CD45 and Basigin (CD147) Are Functional Ligands for Galectin-8 on Human Leukocytes
Abstract
1. Introduction
2. Materials and Methods
2.1. Membrane Preparation
2.2. Coupling of Gal-8 to Affi-Gel-10 Beads
2.3. Gal-8 Affinity Chromatography of HL-60 Cell Membrane Preparation or Intact Cells
2.4. Gal-8 Affinity Chromatography of HL-60 Cells After Cell Surface Biotinylation
2.5. Glyco-Enzymatic Treatment of HL-60 Cells
2.6. Gal-8 Binding Assay
2.7. SDS-PAGE and Gal-8 Blotting
2.8. Mass Spectrometry for Proteomics
2.9. Immunoblotting
2.10. Cell Staining
2.11. RT-PCR
2.12. Confocal Microscopy
2.13. CD45 Inhibition
2.14. Inhibition of Glycosylation
3. Results
3.1. Gal-8 Binds Specifically to a Subset of HL-60 Cell Surface Glycoproteins in a Carbohydrate-Dependent Fashion
3.2. Cell Surface Glycoproteins Captured on Immobilized Gal-8C Are a Subset of the Total Glycoproteins Captured by Immobilized Gal-8
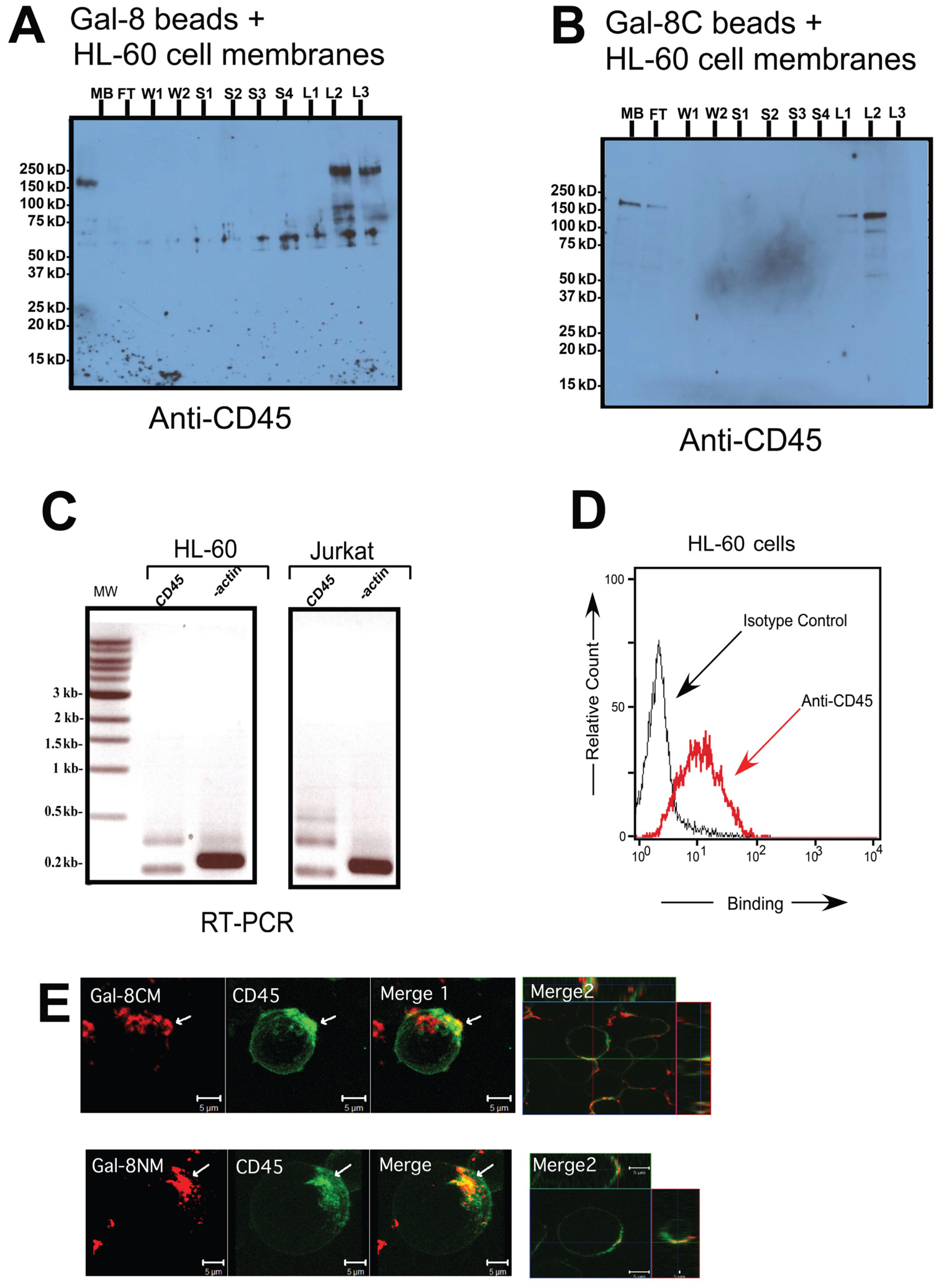
3.3. CD45 Is Involved in Non-Apoptotic PS Exposure Induced by Gal-8
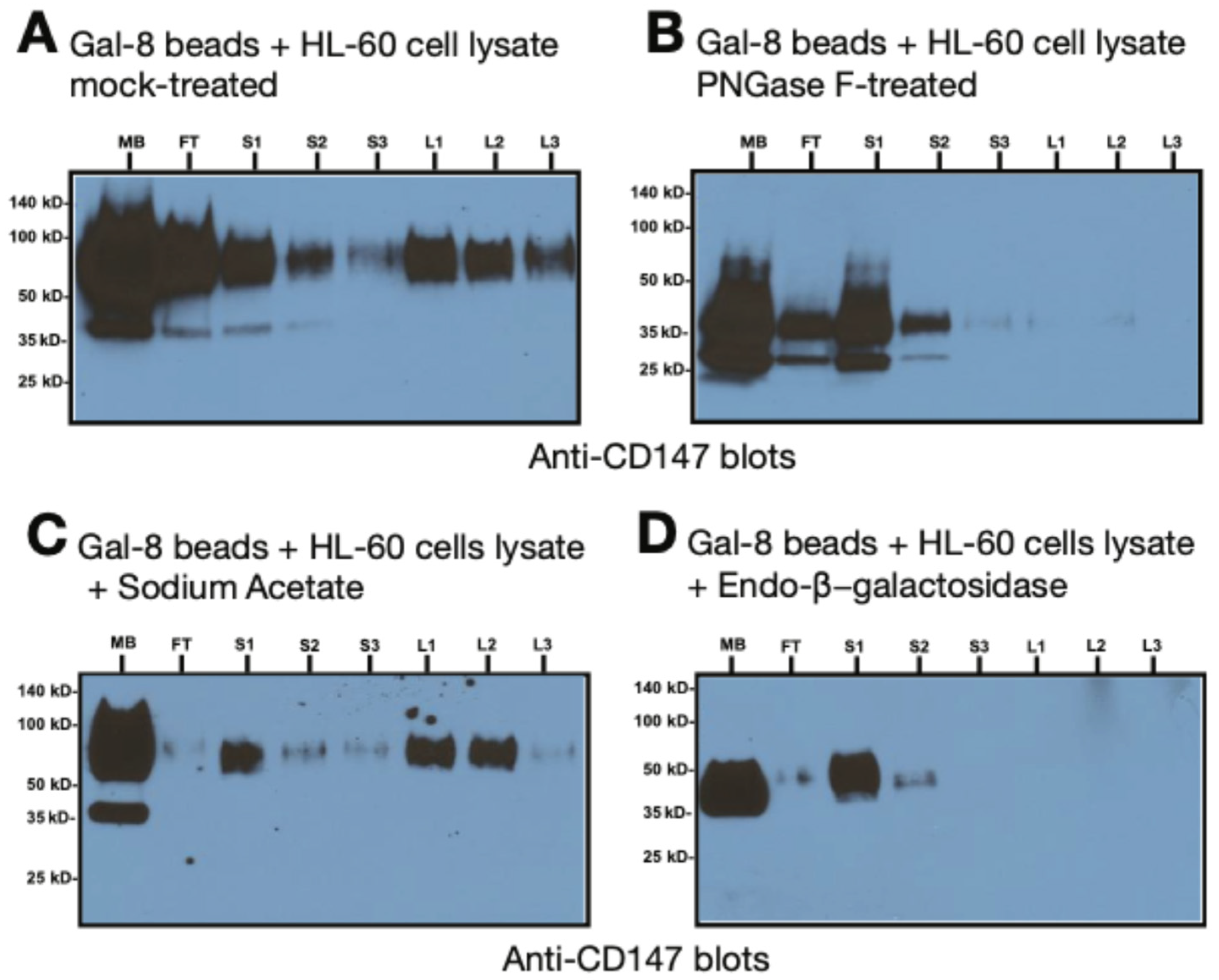
3.4. Gal-8 Binding to Basigin (CD147) Depends on the polyLacNAc Moieties on N-Glycans of CD147
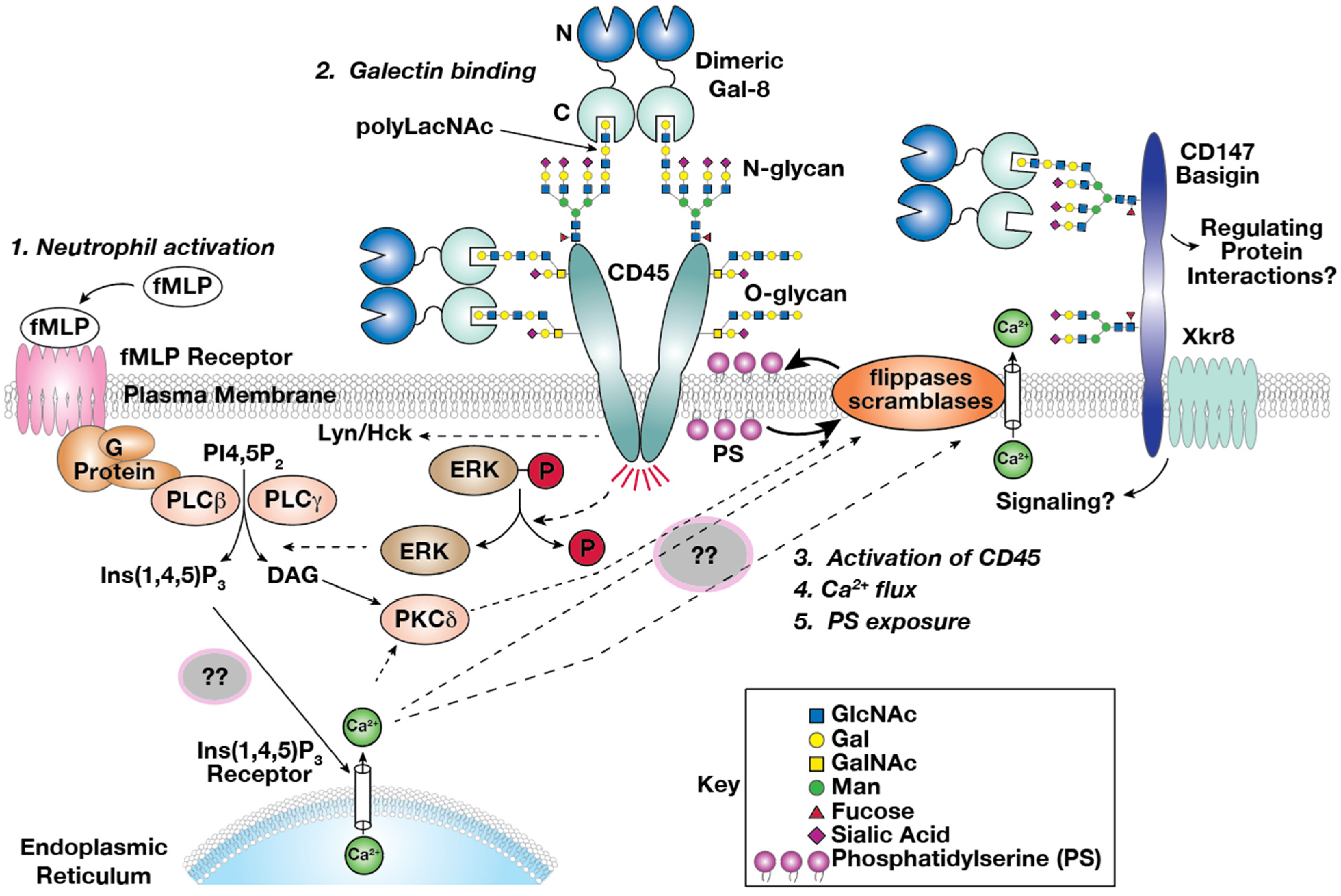
3.5. Gal-8 Binds to Basigin (CD147) on Both Domains and Colocalized on HL-60 Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Benzyl-α GalNAc | Benzyl 2-acetamido-2-deoxy-α-D-galactopyranoside |
| BSA | bovine serum albumin |
| CD45I | CD45 Inhibitor |
| CHO | Chinese hamster ovary |
| CRD | carbohydrate recognition domain |
| DMSO | dimethyl sulfoxide |
| Gal-8 | human Galectin-8 |
| Gal-8C | C-terminal domain of Gal-8 |
| Gal-8N | N-terminal domain of Gal-8 |
| Gal-8CM | R233H mutant of Gal-8 |
| Gal-8NM | R69H mutant of Gal-8 |
| HL-60 | Human promyelocytic leukemia cell |
| PBS | Phosphate buffer saline |
| PBS-TX | Phosphate buffer saline with 1.5% Triton X-100 |
| PI | propidium iodide |
| polyLacNAc | poly-N-acetyllactosamine (-3Galβ1-4GlcNAcβ1-)n |
| PS | phosphatidylserine |
| PTP | Protein Tyrosine Phosphatase |
| Strep-Alexa | Streptavidin-Alexa Fluor labeled |
| Strep-HRP | Streptavidin-horseradish peroxidase |
| TBS | Tris buffer saline |
| TBST | Tris buffer saline with 0.1% Tween |
Appendix A
References
- Liu, F.T.; Stowell, S.R. The role of galectins in immunity and infection. Nat. Rev. Immunol. 2023, 23, 479–494. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Liu, Q.; Wang, D.; Wang, X.; Pan, Z.; Han, B.; He, G. Multifaceted roles of Galectins: From carbohydrate binding to targeted cancer therapy. Biomark. Res. 2025, 13, 49. [Google Scholar] [CrossRef] [PubMed]
- Cummings, R.D.; Rabinovich, G.A.; Stowell, S.R.; Vasta, G. Galectins. In Essentials of Glycobiology, 4th ed.; Varki, A., Cummings, R.D., Esko, J.D., Stanley, P., Hart, G.W., Aebi, M., Mohnen, D., Kinoshita, T., Packer, N.H., Prestegard, J.H., et al., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2022. [Google Scholar]
- Stowell, S.R.; Arthur, C.M.; Dias-Baruffi, M.; Rodrigues, L.C.; Gourdine, J.P.; Heimburg-Molinaro, J.; Ju, T.; Molinaro, R.J.; Rivera-Marrero, C.; Xia, B.; et al. Innate immune lectins kill bacteria expressing blood group antigen. Nat. Med. 2010, 16, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Barondes, S.H.; Cooper, D.N.; Gitt, M.A.; Leffler, H. Galectins. Structure and function of a large family of animal lectins. J. Biol. Chem. 1994, 269, 20807–20810. [Google Scholar] [CrossRef]
- Stowell, S.R.; Karmakar, S.; Arthur, C.M.; Ju, T.; Rodrigues, L.C.; Riul, T.B.; Dias-Baruffi, M.; Miner, J.; McEver, R.P.; Cummings, R.D. Galectin-1 induces reversible phosphatidylserine exposure at the plasma membrane. Mol. Biol. Cell 2009, 20, 1408–1418. [Google Scholar] [CrossRef]
- Stowell, S.R.; Qian, Y.; Karmakar, S.; Koyama, N.S.; Dias-Baruffi, M.; Leffler, H.; McEver, R.P.; Cummings, R.D. Differential roles of galectin-1 and galectin-3 in regulating leukocyte viability and cytokine secretion. J. Immunol. 2008, 180, 3091–3102. [Google Scholar] [CrossRef]
- Stowell, S.R.; Arthur, C.M.; Slanina, K.A.; Horton, J.R.; Smith, D.F.; Cummings, R.D. Dimeric Galectin-8 induces phosphatidylserine exposure in leukocytes through polylactosamine recognition by the C-terminal domain. J. Biol. Chem. 2008, 283, 20547–20559. [Google Scholar] [CrossRef]
- Karmakar, S.; Stowell, S.R.; Cummings, R.D.; McEver, R.P. Galectin-1 signaling in leukocytes requires expression of complex-type N-glycans. Glycobiology 2008, 18, 770–778. [Google Scholar] [CrossRef]
- Stowell, S.R.; Karmakar, S.; Stowell, C.J.; Dias-Baruffi, M.; McEver, R.P.; Cummings, R.D. Human galectin-1, -2, and -4 induce surface exposure of phosphatidylserine in activated human neutrophils but not in activated T cells. Blood 2007, 109, 219–227. [Google Scholar] [CrossRef]
- Arthur, C.M.; Rodrigues, L.C.; Baruffi, M.D.; Sullivan, H.C.; Cummings, R.D.; Stowell, S.R. Detection of phosphatidylserine exposure on leukocytes following treatment with human galectins. Methods Mol. Biol. 2015, 1207, 185–200. [Google Scholar] [CrossRef]
- Collins, S.J. The HL-60 promyelocytic leukemia cell line: Proliferation, differentiation, and cellular oncogene expression. Blood 1987, 70, 1233–1244. [Google Scholar] [CrossRef]
- Nagata, S.; Suzuki, J.; Segawa, K.; Fujii, T. Exposure of phosphatidylserine on the cell surface. Cell Death Differ. 2016, 23, 952–961. [Google Scholar] [CrossRef] [PubMed]
- Rabinovich, G.A.; Toscano, M.A. Turning ‘sweet’ on immunity: Galectin-glycan interactions in immune tolerance and inflammation. Nat. Rev. Immunol. 2009, 9, 338–352. [Google Scholar] [CrossRef] [PubMed]
- Robinson, B.S.; Arthur, C.M.; Evavold, B.; Roback, E.; Kamili, N.A.; Stowell, C.S.; Vallecillo-Zúniga, M.L.; Van Ry, P.M.; Dias-Baruffi, M.; Cummings, R.D.; et al. The Sweet-Side of Leukocytes: Galectins as Master Regulators of Neutrophil Function. Front. Immunol. 2019, 10, 1762. [Google Scholar] [CrossRef] [PubMed]
- Thijssen, V.L.; Hulsmans, S.; Griffioen, A.W. The galectin profile of the endothelium: Altered expression and localization in activated and tumor endothelial cells. Am. J. Pathol. 2008, 172, 545–553. [Google Scholar] [CrossRef] [PubMed]
- Souchak, J.; Mohammed, N.B.B.; Lau, L.S.; Dimitroff, C.J. The role of galectins in mediating the adhesion of circulating cells to vascular endothelium. Front. Immunol. 2024, 15, 1395714. [Google Scholar] [CrossRef]
- Karlsson, M.; Zhang, C.; Mear, L.; Zhong, W.; Digre, A.; Katona, B.; Sjostedt, E.; Butler, L.; Odeberg, J.; Dusart, P.; et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. 2021, 7, eabh2169. [Google Scholar] [CrossRef]
- Troncoso, M.F.; Elola, M.T.; Blidner, A.G.; Sarrias, L.; Espelt, M.V.; Rabinovich, G.A. The universe of galectin-binding partners and their functions in health and disease. J. Biol. Chem. 2023, 299, 105400. [Google Scholar] [CrossRef]
- Bhakta, S.B.; Lundgren, S.M.; Sesti, B.N.; Flores, B.A.; Akdogan, E.; Collins, S.R.; Mercer, F. Neutrophil-like cells derived from the HL-60 cell-line as a genetically-tractable model for neutrophil degranulation. PLoS ONE 2024, 19, e0297758. [Google Scholar] [CrossRef]
- Collins, S.J.; Gallo, R.C.; Gallagher, R.E. Continuous growth and differentiation of human myeloid leukaemic cells in suspension culture. Nature 1977, 270, 347–349. [Google Scholar] [CrossRef]
- Malavez-Cajigas, S.J.; Marini-Martinez, F.I.; Lacourt-Ventura, M.; Rosario-Pacheco, K.J.; Ortiz-Perez, N.M.; Velazquez-Perez, B.; De Jesús-Rojas, W.; Chertow, D.S.; Strich, J.R.; Ramos-Benítez, M.J. HL-60 cells as a valuable model to study LPS-induced neutrophil extracellular traps release. Heliyon 2024, 10, e36386. [Google Scholar] [CrossRef]
- Thomas, M.L. The leukocyte common antigen family. Annu. Rev. Immunol. 1989, 7, 339–369. [Google Scholar] [CrossRef]
- Elliott, J.I.; Surprenant, A.; Marelli-Berg, F.M.; Cooper, J.C.; Cassady-Cain, R.L.; Wooding, C.; Linton, K.; Alexander, D.R.; Higgins, C.F. Membrane phosphatidylserine distribution as a non-apoptotic signalling mechanism in lymphocytes. Nat. Cell Biol. 2005, 7, 808–816. [Google Scholar] [CrossRef] [PubMed]
- Priglinger, C.S.; Szober, C.M.; Priglinger, S.G.; Merl, J.; Euler, K.N.; Kernt, M.; Gondi, G.; Behler, J.; Geerlof, A.; Kampik, A.; et al. Galectin-3 induces clustering of CD147 and integrin-beta1 transmembrane glycoprotein receptors on the RPE cell surface. PLoS ONE 2013, 8, e70011. [Google Scholar] [CrossRef] [PubMed]
- Blenda, A.V.; Kamili, N.A.; Wu, S.-C.; Abel, W.F.; Ayona, D.; Gerner-Smidt, C.; Ho, A.D.; Benian, G.M.; Cummings, R.D.; Arthur, C.M.; et al. Galectin-9 recognizes and exhibits antimicrobial activity toward microbes expressing blood group–like antigens. J. Biol. Chem. 2022, 298. [Google Scholar] [CrossRef] [PubMed]
- Blum, H.; Beier, H.; Gross, H.J. Improved silver staining of plant proteins, RNA and DNA in polyacrylamide gels. Electrophoresis 1987, 8, 93–99. [Google Scholar] [CrossRef]
- Seyfried, N.T.; Gozal, Y.M.; Donovan, L.E.; Herskowitz, J.H.; Dammer, E.B.; Xia, Q.; Ku, L.; Chang, J.; Duong, D.M.; Rees, H.D.; et al. Quantitative analysis of the detergent-insoluble brain proteome in frontotemporal lobar degeneration using SILAC internal standards. J. Proteome Res. 2012, 11, 2721–2738. [Google Scholar] [CrossRef]
- Elias, J.E.; Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 2007, 4, 207–214. [Google Scholar] [CrossRef]
- Barsnes, H.; Vizcaino, J.A.; Eidhammer, I.; Martens, L. PRIDE Converter: Making proteomics data-sharing easy. Nat. Biotechnol. 2009, 27, 598–599. [Google Scholar] [CrossRef]
- Stanton, T.; Boxall, S.; Hirai, K.; Dawes, R.; Tonks, S.; Yasui, T.; Kanaoka, Y.; Yuldasheva, N.; Ishiko, O.; Bodmer, W.; et al. A high-frequency polymorphism in exon 6 of the CD45 tyrosine phosphatase gene (PTPRC) resulting in altered isoform expression. Proc. Natl. Acad. Sci. USA 2003, 100, 5997–6002. [Google Scholar] [CrossRef]
- Lynch, K.W.; Weiss, A. A model system for activation-induced alternative splicing of CD45 pre-mRNA in T cells implicates protein kinase C and Ras. Mol. Cell Biol. 2000, 20, 70–80. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Nishi, N.; Shoji, H.; Seki, M.; Itoh, A.; Miyanaka, H.; Yuube, K.; Hirashima, M.; Nakamura, T. Galectin-8 modulates neutrophil function via interaction with integrin αM. Glycobiology 2003, 13, 755–763. [Google Scholar] [CrossRef]
- Si, Y.; Zhu, J.; Sayed, H.; Mayo, K.H.; Zhou, Y.; Tai, G.; Su, J. CD98hc, a novel of galectin-8 receptor, binds to galectin-8 in an N-glycosylation-dependent manner. Acta Biochim. Biophys. Sin. 2025, 57, 749–757. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, S.; Shinkai, H.; Deutzmann, R.; Paulsson, M.; Timpl, R. Structure and distribution of N-linked oligosaccharide chains on various domains of mouse tumour laminin. Biochem. J. 1988, 252, 453–461. [Google Scholar] [CrossRef] [PubMed]
- Knibbs, R.N.; Perini, F.; Goldstein, I.J. Structure of the major concanavalin A reactive oligosaccharides of the extracellular matrix component laminin. Biochemistry 1989, 28, 6379–6392. [Google Scholar] [CrossRef]
- Stanley, P.; Sundaram, S.; Sallustio, S. A subclass of cell surface carbohydrates revealed by a CHO mutant with two glycosylation mutations. Glycobiology 1991, 1, 307–314. [Google Scholar] [CrossRef]
- Osset, M.; Piñol, M.; Fallon, M.J.; de Llorens, R.; Cuchillo, C.M. Interference of the carbohydrate moiety in coomassie brilliant blue R-250 protein staining. Electrophoresis 1989, 10, 271–273. [Google Scholar] [CrossRef]
- Takaishi, M.; Makino, T.; Morohashi, M.; Huh, N.-H. Identification of Human Hornerin and Its Expression in Regenerating and Psoriatic Skin. J. Biol. Chem. 2005, 280, 4696–4703. [Google Scholar] [CrossRef]
- Okamoto, T.; Hattori, M.; Katsube, Y.; Ota, J.; Asanuma, K.; Usuda, H.; Wada, K.; Suzuki, K.; Nikai, T. Hornerin expressed on endothelial cells via interacting with thrombomodulin modulates vascular inflammation and angiogenesis. Biochim. Biophys. Acta (BBA) Mol. Cell Res. 2025, 1872, 119891. [Google Scholar] [CrossRef]
- Gutknecht, M.F.; Seaman, M.E.; Ning, B.; Cornejo, D.A.; Mugler, E.; Antkowiak, P.F.; Moskaluk, C.A.; Hu, S.; Epstein, F.H.; Kelly, K.A. Identification of the S100 fused-type protein hornerin as a regulator of tumor vascularity. Nat. Commun. 2017, 8, 552. [Google Scholar] [CrossRef]
- Fleming, J.M.; Ginsburg, E.; Oliver, S.D.; Goldsmith, P.; Vonderhaar, B.K. Hornerin, an S100 family protein, is functional in breast cells and aberrantly expressed in breast cancer. BMC Cancer 2012, 12, 266. [Google Scholar] [CrossRef]
- Cotter, K.; Stransky, L.; McGuire, C.; Forgac, M. Recent Insights into the Structure, Regulation, and Function of the V-ATPases. Trends Biochem. Sci. 2015, 40, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Eaton, A.F.; Merkulova, M.; Brown, D. The H+-ATPase (V-ATPase): From proton pump to signaling complex in health and disease. Am. J. Physiol. Cell Physiol. 2021, 320, C392–C414. [Google Scholar] [CrossRef] [PubMed]
- Taetle, R.; Ostergaard, H.; Smedsrud, M.; Trowbridge, I. Regulation of CD45 expression in human leukemia cells. Leukemia 1991, 5, 309–314. [Google Scholar] [PubMed]
- Hermiston, M.L.; Xu, Z.; Weiss, A. CD45: A Critical Regulator of Signaling Thresholds in Immune Cells. Annu. Rev. Immunol. 2003, 21, 107–137. [Google Scholar] [CrossRef]
- Zick, Y. Galectin-8, cytokines, and the storm. Biochem. Soc. Trans. 2022, 50, 135–149. [Google Scholar] [CrossRef]
- Anderson, N.G.; Maller, J.L.; Tonks, N.K.; Sturgill, T.W. Requirement for integration of signals from two distinct phosphorylation pathways for activation of MAP kinase. Nature 1990, 343, 651–653. [Google Scholar] [CrossRef]
- Tribulatti, M.V.; Cattaneo, V.; Hellman, U.; Mucci, J.; Campetella, O. Galectin-8 provides costimulatory and proliferative signals to T lymphocytes. J. Leukoc. Biol. 2009, 86, 371–380. [Google Scholar] [CrossRef]
- Romaniuk, M.A.; Tribulatti, M.V.; Cattaneo, V.; Lapponi, M.J.; Molinas, F.C.; Campetella, O.; Schattner, M. Human platelets express and are activated by galectin-8. Biochem. J. 2010, 432, 535–547. [Google Scholar] [CrossRef]
- Alge-Priglinger, C.S.; Andre, S.; Schoeffl, H.; Kampik, A.; Strauss, R.W.; Kernt, M.; Gabius, H.J.; Priglinger, S.G. Negative regulation of RPE cell attachment by carbohydrate-dependent cell surface binding of galectin-3 and inhibition of the ERK-MAPK pathway. Biochimie 2011, 93, 477–488. [Google Scholar] [CrossRef]
- Elbein, A.D.; Tropea, J.E.; Mitchell, M.; Kaushal, G.P. Kifunensine, a potent inhibitor of the glycoprotein processing mannosidase I. J. Biol. Chem. 1990, 265, 15599–15605. [Google Scholar] [CrossRef]
- Huang, J.; Byrd, J.C.; Yoon, W.H.; Kim, Y.S. Effect of benzyl-alpha-GalNAc, an inhibitor of mucin glycosylation, on cancer-associated antigens in human colon cancer cells. Oncol. Res. 1992, 4, 507–515. [Google Scholar]
- Walcheck, B.; Leppanen, A.; Cummings, R.D.; Knibbs, R.N.; Stoolman, L.M.; Alexander, S.R.; Mattila, P.E.; McEver, R.P. The monoclonal antibody CHO-131 binds to a core 2 O-glycan terminated with sialyl-Lewis x, which is a functional glycan ligand for P-selectin. Blood 2002, 99, 4063–4069. [Google Scholar] [CrossRef] [PubMed]
- van Heyningen, S. Cholera toxin. Biol. Rev. Camb. Philos. Soc. 1977, 52, 509. [Google Scholar] [CrossRef] [PubMed]
- Baenziger, J.U.; Fiete, D. Structural determinants of concanavalin A specificity for oligosaccharides. J. Biol. Chem. 1979, 254, 2400–2407. [Google Scholar] [CrossRef] [PubMed]
- Cummings, R.D.; Kornfeld, S. Characterization of the structural determinants required for the high affinity interaction of asparagine-linked oligosaccharides with immobilized Phaseolus vulgaris leukoagglutinating and erythroagglutinating lectins. J. Biol. Chem. 1982, 257, 11230–11234. [Google Scholar] [CrossRef]
- Plummer, T.H., Jr.; Elder, J.H.; Alexander, S.; Phelan, A.W.; Tarentino, A.L. Demonstration of peptide:N-glycosidase F activity in endo-beta-N-acetylglucosaminidase F preparations. J. Biol. Chem. 1984, 259, 10700–10704. [Google Scholar] [CrossRef]
- Tang, W.; Chang, S.B.; Hemler, M.E. Links between CD147 function, glycosylation, and caveolin-1. Mol. Biol. Cell 2004, 15, 4043–4050. [Google Scholar] [CrossRef]
- Mitsui, Y.; Yamada, K.; Hara, S.; Kinoshita, M.; Hayakawa, T.; Kakehi, K. Comparative studies on glycoproteins expressing polylactosamine-type N-glycans in cancer cells. J. Pharm. Biomed. Anal. 2012, 70, 718–726. [Google Scholar] [CrossRef]
- Togayachi, A.; Tomioka, A.; Fujita, M.; Sukegawa, M.; Noro, E.; Takakura, D.; Miyazaki, M.; Shikanai, T.; Narimatsu, H.; Kaji, H. Identification of Poly-N-Acetyllactosamine-Carrying Glycoproteins from HL-60 Human Promyelocytic Leukemia Cells Using a Site-Specific Glycome Analysis Method, Glyco-RIDGE. J. Am. Soc. Mass. Spectrom. 2018, 29, 1138–1152. [Google Scholar] [CrossRef] [PubMed]
- Piedfer, M.; Dauzonne, D.; Tang, R.; N’Guyen, J.; Billard, C.; Bauvois, B. Aminopeptidase-N/CD13 is a potential proapoptotic target in human myeloid tumor cells. FASEB J. 2011, 25, 2831–2842. [Google Scholar] [CrossRef]
- Mina-Osorio, P. The moonlighting enzyme CD13: Old and new functions to target. Trends Mol. Med. 2008, 14, 361–371. [Google Scholar] [CrossRef]
- Sato, T.; Furukawa, K.; Autero, M.; Gahmberg, C.G.; Kobata, A. Structural study of the sugar chains of human leukocyte common antigen CD45. Biochemistry 1993, 32, 12694–12704. [Google Scholar] [CrossRef]
- Earl, L.A.; Bi, S.; Baum, L.G. N- and O-glycans modulate galectin-1 binding, CD45 signaling, and T cell death. J. Biol. Chem. 2010, 285, 2232–2244. [Google Scholar] [CrossRef]
- Ferguson, B.V.; Ostergaard, H.L. CD45 regulates thymocyte survival during development in fetal thymic organ culture. Immunobiology 2010, 215, 458–465. [Google Scholar] [CrossRef]
- Gao, H.; Henderson, A.; Flynn, D.C.; Landreth, K.S.; Ericson, S.G. Effects of the protein tyrosine phosphatase CD45 on FcgammaRIIa signaling and neutrophil function. Exp. Hematol. 2000, 28, 1062–1070. [Google Scholar] [CrossRef]
- Panchal, R.G.; Ulrich, R.L.; Bradfute, S.B.; Lane, D.; Ruthel, G.; Kenny, T.A.; Iversen, P.L.; Anderson, A.O.; Gussio, R.; Raschke, W.C.; et al. Reduced expression of CD45 protein-tyrosine phosphatase provides protection against anthrax pathogenesis. J. Biol. Chem. 2009, 284, 12874–12885. [Google Scholar] [CrossRef]
- Chen, I.J.; Chen, H.L.; Demetriou, M. Lateral compartmentalization of T cell receptor versus CD45 by galectin-N-glycan binding and microfilaments coordinate basal and activation signaling. J. Biol. Chem. 2007, 282, 35361–35372. [Google Scholar] [CrossRef]
- Haston, W.S.; Maggs, A.F. Evidence for membrane differentiation in polarised leucocytes: The distribution of surface antigens analysed with Ig-gold labelling. J. Cell Sci. 1990, 95 Pt 3, 471–479. [Google Scholar] [CrossRef]
- Seveau, S.; Eddy, R.J.; Maxfield, F.R.; Pierini, L.M. Cytoskeleton-dependent membrane domain segregation during neutrophil polarization. Mol. Biol. Cell 2001, 12, 3550–3562. [Google Scholar] [CrossRef] [PubMed]
- Cattaneo, V.; Tribulatti, M.V.; Campetella, O. Galectin-8 tandem-repeat structure is essential for T-cell proliferation but not for co-stimulation. Biochem. J. 2011, 434, 153–160. [Google Scholar] [CrossRef] [PubMed]
- Pulido, R.; Lacal, P.; Mollinedo, F.; Sanchez-Madrid, F. Biochemical and antigenic characterization of CD45 polypeptides expressed on plasma membrane and internal granules of human neutrophils. FEBS Lett. 1989, 249, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Kirk, P.; Wilson, M.C.; Heddle, C.; Brown, M.H.; Barclay, A.N.; Halestrap, A.P. CD147 is tightly associated with lactate transporters MCT1 and MCT4 and facilitates their cell surface expression. EMBO J. 2000, 19, 3896–3904. [Google Scholar] [CrossRef]
- Wilson, M.C.; Meredith, D.; Fox, J.E.; Manoharan, C.; Davies, A.J.; Halestrap, A.P. Basigin (CD147) is the target for organomercurial inhibition of monocarboxylate transporter isoforms 1 and 4: The ancillary protein for the insensitive MCT2 is EMBIGIN (gp70). J. Biol. Chem. 2005, 280, 27213–27221. [Google Scholar] [CrossRef]
- Feral, C.C.; Nishiya, N.; Fenczik, C.A.; Stuhlmann, H.; Slepak, M.; Ginsberg, M.H. CD98hc (SLC3A2) mediates integrin signaling. Proc. Natl. Acad. Sci. USA 2005, 102, 355–360. [Google Scholar] [CrossRef]
- Kasinrerk, W.; Fiebiger, E.; Stefanova, I.; Baumruker, T.; Knapp, W.; Stockinger, H. Human leukocyte activation antigen M6, a member of the Ig superfamily, is the species homologue of rat OX-47, mouse basigin, and chicken HT7 molecule. J. Immunol. 1992, 149, 847–854. [Google Scholar] [CrossRef]
- Kato, N.; Yuzawa, Y.; Kosugi, T.; Hobo, A.; Sato, W.; Miwa, Y.; Sakamoto, K.; Matsuo, S.; Kadomatsu, K. The E-selectin ligand basigin/CD147 is responsible for neutrophil recruitment in renal ischemia/reperfusion. J. Am. Soc. Nephrol. 2009, 20, 1565–1576. [Google Scholar] [CrossRef]
- Fukuda, M.; Matsumura, G. Endo-beta-galactosidase of Escherichia freundii. Hydrolysis of pig colonic mucin and milk oligosaccharides by endoglycosidic action. Biochem. Biophys. Res. Commun. 1975, 64, 465–471. [Google Scholar] [CrossRef]
- Rillahan, C.D.; Antonopoulos, A.; Lefort, C.T.; Sonon, R.; Azadi, P.; Ley, K.; Dell, A.; Haslam, S.M.; Paulson, J.C. Global metabolic inhibitors of sialyl- and fucosyltransferases remodel the glycome. Nat. Chem. Biol. 2012, 8, 661–668. [Google Scholar] [CrossRef]
- Lee, N.; Wang, W.C.; Fukuda, M. Granulocytic differentiation of HL-60 cells is associated with increase of poly-N-acetyllactosamine in Asn-linked oligosaccharides attached to human lysosomal membrane glycoproteins. J. Biol. Chem. 1990, 265, 20476–20487. [Google Scholar] [CrossRef] [PubMed]
- Mizoguchi, A.; Takasaki, S.; Maeda, S.; Kobata, A. Changes in asparagine-linked sugar chains of human promyelocytic leukemic cells (HL-60) during monocytoid differentiation and myeloid differentiation. Appearance of high mannose-type oligosaccharides in neutral fraction. J. Biol. Chem. 1984, 259, 11943–11948. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, P.P.; McEver, R.P.; Cummings, R.D. Structures of the O-glycans on P-selectin glycoprotein ligand-1 from HL-60 cells. J. Biol. Chem. 1996, 271, 18732–18742. [Google Scholar] [CrossRef]
- Fossum, S.; Mallett, S.; Barclay, A.N. The MRC OX-47 antigen is a member of the immunoglobulin superfamily with an unusual transmembrane sequence. Eur. J. Immunol. 1991, 21, 671–679. [Google Scholar] [CrossRef] [PubMed]
- Spring, F.A.; Holmes, C.H.; Simpson, K.L.; Mawby, W.J.; Mattes, M.J.; Okubo, Y.; Parsons, S.F. The Oka blood group antigen is a marker for the M6 leukocyte activation antigen, the human homolog of OX-47 antigen, basigin and neurothelin, an immunoglobulin superfamily molecule that is widely expressed in human cells and tissues. Eur. J. Immunol. 1997, 27, 891–897. [Google Scholar] [CrossRef]
- Biswas, C.; Zhang, Y.; DeCastro, R.; Guo, H.; Nakamura, T.; Kataoka, H.; Nabeshima, K. The human tumor cell-derived collagenase stimulatory factor (renamed EMMPRIN) is a member of the immunoglobulin superfamily. Cancer Res. 1995, 55, 434–439. [Google Scholar]
- Yurchenko, V.; O’Connor, M.; Dai, W.W.; Guo, H.; Toole, B.; Sherry, B.; Bukrinsky, M. CD147 is a signaling receptor for cyclophilin B. Biochem. Biophys. Res. Commun. 2001, 288, 786–788. [Google Scholar] [CrossRef]
- Pushkarsky, T.; Zybarth, G.; Dubrovsky, L.; Yurchenko, V.; Tang, H.; Guo, H.; Toole, B.; Sherry, B.; Bukrinsky, M. CD147 facilitates HIV-1 infection by interacting with virus-associated cyclophilin A. Proc. Natl. Acad. Sci. USA 2001, 98, 6360–6365. [Google Scholar] [CrossRef]
- Berditchevski, F.; Chang, S.; Bodorova, J.; Hemler, M.E. Generation of monoclonal antibodies to integrin-associated proteins. Evidence that alpha3beta1 complexes with EMMPRIN/basigin/OX47/M6. J. Biol. Chem. 1997, 272, 29174–29180. [Google Scholar] [CrossRef]
- Sakuragi, T.; Kanai, R.; Tsutsumi, A.; Narita, H.; Onishi, E.; Nishino, K.; Miyazaki, T.; Baba, T.; Kosako, H.; Nakagawa, A.; et al. The tertiary structure of the human Xkr8-Basigin complex that scrambles phospholipids at plasma membranes. Nat. Struct. Mol. Biol. 2021, 28, 825–834. [Google Scholar] [CrossRef]
- Suzuki, J.; Denning, D.P.; Imanishi, E.; Horvitz, H.R.; Nagata, S. Xk-related protein 8 and CED-8 promote phosphatidylserine exposure in apoptotic cells. Science 2013, 341, 403–406. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, J.; Imanishi, E.; Nagata, S. Xkr8 phospholipid scrambling complex in apoptotic phosphatidylserine exposure. Proc. Natl. Acad. Sci. USA 2016, 113, 9509–9514. [Google Scholar] [CrossRef] [PubMed]
- Levy, Y.; Arbel-Goren, R.; Hadari, Y.R.; Eshhar, S.; Ronen, D.; Elhanany, E.; Geiger, B.; Zick, Y. Galectin-8 functions as a matricellular modulator of cell adhesion. J. Biol. Chem. 2001, 276, 31285–31295. [Google Scholar] [CrossRef] [PubMed]
- Hadari, Y.R.; Arbel-Goren, R.; Levy, Y.; Amsterdam, A.; Alon, R.; Zakut, R.; Zick, Y. Galectin-8 binding to integrins inhibits cell adhesion and induces apoptosis. J. Cell Sci. 2000, 113 Pt 13, 2385–2397. [Google Scholar] [CrossRef]
- McCall, M.N.; Uppal, K.; Jaffee, H.A.; Zilliox, M.J.; Irizarry, R.A. The Gene Expression Barcode: Leveraging public data repositories to begin cataloging the human and murine transcriptomes. Nucleic Acids Res. 2011, 39, D1011–D1015. [Google Scholar] [CrossRef]
- Marino, G.; Kroemer, G. Mechanisms of apoptotic phosphatidylserine exposure. Cell Res. 2013, 23, 1247–1248. [Google Scholar] [CrossRef]
- Karmakar, S.; Cummings, R.D.; McEver, R.P. Contributions of Ca2+ to galectin-1-induced exposure of phosphatidylserine on activated neutrophils. J. Biol. Chem. 2005, 280, 28623–28631. [Google Scholar] [CrossRef]






| Total Peptide Count | Spectral Counts | Total Peptide Count | Spectral Counts | Annotation | Mass | Gene | Role/Localization |
|---|---|---|---|---|---|---|---|
| C Domain | C Domain | Full Length | Full Length | (kD) | |||
| 31 | 58 | 43 | 71 | aminopeptidase N precursor [Homo sapiens]—CD13 | 109 | ANPEP | Ectoproteinase/Membrane Raft |
| 8 | 16 | 28 | 36 | receptor-type tyrosine-protein phosphatase C isoform 2 precursor [Homo sapiens]—CD45 | 131 | PTPRC | Phosphatase, signaling/Membrane |
| 4 | 4 | 21 | 24 | transferrin receptor protein 1 [Homo sapiens]—CD71 | 85 | TFRC | Transporter/Membrane |
| 17 | 21 | 21 | 27 | 4F2 cell-surface antigen heavy chain isoform c [Homo sapiens]—CD98 | 68 | SLC3A2 | Transporter/Membrane |
| 14 | 15 | 17 | 23 | leucyl-cystinyl aminopeptidase isoform 1 [Homo sapiens] | 117 | LNPEP | Ectoproteinase/Membrane Raft |
| 2 | 2 | 17 | 20 | carboxypeptidase D isoform 1 precursor [Homo sapiens] | 153 | CPD | Ectoproteinase/Membrane |
| 10 | 23 | 23 | 31 | integrin alpha-5 precursor [Homo sapiens]—CD51 | 114 | ITGA5 | Signaling/Membrane |
| 10 | 16 | 7 | 11 | integrin alpha-4 precursor [Homo sapiens]—CD49d | 115 | ITGA4 | Signaling/Membrane |
| 1 | 1 | 9 | 15 | sodium bicarbonate cotransporter 3 isoform c [Homo sapiens]—Na+/HCO3—cotransporter | 123 | SLC4A7 | Transporter/Membrane |
| 9 | 16 | 10 | 16 | integrin beta-1 isoform 1A precursor [Homo sapiens]—CD29 | 88 | ITGB1 | Signaling/Membrane |
| 3 | 3 | 10 | 15 | neutral amino acid transporter B(0) isoform 1 [Homo sapiens] | 57 | SLC1A5 | Transporter/Membrane |
| 6 | 9 | 8 | 13 | basigin isoform 2 [Homo sapiens]—CD147 | 29 | BSG | Signaling/Membrane |
| 1 | 1 | 8 | 9 | cytochrome b-245 heavy chain [Homo sapiens] | 65 | CYBB | Transporter/Membrane |
| 0 | 0 | 20 | 31 | receptor-type tyrosine-protein phosphatase alpha isoform 1 precursor [Homo sapiens] | 91 | PTPRA | Phosphatase, signaling/Membrane |
| 0 | 0 | 18 | 22 | sodium/potassium-transporting ATPase subunit alpha-1 isoform c [Homo sapiens] ATPase/Na+/K+ | 113 | ATP1A1 | Transporter/Membrane |
| 0 | 0 | 20 | 24 | hornerin [Homo sapiens] | 282 | HRNR | Unknown/Cytoplasmic Granule |
| 0 | 0 | 11 | 11 | V-type proton ATPase 116 kDa subunit a isoform 1 isoform c [Homo sapiens]—ATPase/H+ | 96 | ATP6V0A1 | Transporter/Lysosome |
| 0 | 0 | 9 | 11 | multidrug resistance-associated protein 4 isoform 1 [Homo sapiens]—ATP-binding cassette 4 | 149 | ABCC4 | Transporter/Membrane |
| 0 | 0 | 11 | 11 | multidrug resistance-associated protein 1 [Homo sapiens]—ATP-binding cassette 1 | 171 | ABCC1 | Transporter/Membrane |
| 0 | 0 | 10 | 18 | leukosialin precursor [Homo sapiens]—CD43 | 40 | SPN | Signaling/Membrane |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gourdine, J.-P.F.; Nava, P.; Noll, A.J.; Duong, D.M.; Seyfried, N.T.; Cummings, R.D. CD45 and Basigin (CD147) Are Functional Ligands for Galectin-8 on Human Leukocytes. Biomolecules 2025, 15, 1243. https://doi.org/10.3390/biom15091243
Gourdine J-PF, Nava P, Noll AJ, Duong DM, Seyfried NT, Cummings RD. CD45 and Basigin (CD147) Are Functional Ligands for Galectin-8 on Human Leukocytes. Biomolecules. 2025; 15(9):1243. https://doi.org/10.3390/biom15091243
Chicago/Turabian StyleGourdine, Jean-Philippe F., Porfirio Nava, Alexander J. Noll, Duc M. Duong, Nicholas T. Seyfried, and Richard D. Cummings. 2025. "CD45 and Basigin (CD147) Are Functional Ligands for Galectin-8 on Human Leukocytes" Biomolecules 15, no. 9: 1243. https://doi.org/10.3390/biom15091243
APA StyleGourdine, J.-P. F., Nava, P., Noll, A. J., Duong, D. M., Seyfried, N. T., & Cummings, R. D. (2025). CD45 and Basigin (CD147) Are Functional Ligands for Galectin-8 on Human Leukocytes. Biomolecules, 15(9), 1243. https://doi.org/10.3390/biom15091243







