Critical Assessment of Methods to Quantify Biofilm Growth and Evaluate Antibiofilm Activity of Host Defence Peptides
Abstract
1. Introduction
2. Methods and Materials
2.1. Source of Synthetic Peptides and Chemical Reagents
2.2. Bacterial Strains and Growth Conditions
2.3. Biofilm Growth in Glass Tubes
2.4. Biofilm Growth in Flow Cells
2.5. Microtitre Biofilm Inhibition Assay
2.6. Microtitre Biofilm Eradication Assay
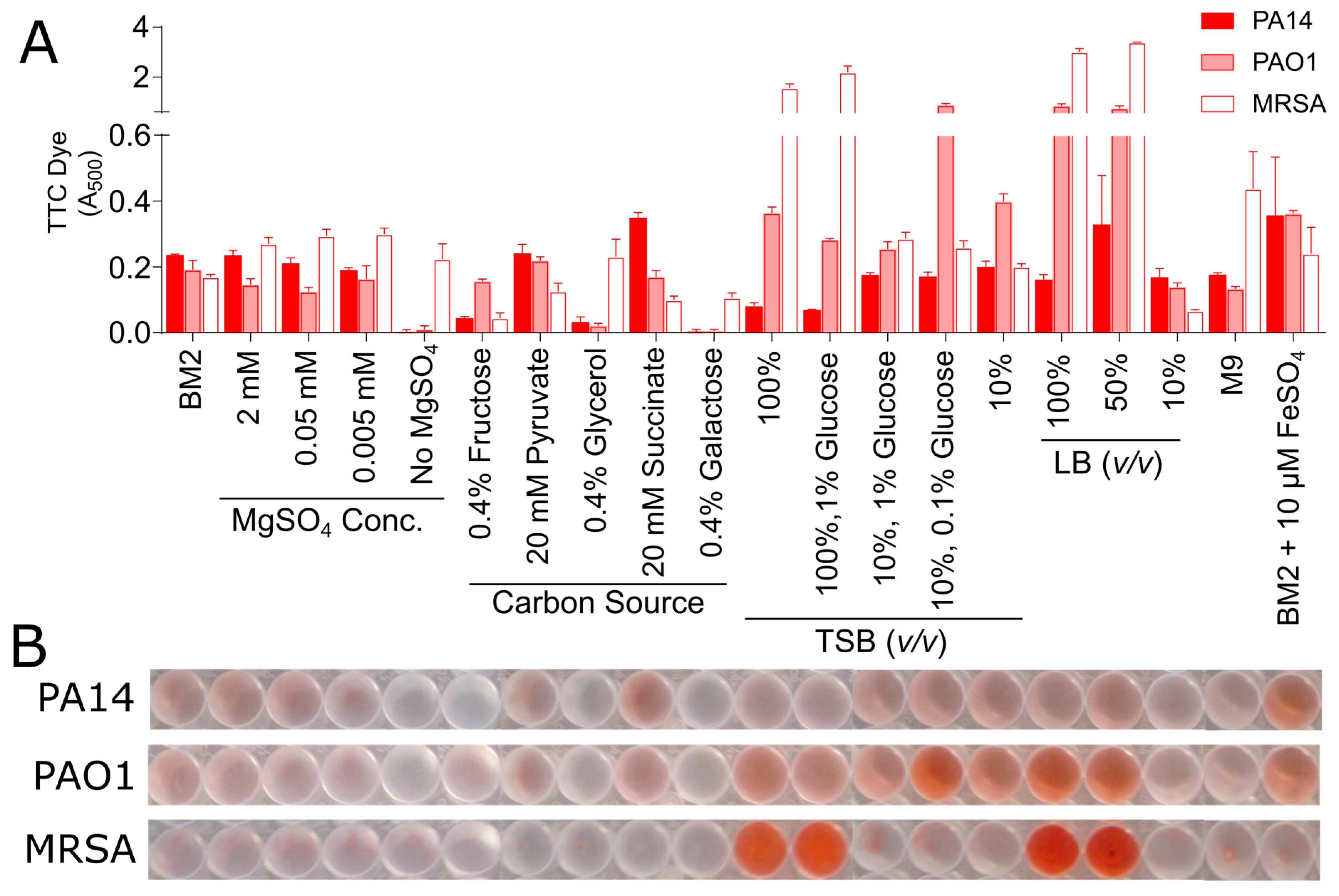
2.7. Effect of Media on Triphenyl Tetrazolium Chloride Metabolism
2.8. Quantification of Adhered Biofilm Cells in Microtitre Plates
3. Results
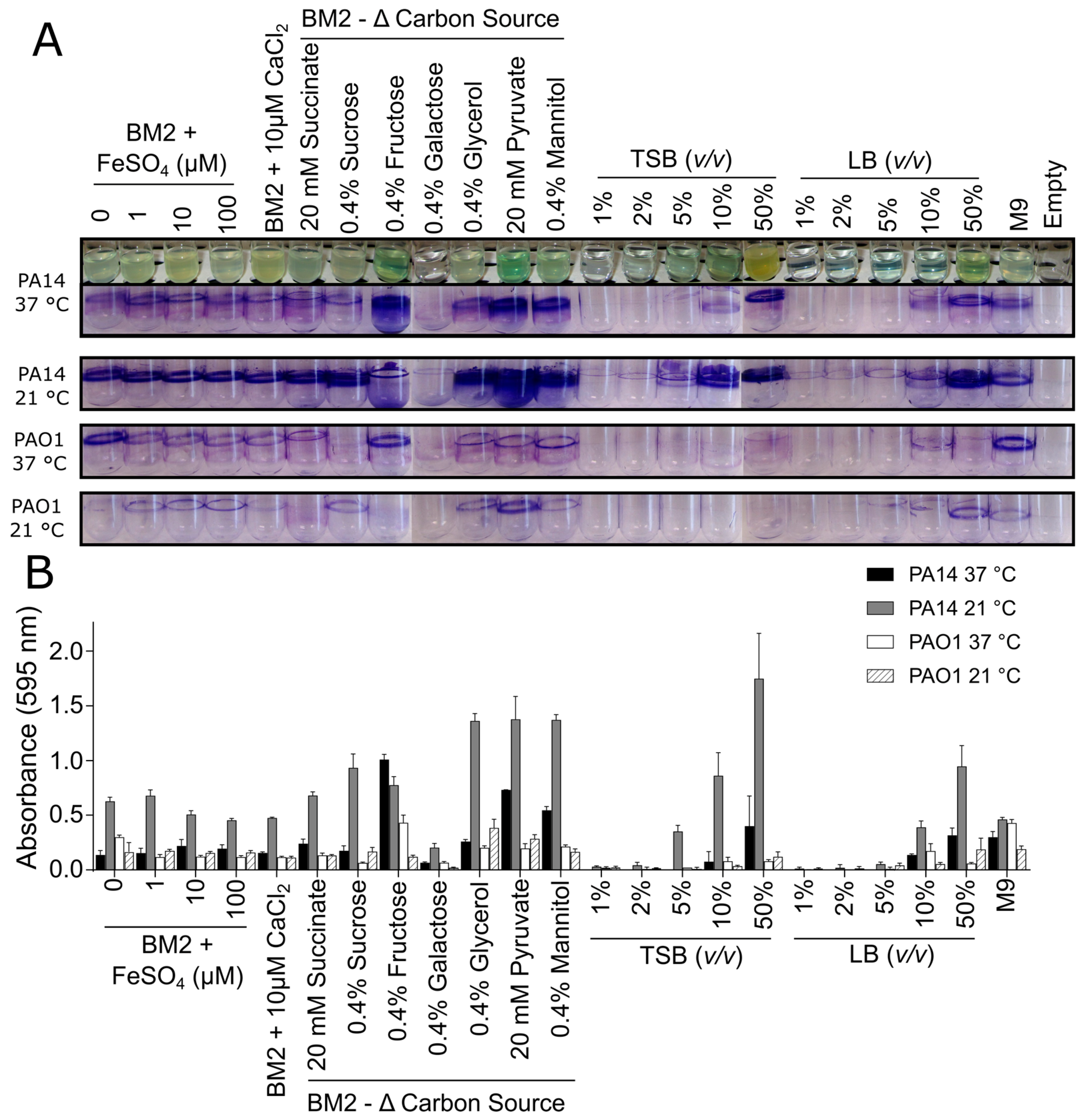
3.1. Biofilm Growth in Glass Tubes
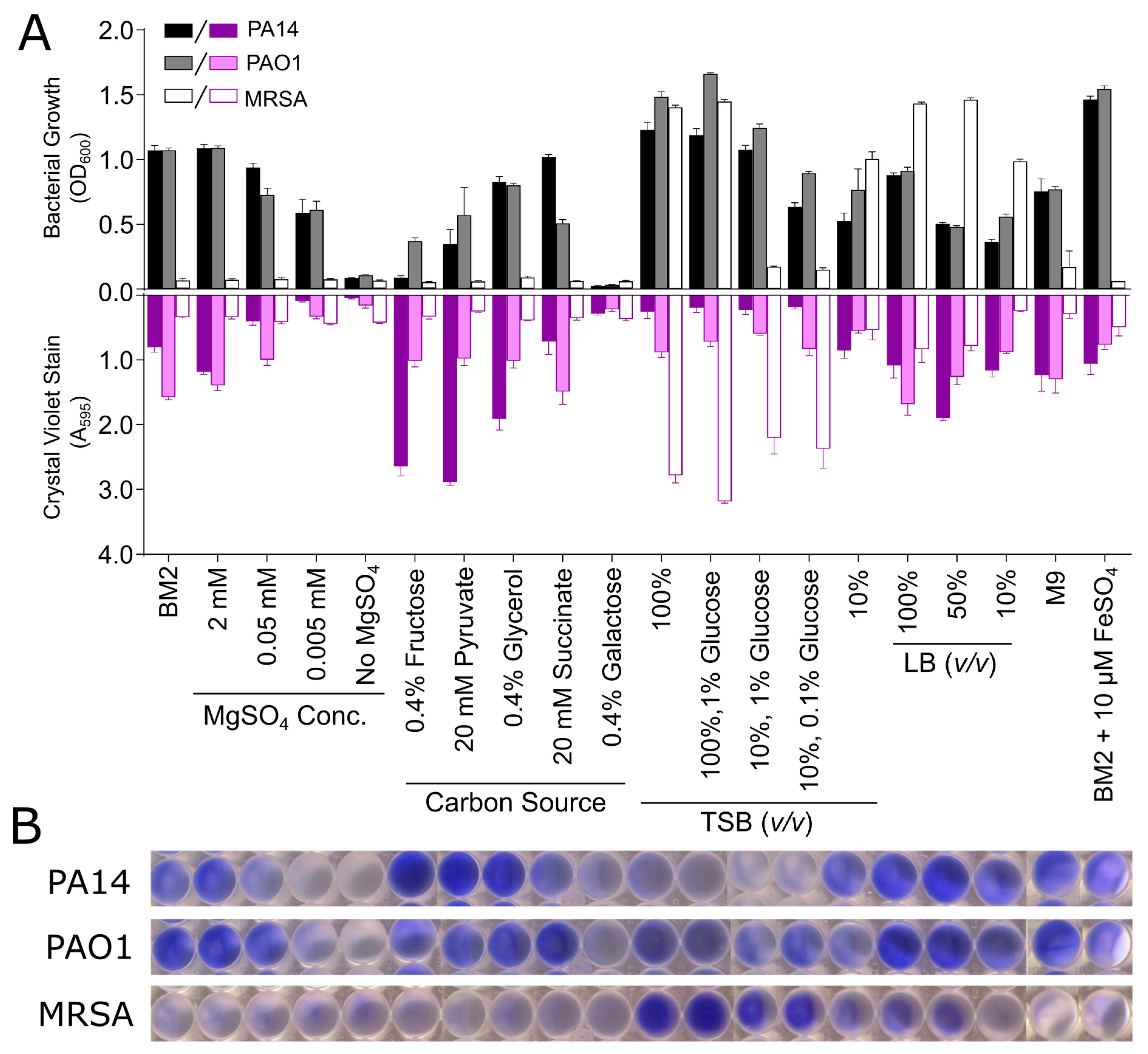
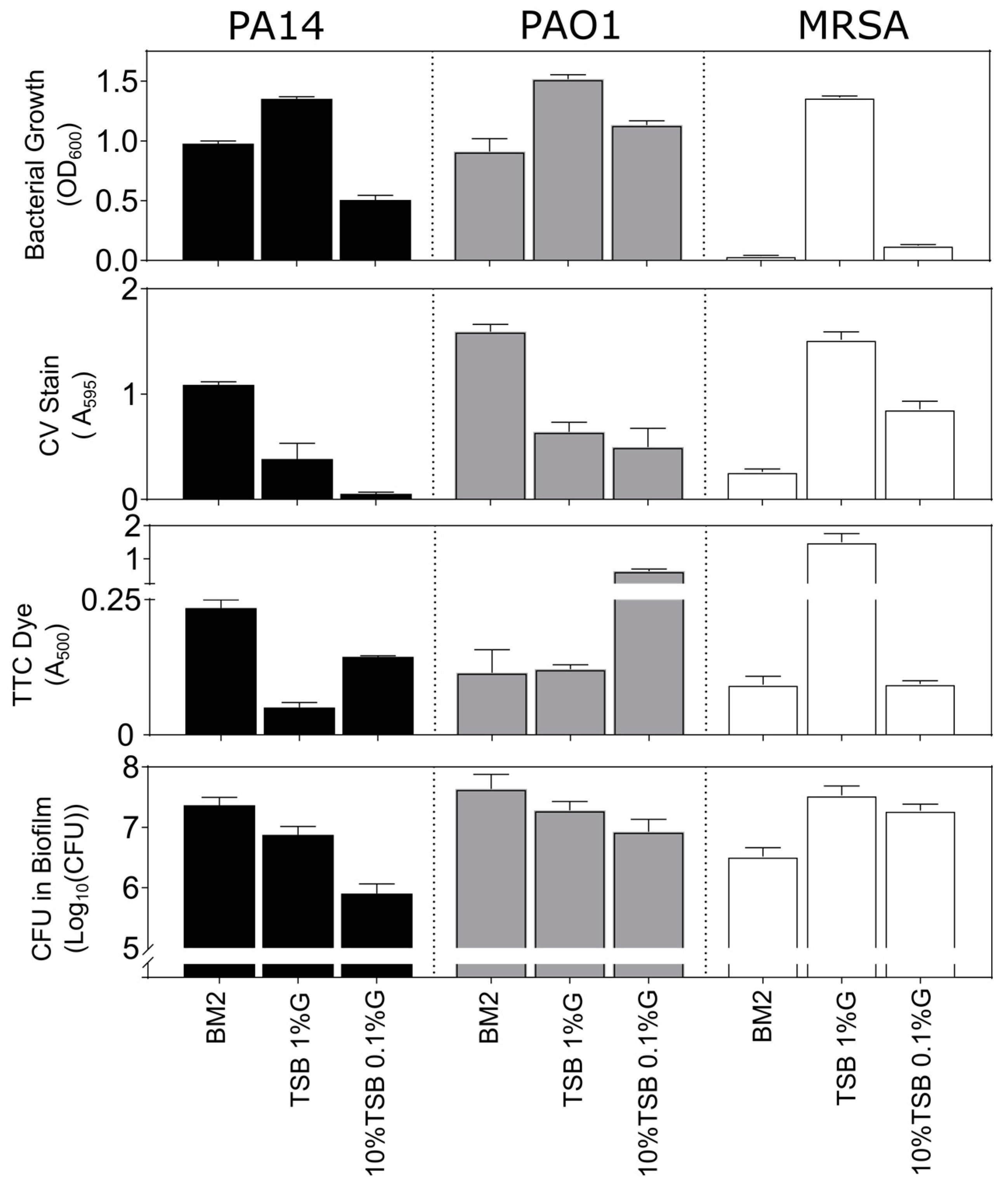
3.2. Biofilm Growth in Microtitre Plates
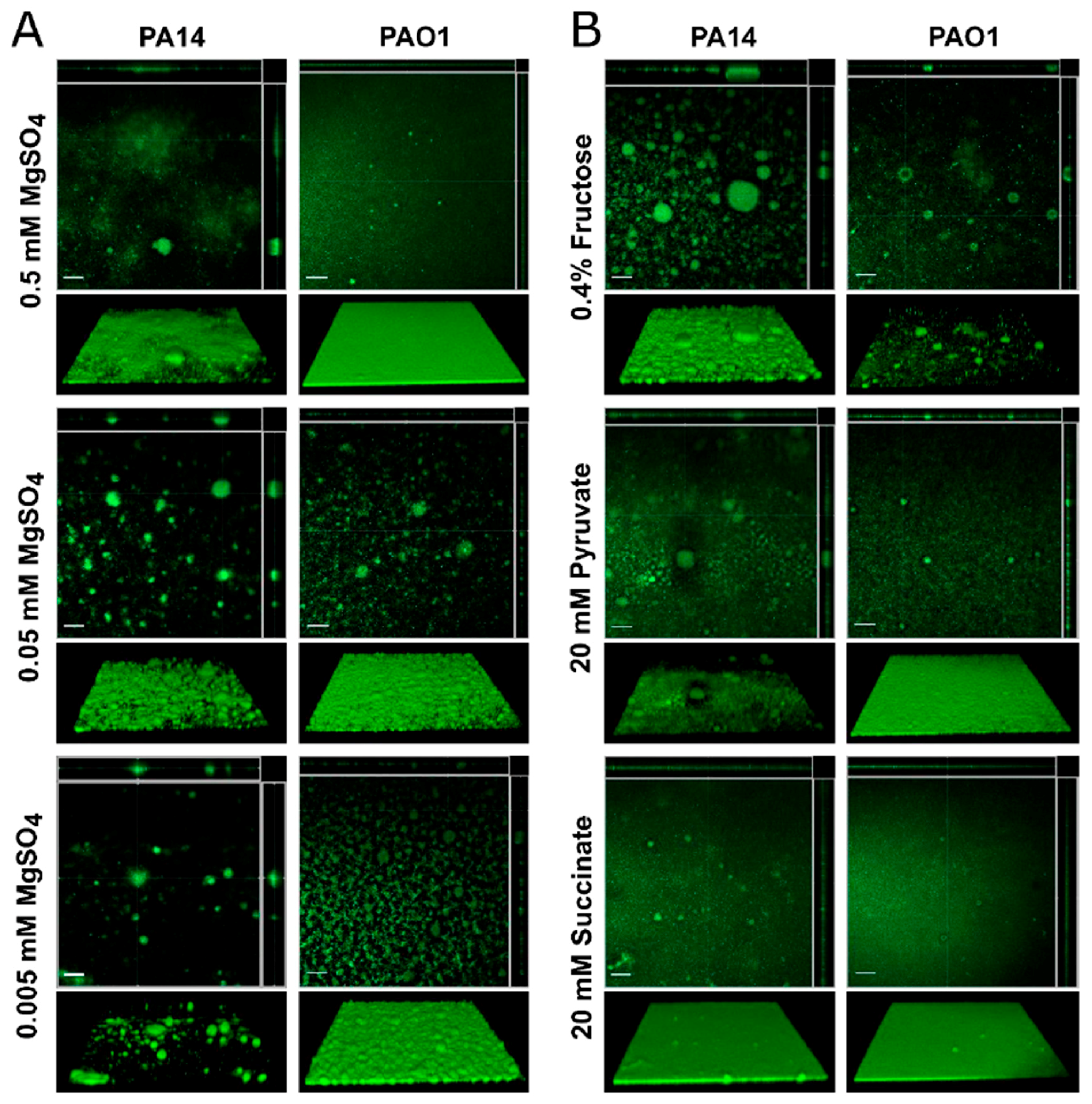
3.3. Effect of Media on Pseudomonas aeruginosa Biofilms Grown under Flow Conditions
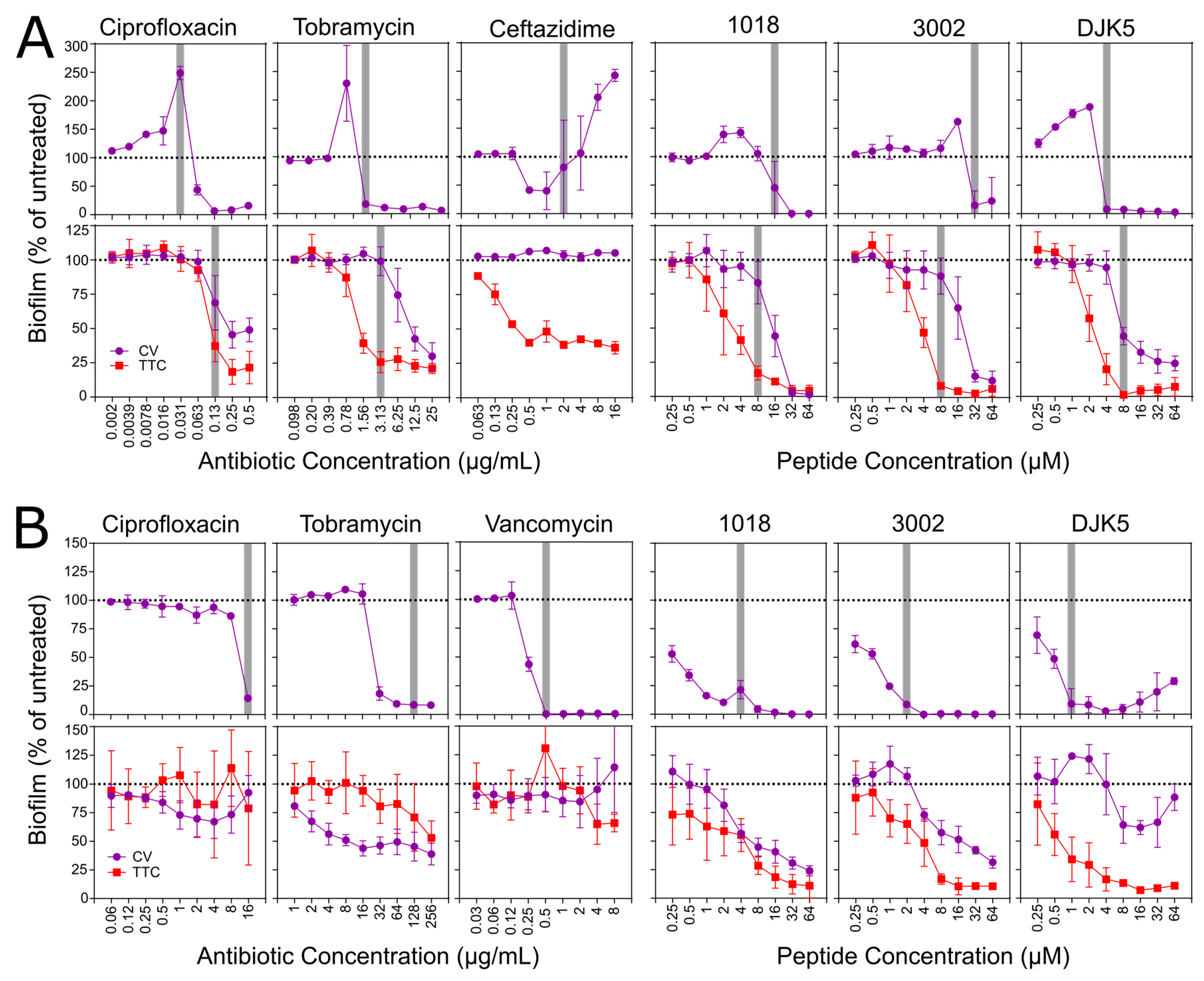
3.4. Biofilm Inhibition and Eradication Assays in Microtitre Plates
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hall, C.W.; Mah, T.-F. Molecular mechanisms of biofilm-based antibiotic resistance and tolerance in pathogenic bacteria. FEMS Microbiol. Rev. 2017, 41, 276–301. [Google Scholar] [CrossRef] [PubMed]
- Neu, L.; Bänziger, C.; Proctor, C.R.; Zhang, Y.; Liu, W.-T.; Hammes, F. Ugly ducklings—The dark side of plastic materials in contact with potable water. NPJ Biofilms Microbiomes 2018, 4, 7. [Google Scholar] [CrossRef] [PubMed]
- Cardinale, M.; Kaiser, D.; Lueders, T.; Schnell, S.; Egert, M. Microbiome analysis and confocal microscopy of used kitchen sponges reveal massive colonization by Acinetobacter, Moraxella and Chryseobacterium species. Sci. Rep. 2017, 7, 5791. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Costerton, J.W.; Stoodley, P. Bacterial biofilms: From the Natural environment to infectious diseases. Nat. Rev. Microbiol. 2004, 2, 95–108. [Google Scholar] [CrossRef] [PubMed]
- Jefferson, K.K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 2004, 236, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Costerton, J.W.; Stewart, P.S.; Greenberg, E.P. Bacterial biofilms: A common cause of persistent infections. Science 1999, 284, 1318–1322. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente-Núñez, C.; Reffuveille, F.; Fernández, L.; Hancock, R.E. Bacterial biofilm development as a multicellular adaptation: Antibiotic resistance and new therapeutic strategies. Curr. Opin. Microbiol. 2013, 16, 580–589. [Google Scholar] [CrossRef] [PubMed]
- Høiby, N.; Bjarnsholt, T.; Givskov, M.; Molin, S.; Ciofu, O. Antibiotic resistance of bacterial biofilms. Int. J. Antimicrob. Agents 2010, 35, 322–332. [Google Scholar] [CrossRef] [PubMed]
- Wolcott, R.D.; Rhoads, D.D.; Bennett, M.E.; Wolcott, B.M.; Gogokhia, L.; Costerton, J.W.; Dowd, S.E. Chronic wounds and the medical biofilm paradigm. J. Wound Care 2010, 19, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, X.; Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucleic Acids Res. 2015, 44, D1087–D1093. [Google Scholar] [CrossRef] [PubMed]
- Hilpert, K.; Volkmer-Engert, R.; Walter, T.; Hancock, R.E.W. High-throughput generation of small antibacterial peptides with improved activity. Nat. Biotechnol. 2005, 23, 1008–1012. [Google Scholar] [CrossRef] [PubMed]
- Jenssen, H.; Fjell, C.D.; Cherkasov, A.; Hancock, R.E.W. QSAR modeling and computer-aided design of antimicrobial peptides. J. Pept. Sci. 2008, 14, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Cherkasov, A.; Hilpert, K.; Jenssen, H.; Fjell, C.D.; Waldbrook, M.; Mullaly, S.C.; Volkmer, R.; Hancock, R.E.W. Use of artificial intelligence in the design of small peptide antibiotics effective against a broad spectrum of highly antibiotic-resistant superbugs. ACS Chem. Biol. 2009, 4, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Overhage, J.; Campisano, A.; Bains, M.; Torfs, E.C.; Rehm, B.H.; Hancock, R.E.W. Human host defense peptide LL-37 prevents bacterial biofilm formation. Infect. Immun. 2008, 76, 4176–4182. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente-Núñez, C.; Reffuveille, F.; Haney, E.F.; Straus, S.K.; Hancock, R.E.W. Broad-spectrum anti-biofilm peptide that targets a cellular stress response. PLoS Pathog. 2014, 10, e1004152. [Google Scholar] [CrossRef] [PubMed]
- Haney, E.F.; Brito-Sánchez, Y.; Trimble, M.J.; Mansour, S.C.; Cherkasov, A.; Hancock, R.E.W. Computer-aided discovery of peptides that specifically attack bacterial biofilms. Sci. Rep. 2018, 8, 1871. [Google Scholar] [CrossRef] [PubMed]
- Spencer, J.J.; Pitts, R.E.; Pearson, R.A.; King, L.B. The effects of antimicrobial peptides WAM-1 and LL-37 on multidrug-resistant Acinetobacter baumannii. Pathog. Dis. 2018, 76, fty007. [Google Scholar] [CrossRef] [PubMed]
- de Breij, A.; Riool, M.; Cordfunke, R.A.; Malanovic, N.; de Boer, L.; Koning, R.I.; Ravensbergen, E.; Franken, M.; van der Heijde, T.; Boekema, B.K.; et al. The antimicrobial peptide SAAP-148 combats drug-resistant bacteria and biofilms. Sci. Transl. Med. 2018, 10, eaan4044. [Google Scholar] [CrossRef] [PubMed]
- Almaaytah, A.; Qaoud, M.T.; Khalil Mohammed, G.; Abualhaijaa, A.; Knappe, D.; Hoffmann, R.; Al-Balas, Q. Antimicrobial and antibiofilm activity of UP-5, an ultrashort antimicrobial peptide designed using only arginine and biphenylalanine. Pharmaceuticals 2018, 11, 3. [Google Scholar] [CrossRef] [PubMed]
- Haney, E.F.; Mansour, S.C.; Hilchie, A.L.; de la Fuente-Núñez, C.; Hancock, R.E.W. High throughput screening methods for assessing antibiofilm and immunomodulatory activities of synthetic peptides. Peptides 2015, 71, 276–285. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente-Núñez, C.; Korolik, V.; Bains, M.; Nguyen, U.; Breidenstein, E.B.; Horsman, S.; Lewenza, S.; Burrows, L.; Hancock, R.E.W. Inhibition of bacterial biofilm formation and swarming motility by a small synthetic cationic peptide. Antimicrob. Agents Chemother. 2012, 56, 2696–2704. [Google Scholar] [CrossRef] [PubMed]
- Wiegand, I.; Hilpert, K.; Hancock, R.E. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 2008, 3, 163–175. [Google Scholar] [CrossRef] [PubMed]
- Moskowitz, S.M.; Foster, J.M.; Emerson, J.; Burns, J.L. Clinically feasible biofilm susceptibility assay for isolates of Pseudomonas aeruginosa from patients with cystic fibrosis. J. Clin. Microbiol. 2004, 42, 1915–1922. [Google Scholar] [CrossRef] [PubMed]
- Mah, T.-F. Establishing the minimal bactericidal concentration of an antimicrobial agent for planktonic cells (MBC-P) and biofilm cells (MBC-B). J. Vis. Exp. 2014. [Google Scholar] [CrossRef] [PubMed]
- Macia, M.D.; Rojo-Molinero, E.; Oliver, A. Antimicrobial susceptibility testing in biofilm-growing bacteria. Clin. Microbiol. Infect. 2014, 20, 981–990. [Google Scholar] [CrossRef] [PubMed]
- Olson, M.E.; Ceri, H.; Morck, D.W.; Buret, A.G.; Read, R.R. Biofilm bacteria: Formation and comparative susceptibility to antibiotics. Can. J. Vet. Res. 2002, 66, 86–92. [Google Scholar] [PubMed]
- Azeredo, J.; Azevedo, N.F.; Briandet, R.; Cerca, N.; Coenye, T.; Costa, A.R.; Desvaux, M.; Di Bonaventura, G.; Hébraud, M.; Jaglic, Z.; et al. Critical review on biofilm methods. Crit. Rev. Microbiol. 2017, 43, 313–351. [Google Scholar] [CrossRef] [PubMed]
- Goeres, D.M.; Loetterle, L.R.; Hamilton, M.A.; Murga, R.; Kirby, D.W.; Donlan, R.M. Statistical assessment of a laboratory method for growing biofilms. Microbiology 2005, 151, 757–762. [Google Scholar] [CrossRef] [PubMed]
- Coenye, T.; Prijck, K.D.; Wever, B.D.; Nelis, H.J. Use of the modified Robbins device to study the in vitro biofilm removal efficacy of NitrAdineTM, a novel disinfecting formula for the maintenance of oral medical devices. J. Appl. Microbiol. 2008, 105, 733–740. [Google Scholar] [CrossRef] [PubMed]
- Nickel, J.C.; Wright, J.B.; Ruseska, I.; Marrie, T.J.; Whitfield, C.; Costerton, J.W. Antibiotic resistance of Pseudomonas aeruginosa colonizing a urinary catheter in vitro. Eur. J. Clin. Microbiol. 1985, 4, 213–218. [Google Scholar] [CrossRef] [PubMed]
- Millar, M.R.; Linton, C.J.; Sherriff, A. Use of a continuous culture system linked to a modified Robbins device or flow cell to study attachment of bacteria to surfaces. In Methods in Enzymology; Doyle, R.J., Ed.; Microbial Growth in Biofilms—Part B: Special Environments and Physicochemical Aspects; Academic Press: Cambridge, MA, USA, 2001; Volume 337, pp. 43–62. [Google Scholar]
- Ceri, H.; Olson, M.E.; Stremick, C.; Read, R.R.; Morck, D.; Buret, A. The Calgary Biofilm Device: New technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J. Clin. Microbiol. 1999, 37, 1771–1776. [Google Scholar] [PubMed]
- Chavant, P.; Gaillard-Martinie, B.; Talon, R.; Hébraud, M.; Bernardi, T. A new device for rapid evaluation of biofilm formation potential by bacteria. J. Microbiol. Methods 2007, 68, 605–612. [Google Scholar] [CrossRef] [PubMed]
- Junka, A.F.; Janczura, A.; Smutnicka, D.; Mączyńska, B.; Anna, S.; Nowicka, J.; Bartoszewicz, M.; Gościniak, G. Use of the real time xCelligence system for purposes of medical microbiology. Pol. J. Microbiol. 2012, 61, 191–197. [Google Scholar] [PubMed]
- Ommen, P.; Zobek, N.; Meyer, R.L. Quantification of biofilm biomass by staining: Non-toxic safranin can replace the popular crystal violet. J. Microbiol. Methods 2017, 141, 87–89. [Google Scholar] [CrossRef] [PubMed]
- Brown, H.L.; van Vliet, A.H.M.; Betts, R.P.; Reuter, M. Tetrazolium reduction allows assessment of biofilm formation by Campylobacter jejuni in a food matrix model. J. Appl. Microbiol. 2013, 115, 1212–1221. [Google Scholar] [CrossRef] [PubMed]
- Sabaeifard, P.; Abdi-Ali, A.; Soudi, M.R.; Dinarvand, R. Optimization of tetrazolium salt assay for Pseudomonas aeruginosa biofilm using microtiter plate method. J. Microbiol. Methods 2014, 105, 134–140. [Google Scholar] [CrossRef] [PubMed]
- Dalecki, A.G.; Crawford, C.L.; Wolschendorf, F. Targeting biofilm associated Staphylococcus aureus using resazurin based drug-susceptibility assay. J. Vis. Exp. 2016. [Google Scholar] [CrossRef] [PubMed]
- Mariscal, A.; Lopez-Gigosos, R.M.; Carnero-Varo, M.; Fernandez-Crehuet, J. Fluorescent assay based on resazurin for detection of activity of disinfectants against bacterial biofilm. Appl. Microbiol. Biotechnol. 2009, 82, 773–783. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A.; Kolter, R. Initiation of biofilm formation in Pseudomonas fluorescens WCS365 proceeds via multiple, convergent signalling pathways: A genetic analysis. Mol. Microbiol. 1998, 28, 449–461. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, G.A. Microtiter dish biofilm formation assay. J. Vis. Exp. 2011, 47, e2437. [Google Scholar] [CrossRef] [PubMed]
- Beveridge, T.J. Use of the gram stain in microbiology. Biotech. Histochem. 2001, 76, 111–118. [Google Scholar] [CrossRef] [PubMed]
- Kwasny, S.M.; Opperman, T.J. Static biofilm cultures of Gram-positive pathogens grown in a microtiter format used for anti-biofilm drug discovery. Curr. Protoc. Pharmacol. 2010, 50. [Google Scholar] [CrossRef]
- Cold Spring Harbor Protocols. M9 recipe. Cold Spring Harb. Protoc. 2006, 2006. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, H.; Lewenza, S. Magnesium limitation is an environmental trigger of the Pseudomonas aeruginosa biofilm lifestyle. PLoS ONE 2011, 6, e23307. [Google Scholar] [CrossRef] [PubMed]
- Song, B.; Leff, L.G. Influence of magnesium ions on biofilm formation by Pseudomonas fluorescens. Microbiol. Res. 2006, 161, 355–361. [Google Scholar] [CrossRef] [PubMed]
- Hoffman, L.R.; D’Argenio, D.A.; MacCoss, M.J.; Zhang, Z.; Jones, R.A.; Miller, S.I. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 2005, 436, 1171–1175. [Google Scholar] [CrossRef] [PubMed]
- Junker, L.M.; Clardy, J. High-throughput screens for small-molecule inhibitors of Pseudomonas aeruginosa biofilm development. Antimicrob. Agents Chemother. 2007, 51, 3582–3590. [Google Scholar] [CrossRef] [PubMed]
- Benoit, M.R.; Conant, C.G.; Ionescu-Zanetti, C.; Schwartz, M.; Matin, A. New device for high-throughput viability screening of flow biofilms. Appl. Environ. Microbiol. 2010, 76, 4136–4142. [Google Scholar] [CrossRef] [PubMed]
- Crémet, L.; Corvec, S.; Batard, E.; Auger, M.; Lopez, I.; Pagniez, F.; Dauvergne, S.; Caroff, N. Comparison of three methods to study biofilm formation by clinical strains of Escherichia coli. Diagn. Microbiol. Infect. Dis. 2013, 75, 252–255. [Google Scholar] [CrossRef] [PubMed]
- Weerasekera, M.M.; Wijesinghe, G.K.; Jayarathna, T.A.; Gunasekara, C.P.; Fernando, N.; Kottegoda, N.; Samaranayake, L.P. Culture media profoundly affect Candida albicans and Candida tropicalis growth, adhesion and biofilm development. Mem. Inst. Oswaldo Cruz 2016, 111, 697–702. [Google Scholar] [CrossRef] [PubMed]
- Olivares, E.; Badel-Berchoux, S.; Provot, C.; Jaulhac, B.; Prévost, G.; Bernardi, T.; Jehl, F. The BioFilm Ring test: A rapid method for routine analysis of Pseudomonas aeruginosa biofilm formation kinetics. J. Clin. Microbiol. 2016, 54, 657–661. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, D.; Hidalgo-Cantabrana, C.; Rodríguez, A.; García, P.; Ruas-Madiedo, P. Monitoring in real time the formation and removal of biofilms from clinical related pathogens using an impedance-based technology. PLoS ONE 2016, 11, e0163966. [Google Scholar] [CrossRef] [PubMed]
- Harrison, J.J.; Stremick, C.A.; Turner, R.J.; Allan, N.D.; Olson, M.E.; Ceri, H. Microtiter susceptibility testing of microbes growing on peg lids: A miniaturized biofilm model for high-throughput screening. Nat. Protoc. 2010, 5, 1236–1254. [Google Scholar] [CrossRef] [PubMed]
- Stepanović, S.; Vuković, D.; Dakić, I.; Savić, B.; Švabić-Vlahović, M. A modified microtiter-plate test for quantification of staphylococcal biofilm formation. J. Microbiol. Methods 2000, 40, 175–179. [Google Scholar] [CrossRef]
- Terraf, M.C.L.; Juárez Tomás, M.S.; Nader-Macías, M.E.F.; Silva, C. Screening of biofilm formation by beneficial vaginal lactobacilli and influence of culture media components. J. Appl. Microbiol. 2012, 113, 1517–1529. [Google Scholar] [CrossRef] [PubMed]
- Combrouse, T.; Sadovskaya, I.; Faille, C.; Kol, O.; Guérardel, Y.; Midelet-Bourdin, G. Quantification of the extracellular matrix of the Listeria monocytogenes biofilms of different phylogenic lineages with optimization of culture conditions. J. Appl. Microbiol. 2013, 114, 1120–1131. [Google Scholar] [CrossRef] [PubMed]
- Adamus-Białek, W.; Kubiak, A.; Czerwonka, G. Analysis of uropathogenic Escherichia coli biofilm formation under different growth conditions. Acta Biochim. Pol. 2015, 62, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Waldrop, R.; McLaren, A.; Calara, F.; McLemore, R. Biofilm growth has a threshold response to glucose in vitro. Clin. Orthop. 2014, 472, 3305–3310. [Google Scholar] [CrossRef] [PubMed]
- Extremina, C.I.; Costa, L.; Aguiar, A.I.; Peixe, L.; Fonseca, A.P. Optimization of processing conditions for the quantification of enterococci biofilms using microtitre-plates. J. Microbiol. Methods 2011, 84, 167–173. [Google Scholar] [CrossRef] [PubMed]
- Peeters, E.; Nelis, H.J.; Coenye, T. Comparison of multiple methods for quantification of microbial biofilms grown in microtiter plates. J. Microbiol. Methods 2008, 72, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Hassan, A.; Usman, J.; Kaleem, F.; Omair, M.; Khalid, A.; Iqbal, M. Evaluation of different detection methods of biofilm formation in the clinical isolates. Braz. J. Infect. Dis. 2011, 15, 305–311. [Google Scholar] [CrossRef]
- Knobloch, J.K.-M.; Horstkotte, M.A.; Rohde, H.; Mack, D. Evaluation of different detection methods of biofilm formation in Staphylococcus aureus. Med. Microbiol. Immunol. 2002, 191, 101–106. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Haney, E.F.; Trimble, M.J.; Cheng, J.T.; Vallé, Q.; Hancock, R.E.W. Critical Assessment of Methods to Quantify Biofilm Growth and Evaluate Antibiofilm Activity of Host Defence Peptides. Biomolecules 2018, 8, 29. https://doi.org/10.3390/biom8020029
Haney EF, Trimble MJ, Cheng JT, Vallé Q, Hancock REW. Critical Assessment of Methods to Quantify Biofilm Growth and Evaluate Antibiofilm Activity of Host Defence Peptides. Biomolecules. 2018; 8(2):29. https://doi.org/10.3390/biom8020029
Chicago/Turabian StyleHaney, Evan F., Michael J. Trimble, John T. Cheng, Quentin Vallé, and Robert E.W. Hancock. 2018. "Critical Assessment of Methods to Quantify Biofilm Growth and Evaluate Antibiofilm Activity of Host Defence Peptides" Biomolecules 8, no. 2: 29. https://doi.org/10.3390/biom8020029
APA StyleHaney, E. F., Trimble, M. J., Cheng, J. T., Vallé, Q., & Hancock, R. E. W. (2018). Critical Assessment of Methods to Quantify Biofilm Growth and Evaluate Antibiofilm Activity of Host Defence Peptides. Biomolecules, 8(2), 29. https://doi.org/10.3390/biom8020029




