Pre-mRNA Splicing in Plants: In Vivo Functions of RNA-Binding Proteins Implicated in the Splicing Process
Abstract
:1. Introduction
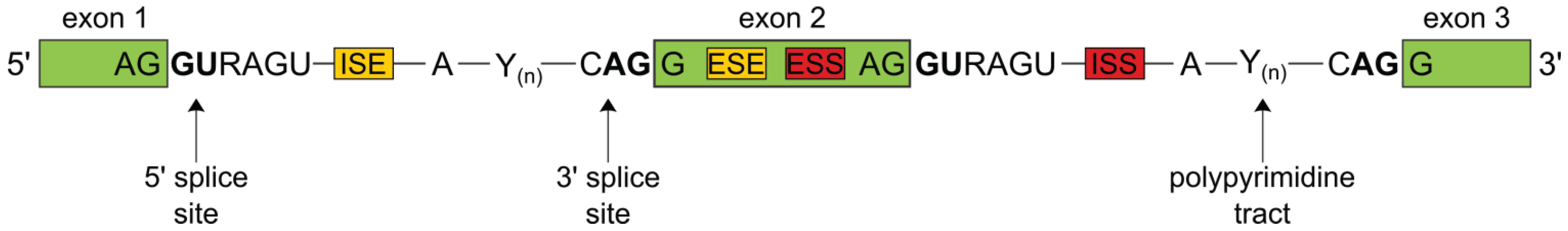
1.1. The Splicing Process
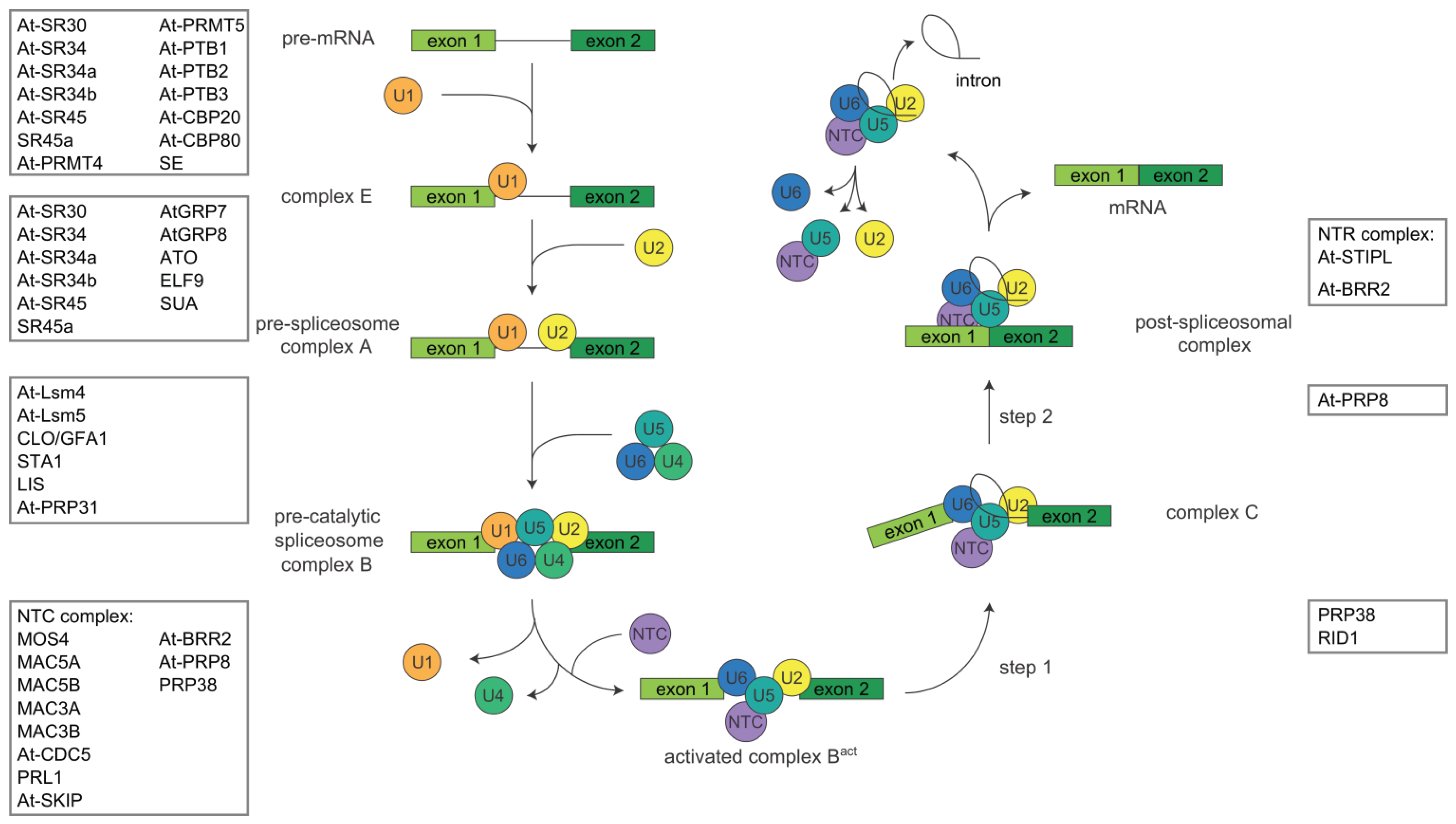
1.2. Alternative Splicing
1.3. Regulation of Splicing
1.4. Pre-mRNA Splicing in Plants
2. Proteins Involved in Pre-mRNA Splicing in Arabidopsis thaliana
2.1. SR Proteins
2.2. hnRNPs
2.2.1. Polypyrimidine Tract Binding Proteins
2.2.2. Glycine-Rich RNA Binding Proteins
2.3. Spliceosomal Proteins
2.3.1. General snRNP Proteins
2.3.2. U snRNP Specific Proteins
2.3.3. The Nineteen Complex
2.3.4. The Nineteen Related Complex
2.4. RNA Export Factors
2.5. Miscellaneous Other RNA Binding Proteins
3. Factors with a Dual Role in Splicing of Pre-mRNAs and microRNA Precursors
4. Conclusions/Outlook
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Berget, S.M.; Moore, C.; Sharp, P.A. Spliced segments at the 5' terminus of adenovirus 2 late mRNA. Proc. Natl. Acad. Sci. USA 1977, 74, 3171–3175. [Google Scholar] [CrossRef] [PubMed]
- Chow, L.T.; Gelinas, R.E.; Broker, T.R.; Roberts, R.J. An amazing sequence arrangement at the 5' ends of adenovirus 2 messenger RNA. Cell 1977, 12, 1–8. [Google Scholar] [CrossRef]
- Wahl, M.C.; Will, C.L.; Lührmann, R. The spliceosome: Design principles of a dynamic RNP machine. Cell 2009, 136, 701–718. [Google Scholar] [CrossRef] [PubMed]
- Will, C.L.; Lührmann, R. Spliceosome Structure and Function. Cold Spring Harbor Perspect. Biol. 2010. [Google Scholar] [CrossRef] [PubMed]
- Matera, A.G.; Wang, Z. A day in the life of the spliceosome. Nat. Rev. Mol. Cell Biol. 2014, 15, 108–121. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Moore, M.J. Spliceosomes. Curr. Biol. 2015, 25, R181–R183. [Google Scholar] [CrossRef] [PubMed]
- Tharun, S. Roles of eukaryotic LSM proteins in the regulation of mRNA function. Int. Rev. Cell. Mol. Biol. 2009, 272, 149–189. [Google Scholar] [PubMed]
- Cordin, O.; Beggs, J.D. RNA helicases in splicing. RNA Biol. 2013, 10, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Blencowe, B.J. Alternative splicing: New insights from global analyses. Cell 2006, 126, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.W.J.; Valcarel, J. Alternative pre-mRNA splicing: The logic of combinatorial control. Trends Biochem. Sci. 2000, 25, 381–388. [Google Scholar] [CrossRef]
- Nilsen, T.W.; Graveley, B.R. Expansion of the eukaryotic proteome by alternative splicing. Nature 2010, 463, 457–463. [Google Scholar] [CrossRef] [PubMed]
- McGlincy, N.J.; Smith, C.W. Alternative splicing resulting in nonsense-mediated mRNA decay: What is the meaning of nonsense? Trends Biochem. Sci. 2008, 33, 385–393. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Kan, J.L.; Green, M.R. Arginine-serine-rich domains bound at splicing enhancers contact the branchpoint to promote prespliceosome assembly. Mol. Cell 2004, 13, 367–376. [Google Scholar] [CrossRef]
- Han, S.P.; Tang, Y.H.; Smith, R. Functional diversity of the hnRNPs: Past, present and perspectives. Biochem. J. 2010, 430, 379–392. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Xiao, X.; Zhang, J.; Choudhury, R.; Robertson, A.; Li, K.; Ma, M.; Burge, C.B.; Wang, Z. A complex network of factors with overlapping affinities repress splicing through intronic elements. Nat. Struct. Mol. Biol. 2013, 20, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Ule, J.; Stefani, G.; Mele, A.; Ruggiu, M.; Wang, X.; Taneri, B.; Gaasterland, T.; Blencowe, B.J.; Darnell, R.B. An RNA map predicting NOVA-dependent splicing regulation. Nature 2006, 444, 580–586. [Google Scholar] [CrossRef] [PubMed]
- Tazi, J.; Bakkour, N.; Stamm, S. Alternative splicing and disease. Biochim. Biophys. Acta Mol. Basis Dis. 2009, 1792, 14–26. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.M.; Zack, D.J. Alternative splicing and retinal degeneration. Clin. Genet. 2013, 84, 142–149. [Google Scholar] [CrossRef] [PubMed]
- Ladomery, M. Aberrant alternative splicing is another hallmark of cancer. Int. J. Cell Biol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S. Alternative splicing of pre-messenger RNAs in plants in the genomic Era. Ann. Rev. Plant Biol. 2007, 58, 267–294. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S.N.; Marquez, Y.; Kalyna, M.; Barta, A. Complexity of the alternative splicing landscape in plants. Plant Cell 2013, 25, 3657–3683. [Google Scholar] [CrossRef] [PubMed]
- Filichkin, S.A.; Priest, H.D.; Givan, S.A.; Shen, R.; Bryant, D.W.; Fox, S.E.; Wong, W.K.; Mockler, T.C. Genome-wide mapping of alternative splicing in Arabidopsis thaliana. Genome Res. 2010, 20, 45–58. [Google Scholar] [CrossRef] [PubMed]
- Syed, N.H.; Kalyna, M.; Marquez, Y.; Barta, A.; Brown, J.W.S. Alternative splicing in plants-coming of age. Trends Plant Sci. 2012, 17, 616–623. [Google Scholar] [CrossRef] [PubMed]
- Staiger, D.; Brown, J.W.S. Alternative splicing at the intersection of biological timing, development, and stress responses. Plant Cell 2013, 25, 3640–3656. [Google Scholar] [CrossRef] [PubMed]
- Staiger, D.; Korneli, C.; Lummer, M.; Navarro, L. Emerging role for RNA-based regulation in plant immunity. New Phytol. 2013, 197, 394–404. [Google Scholar] [CrossRef] [PubMed]
- Loraine, A.; McCormick, S.; Estrada, A.; Patel, K.; Qin, P. High-throughput sequencing of Arabidopsis thaliana pollen cDNA uncovers novel transcription and alternative splicing. Plant Physiol. 2013, 162, 1092–1109. [Google Scholar] [CrossRef] [PubMed]
- Staiger, D. Shaping the Arabidopsis transcriptome through alternative splicing. Adv. Bot. 2015. [Google Scholar] [CrossRef]
- Marquez, Y.; Brown, J.W.S.; Simpson, C.G.; Barta, A.; Kalyna, M. Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis. Genome Res. 2012, 22, 1184–1195. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.W.S.; Feix, G.; Frendewey, D. Accurate in vitro splicing of two pre-mRNA plant introns in a HeLa cell nuclear extract. Embo J. 1986, 5, 2749–2758. [Google Scholar] [PubMed]
- Barta, A.; Sommergruber, K.; Thompson, D.; Hartmuth, K.; Matzke, M.; Matzke, A.M. The expression of a nopaline synthase—Human growth hormone chimaeric gene in transformed tobacco and sunflower callus tissue. Plant Mol. Biol. 1986, 6, 347–357. [Google Scholar] [CrossRef] [PubMed]
- Wiebauer, K.; Herrero, J.J.; Filipowicz, W. Nuclear pre-mRNA processing in plants: Distinct modes of 3'-splice-site selection in plants and animals. Mol. Cell. Biol. 1988, 8, 2042–2051. [Google Scholar] [PubMed]
- Iwata, H.; Gotoh, O. Comparative analysis of information contents relevant to recognition of introns in many species. BMC Genomics 2011. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.B.; Brendel, V. The ASRG database: Identification and survey of Arabidopsis thaliana genes involved in pre-mRNA splicing. Genome Biol. 2004. [Google Scholar] [CrossRef]
- Koncz, C.; deJong, F.; Villacorta, N.; Szakonyi, D.; Koncz, Z. The spliceosome-activating complex: Molecular mechanisms underlying the function of a pleiotropic regulator. Front. Plant Sci. 2012. [Google Scholar] [CrossRef] [PubMed]
- Reddy, A.S. Plant serine/arginine-rich proteins and their role in pre-mRNA splicing. Trends Plant Sci. 2004, 9, 541–547. [Google Scholar] [CrossRef] [PubMed]
- Lopato, S.; Mayeda, A.; Krainer, A.R.; Barta, A. Pre-mRNA splicing in plants: Characterization of Ser/Arg splicing factors. Proc. Natl. Acad. Sci. USA 1996, 93, 3074–3079. [Google Scholar] [CrossRef] [PubMed]
- Lazar, G.; Schaal, T.; Maniatis, T.; Goodman, H.M. Identification of a plant serine-arginine-rich protein similar to the mammalian splicing factor SF2/ASF. Proc. Natl. Acad. Sci. USA 1995, 92, 7672–7676. [Google Scholar] [CrossRef] [PubMed]
- Barta, A.; Kalyna, M.; Reddy, A.S. Implementing a rational and consistent nomenclature for Serine/Arginine-rich protein splicing factors (SR Proteins) in plants. Plant Cell 2010, 22, 2926–2929. [Google Scholar] [CrossRef] [PubMed]
- Duque, P. A role for SR proteins in plant stress responses. Plant Signal. Behav. 2011, 6, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Richardson, D.N.; Rogers, M.F.; Labadorf, A.; Ben-Hur, A.; Guo, H.; Paterson, A.H.; Reddy, A.S. Comparative analysis of Serine/Arginine-rich proteins across 27 eukaryotes: Insights into sub-family classification and extent of alternative splicing. PLoS ONE 2011, 6, e24542. [Google Scholar] [CrossRef] [PubMed]
- Kalyna, M.; Lopato, S.; Voronin, V.; Barta, A. Evolutionary conservation and regulation of particular alternative splicing events in plant SR proteins. Nucleic Acids Res. 2006, 34, 4395–4405. [Google Scholar] [CrossRef] [PubMed]
- Lopato, S.; Forstner, C.; Kalyna, M.; Hilscher, J.; Langhammer, U.; Indrapichate, K.; Lorkovic, Z.J.; Barta, A. Network of interactions of a novel plant-specific Arg/Ser-rich protein, atRSZ33, with atSC35-like splicing factors. J. Biol. Chem. 2002, 277, 39989–39998. [Google Scholar] [CrossRef] [PubMed]
- Kalyna, M.; Lopato, S.; Barta, A. Ectopic expression of atRSZ33 reveals its function in splicing and causes pleiotropic changes in development. Mol. Biol. Cell 2003, 14, 3565–3577. [Google Scholar] [CrossRef] [PubMed]
- Lopato, S.; Kalyna, M.; Dorner, S.; Kobayashi, R.; Krainer, A.R.; Barta, A. atSRp30, one of two SF2/ASF-like proteins from Arabidopsis thaliana, regulates splicing of specific plant genes. Genes Dev. 1999, 13, 987–1001. [Google Scholar] [CrossRef] [PubMed]
- Petrillo, E.; Godoy Herz, M.A.; Fuchs, A.; Reifer, D.; Fuller, J.; Yanovsky, M.J.; Simpson, C.; Brown, J.W.S.; Barta, A.; Kalyna, M.; et al. A Chloroplast Retrograde Signal Regulates Nuclear Alternative Splicing. Science 2014, 344, 427–430. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Du, B.; Liu, D.; Qi, X. Splicing factor SR34b mutation reduces cadmium tolerance in Arabidopsis by regulating iron-regulated transporter 1 gene. Biochem. Biophys. Res. Commun. 2014, 455, 312–317. [Google Scholar] [CrossRef] [PubMed]
- Lazar, G.; Goodman, H. The Arabidopsis splicing factor SR1 is regulated by alternative splicing. Plant Mol. Biol. 2000, 42, 571–581. [Google Scholar] [CrossRef] [PubMed]
- Golovkin, M.; Reddy, A.S. The plant U1 small nuclear ribonucleoprotein particle 70K protein interacts with two novel Serine/Arginine-rich proteins. Plant Cell 1998, 10, 1637–1648. [Google Scholar] [PubMed]
- Lopato, S.; Waigmann, E.; Barta, A. Characterization of a novel arginine/serine-rich splicing factor in Arabidopsis. Plant Cell 1996, 8, 2255–2264. [Google Scholar] [CrossRef] [PubMed]
- Palusa, S.G.; Reddy, A.S. Extensive coupling of alternative splicing of pre-mRNAs of Serine/Arginine (SR) genes with nonsense-mediated decay. New Phytol. 2010, 185, 83–89. [Google Scholar] [CrossRef] [PubMed]
- Kalyna, M.; Simpson, C.G.; Syed, N.H.; Lewandowska, D.; Marquez, Y.; Kusenda, B.; Marshall, J.; Fuller, J.; Cardle, L.; McNicol, J.; et al. Alternative splicing and nonsense-mediated decay modulate expression of important regulatory genes in Arabidopsis. Nucleic Acids Res. 2012, 40, 2454–2469. [Google Scholar] [CrossRef] [PubMed]
- Iida, K.; Go, M. Survey of conserved alternative splicing events of mRNAs encoding SR Proteins in land plants. Mol. Biol. Evol. 2006, 23, 1085–1094. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.; Palusa, S.G.; Prasad, K.V.S.K.; Ali, G.S.; Surabhi, G.-K.; Ben-Hur, A.; Abdel-Ghany, S.E.; Reddy, A.S.N. Identification of an intronic splicing regulatory element involved in autoregulation of alternative splicing of the SCL33 pre-mRNA. Plant J. 2012, 72, 935–946. [Google Scholar] [PubMed]
- Lykke-Andersen, J.; Shu, M.D.; Steitz, J.A. Communication of the position of exon-exon junctions to the mRNA surveillance machinery by the protein RNPS1. Science 2001, 293, 1836–1839. [Google Scholar] [CrossRef] [PubMed]
- Ali, G.S.; Palusa, S.G.; Golovkin, M.; Prasad, J.; Manley, J.L.; Reddy, A.S.N. Regulation of plant developmental processes by a novel splicing factor. PLoS ONE 2007, 2, e471. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.N.; Mount, S.M. Two alternatively spliced isoforms of the Arabidopsis thaliana SR45 protein have distinct roles during normal plant development. Plant Physiol. 2009, 150, 1450–1458. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-N.; Mo, C.; Garrett, W.M.; Cooper, B. Phosphothreonine 218 is required for the function of SR45.1 in regulating flower petal development in Arabidopsis. Plant Signal. Behav. 2014. [Google Scholar] [CrossRef]
- Carvalho, R.F.; Carvalho, S.D.; Duque, P. The plant-specific SR45 protein negatively regulates glucose and ABA signaling during early seedling development in Arabidopsis. Plant Physiol. 2010, 154, 772–783. [Google Scholar] [CrossRef] [PubMed]
- Filichkin, S.A.; Cumbie, J.S.; Dharmawadhana, J.P.; Jaiswal, P.; Chang, J.H.; Palusa, S.G.; Reddy, A.S.N.; Megraw, M.; Mockler, T.C. Environmental stresses modulate abundance and timing of alternatively spliced circadian transcripts in Arabidopsis. Mol. Plant 2015, 8, 207–227. [Google Scholar] [CrossRef] [PubMed]
- Day, I.S.; Golovkin, M.; Palusa, S.G.; Link, A.; Ali, G.S.; Thomas, J.; Richardson, D.N.; Reddy, A.S.N. Interactions of SR45, an SR-like protein, with spliceosomal proteins and an intronic sequence: Insights into regulated splicing. Plant J. 2012, 71, 936–947. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente van Bentem, S.; Anrather, D.; Roitinger, E.; Djamei, A.; Hufnagl, T.; Barta, A.; Csaszar, E.; Dohnal, I.; Lecourieux, D.; Hirt, H. Phosphoproteomics reveals extensive in vivo phosphorylation of Arabidopsis proteins involved in RNA metabolism. Nucleic Acids Res. 2006, 34, 3267–3278. [Google Scholar] [CrossRef] [PubMed]
- Feilner, T.; Hultschig, C.; Lee, J.; Meyer, S.; Immink, R.G.; Koenig, A.; Possling, A.; Seitz, H.; Beveridge, A.; Scheel, D.; et al. High throughput identification of potential Arabidopsis mitogen-activated protein kinases substrates. Mol. Cell. Proteomics 2005, 4, 1558–1568. [Google Scholar] [CrossRef] [PubMed]
- Savaldi-Goldstein, S.; Aviv, D.; Davydov, O.; Fluhr, R. Alternative splicing modulation by a LAMMER Kinase impinges on developmental and transcriptome expression. Plant Cell 2003, 15, 926–938. [Google Scholar] [CrossRef] [PubMed]
- Wachter, A.; Rühl, C.; Stauffer, E. The role of polypyrimidine tract-binding proteins and other hnRNP proteins in plant splicing regulation. Front. Plant Sci. 2012. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo, J.M.; Majós, N.; Bonnal, S.; Martínez, C.; Castelo, R.; Guigó, R.; Bilbao, D.; Valcárcel, J. Regulation of fas alternative splicing by antagonistic effects of TIA-1 and PTB on exon definition. Mol. Cell 2005, 19, 475–484. [Google Scholar] [CrossRef] [PubMed]
- Boutz, P.L.; Stoilov, P.; Li, Q.; Lin, C.H.; Chawla, G.; Ostrow, K.; Shiue, L.; Ares, M., Jr.; Black, D.L. A post-transcriptional regulatory switch in polypyrimidine tract-binding proteins reprograms alternative splicing in developing neurons. Genes Dev. 2007, 21, 1636–1652. [Google Scholar] [CrossRef] [PubMed]
- Spellman, R.; Llorian, M.; Smith, C.W. Crossregulation and functional redundancy between the splicing regulator PTB and its paralogs nPTB and ROD1. Mol. Cell 2007, 27, 420–434. [Google Scholar] [CrossRef] [PubMed]
- Stauffer, E.; Westermann, A.; Wagner, G.; Wachter, A. Polypyrimidine tract-binding protein homologues from Arabidopsis underlie regulatory circuits based on alternative splicing and downstream control. Plant J. 2010, 64, 243–255. [Google Scholar] [CrossRef] [PubMed]
- Rühl, C.; Stauffer, E.; Kahles, A.; Wagner, G.; Drechsel, G.; Rätsch, G.; Wachter, A. Polypyrimidine tract binding protein homologs from arabidopsis are key regulators of alternative splicing with implications in fundamental developmental processes. Plant Cell 2012, 24, 4360–4375. [Google Scholar] [CrossRef] [PubMed]
- Simpson, C.G.; Lewandowska, D.; Liney, M.; Davidson, D.; Chapman, S.; Fuller, J.; McNicol, J.; Shaw, P.; Brown, J.W.S. Arabidopsis PTB1 and PTB2 proteins negatively regulate splicing of a mini-exon splicing reporter and affect alternative splicing of endogenous genes differentially. New Phytol. 2014, 203, 424–436. [Google Scholar] [CrossRef] [PubMed]
- Penfield, S.; Josse, E.M.; Halliday, K.J. A role for an alternative splice variant of PIF6 in the control of Arabidopsis primary seed dormancy. Plant Mol. Biol. 2010, 73, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Mangeon, A.; Junqueira, R.M.; Sachetto-Martins, G. Functional diversity of the plant glycine-rich proteins superfamily. Plant Signal. Behav. 2010, 5, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Ciuzan, O.; Hancock, J.; Pamfil, D.; Wilson, I.; Ladomery, M. The evolutionarily conserved multifunctional glycine-rich RNA-binding proteins play key roles in development and stress adaptation. Physiol. Plant. 2015, 153, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, C.D.; Kreps, J.A.; Simon, A.E. Genes encoding glycine-rich Arabidopsis thaliana proteins with RNA-binding motifs are influenced by cold treatment and an endogenous circadian rhythm. Plant Physiol. 1994, 104, 1015–1025. [Google Scholar] [CrossRef] [PubMed]
- Staiger, D.; Heintzen, C. The circadian system of Arabidopsis thaliana: Forward and reverse genetic approaches. Chronobiol. Int. 1999, 16, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Staiger, D.; Zecca, L.; Wieczorek Kirk, D.A.; Apel, K.; Eckstein, L. The circadian clock regulated RNA-binding protein AtGRP7 autoregulates its expression by influencing alternative splicing of its own pre-mRNA. Plant J. 2003, 33, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Schöning, J.C.; Streitner, C.; Meyer, I.M.; Gao, Y.; Staiger, D. Reciprocal regulation of glycine-rich RNA-binding proteins via an interlocked feedback loop coupling alternative splicing to nonsense-mediated decay in Arabidopsis. Nucleic Acids Res. 2008, 36, 6977–6987. [Google Scholar] [CrossRef] [PubMed]
- Simpson, C.G.; Fuller, J.; Maronova, M.; Kalyna, M.; Davidson, D.; McNicol, J.; Barta, A.; Brown, J.W. Monitoring changes in alternative precursor messenger RNA splicing in multiple gene transcripts. Plant J. 2008, 53, 1035–1048. [Google Scholar] [CrossRef] [PubMed]
- Streitner, C.; Köster, T.; Simpson, C.G.; Shaw, P.; Danisman, S.; Brown, J.W.S.; Staiger, D. An hnRNP-like RNA-binding protein affects alternative splicing by in vivo interaction with target transcripts in Arabidopsis thaliana. Nucleic Acids Res. 2012, 40, 11240–11255. [Google Scholar] [CrossRef] [PubMed]
- Köster, T.; Haas, M.; Staiger, D. The RIPper case: Identification of RNA-binding protein targets by RNA immunoprecipitation. Methods Mol. Biol. 2014, 1158, 107–121. [Google Scholar] [PubMed]
- Nicaise, V.; Joe, A.; Jeong, B.; Korneli, C.; Boutrot, F.; Wested, I.; Staiger, D.; Alfano, J.R.; Zipfel, C. Pseudomonas HopU1 affects interaction of plant immune receptor mRNAs to the RNA-binding protein GRP7. EMBO J. 2013, 32, 701–712. [Google Scholar] [CrossRef] [PubMed]
- Hackmann, C.; Korneli, C.; Kutyniok, M.; Köster, T.; Wiedenlübbert, M.; Müller, C.; Staiger, D. Salicylic acid-dependent and -independent impact of an RNA-binding protein on plant immunity. Plant Cell Environ. 2014, 37, 696–706. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.Q.; Guo, M.; Jeong, B.R.; Tian, F.; Elthon, T.E.; Cerny, R.L.; Staiger, D.; Alfano, J.R. A type III effector ADP-ribosylates RNA-binding proteins and quells plant immunity. Nature 2007, 447, 284–288. [Google Scholar] [CrossRef] [PubMed]
- Jeong, B.-R.; Lin, Y.; Joe, A.; Guo, M.; Korneli, C.; Yang, H.; Wang, P.; Yu, M.; Cerny, R.L.; Staiger, D.; et al. Structure function analysis of an ADP-ribosyltransferase type III effector and its RNA-binding target in plant immunity. J. Biol. Chem. 2011, 286, 43272–43281. [Google Scholar] [CrossRef] [PubMed]
- Swaraz, A.M.; Park, Y.D.; Hur, Y. Knock-out mutations of Arabidopsis SmD3-b induce pleotropic phenotypes through altered transcript splicing. Plant Sci. 2011, 180, 661–671. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zhang, S.; Zhang, Y.; Wang, X.; Li, D.; Li, Q.; Yue, M.; Li, Q.; Zhang, Y.E.; Xu, Y.; et al. Arabidopsis floral initiator SKB1 confers high salt tolerance by regulating transcription and pre-mRNA splicing through altering histone H4R3 and small nuclear ribonucleoprotein LSM4 methylation. Plant Cell 2011, 23, 396–411. [Google Scholar] [CrossRef] [PubMed]
- Perez-Santángelo, S.; Mancini, E.; Francey, L.J.; Schlaen, R.G.; Chernomoretz, A.; Hogenesch, J.B.; Yanovsky, M.J. Role for LSM genes in the regulation of circadian rhythms. Proc. Natl. Acad. Sci. USA 2014, 111, 15166–15171. [Google Scholar] [CrossRef] [PubMed]
- Golisz, A.; Sikorski, P.J.; Kruszka, K.; Kufel, J. Arabidopsis thaliana LSM proteins function in mRNA splicing and degradation. Nucleic Acids Res. 2013, 41, 6232–6249. [Google Scholar] [CrossRef] [PubMed]
- Cui, P.; Zhang, S.; Ding, F.; Ali, S.; Xiong, L. Dynamic regulation of genome-wide pre-mRNA splicing and stress tolerance by the Sm-like protein LSm5 in Arabidopsis. Genome Biol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Meister, G.; Eggert, C.; Buhler, D.; Brahms, H.; Kambach, C.; Fischer, U. Methylation of Sm proteins by a complex containing PRMT5 and the putative U snRNP assembly factor pICln. Curr. Biol. 2001, 11, 1990–1994. [Google Scholar] [CrossRef]
- Friesen, W.J.; Paushkin, S.; Wyce, A.; Massenet, S.; Pesiridis, G.S.; van Duyne, G.; Rappsilber, J.; Mann, M.; Dreyfuss, G. The methylosome, a 20S complex containing JBP1 and pICln, produces dimethylarginine-modified Sm proteins. Mol. Cell Biol. 2001, 21, 8289–8300. [Google Scholar] [CrossRef] [PubMed]
- Pei, Y.; Niu, L.; Lu, F.; Liu, C.; Zhai, J.; Kong, X.; Cao, X. Mutations in the Type II protein arginine methyltransferase AtPRMT5 result in pleiotropic developmental defects in Arabidopsis. Plant Physiol. 2007, 144, 1913–1923. [Google Scholar] [CrossRef] [PubMed]
- Hernando, C.E.; Sanchez, S.E.; Mancini, E.; Yanovsky, M.J. Genome wide comparative analysis of the effects of PRMT5 and PRMT4/CARM1 arginine methyltransferases on the Arabidopsis thaliana transcriptome. BMC Genomics 2015. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, S.E.; Petrillo, E.; Beckwith, E.J.; Zhang, X.; Rugnone, M.L.; Hernando, C.E.; Cuevas, J.C.; Godoy Herz, M.A.; Depetris-Chauvin, A.; Simpson, C.G.; et al. A methyl transferase links the circadian clock to the regulation of alternative splicing. Nature 2010, 468, 112–116. [Google Scholar] [CrossRef] [PubMed]
- Hong, S.; Song, H.R.; Lutz, K.; Kerstetter, R.A.; Michael, T.P.; McClung, C.R. Type II protein arginine methyltransferase 5 (PRMT5) is required for circadian period determination in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2010, 107, 21211–21216. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Gu, L.; Liu, C.; Lu, T.; Lu, F.; Lu, Z.; Cui, P.; Pei, Y.; Wang, B.; Hu, S.; et al. Arginine methylation mediated by the Arabidopsis homolog of PRMT5 is essential for proper pre-mRNA splicing. Proc. Natl. Acad. Sci. USA 2010, 107, 19114–19119. [Google Scholar] [CrossRef] [PubMed]
- Cheng, D.; Côté, J.; Shaaban, S.; Bedford, M.T. The arginine methyltransferase CARM1 regulates the coupling of transcription and mRNA processing. Mol. Cell 2007, 25, 71–83. [Google Scholar] [CrossRef] [PubMed]
- Niu, L.; Zhang, Y.; Pei, Y.; Liu, C.; Cao, X. Redundant requirement for a pair of PROTEIN ARGININE METHYLTRANSFERASE4 homologs for the proper regulation of Arabidopsis flowering time. Plant Physiol. 2008, 148, 490–503. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.H.; Kapoor, A.; Zhu, J.; Zhu, J.K. STABILIZED1, a stress-upregulated nuclear protein, is required for pre-mRNA splicing, mRNA turnover, and stress tolerance in Arabidopsis. Plant Cell 2006, 18, 1736–1749. [Google Scholar] [CrossRef] [PubMed]
- Thalhammer, A.; Bryant, G.; Sulpice, R.; Hincha, D.K. Disordered cold regulated15 proteins protect chloroplast membranes during freezing through binding and folding, but do not stabilize chloroplast enzymes in vivo. Plant Physiol. 2014, 166, 190–201. [Google Scholar] [CrossRef] [PubMed]
- Du, J.-L.; Zhang, S.-W.; Huang, H.-W.; Cai, T.; Li, L.; Chen, S.; He, X.-J. The splicing factor PRP31 is involved in transcriptional gene silencing and stress response in Arabidopsis. Mol. Plant 2015, 8, 1053–1068. [Google Scholar] [CrossRef] [PubMed]
- Gross-Hardt, R.; Kägi, C.; Baumann, N.; Moore, J.M.; Baskar, R.; Gagliano, W.B.; Jürgens, G.; Grossniklaus, U. LACHESIS restricts gametic cell fate in the female gametophyte of Arabidopsis. PLoS Biol. 2007, 5, e47. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Yuan, L.; Liu, N.Y.; Shi, D.Q.; Liu, J.; Yang, W.C. GAMETOPHYTIC FACTOR 1, involved in pre-mRNA splicing, is essential for megagametogenesis and embryogenesis in Arabidopsis. J. Integr. Plant Biol. 2009, 51, 261–271. [Google Scholar] [CrossRef] [PubMed]
- Moll, C.; von Lyncker, L.; Zimmermann, S.; Kägi, C.; Baumann, N.; Twell, D.; Grossniklaus, U.; Groß-Hardt, R. CLO/GFA1 and ATO are novel regulators of gametic cell fate in plants. Plant J. 2008, 56, 913–921. [Google Scholar] [CrossRef] [PubMed]
- Hornyik, C.; Duc, C.; Rataj, K.; Terzi, L.C.; Simpson, G.G. Alternative polyadenylation of antisense RNAs and flowering time control. Biochem. Soc. Trans. 2010, 38, 1077–1081. [Google Scholar] [CrossRef] [PubMed]
- Pena, V.; Rozov, A.; Fabrizio, P.; Lührmann, R.; Wahl, M.C. Structure and function of an RNase H domain at the heart of the spliceosome. EMBO J. 2008, 27, 2929–2940. [Google Scholar] [CrossRef] [PubMed]
- Grainger, R.J.; Beggs, J.D. Prp8 protein: At the heart of the spliceosome. RNA 2005, 11, 533–557. [Google Scholar] [CrossRef] [PubMed]
- Marquardt, S.; Raitskin, O.; Wu, Z.; Liu, F.; Sun, Q.; Dean, C. Functional consequences of splicing of the antisense transcript COOLAIR on FLC transcription. Mol. Cell 2014, 54, 156–165. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Tian, Q.; Hou, Z.; Mucha, M.; Aukerman, M.; Olsen, O.A. The Arabidopsis thaliana AT PRP39-1 gene, encoding a tetratricopeptide repeat protein with similarity to the yeast pre-mRNA processing protein PRP39, affects flowering time. Plant Cell Rep. 2007, 26, 1357–1366. [Google Scholar] [CrossRef] [PubMed]
- Song, H.-R.; Song, J.-D.; Cho, J.-N.; Amasino, R.M.; Noh, B.; Noh, Y.-S. The RNA binding protein ELF9 directly reduces SUPPRESSOR OF OVEREXPRESSION OF CO1 transcript levels in Arabidopsis, possibly via nonsense-mediated mRNA decay. Plant Cell 2009, 21, 1195–1211. [Google Scholar] [CrossRef] [PubMed]
- Sugliani, M.; Brambilla, V.; Clerkx, E.J.; Koornneef, M.; Soppe, W.J. The conserved splicing factor SUA controls alternative splicing of the developmental regulator ABI3 in Arabidopsis. Plant Cell 2010, 22, 1936–1946. [Google Scholar] [CrossRef] [PubMed]
- Hogg, R.; McGrail, J.C.; O’Keefe, R.T. The function of the NineTeen Complex (NTC) in regulating spliceosome conformations and fidelity during pre-mRNA splicing. Biochem. Soc. Trans. 2010, 38, 1110–1115. [Google Scholar] [CrossRef] [PubMed]
- Palma, K.; Zhao, Q.; Cheng, Y.T.; Bi, D.; Monaghan, J.; Cheng, W.; Zhang, Y.; Li, X. Regulation of plant innate immunity by three proteins in a complex conserved across the plant and animal kingdoms. Genes Dev. 2007, 21, 1484–1493. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Goritschnig, S.; Dong, X.; Li, X. A gain-of-function mutation in a plant disease resistance gene leads to constitutive activation of downstream signal transduction pathways in suppressor of Npr1-1, constitutive 1. Plant Cell 2003, 15, 2636–2646. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, J.; Xu, F.; Gao, M.; Zhao, Q.; Palma, K.; Long, C.; Chen, S.; Zhang, Y.; Li, X. Two Prp19-like U-box proteins in the MOS4-associated complex play redundant roles in plant innate immunity. PLoS Pathog. 2009, 5, e1000526. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Xu, S.; Wiermer, M.; Zhang, Y.; Li, X. The cyclin L homolog MOS12 and the MOS4-associated complex are required for proper splicing of plant resistance genes. Plant J. 2012, 70, 916–928. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Liu, Y.; Ding, P.; Li, Y.; Kong, Q.; Zhang, Y. Splicing of receptor-like kinase-encoding SNC4 and CERK1 is regulated by two conserved splicing factors that are required for plant immunity. Mol. Plant 2014, 7, 1766–1775. [Google Scholar] [CrossRef] [PubMed]
- Monaghan, J.; Xu, F.; Xu, S.; Zhang, Y.; Li, X. Two putative RNA-binding proteins function with unequal genetic redundancy in the MOS4-associated complex. Plant Physiol. 2010, 154, 1783–1793. [Google Scholar] [CrossRef] [PubMed]
- Rasche, N.; Dybkov, O.; Schmitzova, J.; Akyildiz, B.; Fabrizio, P.; Luhrmann, R. Cwc2 and its human homologue RBM22 promote an active conformation of the spliceosome catalytic centre. EMBO J. 2012, 31, 1591–1604. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wu, F.; Xie, Q.; Wang, H.; Wang, Y.; Yue, Y.; Gahura, O.; Ma, S.; Liu, L.; Cao, Y.; et al. SKIP Is a Component of the Spliceosome Linking Alternative Splicing and the Circadian Clock in Arabidopsis. Plant Cell 2012, 24, 3278–3295. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.; Li, J.; Gao, Z.; Lu, Y.; Yu, J.; Zheng, Q.; Yan, S.; Zhang, W.; He, H.; Ma, L.; et al. SKIP confers osmotic tolerance during salt stress by controlling alternative gene splicing in Arabidopsis. Mol. Plant 2015, 8, 1038–1052. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.A.; Williams, B.A.; McNicol, J.; Simpson, C.G.; Brown, J.W.S.; Harmer, S.L. Mutation of Arabidopsis SPLICEOSOMAL TIMEKEEPER LOCUS1 causes circadian clock defects. Plant Cell 2012, 24, 4907–4916. [Google Scholar] [CrossRef] [PubMed]
- Dias, A.P.; Dufu, K.; Lei, H.; Reed, R. A role for TREX components in the release of spliced mRNA from nuclear speckle domains. Nat. Commun. 2010. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Zhou, X.; Wen, C.-K. HYPER RECOMBINATION1 of the THO/TREX complex plays a role in controlling transcription of the REVERSION-TO-ETHYLENE SENSITIVITY1 gene in Arabidopsis. PLoS Genet. 2015, 11, e1004956. [Google Scholar] [CrossRef] [PubMed]
- Furumizu, C.; Tsukaya, H.; Komeda, Y. Characterization of EMU, the Arabidopsis homolog of the yeast THO complex member HPR1. RNA 2010, 16, 1809–1817. [Google Scholar] [CrossRef] [PubMed]
- Francisco-Mangilet, A.G.; Karlsson, P.; Kim, M.-H.; Eo, H.J.; Oh, S.A.; Kim, J.H.; Kulcheski, F.R.; Park, S.K.; Manavella, P.A. THO2, core member of the THO/TREX complex, is required for micro RNA production in Arabidopsis. Plant J. 2015, 82, 1018–1029. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Cui, P.; Chen, H.; Ali, S.; Zhang, S.; Xiong, L. A KH-domain RNA-binding protein interacts with FIERY2/CTD phosphatase-like 1 and splicing factors and is important for pre-mRNA splicing in Arabidopsis. PLoS Genet. 2013, 9, e1003875. [Google Scholar] [CrossRef] [PubMed]
- Guan, Q.; Wu, J.; Zhang, Y.; Jiang, C.; Liu, R.; Chai, C.; Zhu, J. A DEAD box RNA helicase is critical for pre-mRNA splicing, cold-responsive gene regulation, and cold tolerance in Arabidopsis. Plant Cell 2013, 25, 342–356. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, N.; Yoshimura, K.; Kimura, A.; Yabuta, Y.; Shigeoka, S. Differential expression of alternatively spliced mRNAs of Arabidopsis SR protein homologues, atSR30 and atSR45a, in response to environmental stress. Plant Cell Physiol. 2007, 48, 1036–1049. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, N.; Kimura, A.; Yoshimura, K.; Shigeoka, S. Plant-specific SR-related protein atSR45a interacts with spliceosomal proteins in plant nucleus. Plant Mol. Biol. 2009, 70, 241–252. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, T.; Kanno, T.; Liang, S.-C.; Chen, P.-Y.; Liao, W.-W.; Lin, W.-D.; Matzke, A.J.M.; Matzke, M. An Rtf2 domain-containing protein influences pre-mrna splicing and is essential for embryonic development in Arabidopsis thaliana. Genetics 2015, 200, 523–535. [Google Scholar] [CrossRef] [PubMed]
- Ohtani, M.; Demura, T.; Sugiyama, M. Arabidopsis ROOT INITIATION DEFECTIVE1, a DEAH-box RNA helicase involved in pre-mRNA splicing, is essential for plant development. Plant Cell 2013, 25, 2056–2069. [Google Scholar] [CrossRef] [PubMed]
- Raczynska, K.D.; Simpson, C.G.; Ciesiolka, A.; Szewc, L.; Lewandowska, D.; McNicol, J.; Szweykowska-Kulinska, Z.; Brown, J.W.; Jarmolowski, A. Involvement of the nuclear cap-binding protein complex in alternative splicing in Arabidopsis thaliana. Nucleic Acids Res. 2010, 38, 265–278. [Google Scholar] [CrossRef] [PubMed]
- Laubinger, S.; Sachsenberg, T.; Zeller, G.; Busch, W.; Lohmann, J.U.; Ratsch, G.; Weigel, D. Dual roles of the nuclear cap-binding complex and SERRATE in pre-mRNA splicing and microRNA processing in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2008, 105, 8795–8800. [Google Scholar] [CrossRef] [PubMed]
- Rogers, K.; Chen, X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 2013, 25, 2383–2399. [Google Scholar] [CrossRef] [PubMed]
- Brodersen, P.; Sakvarelidze-Achard, L.; Bruun-Rasmussen, M.; Dunoyer, P.; Yamamoto, Y.Y.; Sieburth, L.; Voinnet, O. Widespread translational inhibition by plant miRNAs and siRNAs. Science 2008, 320, 1185–1190. [Google Scholar] [CrossRef] [PubMed]
- Ren, G.; Xie, M.; Dou, Y.; Zhang, S.; Zhang, C.; Yu, B. Regulation of miRNA abundance by RNA binding protein TOUGH in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 12817–12821. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Yang, J.-Y.; Xu, J.; Jang, I.-C.; Prigge, M.J.; Chua, N.-H. Two CAP BINDING PROTEINS CBP20 and CBP80 are involved in processing primary microRNAs. Plant Cell Physiol. 2008, 49, 1634–1644. [Google Scholar] [CrossRef] [PubMed]
- Gregory, B.D.; O’Malley, R.C.; Lister, R.; Urich, M.A.; Tonti-Filippini, J.; Chen, H.; Millar, A.H.; Ecker, J.R. A link between RNA metabolism and silencing affecting Arabidopsis development. Dev. Cell 2008, 14, 854–866. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, Y.; Takashi, Y.; Watanabe, Y. The interaction between DCL1 and HYL1 is important for efficient and precise processing of pri-miRNA in plant microRNA biogenesis. RNA 2006, 12, 206–212. [Google Scholar] [CrossRef] [PubMed]
- Manavella, P.A.; Hagmann, J.; Ott, F.; Laubinger, S.; Franz, M.; Macek, B.; Weigel, D. Fast-forward genetics identifies plant CPL phosphatases as regulators of miRNA processing factor HYL1. Cell 2012, 151, 859–870. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Bi, L.; Zheng, B.; Ji, L.; Chevalier, D.; Agarwal, M.; Ramachandran, V.; Li, W.; Lagrange, T.; Walker, J.C.; et al. The FHA domain proteins DAWDLE in Arabidopsis and SNIP1 in humans act in small RNA biogenesis. Proc. Natl. Acad. Sci. USA 2008, 105, 10073–10078. [Google Scholar] [CrossRef] [PubMed]
- Dziembowski, A.; Ventura, A.P.; Rutz, B.; Caspary, F.; Faux, C.; Halgand, F.; Laprévote, O.; Séraphin, B. Proteomic analysis identifies a new complex required for nuclear pre-mRNA retention and splicing. EMBO J. 2004, 23, 4847–4856. [Google Scholar] [CrossRef] [PubMed]
- Raczynska, K.D.; Stepien, A.; Kierzkowski, D.; Kalak, M.; Bajczyk, M.; McNicol, J.; Simpson, C.G.; Szweykowska-Kulinska, Z.; Brown, J.W.S.; Jarmolowski, A. The SERRATE protein is involved in alternative splicing in Arabidopsis thaliana. Nucleic Acids Res. 2013, 42, 1224–1244. [Google Scholar] [CrossRef] [PubMed]
- Köster, T.; Meyer, K.; Weinholdt, C.; Smith, L.M.; Lummer, M.; Speth, C.; Grosse, I.; Weigel, D.; Staiger, D. Regulation of pri-miRNA processing by the hnRNP-like protein AtGRP7 in Arabidopsis. Nucleic Acids Res. 2014, 42, 9925–9936. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Xie, M.; Ren, G.; Yu, B. CDC5, a DNA binding protein, positively regulates posttranscriptional processing and/or transcription of primary microRNA transcripts. Proc. Natl. Acad. Sci. USA 2013, 110, 17588–17593. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Liu, Y.; Yu, B. PRL1, an RNA-binding protein, positively regulates the accumulation of miRNAs and siRNAs in Arabidopsis. PLoS Genet. 2014, 10, e1004841. [Google Scholar] [CrossRef] [PubMed]
- Ben Chaabane, S.; Liu, R.; Chinnusamy, V.; Kwon, Y.; Park, J.-H.; Kim, S.Y.; Zhu, J.-K.; Yang, S.W.; Lee, B.-H. STA1, an Arabidopsis pre-mRNA processing factor 6 homolog, is a new player involved in miRNA biogenesis. Nucleic Acids Res. 2013, 41, 1984–1997. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, F.; Marnef, A.; Cheung, M.-K.; Wilson, I.; Hancock, J.; Staiger, D.; Ladomery, M. A proteomic analysis of oligo(dT)-bound mRNP containing oxidative stress-induced Arabidopsis thaliana RNA-binding proteins ATGRP7 and ATGRP8. Mol. Biol. Rep. 2010, 37, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Castello, A.; Fischer, B.; Eichelbaum, K.; Horos, R.; Beckmann, B.M.; Strein, C.; Davey, N.E.; Humphreys, D.T.; Preiss, T.; Steinmetz, L.M.; et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 2012, 149, 1393–1406. [Google Scholar] [CrossRef] [PubMed]
- Lorkovic, Z.J.; Barta, A. Genome analysis: RNA recognition motif (RRM) and K homology (KH) domain RNA-binding proteins from the flowering plant Arabidopsis thaliana. Nucleic Acids Res. 2002, 30, 623–635. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Meyer, K.; Koester, T.; Staiger, D. Pre-mRNA Splicing in Plants: In Vivo Functions of RNA-Binding Proteins Implicated in the Splicing Process. Biomolecules 2015, 5, 1717-1740. https://doi.org/10.3390/biom5031717
Meyer K, Koester T, Staiger D. Pre-mRNA Splicing in Plants: In Vivo Functions of RNA-Binding Proteins Implicated in the Splicing Process. Biomolecules. 2015; 5(3):1717-1740. https://doi.org/10.3390/biom5031717
Chicago/Turabian StyleMeyer, Katja, Tino Koester, and Dorothee Staiger. 2015. "Pre-mRNA Splicing in Plants: In Vivo Functions of RNA-Binding Proteins Implicated in the Splicing Process" Biomolecules 5, no. 3: 1717-1740. https://doi.org/10.3390/biom5031717
APA StyleMeyer, K., Koester, T., & Staiger, D. (2015). Pre-mRNA Splicing in Plants: In Vivo Functions of RNA-Binding Proteins Implicated in the Splicing Process. Biomolecules, 5(3), 1717-1740. https://doi.org/10.3390/biom5031717






