish genomes. Gene 2006, 370, 75–82. [Google Scholar] [CrossRef] [PubMed]
- Iwamatsu, T. Stages of normal development in the medaka Oryzias latipes. Mech. Dev. 2004, 12, 605–618. [Google Scholar] [CrossRef] [PubMed]
- Oxendine, S.L.; Cowden, J.; Hinton, D.E.; Padilla, S. Adapting the medaka embryo assay to a high-throughput approach for developmental toxicity testing. Neuro Toxicol. 2006, 27, 840–845. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, J.; Hofsten, J.V.; Olsson, P.E. Generating transparent zebrafish: A refined method to improve detection of gene expression during embryonic development. Mar. Biotechnol. 2001, 3, 522–527. [Google Scholar] [CrossRef] [PubMed]
- Sandelin, A.; Alkema, W.; Engstrom, P.; Wasserman, W.W.; Lenhard, B. JASPAR: An open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. 2004, 32, D91–D94. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Kaufmann, M.; Morgenstern, B. DIALIGN-TX: Greedy and Progressive Approaches for Segment-Based Multiple Sequence Alignment. Algorithms Mol Biol. 2008, 3, 6. [Google Scholar] [CrossRef] [PubMed]
- Amemiya, C.T.; Powers, T.P.; Prohaska, S.J.; Grimwood, J.; Schmutz, J.; Dickson, M.; Miyake, T.; Schoenborn, M.A.; Myers, R.M.; Ruddle, F.H.; et al. Complete Hox cluster characterization of the coelacanth provides further evidence for slow evolution of its genome. Proc. Natl. Acad. Sci. USA 2010, 107, 3622–3627. [Google Scholar] [CrossRef] [PubMed]
- Chiu, C.; Dewar, K.; Wagner, G.P.; Takahashi, K.; Ruddle, F.; Ledje, C.; Bartsch, P.; Scemama, J.; Stellwag, E.J.; Fried, C.; et al. Bichir HoxA cluster sequence reveals surpising trends in ray-finned fish genomic evolution. Genome Res. 2004, 14, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Church, D.M.; Goodstadt, L.; Hillier, L.W.; Zody, M.C.; Goldstein, S.; She, X.; Bult, C.J.; Agarwala, R.; Cherry, J.L.; DiCuccio, M.; et al. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 2009, 7, e1000112. [Google Scholar] [CrossRef] [PubMed]
- Hellsten, U.; Harland, R.M.; Gilchrist, M.J.; Hendrix, D.; Jurka, J.; Kapitonov, V.; Ovcharenko, I.; Putnam, N.H.; Shu, S.; Taher, L.; et al. The genome of the western clawed frog Xenopus tropicalis. Science 2010, 328, 633–636. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.B.; Amemiya, C.; Bailey, W.; Kawasaki, K.; Mezey, J.; Miller, W.; Minoshima, S.; Shimizu, N.; Wagner, G.; Ruddle, F. Hox cluster genomics in the horn shark, Heterodontus francisci. Proc. Natl. Acad. Sci. USA 2000, 97, 1655–1660. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.P.; Koh, E.G.L.; Tay, A.; Brenner, S.; Venkatesh, B. Highly conserved syntenic blocks at the vertebrate Hox loci and conserved regulatory elements within and outside Hox gene clusters. Proc. Natl. Acad. Sci. USA 2006, 103, 6994–6999. [Google Scholar] [CrossRef] [PubMed]
- Malaga-Trillo, E.; Meyer, A. Genome duplications and accelerated evolution of Hox genes and cluster architecture in teleost fishes. Am. Zool. 2001, 41, 676–686. [Google Scholar]
- Oulion, S.; Debiais-Thibaud, M.; D’Aubenton-Carafa, Y.; Thermes, C.; da Silva, C.; Bernard-Samain, S.; Gavory, F.; Wincker, P.; Mazan, S.; Casane, D. Evolution of Hox gene clusters in gnathostomes: Insights from a survey of a shark (Scyliorhinus canicula) transcriptome. Mol. Biol. Evol. 2010, 27, 2829–2838. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Raincrow, J.D. Hox Cluster Intergenic Sequence Evolution. Ph.D. Thesis, Rutgers University, NJ, USA, 2010. [Google Scholar]
- Goode, D.K.; Callaway, H.A.; Cerda, G.A.; Lewis, K.E.; Elgar, G. Minor change, major divergence: Divergent functions of highly conserved cis-regulatory elements subsequent to whole genome duplication events. Development 2011, 138, 879–884. [Google Scholar] [CrossRef] [PubMed]
- Santagati, F.; Minoux, M.; Ren, S.Y.; Rijli, F.M. Temporal requirement of Hoxa2 in cranial neural crest skeletal morphogenesis. Development 2005, 132, 4927–4936. [Google Scholar] [CrossRef] [PubMed]
- Scemama, J.L.; Hunter, M.; McCallum, J.; Prince, V.; Stellwag, E. Evolutionary divergence of vertebrate Hoxb2 expression patterns and transcriptional regulatory loci. J. Exp. Zool. B Mol. Evol. Dev. 2002, 294, 285–299. [Google Scholar] [CrossRef] [PubMed]
- Caroll, S.B. Evo-Devo and an expanding evolutionary synthesis: A genetic theory of morphological evolution. Cell 2008, 134, 25–36. [Google Scholar] [CrossRef] [PubMed]
- Capellini, T.D.; di Giacomo, G.; Salsi, V.; Brendolan, A.; Ferretti, E.; Srivastava, D.; Zappavigna, V.; Selleri, L. Pbx1/Pbx2 requirement for distal limb patterning is mediated by the hierarchical control of Hox gene spatial distribution and Shh expression. Development 2006, 133, 2263–2273. [Google Scholar] [CrossRef] [PubMed]
- Kmita, M.; Tarchini, B.; Zakany, J.; Logan, M.; Tabin, C.J.; Duboule, D. Early developmental arrest of mammalian limbs lacking HoxA/HoxD gene function. Nature 2005, 435, 1113–1116. [Google Scholar] [CrossRef] [PubMed]
- Mercader, N.; Leonardo, E.; Azpiazu, N.; Serrano, A.; Morata, G.; Martinez, C.; Torres, M. Conserved regulation of proximodistal limb axis development by Meis1/Hth. Nature 1999, 402, 425–429. [Google Scholar] [PubMed]
- Tamura, K.; Yokouchi, Y.; Kuroiwa, A.; Ide, H. Retinoic acid changes the proximodistal developmental competence and affinity of distal cells in the developing chick limb bud. Dev. Biol. 1997, 188, 224–234. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Yonei-Tamura, S.; Yano, T.; Yokoyama, H.; Ide, H. The autopod: Its formation during limb development. Dev. Growth Differ. 2008, 50, S177–S187. [Google Scholar] [CrossRef] [PubMed]
- Zeller, R. The temporal dynamics of vertebrate limb development, teratogenesis and evolution. Curr. Opin. Genet. Dev. 2010, 20, 384–390. [Google Scholar] [CrossRef] [PubMed]


| Primer | Sequence 5′ to 3′ | 5′ Start Site |
|---|---|---|
| Medaka hoxa2a Genomic Primers | ||
| A2a For | TTATTCCCACAACCCTTTCATTTCG | −2691 |
| A2a Rev | CACACTCAGCCACAATCTCTTCTTC | 1846 |
| Medaka ψhoxa2b Genomic Primers | ||
| A2b For | ACACAGCAGGGGTCAACAATAGGTC | −3093 |
| A2b Rev | ATAGGCAGAGCACGAAAACAAAATG | 3193 |
| Medaka hoxa2a UER(K20-RE5) Forward Primers | ||
| AF1 | GATCGATATCGAACAGGCTGAAATCCACTGAATGC | −1778 |
| AF2 | GATCGATATCGCTTCTAATCTGAGAAGCCAGTGTTTC | −1468 |
| AF3 | GATCGATATCATGTGTTGCGAGGGCACCGAGCTGTC | −1392 |
| AF4 | GATCGATATCGAGTAAGATTGATCGCGCACAGGCTTC | −1354 |
| Medaka hoxa2a UER(K20-RE5) Reverse Primers | ||
| AR1 | GATCGAATTCGTTTGCTGTGGAACAGAGGAAAGAAG | −1247 |
| AR2 | GATCGAATTCTTATATACCAAACAAAGAGTCCTGG | −1303 |
| AR3 | GATCGAATTCTTACTCGCCAAAAGGTCTGACAGCTC | −1348 |
| Medaka ψhoxa2b UER(K20-RE5) Forward Primers | ||
| BF1 | GATCGATATCATGTGCCAACACCCACTCACCCCAG | −1068 |
| BF2 | GATCGATATCCTTCGCTCCGCACCGAGGGCATCCTC | −868 |
| BF3 | GATCGATATCATGTTCTCTAAGGGCAAAGAGCTGTC | −803 |
| BF4 | GATCGATATCTGGAAAGATTGATCACACAGAATACC | −765 |
| Medaka ψhoxa2b UER(K20-RE5) Reverse Primers | ||
| BR1 | GATCGAATTCAAAAAGCTGCAGGAAAAGGAGGGGATC | −671 |
| BR2 | GATCGAATTCCCGGGCTCTGAACAAAAGATTCCTG | −715 |
| BR3 | GATCGAATTCTTTCCAGCCAAGAGCTCTGACAGCTC | −759 |
| Construct | Primer Pairs | Amplicon Length | Construct Schematic | Hindbrain Expression (F0) | CNCC Expression (F0)) | F1s |
|---|---|---|---|---|---|---|
| Medaka hoxa2a UER(K20-RE5) construct design and transgenic analysis | ||||||
| 1 | AF1/AR1 | 531 bp |  | 42/48 (87.5%) | 42/48 (87.5%) | 3 |
| 2 | AF2/AR1 | 221 bp |  | 64/84 (76%) | 56/84 (67%) | 3 |
| 3 | AF3/AR1 | 145 bp |  | 39/49 (80%) | 41/49 (84%) | 0 |
| 4 | AF4/AR1 | 107 bp |  | 7/47 (15%) * | 23/49 (49%) * | 0 |
| 5 | AF1/AR2 | 475 bp |  | 42/50 (84%) | 42/50 (84%) | 4 |
| 6 | AF1/AR3 | 430 bp |  | 0/52 (0%) | 0/52 (0%) | 0 |
| 7 | AF3/AR2 | 89 bp |  | 52/62 (84%) | 52/62 (84%) | 4 |
| Medaka ψhoxa2b UER(K20-RE5) construct design and transgenic analysis | ||||||
| 8 | BF1/BR1 | 397 bp |  | 46/51 (90%) | 46/51 (90%) | 2 |
| 9 | BF2/BR1 | 197 bp |  | 47/52 (90%) | 47/52 (90%) | 4 |
| 10 | BF3/BR1 | 132 bp | 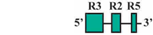 | 33/38 (87%) | 33/38 (87%) | 3 |
| 11 | BF4/BR1 | 94 bp |  | 27/52 (52%) * | 27/52 (52%)* | 0 |
| 12 | BF1/BR2 | 353 bp |  | 33/41 (80%) | 33/41 (80%) | 3 |
| 13 | BF1/BR3 | 309 bp |  | 9/64 (14%) * | 15/64 (23%) * | 0 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Davis, A.; Reubens, M.C.; Stellwag, E.J. Functional and Comparative Genomics of Hoxa2 Gene cis-Regulatory Elements: Evidence for Evolutionary Modification of Ancestral Core Element Activity. J. Dev. Biol. 2016, 4, 15. https://doi.org/10.3390/jdb4020015
Davis A, Reubens MC, Stellwag EJ. Functional and Comparative Genomics of Hoxa2 Gene cis-Regulatory Elements: Evidence for Evolutionary Modification of Ancestral Core Element Activity. Journal of Developmental Biology. 2016; 4(2):15. https://doi.org/10.3390/jdb4020015
Chicago/Turabian StyleDavis, Adam, Michael C. Reubens, and Edmund J. Stellwag. 2016. "Functional and Comparative Genomics of Hoxa2 Gene cis-Regulatory Elements: Evidence for Evolutionary Modification of Ancestral Core Element Activity" Journal of Developmental Biology 4, no. 2: 15. https://doi.org/10.3390/jdb4020015
APA StyleDavis, A., Reubens, M. C., & Stellwag, E. J. (2016). Functional and Comparative Genomics of Hoxa2 Gene cis-Regulatory Elements: Evidence for Evolutionary Modification of Ancestral Core Element Activity. Journal of Developmental Biology, 4(2), 15. https://doi.org/10.3390/jdb4020015





