mgwr: A Python Implementation of Multiscale Geographically Weighted Regression for Investigating Process Spatial Heterogeneity and Scale
Abstract
:1. Introduction
2. Source Code and Datasets
2.1. Source Code and Installation
2.2. Datasets
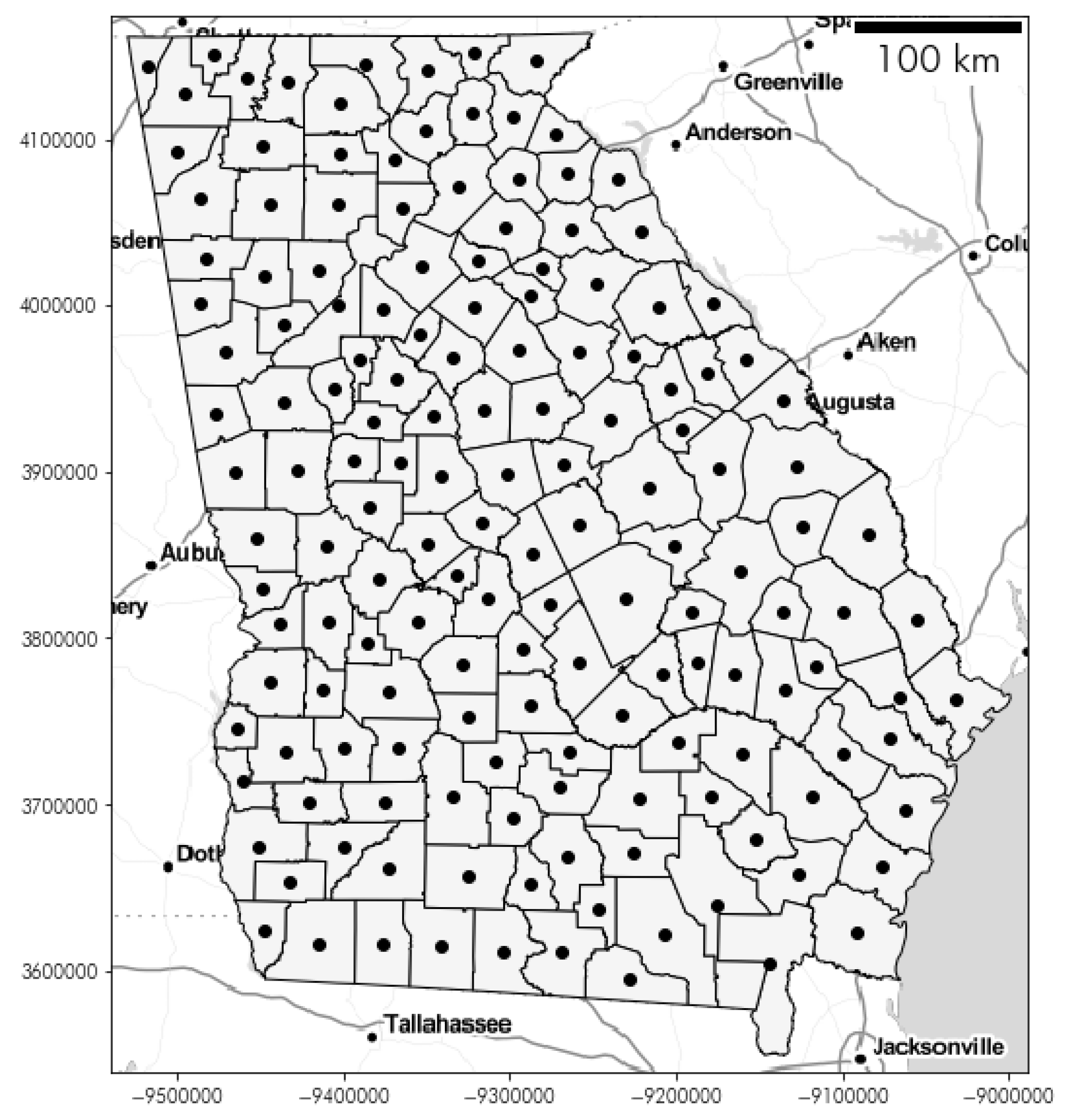
2.2.1. Georgia Dataset
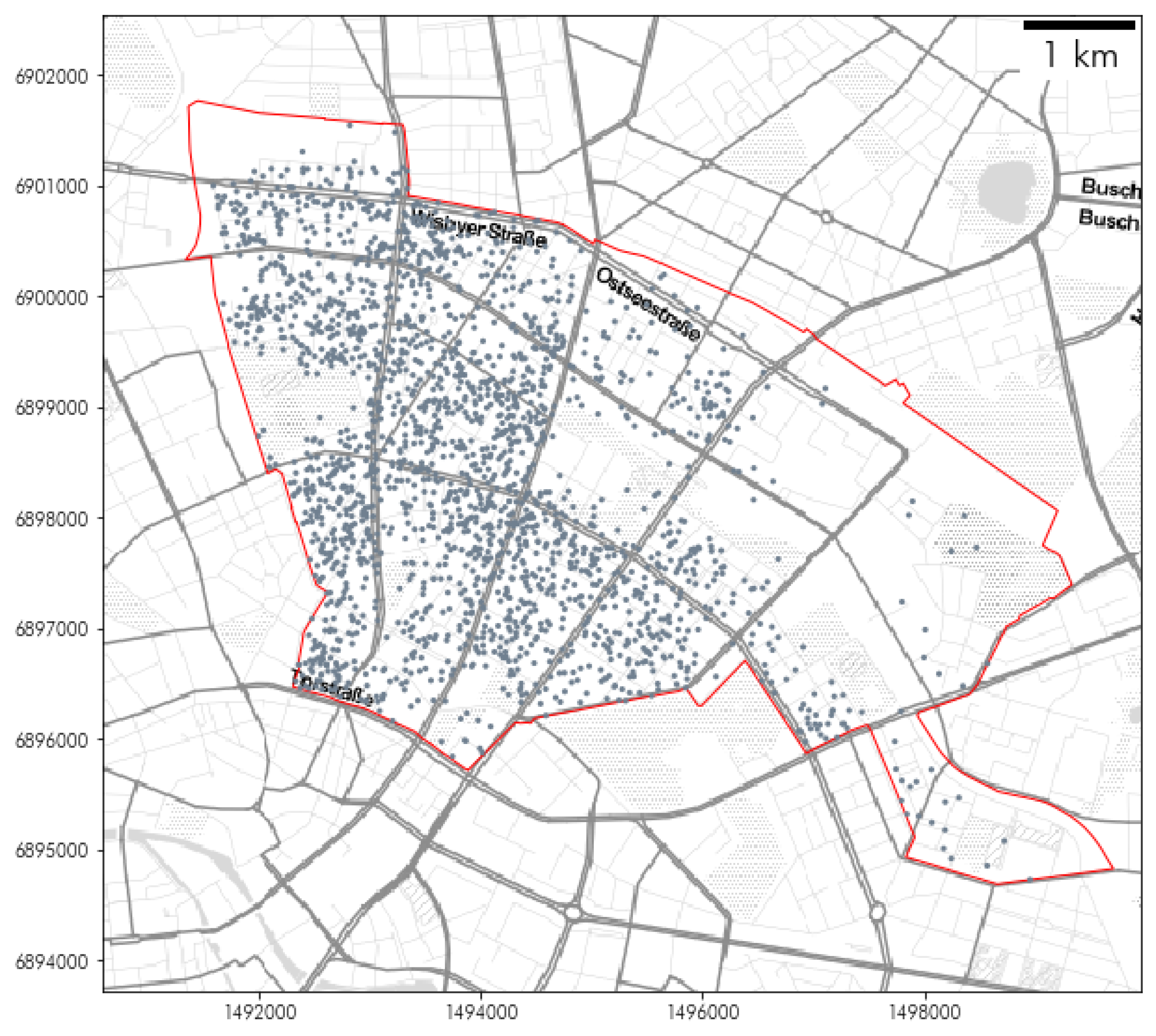
2.2.2. Berlin Airbnb Dataset
3. GWR Functionality
3.1. Distance-Weighting Scheme
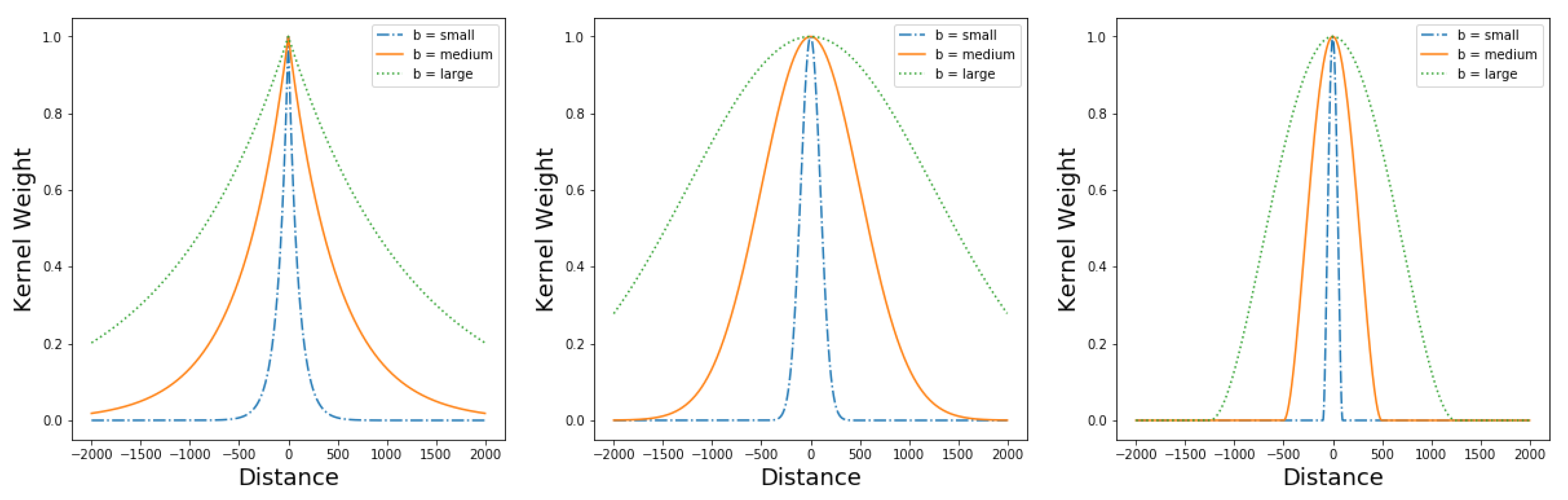
3.1.1. Kernel Functions
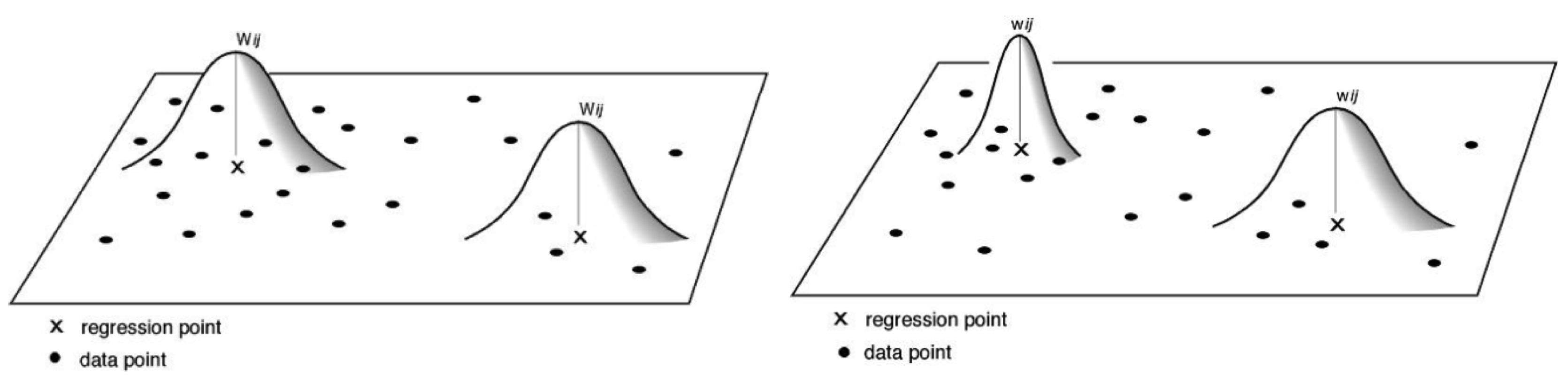
3.1.2. Kernel Types
3.2. Bandwidth Selection
3.3. Model Calibration
3.4. Probability Models
from spglm.family import Poisson, Binomialand then it is necessary to set family = Poisson() or family = Binomial() when instantiating a Sel_BW or GWR object. Generally, it is not necessary to import or specify a Gaussian family object since it is the default behavior across mgwr.
3.5. Model Diagnostics
3.5.1. Model Fit
3.5.2. Inference on Individual Parameter Estimates
3.5.3. Inference on Surface of Parameter Estimates
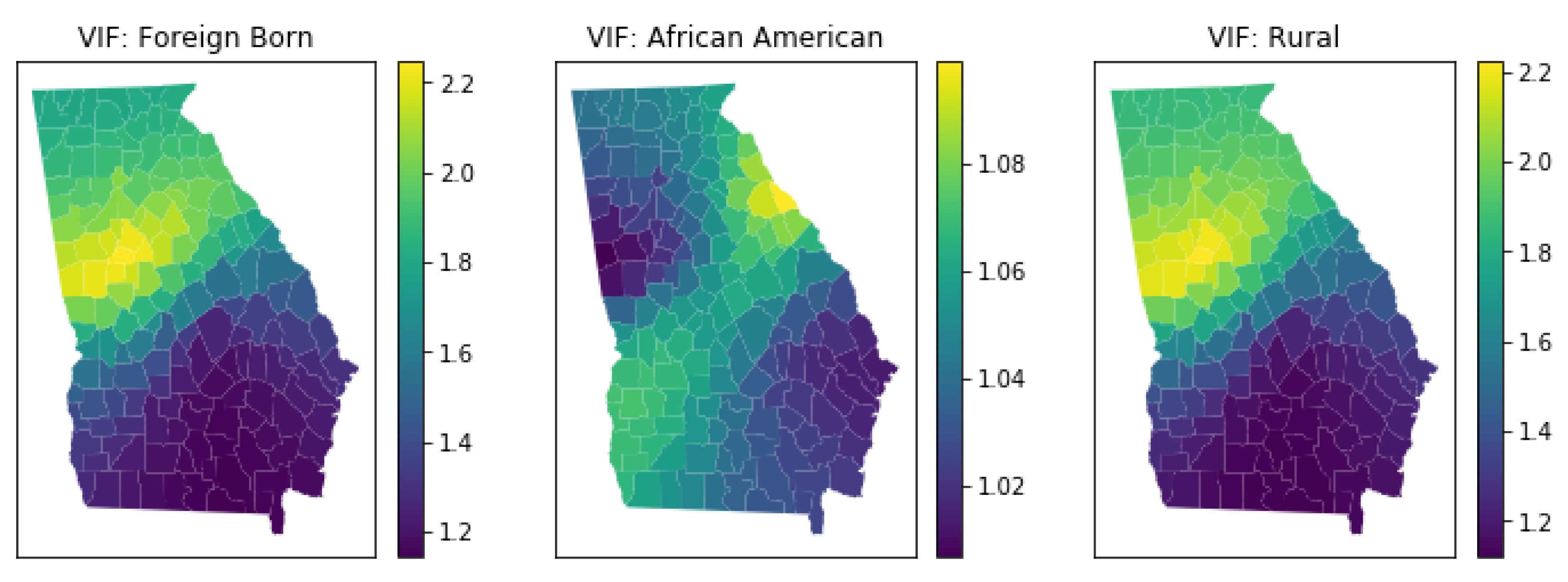
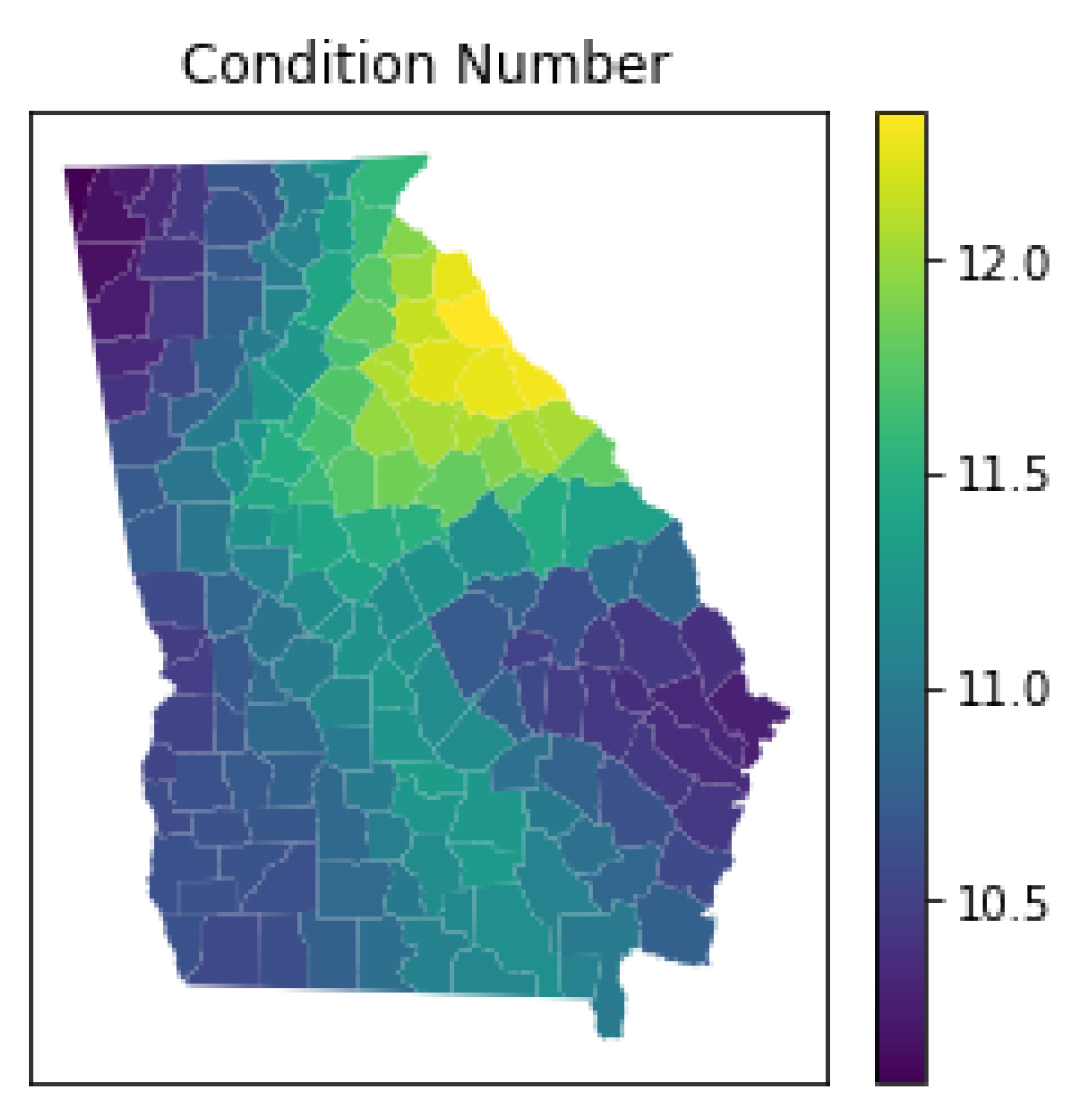
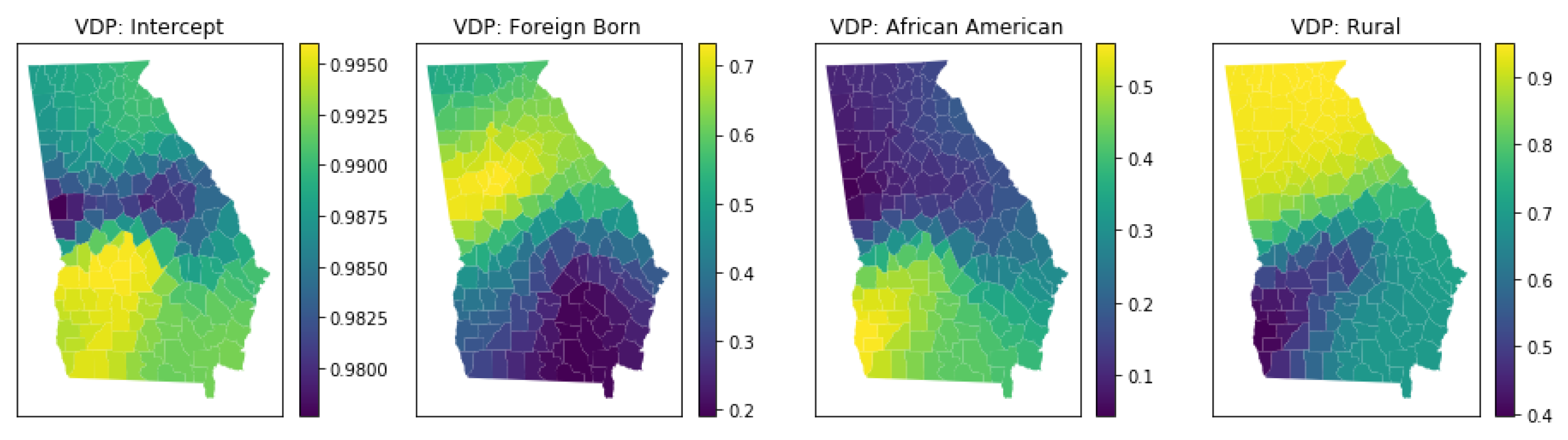
3.5.4. Local Multicollinearity
3.6. Out-of-Sample Spatial Prediction
4. MGWR Functionality
4.1. Standardizing the Variables
4.2. Bandwidth Selection and Model Calibration
4.3. Manually Setting Covariate-Specific Bandwidths
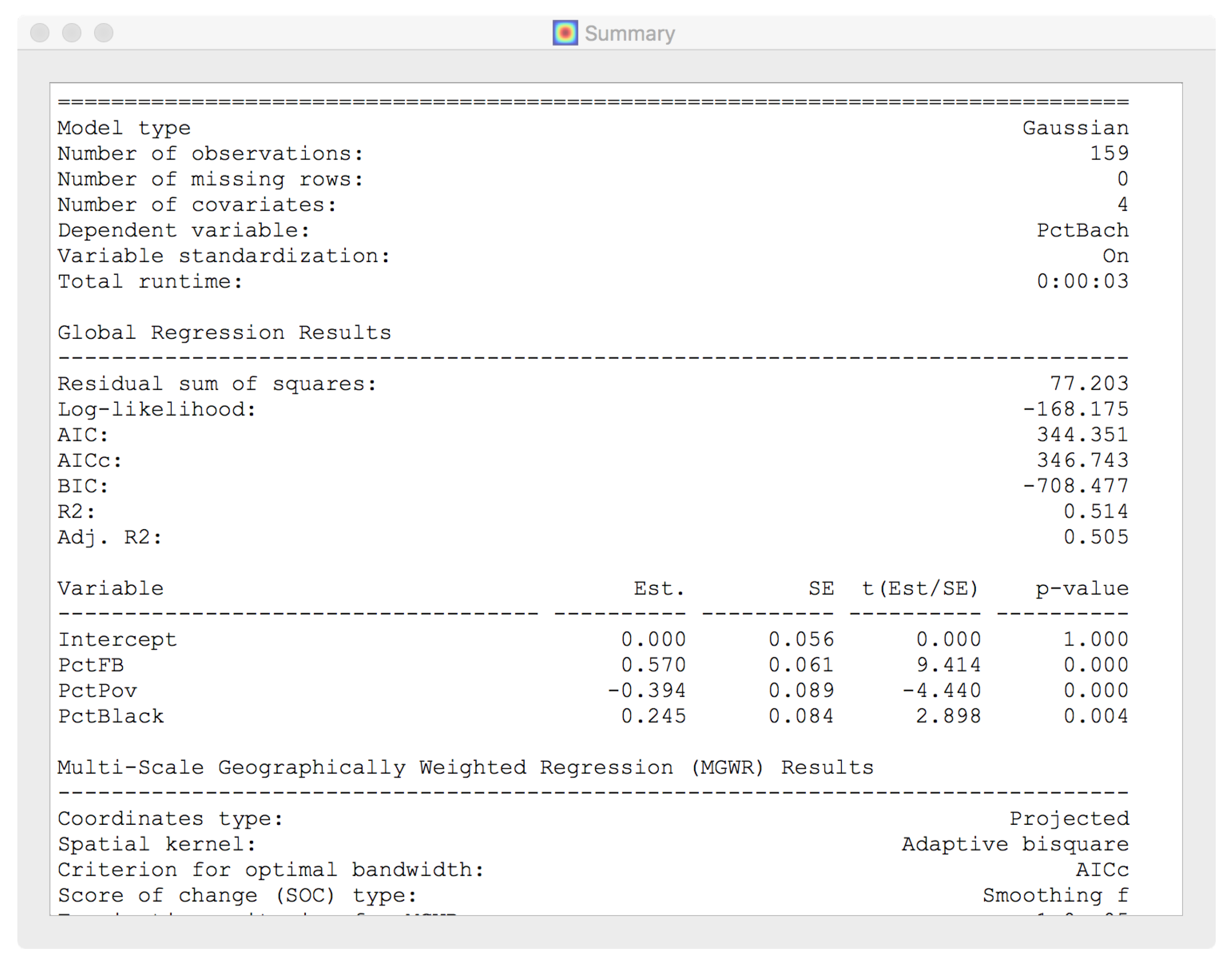
4.4. Model Fit
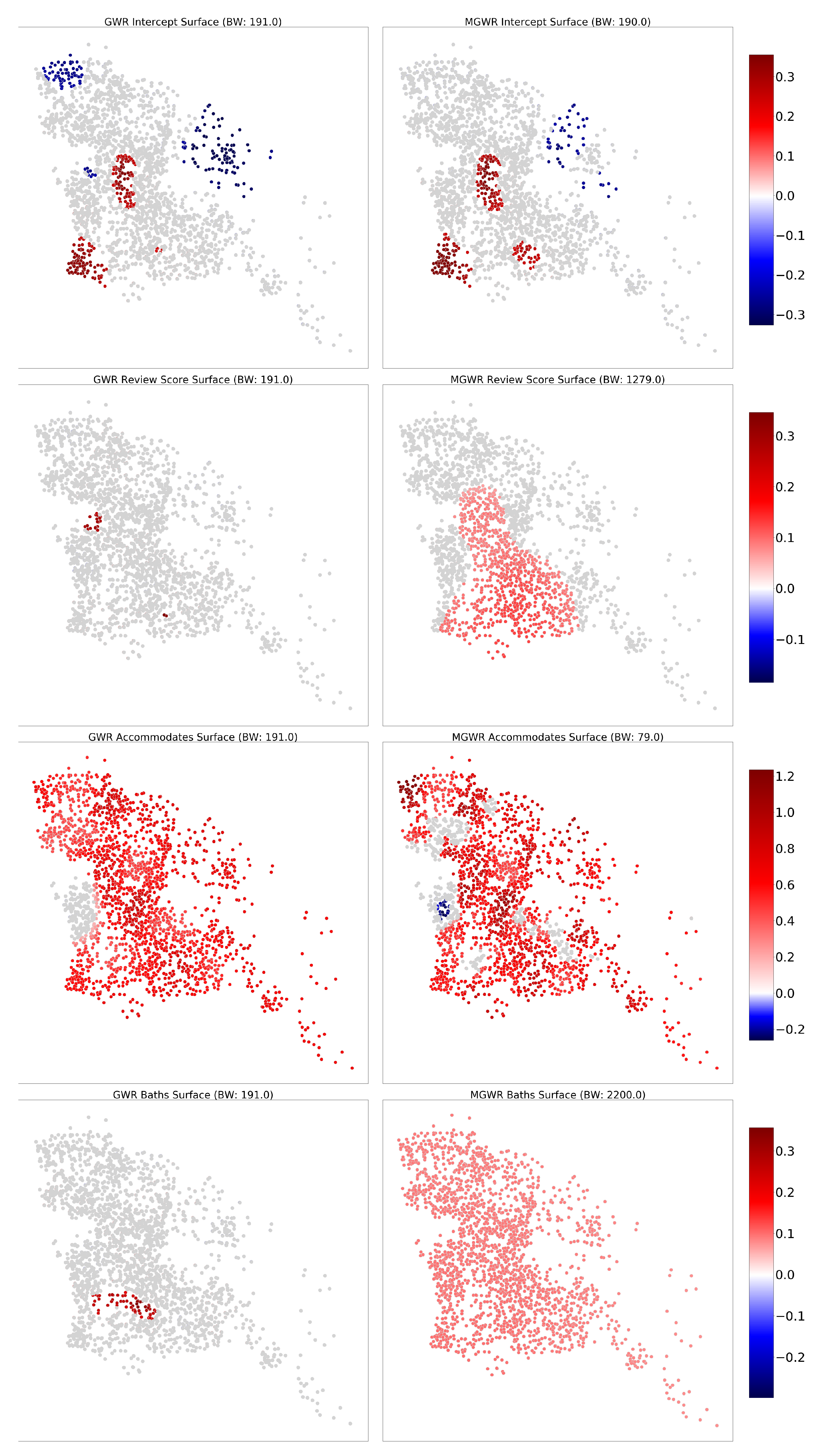
4.5. Inference on Parameter Estimates
4.5.1. The Georgia Dataset
4.5.2. The Berlin Dataset
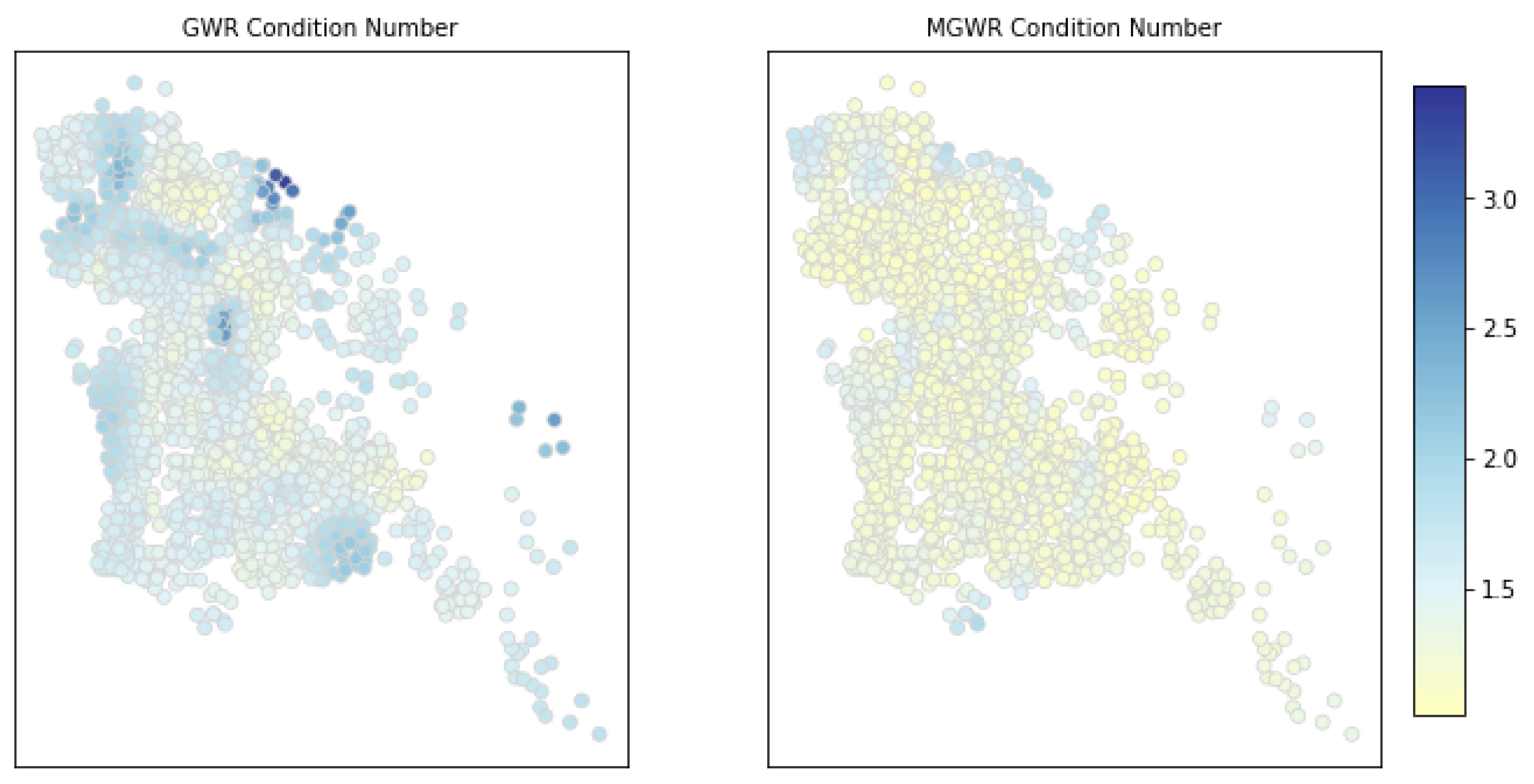
4.6. Local Multicollinearity
5. Additional Features
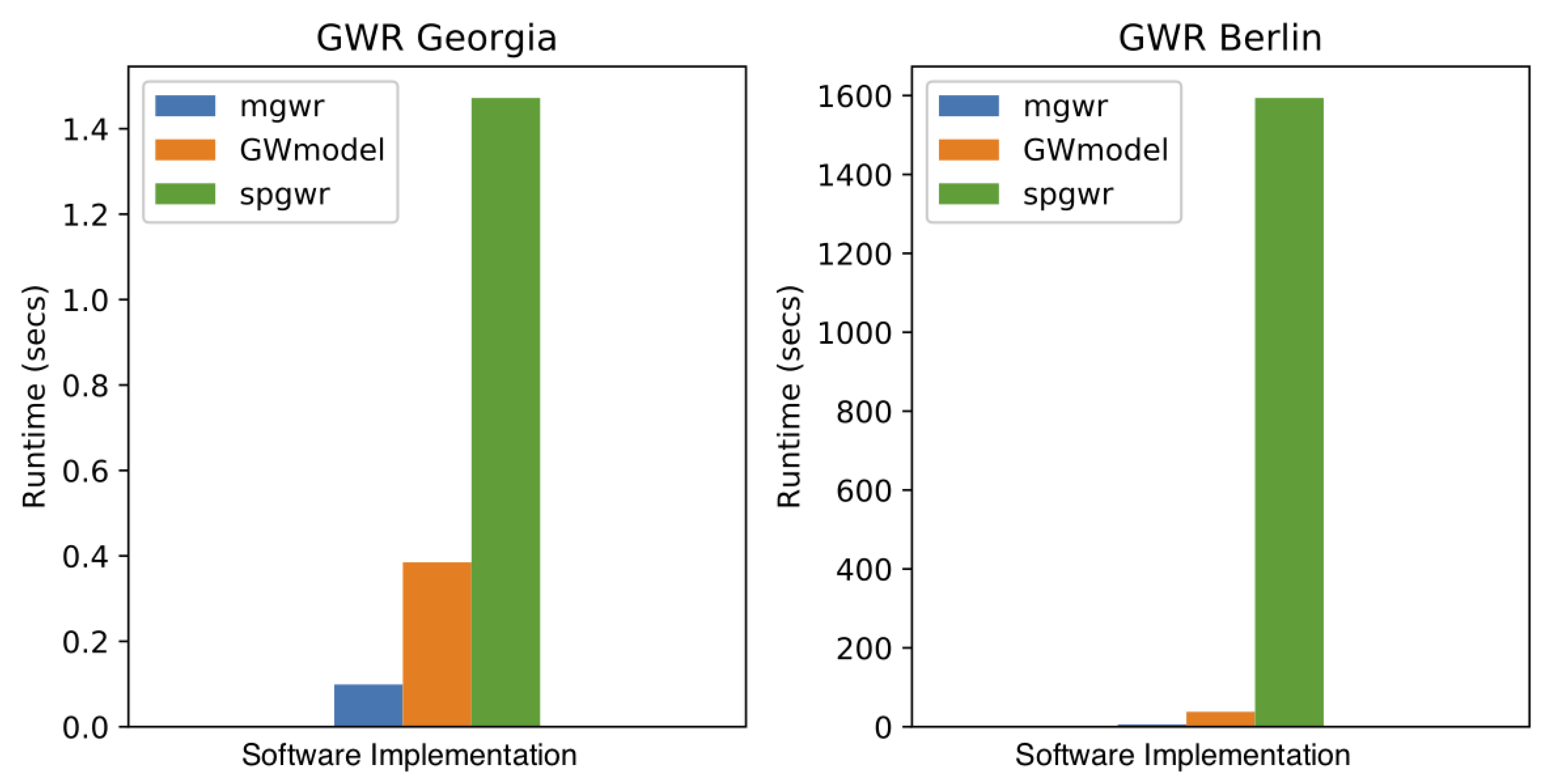
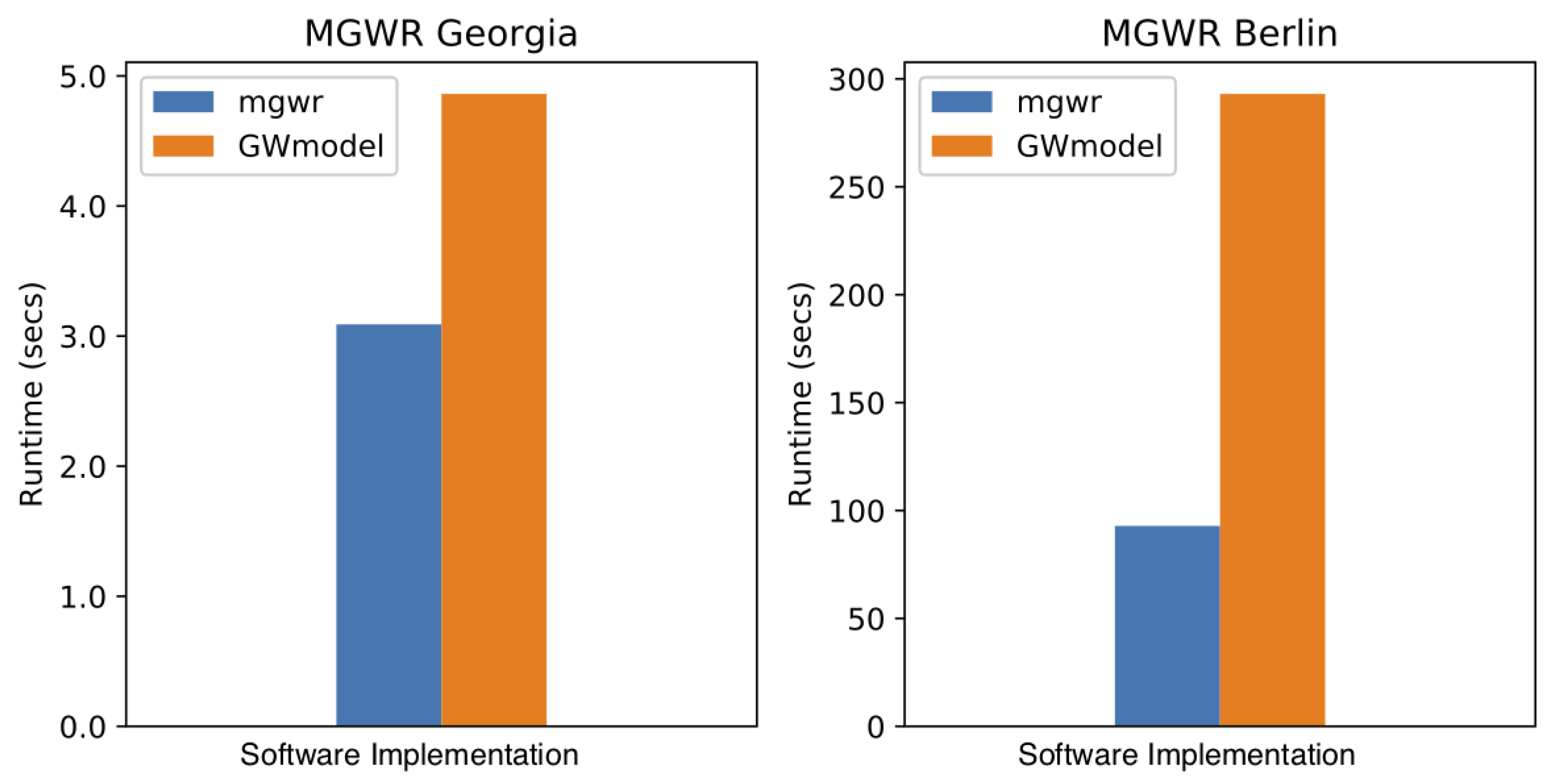
5.1. Computational Efficiency
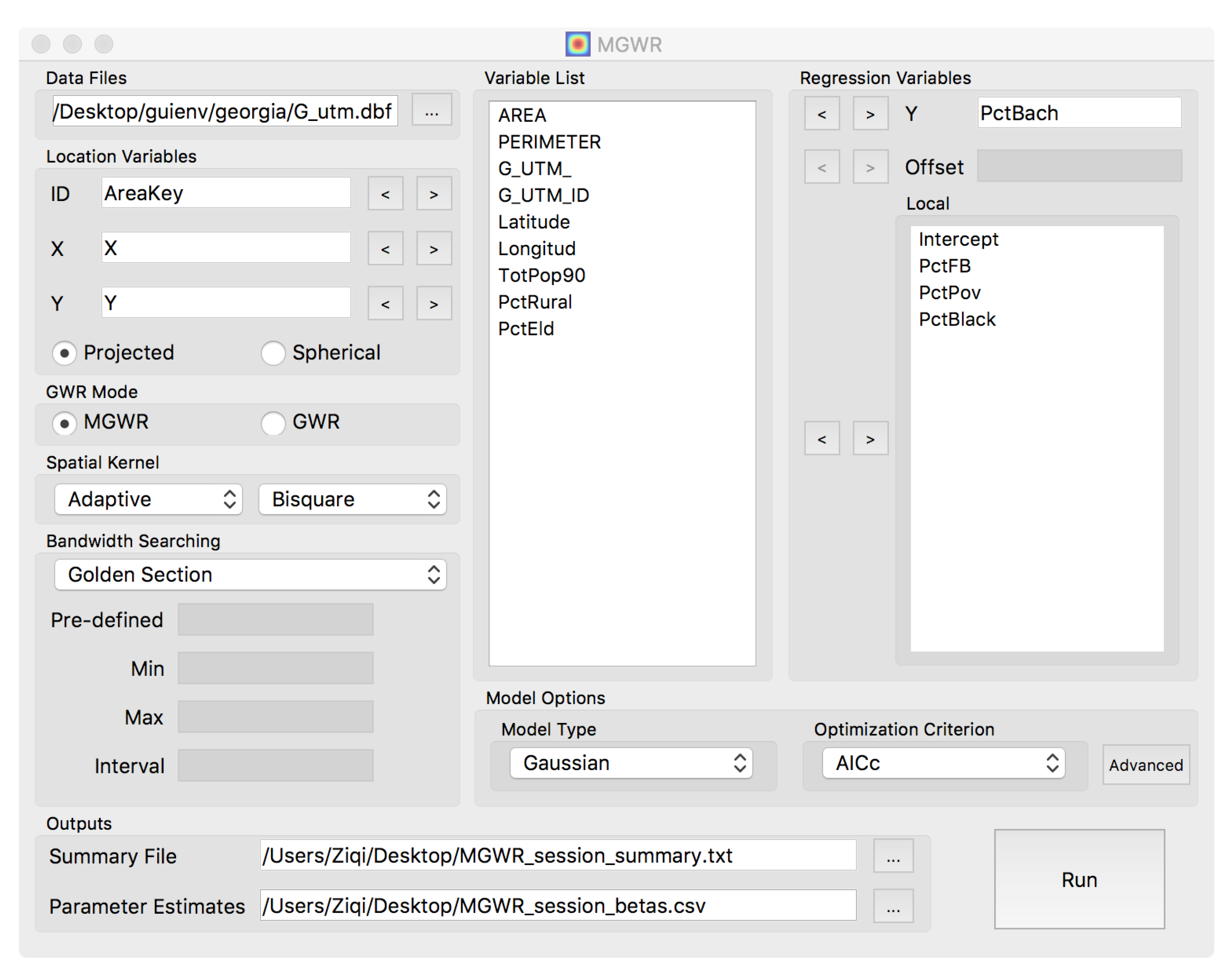
5.2. Accessibility
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Tobler, W.R. A Computer Movie Simulating Urban Growth in the Detroit Region. Econ. Geogr. 1970, 46, 234. [Google Scholar] [CrossRef]
- Fotheringham, A.S.; Brunsdon, C.; Charlton, M. Geographically Weighted Regression: The Analysis of Spatially Varying Relationships; John Wiley & Sons: Hoboken, NJ, USA, 2002. [Google Scholar]
- Fotheringham, A.S.; Yang, W.; Kang, W. Multi-Scale Geographically Weighted Regression. Ann. Am. Assoc. Geogr. 2017, 107, 1247–1265. [Google Scholar]
- Environmental Systems Research Institute (ESRI). ArcMap 10.3 Spatial Analyst Toolbox; ESRI: Redlands, CA, USA, 2018. [Google Scholar]
- Bivand, R.; Yu, D.; Nakaya, T.; Garcia-Lopez, M.A. spgwr: Geographically Weighted Regression, R package version 0.6-32; 2017. [Google Scholar]
- Wheeler, D. gwrr: Fits Geographically Weighted Regression Models with Diagnostic Tools, R package version 0.2-1; 2013. [Google Scholar]
- Yu, H.; Fotheringham, A.S.; Li, Z.; Oshan, T.; Kang, W.; Wolf, L.J. Inference in multiscale geographically weighted regression. Geogr. Anal. 2019. [Google Scholar] [CrossRef]
- Lu, B.; Harris, P.; Charlton, M.; Brundson, C.; Nayaka, T.; Gollini, I. GWmodel: Geographically-Weighted Models, R package version 2.0-5; 2018. [Google Scholar]
- Lu, B.; Brunsdon, C.; Charlton, M.; Harris, P. Geographically weighted regression with parameter-specific distance metrics. Int. J. Geogr. Inf. Sci. 2017, 31, 982–998. [Google Scholar] [CrossRef]
- Li, Z.; Fotheringham, A.S.; Li, W.; Oshan, T. Fast Geographically Weighted Regression (FastGWR): A Scalable Algorithm to Investigate Spatial Process Heterogeneity in Millions of Observations. Int. J. Geogr. Inf. Sci. 2018. [Google Scholar] [CrossRef]
- Griffith, D.A. Spatial-filtering-based contributions to a critique of geographically weighted regression (GWR). Environ. Plan. A 2008, 40, 2751–2769. [Google Scholar] [CrossRef]
- Da Silva, A.R.; Fotheringham, A.S. The Multiple Testing Issue in Geographically Weighted Regression: The Multiple Testing Issue in GWR. Geogr. Anal. 2015. [Google Scholar] [CrossRef]
- Wheeler, D.; Tiefelsdorf, M. Multicollinearity and correlation among local regression coefficients in geographically weighted regression. J. Geogr. Syst. 2005, 7, 161–187. [Google Scholar] [CrossRef]
- Belsey, D.A.; Kuh, E.; Welsch, R.E. Regression Diagnostics: Identifying Influential Data and Sources of Collinearity; Wiley: New York, NY, USA, 1980. [Google Scholar]
- O’brien, R.M. A Caution Regarding Rules of Thumb for Variance Inflation Factors. Qual. Quant. 2007, 41, 673–690. [Google Scholar] [CrossRef]
- Wheeler, D.C. Diagnostic Tools and a Remedial Method for Collinearity in Geographically Weighted Regression. Environ. Plan. A 2007, 39, 2464–2481. [Google Scholar] [CrossRef]
- Fotheringham, A.S.; Oshan, T.M. Geographically weighted regression and multicollinearity: Dispelling the myth. J. Geogr. Syst. 2016, 18, 303–329. [Google Scholar] [CrossRef]
- Oshan, T.M.; Fotheringham, A.S. A Comparison of Spatially Varying Regression Coefficient Estimates Using Geographically Weighted and Spatial-Filter-Based Techniques: A Comparison of Spatially Varying Regression. Geogr. Anal. 2017. [Google Scholar] [CrossRef]
- Murakami, D.; Lu, B.; Harris, P.; Brunsdon, C.; Charlton, M.; Nakaya, T.; Griffith, D.A. The importance of scale in spatially varying coefficient modeling. arXivt 2017, arXiv:1709.08764. [Google Scholar] [CrossRef]
- Harris, P.; Fotheringham, A.S.; Crespo, R.; Charlton, M. The Use of Geographically Weighted Regression for Spatial Prediction: An Evaluation of Models Using Simulated Data Sets. Math. Geosci. 2010, 42, 657–680. [Google Scholar] [CrossRef]
- Lu, B.; Yang, W.; Ge, Y.; Harris, P. Improvements to the calibration of a geographically weighted regression with parameter-specific distance metrics and bandwidths. Comput. Environ. Urban Syst. 2018. [Google Scholar] [CrossRef]
- Comber, A.; Chi, K.; Quang Huy, M.; Nguyen, Q.; Lu, B.; Huu Phe, H.; Harris, P. Distance metric choice can both reduce and induce collinearity in geographically weighted regression. Environ. Plan. B Urban Anal. City Sci. 2018. [Google Scholar] [CrossRef]

| Short Name | Description |
|---|---|
| PctBach | Percentage of the population with a bachelor’s degree or higher |
| PctFB | Percentage of the population that was born in a foreign country |
| PctBlack | Percentage of the population that identifies as African American |
| PctRural | Percentage of the population that is classified as living in a rural area |
| Short Name | Description |
|---|---|
| Log price | Logged price of rental unit |
| Score | Cumulative review score from previous customers for each rental unit |
| Accommodates | Number of individuals a rental unit can accommodate |
| Bathrooms | Number of bathrooms in each rental unit |
| Function | Specification | Input Parameter |
|---|---|---|
| Gaussian | kernel=‘gaussian’ | |
| Exponential | kernel=‘exponential’ | |
| Bi-square | kernel=‘bisquare’ |
| Name | Input Parameter |
|---|---|
| Cross-validation (CV) | criterion=‘CV’ |
| Akaike information criterion (AIC) | criterion = ‘AIC’ |
| Corrected AIC (AICc) | criterion = ‘AICc’ |
| Bayesian information criterion (BIC) | criterion = ‘BIC’ |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oshan, T.M.; Li, Z.; Kang, W.; Wolf, L.J.; Fotheringham, A.S. mgwr: A Python Implementation of Multiscale Geographically Weighted Regression for Investigating Process Spatial Heterogeneity and Scale. ISPRS Int. J. Geo-Inf. 2019, 8, 269. https://doi.org/10.3390/ijgi8060269
Oshan TM, Li Z, Kang W, Wolf LJ, Fotheringham AS. mgwr: A Python Implementation of Multiscale Geographically Weighted Regression for Investigating Process Spatial Heterogeneity and Scale. ISPRS International Journal of Geo-Information. 2019; 8(6):269. https://doi.org/10.3390/ijgi8060269
Chicago/Turabian StyleOshan, Taylor M., Ziqi Li, Wei Kang, Levi J. Wolf, and A. Stewart Fotheringham. 2019. "mgwr: A Python Implementation of Multiscale Geographically Weighted Regression for Investigating Process Spatial Heterogeneity and Scale" ISPRS International Journal of Geo-Information 8, no. 6: 269. https://doi.org/10.3390/ijgi8060269
APA StyleOshan, T. M., Li, Z., Kang, W., Wolf, L. J., & Fotheringham, A. S. (2019). mgwr: A Python Implementation of Multiscale Geographically Weighted Regression for Investigating Process Spatial Heterogeneity and Scale. ISPRS International Journal of Geo-Information, 8(6), 269. https://doi.org/10.3390/ijgi8060269





