Abstract
Mutations in the multidomain protein Leucine-rich-repeat kinase 2 (LRRK2) have been identified as a genetic risk factor for both sporadic and familial Parkinson’s disease (PD). LRRK2 has two enzymatic domains: a RocCOR tandem with GTPase activity and a kinase domain. In addition, LRRK2 has three N-terminal domains: ARM (Armadillo repeat), ANK (Ankyrin repeat), and LRR (Leucine-rich-repeat), and a C-terminal WD40 domain, all of which are involved in mediating protein–protein interactions (PPIs) and regulation of the LRRK2 catalytic core. The PD-related mutations have been found in nearly all LRRK2 domains, and most of them have increased kinase activity and/or decreased GTPase activity. The complex activation mechanism of LRRK2 includes at least intramolecular regulation, dimerization, and membrane recruitment. In this review, we highlight the recent developments in the structural characterization of LRRK2 and discuss these developments from the perspective of the LRRK2 activation mechanism, the pathological role of the PD mutants, and therapeutic targeting.
1. Introduction
Parkinson’s disease (PD), the second most common progressive neurodegenerative disease, affects more than 7 million people worldwide, and it is estimated that this number will be doubled by 2040 [1]. Besides environmental factors, in 10–15% of PD patients, genetic predisposition seems to play a major causative role. Through genome-wide association studies, about 20 PD-associated genes have been identified so far, including alpha-synuclein (SNCA), glucocerebrosidase (GBA), and Leucine-rich-repeat kinase 2 (LRRK2) [2,3]. LRRK2 is a commonly mutated gene that emerges in both sporadic and familial PD [4]. Furthermore, recently, it was found that LRRK2 kinase activity was enhanced in idiopathic PD postmortem brain tissue [5].
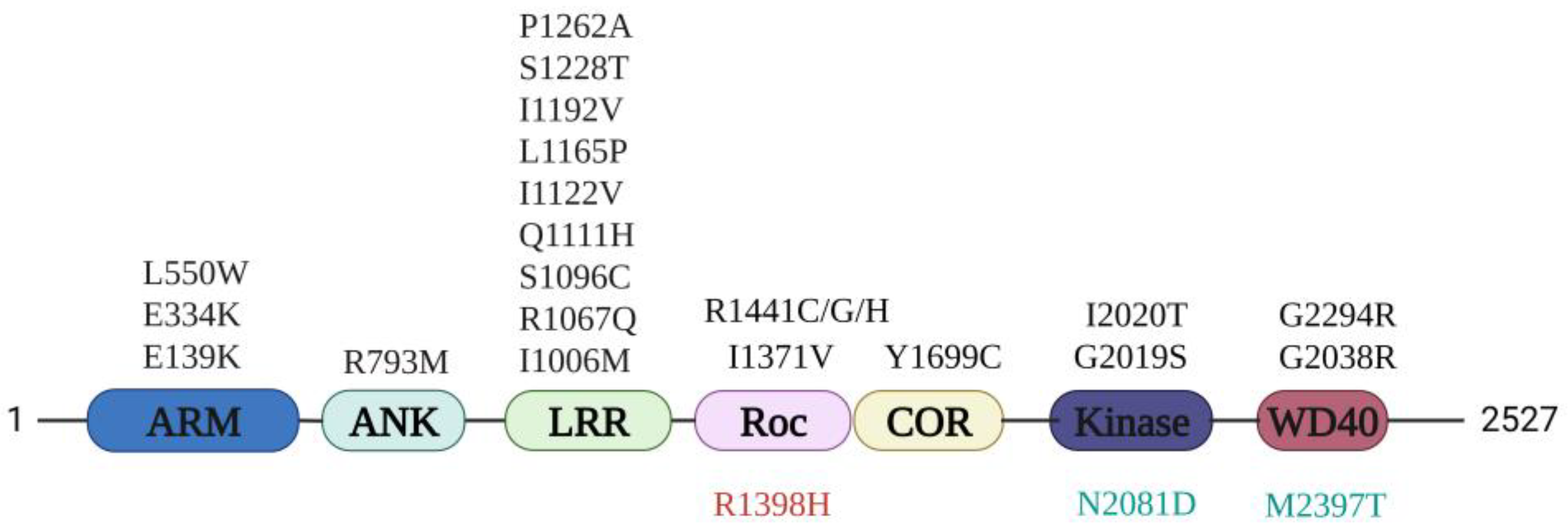
LRRK2 is a large 286 KDa protein that comprises seven domains (Figure 1). The three N-terminal domains, ARM (Armadillo repeat), ANK (Ankyrin repeat), LRR (Leucine-rich-repeat), and the C-terminal WD40 domain are involved in mediating protein–protein interactions (PPIs) and regulation of the LRRK2 catalytic core. The catalytic core consists of a RocCOR tandem, Ras of complex protein (Roc) and C-terminal of Roc (COR), and a kinase domain. The RocCOR domain has GTPase activity and mediates LRRK2 dimerization, while the kinase domain can phosphorylate a subgroup of Rab proteins to control several cellular functions, including intracellular trafficking and autophagy [6,7,8,9]. A multitude of PD-related mutations have been identified over the different LRRK2 domains. The most common mutations, R1441G, Y1699C, and G2019S, are located within the catalytic core and show an increased kinase activity and reduced GTPase activity [10,11,12]. In addition, mutations in LRRK2 have also been linked to other diseases, such as Crohn’s disease (CD) and Hansen’s disease [13,14]. Interestingly, the R1398H variant, which is located within the Roc domain, has been reported to be protective in both PD and CD [15].
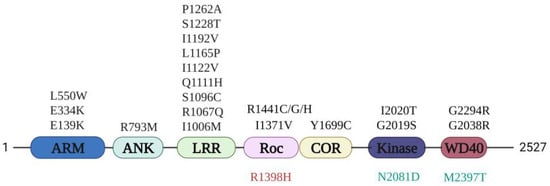
Figure 1.
LRRK2 domain structure and Parkinson’s disease (PD) mutations. Armadillo (ARM), Ankyrin (ANK), Leucine-rich-repeat (LRR), Ras of complex proteins (Roc), C-terminal of Roc (COR), kinase (KIN), and WD40 domains. The PD-related mutations (black), protective mutation (red), and Crohn’s disease (CD) risk factor (green) are indicated.
2. Molecular Structure of LRRK2
2.1. From Bacterial Roco to Human Full-Length LRRK2 Structures
Structural studies on LRRK2 have been ongoing for almost two decades. However, the expression of the separate LRRK2 domains and full-length protein has been a major challenge for a long time. So far, only the crystal structures of the human LRRK2 Roc and WD40 domain have been resolved [16,17]. In addition, X-ray structures are available from homologous Roco family proteins (Table 1). A major breakthrough in the field of structural biology in general and for determining the LRRK2 structure has been the recent developments in electron microscopy (EM). The possibilities related to stable and higher field emission EM, a robust vitrification apparatus, a volta phase plate, faster direct electron detector, and improved data analysing software have resulted in an increase in EM-based structure publications by over 70% in the last three years [18,19,20]. Using these techniques in combination with optimized purification protocols for full-length LRRK2 (flLRRK2) resulted in the first low-resolution negative-stain EM models of LRRK2 in 2016 [21,22]. More recently, the structures of a construct comprising Roc-COR-Kinase-WD40 domains (LRRK2RCKW) and flLRRK2 have been reported by Cryo-EM [23,24,25]. In addition, with Cryo-electron tomography (Cryo-ET), an in situ microtubule-bound full-length LRRK2 structure in cells was determined [26]. Another breakthrough in structural biology is the DeepMind-produced 3D protein structure prediction computational method, AlphaFold2, which makes it possible to predict reliable structures based on multiple sequence alignment and pair representation algorithms in silico [27,28].

Table 1.
Published LRRK2 and Roco structures. Hs (Homo sapiens), Ct (Chlorobium tepidum), Dd (Dictyostelium discoideum), and Mb (Methanosarcina barkeri).
In this review, we summarize the current structural knowledge of LRRK2 in the context of the complex activation mechanism of LRRK2 and discuss the potential mechanism of the disease-linked mutations.
2.2. Kinase Domain Structure
The LRRK2 kinase domain is considered the main output of the protein. It can phosphorylate a group of Rab GTPase family members to regulate several cellular pathways [33]. In addition, auto-phosphorylation of LRRK2 is also important for intramolecular regulation [34,35]. The protein kinase family was discovered by Krebs and Fisher during their study of glycogen phosphorylase in 1956 [36]. Kinase domains catalyze the transfer of γ-phosphate from ATP to a substrate. The kinase family is classified into three groups: Serine/Threonine kinase, tyrosine kinase, and atypical kinase. Based on the kinase domain sequence, LRRK2 belongs to the tyrosine kinase-like kinase (TKL) subfamily, a subfamily of Serine/Threonine kinase, whose members show sequence similarity to tyrosine kinases (TK) but lack TK-specific motifs [37,38].
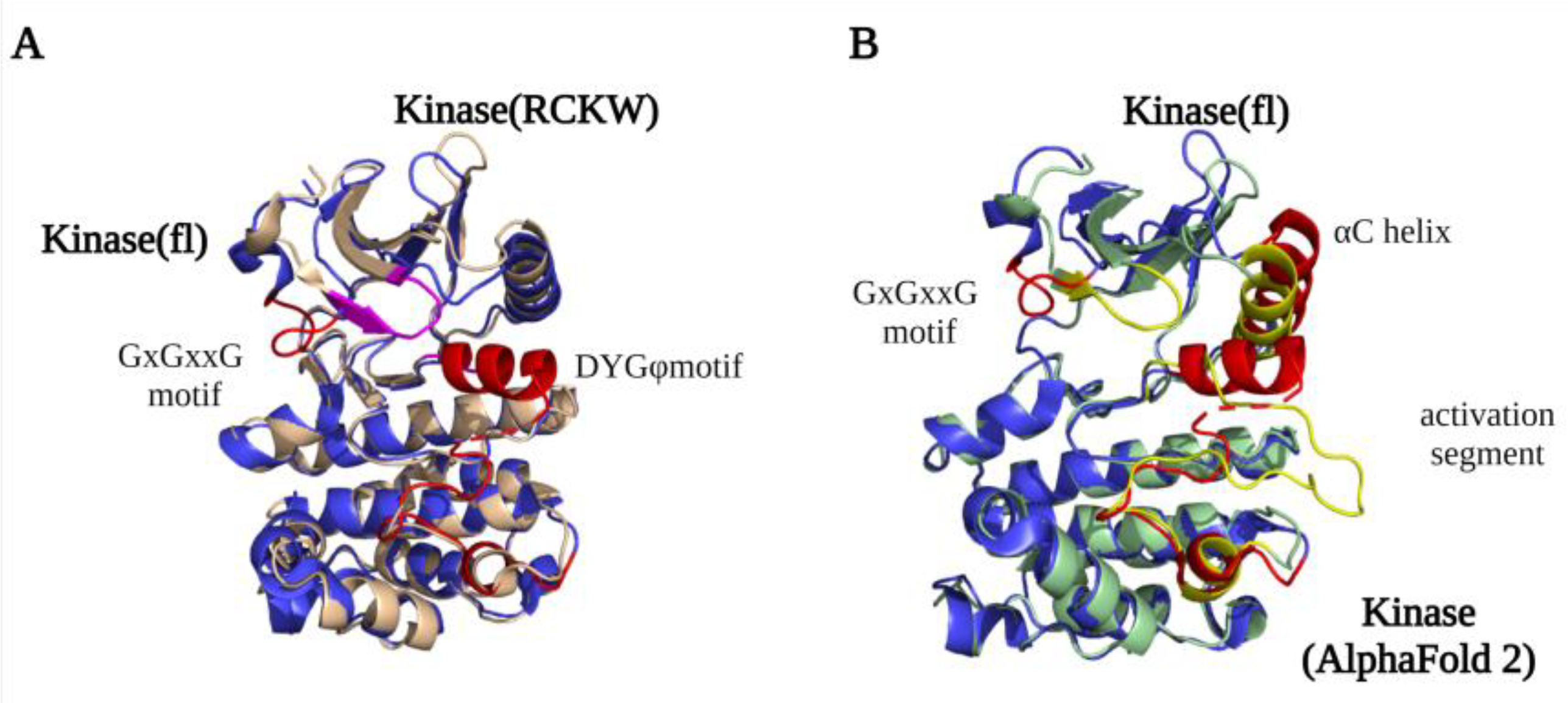
A detailed structural map of the LRRK2 kinase domain first became available after the high-resolution LRRK2RCKW and flLRRK2 Cryo-EM structures were published in 2019 [23,24]. The structures revealed that LRRK2 has a typical kinase structure with a rich β-sheets-formed N-lobe and an α-helix-composed C-lobe. The α-helix in the C-lobe forms the activation segment, which consists of the DFGψ (DYGψ in LRRK2) motif—activation loop—P + 1 loop—APE motif—α-F, and the N-lobe is tightly anchored to the kinase C-lobe by the β4-αC loop [38,39]. However, due to flexibility, the structure of the activation segment has only been partly resolved in the LRRK2RCKW and flLRRK2 structures, while in the flLRRK2 structure, the DYGψ motif forms an unusual helix (Figure 2A). In the activation segment, the DYGψ motif works as a “brake” for LRRK2 conformational changes and activation: the Asp residue from the DFGψ motif binds with Mg2+ to indirectly interact with the oxygen of the β phosphate from ATP, and the P + 1 loop and APE motif provide the docking and interacting sites for substrates peptides, while the αF works as a scaffold that anchors the activation segment in a stable conformation in the kinase inactive state [40]. The two experimentally determined LRRK2 kinase structures both represent an inactive kinase conformation, with the activation segment collapsed on the protein interface in a locked position, thereby hindering substrate or ATP binding. In contrast, the predicted AlphaFold2 structure represents a DYGout active kinase conformation (Figure 2B). The conserved GxGxxG motif in the N-lobe forms most of the ATP binding pocket and is crucial for the stabilization of the phosphate and catalytic activity. Interestingly, the GxGxxG motif adopts a different conformation in the LRRK2RCKW and flLRRK2 (Figure 2A), which might be due to the different concentrations of ATP in the sample preparation.
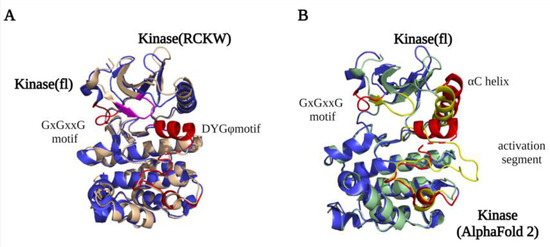
Figure 2.
LRRK2 kinase domain structures. (A) An overlay of the kinase domain from LRRK2RCKW (PDB ID: 6VP7) (light yellow) and flLRRK2 (PDB ID: 7LHW) (blue). The main difference between the structures is the conformation of the GxGxxG motif (highlighted LRRK2RCKW in purple and flLRRK2 in red). (B) An overlay of the kinase domain from flLRRK2 (PDB ID: 7LHW) (blue) and AlphaFold2 model (green) structures. The GxGxxG motif, αC helix, and activation segments are highlighted in red (flLRRK2) and yellow (AlphaFold2), respectively.
By comparing the structures of the LRRK2 kinase domain with those of Braf (B-Raf Proto-Oncogene, Serine/Threonine kinase) and Src (Proto-oncogene tyrosine-protein kinase), the conserved Regulator spine (R-spine) and Catalytic spine (C-spine) were identified [41]. The C-spine has a substrate-capturing, nucleotide-binding, and phosphate transfer function, while the R-spine generally only assembles in kinase active state [42]. However, unexpectedly, the R-spine is also “visible” in the inactive LRRK2RCKW and flLRRK2 structures. The highly conserved Phe in the DFGψ motif is one of the four R-spine residues, and LRRK2 has, instead of a DFGψ motif, an unusual DYGψ motif. Interestingly, mutating DYGψ to DFGψ resulted in an LRRK2 hyperactive phenotype, suggesting that Y2018 is important for stabilizing a kinase inactive conformation [43]. The DYGψ motif thus serves as a conformational switch that regulates LRRK2 activation.
2.3. RocCOR Structure
The LRRK2 RocCOR domain has GTPase hydrolysis activity and, in addition, regulates LRRK2 dimerization [44]. As discussed in more detail below, dimerization of LRRK2 is important for GTPase and kinase activation.
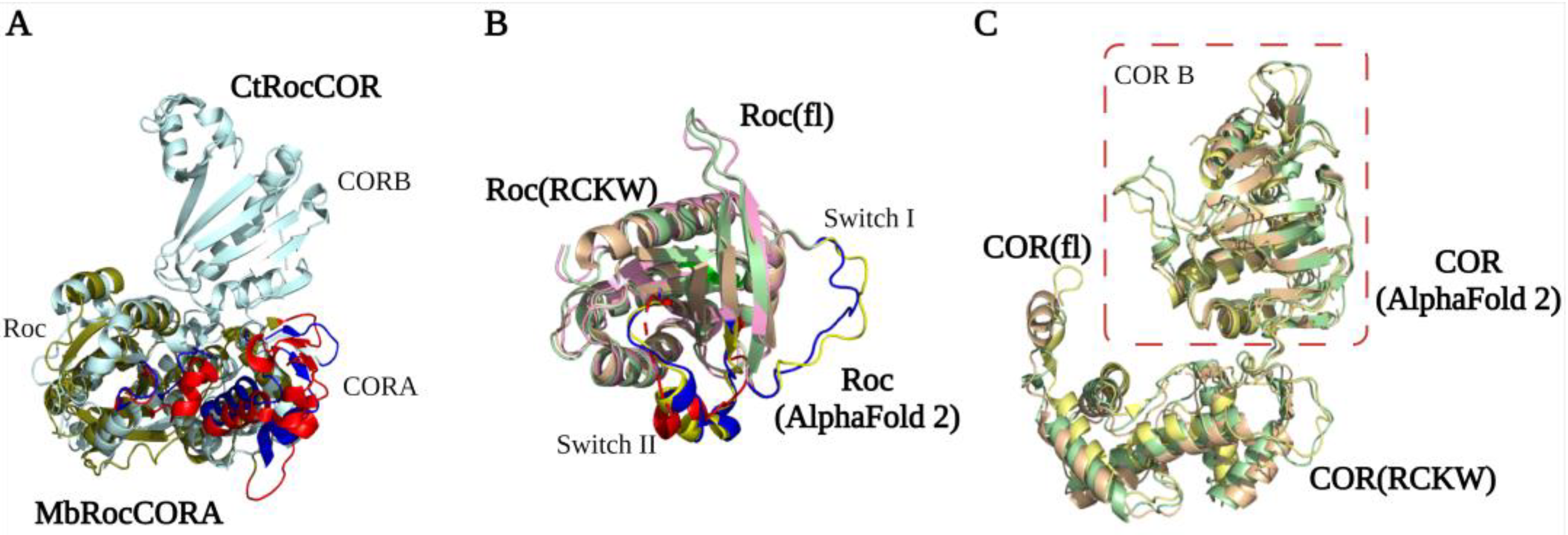
The first LRRK2 Roc crystal structure reported by Deng et al. in 2008 showed that the GDP-bound Roc domain has a typical G domain fold with five α-helix and six β-strands [16,45]. The Roc domain has a highly conserved P-loop (G1), switch I motif (G2), and switch II motif (G3) that together are responsible for Mg2+ and nucleotide binding. The G4-G5 motifs are important for guanine specificity, and are only partly conserved in LRRK2 [46,47,48]. The structure shows that two Roc proteins assemble in a dimeric structure by the interaction of one Roc N-terminal with the C-terminal of the other protomer, thereby forming a pseudo-twofold [16]. However, this unusual dimeric structure is not observed in structures of homologous Roco family proteins nor in any of the high-resolution LRRK2 structures. Furthermore, both an N and C terminal extension of the Roc domain can disrupt the dimerization. Therefore, this structure is most likely caused by a crystallization artifact that was induced by the short flanking boundaries of the construct used [49]. Biochemical and structural characterization of the RocCOR domain of C. tepidum (Ct) Roco has revealed instead that the dimerization is mainly mediated by the C-terminal of the COR domain (CORB) [29]. The full-length CtRoco structure showed that in addition to the tight CORB interaction, the Roc domain is also involved in the dimeric interaction by Roc–Roc, Roc–CORA (N-terminal of the COR domain), and Roc–CORB interactions. These dimeric interfaces are mainly formed by the P-loop, switch II, and a highly conserved Roc dimerization loop between β4 and α3 [32]. An overlay of the CtRoco structures and a monomeric MbRocCORA of Methanosarcina barkeri (Mb) Roco2 structure show there are rearrangements of the P-loop, switch II, and CORA domain, suggesting that they might play an important role during the Roco G-protein cycle [31] (Figure 3A). Initially, it was proposed that Roco proteins form constitutive dimers; however, detailed biophysical and biochemical characterization of the CtRoco proteins revealed a monomer–dimer transition during the G-protein cycle [29,50]. The protein is mainly monomeric in the GTP state, while it forms dimers in the nucleotide-free and GDP-bound states [50]. Furthermore, an analogue of LRRK2 PD-associated mutation stabilizes the dimer and shows decreased GTPase activity [50]. Together, this shows that dimerization is an important step in the Roco activation mechanism.
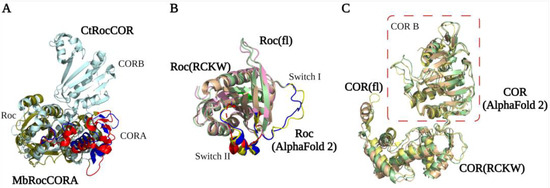
Figure 3.
RocCOR tandem structures. (A) Alignment of the Mb (PDB ID: 4WNR) and Ct (PDB ID: 3DPU) RocCOR domains in dark green and light blue respectively. The rearrangement of P-loop, switch II, and CORA domain are indicated in red (MbRocCORA) and blue (CtRocCOR). (B,C) Alignment of the Roc/COR domain from LRRK2RCKW (PDB ID: 6VP7) (light yellow), flLRRK2 (PDB ID: 7LHW) (pink), and AlphaFold2 (green). The partly and totally missed switch I/II in LRRK2RCKW are highlighted in blue (flLRRK2), red (LRRK2RCKW), and yellow (AlphaFold2), respectively.
The LRRK2RCKW, flLRRK2, and predicted AlphaFold2 RocCOR structures all overlap well, with the exception of some differences observed for the Switch II in the LRRK2RCKW structure (Figure 3B). All three structures show that the COR domain consists of two subdomains that are connected by a flexible linker [23,24]. CORA consists of multiple α-helices and a short three-stranded antiparallel sheet (Figure 3C). The CORB sub-domain consists of four flanked helices, a central seven-stranded antiparallel sheet, and a hairpin motif. Consistent with the bacterial Roco structures, the CORB domain is the main dimerization interface in the flLRRK2 structure, which is formed by two partially overlapped β sheets [24]. In addition, the LRRK2RCKW single-particle and microtubule-associated structures revealed a WD40–WD40 domain interface [23,26]. How these COR–COR and WD40–WD40 interactions contribute to LRRK2 dimerization, and whether the nucleotide-dependent dimerization mechanism of CtRoco is conserved in LRRK2, need to be determined.
2.4. N-Terminal and C-Terminal Scaffold Structure
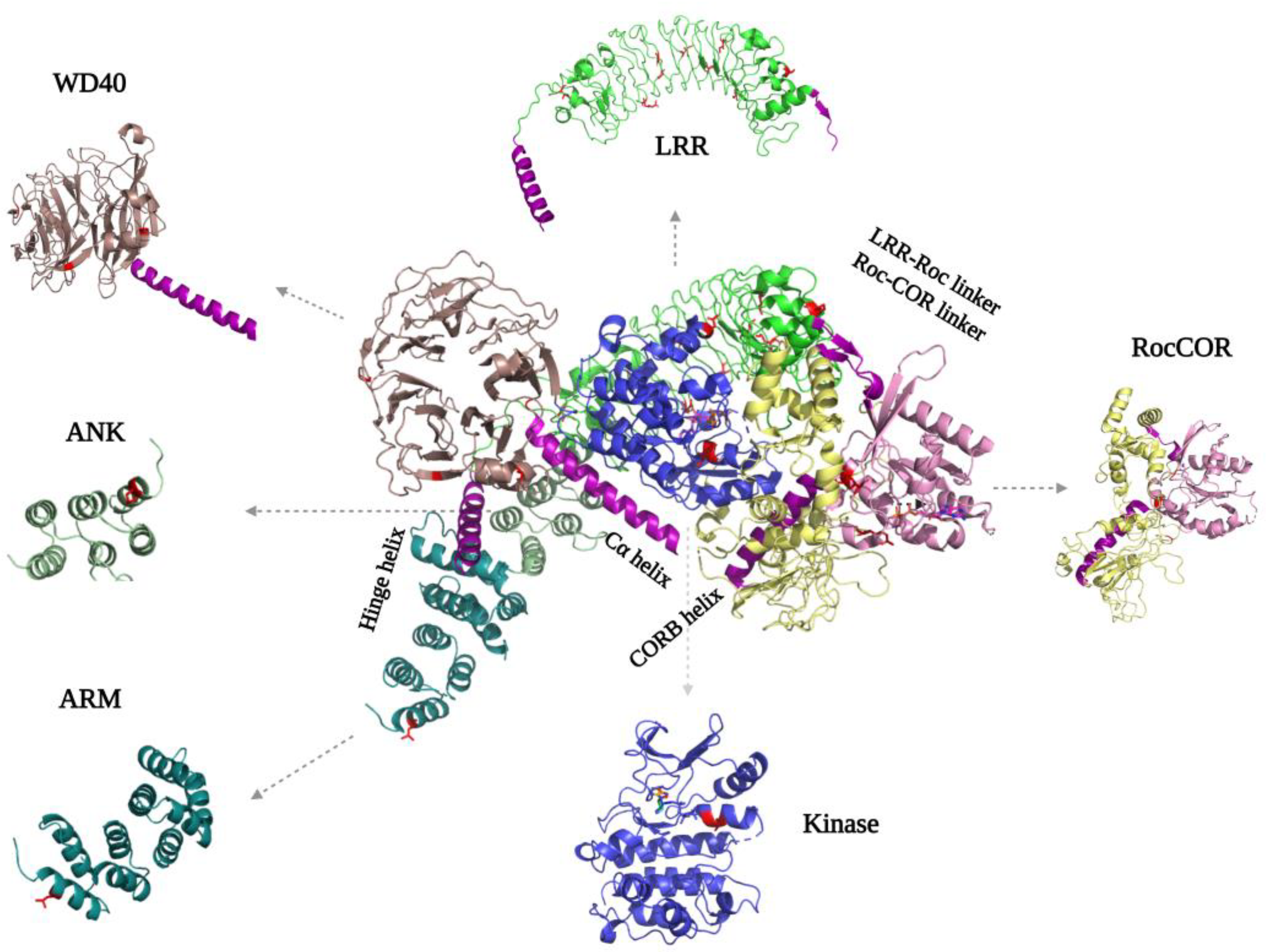
The LRRK2 N-terminal ARM, ANK, LRR, and C-terminal WD40 domains are tandem repeat domains that most likely are involved in PPIs with upstream and downstream regulators [51,52,53]. Recent data suggest that, in addition, these domains have intramolecular interactions that are involved in regulating LRRK2 activity. Despite numerous efforts, structural information about the N-terminal domains remains scarce. Even in the recently published flLRRK2 structure, most of the ARM domain is still not resolved, probably due to its flexibility [24] (Figure 4). The ARM domain that is visible in the structure also does not show the expected typical three-helical structure [51]. The AlphaFold2 model suggests that the LRRK2 ARM domain groove is shallower than in other typical ARM structures, but it does predict 14 repeats in the region of aa49-702 that form a typical three-helix patched solenoid structure, with the shortest helix, H1, perpendicular to the left two antiparallel helices. The H1 and H2 are located at the cylindrical outer surface and are predicted to, together with H3, mediate most of the PPIs [54].
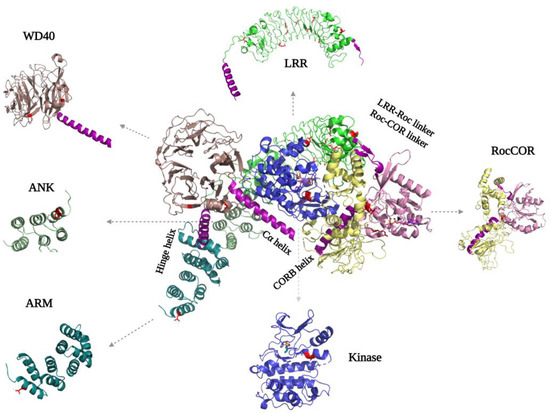
Figure 4.
FlLRRK2 (PDB ID: 7LHW) with the separated domain’s structure on the sides. The helixes that are important for interdomain mediation are highlighted in purple. The PD mutations are indicated in both the full-length structure and separate domains in red, where they are located either in intradomain or interdomain interfaces.
For the ANK domain, it was initially predicted that it consists of seven repeats from aa679-902, while a later study proposed it consists of five repeats from aa690 to aa860 [51,55]. However, both the flLRRK2 and AlphaFold2 structures show that the ANK domain only comprises three typical antiparallel helix repeats (aa705-795) that are connected by a hairpin (Figure 4). The LRRK2 ANK domain forms a typical L-shaped structure with the extended hairpin exposed to the outside to mediate PPIs [56,57,58]. The structures also revealed that the predicted C-terminal segment of the ANK domain (aa795-860) is actually part of the LRR domain, where it functions as a hinge helix that connects the LRR domain with the ANK, ARM, and WD40 domains [24].
The LRR domain of flLRRK2 is folded into a 14 + 1 repeat horse-shaped structure [24]. The beta-strand is located at the concave surface, and it is connected with the convex side α-helix by a loop [24]. After the last strand-helix repeat at the C-terminus of the LRR domain, there is a short linker that crosses the Roc–CORA linker and forms a connection with the Roc domain (Figure 4). In the CtRoco protein, this linker between the LRR and Roc domain is around 40 residues in length and includes a PLxxPPPE motif that is conserved in prokaryotic Roco proteins [29]. This linker in CtRoco was shown to be important for the stability of the Roc–COR construct protein, and the crystal structure showed direct interaction of the linker with the Roc domain switch II motif [29,32]. However, in both the flLRRK2 and AlphaFold2 structures, the linker and LRR domain are far away from the Roc catalytic core.
The crystal structure of the dimeric WD40 domain published by Zhang et al. in 2019 revealed a typical seven blade ring-like structure, each consisting of four anti-parallel β-strands [17] (Figure 4). The WD40–WD40 dimeric interface is formed by the circumference of blades V, IV, and VI, rather than by the popular blade top surfaces. Consistently, this same WD40 dimer interface is present in the microtubule-bound LRRK2 in situ structure and the trimeric LRRK2RCKW EM structure [23,26]. Interestingly, the extended LRRK2RCKW, flLRRK2, and AlphaFold2 structures also show that the C-terminal helix of the WD40 domain directly interacts with the CORB and kinase domain, suggesting that the WD40 domain might regulate the enzymatic activity of LRRK2 [24] (Figure 4).
3. Structural Studies to Understand the LRRK2 Activation Mechanism
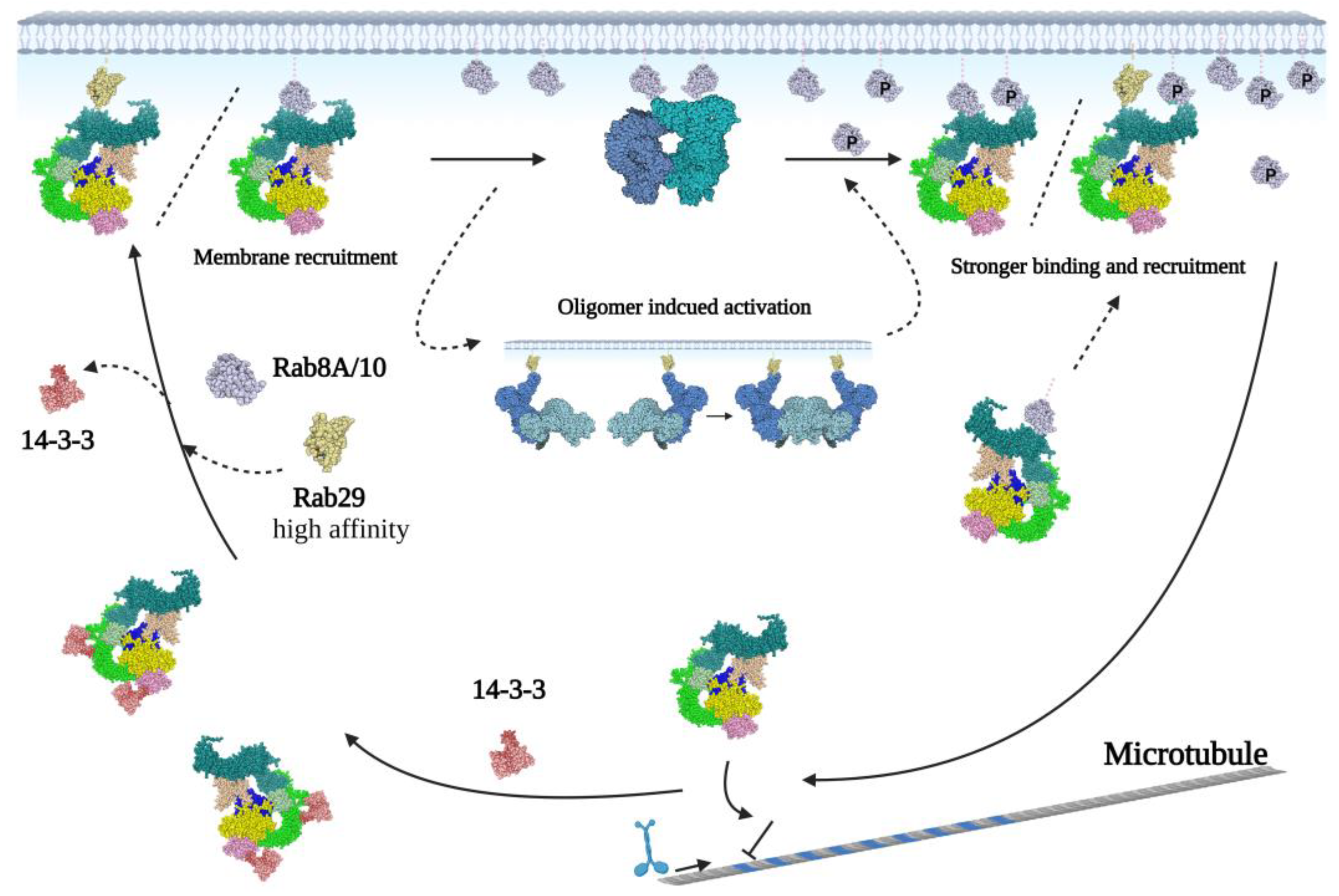
From biochemical and cellular data, it is clear that LRRK2 cycles between a monomeric cytosolic state that has low kinase activity and a high active dimeric membrane bound state [59] (Figure 5). In the cytosol, LRRK2 is most likely GTP-bound, and this monomeric conformation is stabilized by phosphorylation-dependent binding of 14-3-3 to LRRK2 s910 and s935 [60]. The flLRRK2 structures and cross-linking data show that in this inactive state, the WD40 domain is positioned very close to the N-terminal domains and even directly interacts with the N-terminus by the hinge helix (Figure 4). Schmidt et al. hypothesized that the WD40 domain together with the N-terminal domains works as the “brake” of LRRK2 activation by trapping LRRK2 in an inactive monomer state [41]. This hypothesis is also consistent with the active kinase structure and shifted N-terminal conformation in the AlphaFold2 model [41]. The binding of various Rab family proteins to the LRRK2 ARM domain can release this “brake” and induce membrane translocation [41,60,61]. The ARM domain extends to the outside of the LRRK2 dimeric core, far away from the other LRRK2 domains [21,22]. In this conformation, the ARM domain is accessible for binding to Rab proteins, as well as FAS-associated death domain protein (FADD) or with the E3 ubiquitin ligase C-terminus of Hsc 70-interacting protein (CHIP) to induce neuronal death or mediate LRRK2 degradation, respectively [62,63]. The high local concentration of LRRK2 at the membrane most likely induces dimerization of the protein. During dimerization, LRRK2-bound GTP is hydrolyzed and the kinase is fully active for substrate phosphorylation. The recently published Rab29-bound LRRK2 tetramer structure suggests that in addition to dimers, LRRK2 can also form oligomers at the membrane for activation [64]. Furthermore, the phosphorylated-membrane-localized Rab8 and Rab10 can bind to an additional site in the ARM domain, thereby facilitating a much stronger anchor for LRRK2, which results in prolonged LRRK2 activation [65] (Figure 5). From several studies, it is now clear that there is a correlation between GTPase and kinase activity; however, the exact mechanism of this interaction is still not completely understood [66,67]. Multiple studies have shown that Roc–CORA and the kinase domain are not in close proximity and thus cannot directly interact [21]. Instead, a recently published Hydrogen-Deuterium Exchange Mass Spectrometry (HDX-MS) study revealed that the Roc–CORA segment can induce a shift from a compact to an extended conformation during kinase activation [68]. Therefore, the interaction between the Roc and kinase domains may rely on moving the kinase domain to the Roc–CORA domain and simultaneously exposing the CORB domain to allow dimerization [68]. In addition, other domains could also be involved in the communication between the Roc and kinase domain. In the predicted AlphaFold2 model, and flLRRK2 compact and “J” shape structures, an N-terminal–Roc–CORA and a C-terminal CORB–kinase-WD40 axis are visible (Figure 4). The C-terminal CORB–kinase–WD40 axis is connected by a unique α-helix at the C-terminus, which is crucial for the stability of the LRRK2RCKW protein [23]. The two axes cross at the kinase and ANK domain, while the kinase domain is embodied by the solenoid LRR convex side in both the LRRK2 compact and “J” shape structure [21,22,23,24]. This conformation may provide a suitable steric space for the kinase domain and the autophosphorylation clusters from the ANK–LRR and the LRR–Roc linker. It has been shown that autophosphorylation is important for both GTPase function and kinase activity [35].
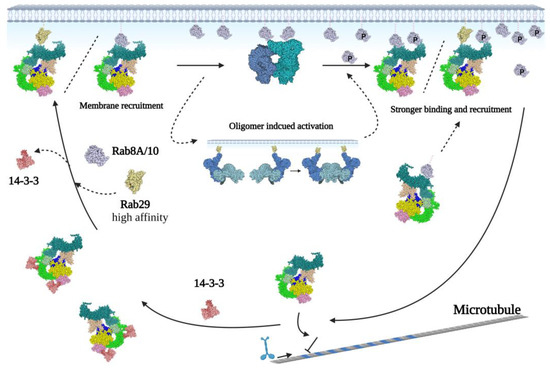
Figure 5.
Proposed model of the LRRK2 activation cycle. LRRK2 is mainly monomeric in the cytosol, a conformation that is stabilized by binding to 14-3-3. Binding of Rab proteins to the ARM domain can induce recruitment of LRRK2 to membranes, where the high concentration of LRRK2 facilitates oligomerization and activation. Subsequently, the phosphorylated substrates, Rab8 and Rab10, can bind to the ARM domain to strengthen the LRRK2 membrane binding, thereby prolonging LRRK2 activation. (LRRK2 is indicated by AlphaFold2 model with: ARM in dark green, ANK in light green, LRR in green, Roc in pink, COR in yellow, kinase in blue, and WD40 in light yellow. The membrane-bounded dimeric/oligomeric protomers are indicated with blue or cyan).
Taken together, this shows that all LRRK2 domains are involved in the activation, phosphorylation, and/or regulation of the cellular localization. However, LRRK2 inactivation, monomerization, and dissociation from the membrane still need to be determined. In the presence of ATP-competitive inhibitors, LRRK2 accumulates in the form of well-ordered filaments on microtubules, where, in turn, it acts as a roadblock for both actin and dynein movement [23]. The mutations within the COR dimeric interface can significantly reduce the microtubule binding [25]. Moreover, a recent microtubule-associated LRRK2RCKW structure revealed that the microtubule binding is mediated by two patches within the Roc domain [25]. However, the accumulation of LRRK2 on microtubules has only been shown in cells that overexpress LRRK2 [69]. Therefore, it is unclear if (and how) this plays a role in PD and targeting of LRRK2.
4. PD Mutation Localization and Potential Pathogenesis
The most common PD mutation, G2019S, is located within the kinase domain and has an increased kinase activity [12] (Figure 1). However, the structural analysis shows no difference between the conformation of the DYGψ motif in the G2019S and WT flLRRK2 structures [24]. Interestingly, the G2019S mutation generates an additional hydrogen bond with residue Q1918 in the αC-helix, which could stabilize the kinase N-lobe and C-lobe in a closed conformation to mediate the kinase activation [41]. This suggests that the G2019S mutation may modify the kinase activity by shifting the inherent active–inactive kinase state equilibrium. The I2020T most likely also stabilizes the kinase domain in an active state via an additional hydrogen bond [41,70]. This mechanism was confirmed by HDX-MS experiments that revealed WT LRRK2 kinase is predominantly in the inactive state, while PD mutations favor the active state [41].
The N2081D mutation is a risk factor for both CD and PD (Figure 1). This mutation has increased kinase activity and decreased acetylation of α-tubulin, which in cells results in an impaired lysosome response and autophagy [15]. Interestingly, N2081 is located in a helix in the kinase domain that is in close proximity to the LRR concave side, suggesting this mutation may alter the kinase activity by controlling the conformation of the N-terminal lid.
Several PD mutations have been identified in the RocCOR tandem domain (Figure 1). Most of these mutations are located in the Roc–COR interface and directly affect the GTPase activity and/or dimerization [68] (Figure 4). As described above, LRRK2 needs to go through a complete monomer–dimer cycle to become completely active. Thus, mutations shifting the equilibrium either to the monomer or dimer will change the LRRK2 activity. The R1441H mutation prolongs the LRRK2 active state and decreases GTPase activity by increasing the nucleotide affinity [49]. So far, there is no evidence that the R1441G/C mutations also affect the GTPase activity. However, it was shown that the R1441 mutations can disrupt the S1444 PKA consensus recognition site, and that 14-3-3 binds to S1444 in a phosphorylation-dependent manner [71]. The R1441 mutations could thus lead to decreased 14-3-3 binding, which, subsequently, can influence LRRK2 translocation and protein stability [72,73]. The LRRK2 Y1699C mutant strengthens the RocCOR intramolecular interaction and weakens LRRK2 dimerization, resulting in decreased GTPase activity [11]. The L487A mutation of CtRoco (analogous to LRRK2 I1371V) is located in the Roc–COR interface and induces stabilization of the dimer, thereby decreasing the GTPase activity [74]. The protective variant located in the RocCOR domain for both PD and CD, R1398H, also shows increased RocCOR dimerization and GTP hydrolysis [15,75].
The G2385R PD mutation is mainly found within the Chinese Han and Korean populations [17,76,77]. It is located in the WD40 domain and affects both the GTPase and kinase activity (Figure 1). Two potential mechanisms for G2385R have been proposed. The first hypothesis is that the G2385R disturbs the WD40 domain interface. The WD40 domain crystal structure indeed shows that G2385R mutation is located in the dimerization interface, and biochemical data suggest that this mutation disturbs LRRK2 dimerization [17]. However, in the flLRRK2 and AlphaFold2 structures, residue G2385 is not situated within the dimer interface, but instead faces the N-terminus. The second proposed mechanism states that the introduced positive charged Arginine disrupts the interaction with proteins, such as 14-3-3, Heat shock protein 90 (Hsp90), and Synaptic Vesicle (SV) proteins [78,79]. These impaired interactions could affect LRRK2 stability and localization, and thereby potentially lead to neurodegeneration. The same mechanism most likely also holds true for a less common WD40 domain-linked PD-mutation, G2294R, which affects the backbone of the WD40 β-propeller and reduces LRRK2 stability [80]. The CD-related mutation M2397T, which is localized at an inserted α-helix in the WD40 propeller blade, prolongs the LRRK2 half-life [81].
Many rare PD variants have been identified in the LRRK2 N-terminus (Figure 1), but the pathogenesis of these variants remains to be determined. However, mechanistically, these mutants most likely change the LRRK2 activation cycle by influencing PPIs. The E139K within the ARM domain could alter LRRK2 binding with Dynamin-related protein 1 (DRP1) and decrease LRRK2 binding with synuclein vesicles to promote vesicle fusion [82,83]. The E334K mutation could affect the ARM domain charge interaction for PPIs, and the mutation L550W could affect FADD binding to induce neuronal cell death [62]. There are also plenty of rare PD mutations in the LRR domain, such as I1006M, R1067Q, S1096C, Q1111H, I1122V, L1165P, I1192V, S1228T, and P1262A [84,85]. Mapping these mutations on the flLRRK2 structure shows that they are either at the convex or concave surface (Figure 4). Those mutations thus likely affect either intramolecular interactions (convex interfaces) or intermolecular interactions (concave surfaces) [86,87]. The only putative PD mutation in the ANK domain is R793M, which is located in the groove surface and close to the LRR-inserted hinge-helix, kinase activation loop, and the WD40 C-terminal helix (Figure 4).
Altogether, these studies show that, although almost all mutants result in increased kinase activity and reduced GTPase activity, the various mutations affect different steps in the activation mechanism.
5. Perspective
Recent developments in structural biology have provided key insights into the LRRK2 activation mechanism and have helped explain the mechanistic effect of important PD LRRK2 mutations. These structures also have revealed that various interfaces on LRRK2 are important for intermolecular and intramolecular interactions, dimerization, and activation. Importantly, these interfaces are potential targets to block LRRK2 activity [88,89]. There are still several important questions to be answered. Multiple studies have shown that LRRK2 cycles between an inactive cytosolic monomeric and active dimer/oligomeric membrane-bound state. However, so far, all high-resolution structures that have been solved are either from LRRK2 in solution or LRRK2 bound to microtubules. Furthermore, since the structure of the N-terminal domain has not been fully resolved, the binding mode of PPIs, and their role in membrane recruitment and intramolecular regulation, are not completely understood. It also remains to be determined which domains are involved in membrane binding and recruitment, how membrane-bound LRRK2 undergoes dimerization, how the cross-talk between the Roc and kinase domain is regulated, how the dynamic kinase conformation changes during substrate loading and phosphorylation, and how exactly the PD mutations affect these different steps in the activation mechanism. Detailed biochemical, biophysical, and structural analysis of LRRK2 interactions with PPIs and structures of the different activation states of LRRK2 bound to membranes will be instrumental for answering these questions.
Author Contributions
Conceptualization, X.Z. and A.K.; writing—original draft preparation, X.Z. and A.K.; writing—review and editing, X.Z. and A.K.; visualization, X.Z. and A.K.; supervision, A.K. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Chinese Scholarship Council for (X.Z 201806250031).
Data Availability Statement
Not applicable.
Acknowledgments
We want to thank Maarten Linskens for carefully reading the manuscript. All figures were created with BioRender.com (accessed on 21 February 2023).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Dorsey, E.; Sherer, T.; Okun, M.; Bloem, B. The Rise of Parkinson’s Disease. Am. Sci. 2020, 108, 176. [Google Scholar] [CrossRef]
- Lee, F.J.S.; Liu, F. Genetic Factors Involved in the Pathogenesis of Parkinson’s Disease. Brain Res. Rev. 2008, 58, 354–364. [Google Scholar] [CrossRef] [PubMed]
- Day, J.O.; Mullin, S. The Genetics of Parkinson’s Disease and Implications for Clinical Practice. Genes 2021, 12, 1006. [Google Scholar] [CrossRef]
- Chandler, R.J.; Cogo, S.; Lewis, P.A.; Kevei, E. Modelling the Functional Genomics of Parkinson’s Disease in Caenorhabditis Elegans: LRRK2 and Beyond. Biosci. Rep. 2021, 41, BSR20203672. [Google Scholar] [CrossRef]
- Di Maio, R.; Hoffman, E.K.; Rocha, E.M.; Keeney, M.T.; Sanders, L.H.; De Miranda, B.R.; Zharikov, A.; Van Laar, A.; Stepan, A.F.; Lanz, T.A.; et al. LRRK2 Activation in Idiopathic Parkinson’s Disease. Sci. Transl. Med. 2018, 10, 451. [Google Scholar] [CrossRef] [PubMed]
- Li, J.Q.; Tan, L.; Yu, J.T. The Role of the LRRK2 Gene in Parkinsonism. Mol. Neurodegener. 2014, 9, 47. [Google Scholar] [CrossRef]
- Steger, M.; Tonelli, F.; Ito, G.; Davies, P.; Trost, M.; Vetter, M.; Wachter, S.; Lorentzen, E.; Duddy, G.; Wilson, S.; et al. Phosphoproteomics Reveals That Parkinson’s Disease Kinase LRRK2 Regulates a Subset of Rab GTPases. Elife 2016, 5. [Google Scholar] [CrossRef]
- Taylor, M.; Alessi, D.R. Advances in Elucidating the Function of Leucine-Rich Repeat Protein Kinase-2 in Normal Cells and Parkinson’s Disease. Curr. Opin. Cell Biol. 2020, 63, 102–113. [Google Scholar] [CrossRef]
- Madureira, M.; Connor-Robson, N.; Wade-Martins, R. LRRK2: Autophagy and Lysosomal Activity. Front. Neurosci. 2020, 14, 498. [Google Scholar] [CrossRef]
- Lewis, P.A.; Greggio, E.; Beilina, A.; Jain, S.; Baker, A.; Cookson, M.R. The R1441C Mutation of LRRK2 Disrupts GTP Hydrolysis. Biochem. Biophys. Res. Commun. 2007, 357, 668–671. [Google Scholar] [CrossRef]
- Daniẽls, V.; Vancraenenbroeck, R.; Law, B.M.H.; Greggio, E.; Lobbestael, E.; Gao, F.; De Maeyer, M.; Cookson, M.R.; Harvey, K.; Baekelandt, V.; et al. Insight into the Mode of Action of the LRRK2 Y1699C Pathogenic Mutant. J. Neurochem. 2011, 116, 304–315. [Google Scholar] [CrossRef] [PubMed]
- West, A.B.; Moore, D.J.; Choi, C.; Andrabi, S.A.; Li, X.; Dikeman, D.; Biskup, S.; Zhang, Z.; Lim, K.L.; Dawson, V.L.; et al. Parkinson’s Disease-Associated Mutations in LRRK2 Link Enhanced GTP-Binding and Kinase Activities to Neuronal Toxicity. Hum. Mol. Genet. 2007, 16, 223–232. [Google Scholar] [CrossRef] [PubMed]
- Van Limbergen, J.; Wilson, D.C.; Satsangi, J. The Genetics of Crohn’s Disease. Annu. Rev. Genomics Hum. Genet. 2009, 10, 89–116. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.-R.; Huang, W.; Chen, S.-M.; Sun, L.-D.; Liu, H.; Li, Y.; Cui, Y.; Yan, X.-X.; Yang, H.-T.; Yang, R.-D.; et al. Genomewide Association Study of Leprosy. N. Engl. J. Med. 2009, 361, 2609–2618. [Google Scholar] [CrossRef] [PubMed]
- Hui, K.Y.; Fernandez-Hernandez, H.; Hu, J.; Schaffner, A.; Pankratz, N.; Hsu, N.Y.; Chuang, L.S.; Carmi, S.; Villaverde, N.; Li, X.; et al. Functional Variants in the LRRK2 Gene Confer Shared Effects on Risk for Crohn’s Disease and Parkinson’s Disease. Sci. Transl. Med. 2018, 10, 423. [Google Scholar] [CrossRef] [PubMed]
- Deng, J.; Lewis, P.A.; Greggio, E.; Sluch, E.; Beilina, A.; Cookson, M.R. Structure of the ROC Domain from the Parkinson’s Disease-Associated Leucine-Rich Repeat Kinase 2 Reveals a Dimeric GTPase. Proc. Natl. Acad. Sci. USA 2008, 105, 1499–1504. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Fan, Y.; Ru, H.; Wang, L.; Magupalli, V.G.; Taylor, S.S.; Alessi, D.R.; Wu, H. Crystal Structure of the WD40 Domain Dimer of LRRK2. Proc. Natl. Acad. Sci. USA 2019, 116, 1579–1584. [Google Scholar] [CrossRef]
- Merk, A.; Bartesaghi, A.; Banerjee, S.; Falconieri, V.; Rao, P.; Davis, M.I.; Pragani, R.; Boxer, M.B.; Earl, L.A.; Milne, J.L.S.; et al. Breaking Cryo-EM Resolution Barriers to Facilitate Drug Discovery. Cell 2016, 165, 1698–1707. [Google Scholar] [CrossRef]
- Lyumkis, D. Challenges and Opportunities in Cryo-EM Single-Particle Analysis. J. Biol. Chem. 2019, 294, 5181–5197. [Google Scholar] [CrossRef]
- Benjin, X.; Ling, L. Developments, Applications, and Prospects of Cryo-Electron Microscopy. Protein Sci. 2020, 29, 872–882. [Google Scholar] [CrossRef]
- Guaitoli, G.; Raimondi, F.; Gilsbach, B.K.; Gómez-Llorente, Y.; Deyaert, E.; Renzi, F.; Li, X.; Schaffner, A.; Jagtap, P.K.A.; Boldt, K.; et al. Structural Model of the Dimeric Parkinson’s Protein LRRK2 Reveals a Compact Architecture Involving Distant Interdomain Contacts. Proc. Natl. Acad. Sci. USA 2016, 113, E4357–E4366. [Google Scholar] [CrossRef] [PubMed]
- Sejwal, K.; Chami, M.; Rémigy, H.; Vancraenenbroeck, R.; Sibran, W.; Sütterlin, R.; Baumgartner, P.; McLeod, R.; Chartier-Harlin, M.C.; Baekelandt, V.; et al. Cryo-EM Analysis of Homodimeric Full-Length LRRK2 and LRRK1 Protein Complexes. Sci. Rep. 2017, 7, 8667. [Google Scholar] [CrossRef] [PubMed]
- Deniston, C.K.; Salogiannis, J.; Mathea, S.; Snead, D.M.; Lahiri, I.; Matyszewski, M.; Donosa, O.; Watanabe, R.; Böhning, J.; Shiau, A.K.; et al. Structure of LRRK2 in Parkinson’s Disease and Model for Microtubule Interaction. Nature 2020, 588, 344–349. [Google Scholar] [CrossRef]
- Myasnikov, A.; Zhu, H.; Hixson, P.; Xie, B.; Yu, K.; Pitre, A.; Peng, J.; Sun, J. Structural Analysis of the Full-Length Human LRRK2. Cell 2021, 184, 3519–3527.e10. [Google Scholar] [CrossRef]
- Snead, D.M.; Matyszewski, M.; Dickey, A.M.; Lin, Y.X.; Leschziner, A.E.; Reck-Peterson, S.L. Structural Basis for Parkinson’s Disease-Linked LRRK2’s Binding to Microtubules. Nat. Struct. Mol. Biol. 2022, 29, 1196–1207. [Google Scholar] [CrossRef]
- Watanabe, R.; Buschauer, R.; Böhning, J.; Audagnotto, M.; Lasker, K.; Lu, T.W.; Boassa, D.; Taylor, S.; Villa, E. The In Situ Structure of Parkinson’s Disease-Linked LRRK2. Cell 2020, 182, 1508–1518.e16. [Google Scholar] [CrossRef] [PubMed]
- Skolnick, J.; Gao, M.; Zhou, H.; Singh, S. AlphaFold 2: Why It Works and Its Implications for Understanding the Relationships of Protein Sequence, Structure, and Function. J. Chem. Inf. Model. 2021, 61, 4827–4831. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Gotthardt, K.; Weyand, M.; Kortholt, A.; Van Haastert, P.J.M.; Wittinghofer, A. Structure of the Roc-COR Domain Tandem of C. Tepidum, a Prokaryotic Homologue of the Human LRRK2 Parkinson Kinase. EMBO J. 2008, 27, 2239–2249. [Google Scholar] [CrossRef]
- Gilsbach, B.K.; Ho, F.Y.; Vetter, I.R.; Van Haastert, P.J.M.; Wittinghofer, A.; Kortholt, A. Roco Kinase Structures Give Insights into the Mechanism of Parkinson Disease-Related Leucine-Rich-Repeat Kinase 2 Mutations. Proc. Natl. Acad. Sci. USA 2012, 109, 10322–10327. [Google Scholar] [CrossRef]
- Terheyden, S.; Ho, F.Y.; Gilsba, B.K.; Wittinghofer, A.; Kortholt, A. Revisiting the Roco G-Protein Cycle. Biochem. J. 2015, 465, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Deyaert, E.; Leemans, M.; Singh, R.K.; Gallardo, R.; Steyaert, J.; Kortholt, A.; Lauer, J.; Versées, W. Structure and Nucleotide-Induced Conformational Dynamics of the Chlorobium Tepidum Roco Protein. Biochem. J. 2019, 476, 51–66. [Google Scholar] [CrossRef] [PubMed]
- Steger, M.; Diez, F.; Dhekne, H.S.; Lis, P.; Nirujogi, R.S.; Karayel, O.; Tonelli, F.; Martinez, T.N.; Lorentzen, E.; Pfeffer, S.R.; et al. Systematic Proteomic Analysis of LRRK2-Mediated Rab GTPase Phosphorylation Establishes a Connection to Ciliogenesis. Elife 2017, 6, e31012. [Google Scholar] [CrossRef]
- Liu, Z.; Mobley, J.A.; Delucas, L.J.; Kahn, R.A.; West, A.B. LRRK2 Autophosphorylation Enhances Its GTPase Activity. FASEB J. 2016, 30, 336–347. [Google Scholar] [CrossRef] [PubMed]
- Webber, P.J.; Smith, A.D.; Sen, S.; Renfrow, M.B.; Mobley, J.A.; West, A.B. Autophosphorylation in the Leucine-Rich Repeat Kinase 2 (LRRK2) GTPase Domain Modifies Kinase and GTP-Binding Activities. J. Mol. Biol. 2011, 412, 94–110. [Google Scholar] [CrossRef]
- Krebs, E.G. An Accidental Biochemist. Annu. Rev. Biochem. 1998, 67, xiii–xxxii. [Google Scholar] [CrossRef]
- Manning, G.; Whyte, D.B.; Martinez, R.; Hunter, T.; Sudarsanam, S. The Protein Kinase Complement of the Human Genome. Science 2002, 298, 1912–1934. [Google Scholar] [CrossRef]
- Taylor, S.S.; Kornev, A.P. Protein Kinases: Evolution of Dynamic Regulatory Proteins. Trends Biochem. Sci. 2011, 36, 65–776. [Google Scholar] [CrossRef]
- Shudler, M.; Niv, M.Y. Blockmaster: Partitioning Protein Kinase Structures Using Normal-Mode Analysis. J. Phys. Chem. A 2009, 113, 7528–7534. [Google Scholar] [CrossRef]
- Modi, V.; Dunbrack, R.L. Defining a New Nomenclature for the Structures of Active and Inactive Kinases. Proc. Natl. Acad. Sci. USA 2019, 116, 6818–6827. [Google Scholar] [CrossRef]
- Schmidt, S.H.; Weng, J.H.; Aoto, P.C.; Boassa, D.; Mathea, S.; Silletti, S.; Hu, J.; Wallbott, M.; Komives, E.A.; Knapp, S.; et al. Conformation and Dynamics of the Kinase Domain Drive Subcellular Location and Activation of LRRK2. Proc. Natl. Acad. Sci. USA 2021, 118, e2100844118. [Google Scholar] [CrossRef] [PubMed]
- Taylor, S.S.; Kaila-Sharma, P.; Weng, J.H.; Aoto, P.; Schmidt, S.H.; Knapp, S.; Mathea, S.; Herberg, F.W. Kinase Domain Is a Dynamic Hub for Driving LRRK2 Allostery. Front. Mol. Neurosci. 2020, 13, 538219. [Google Scholar] [CrossRef]
- Schmidt, S.H.; Knape, M.J.; Boassa, D.; Mumdey, N.; Kornev, A.P.; Ellisman, M.H.; Taylor, S.S.; Herberg, F.W. The Dynamic Switch Mechanism That Leads to Activation of LRRK2 Is Embedded in the DFGψ Motif in the Kinase Domain. Proc. Natl. Acad. Sci. USA 2019, 116, 14979–14988. [Google Scholar] [CrossRef] [PubMed]
- Mills, R.D.; Liang, L.Y.; Lio, D.S.S.; Mok, Y.F.; Mulhern, T.D.; Cao, G.; Griffin, M.; Kenche, V.B.; Culvenor, J.G.; Cheng, H.C. The Roc-COR Tandem Domain of Leucine-Rich Repeat Kinase 2 Forms Dimers and Exhibits Conventional Ras-like GTPase Properties. J. Neurochem. 2018, 147, 409–428. [Google Scholar] [CrossRef]
- Paduch, M.; Jeleń, F.; Otlewski, J. Structure of Small G Proteins and Their Regulators. Acta Biochim. Pol. 2001, 48, 829–850. [Google Scholar] [CrossRef] [PubMed]
- Wittinghofer, A.; Vetter, I.R. Structure-Function Relationships of the G Domain, a Canonical Switch Motif. Annu. Rev. Biochem. 2011, 80, 943–971. [Google Scholar] [CrossRef] [PubMed]
- Bourne, H.R.; Sanders, D.A.; McCormick, F. The GTPase Superfamily: Conserved Structure and Molecular Mechanism. Nature 1991, 349, 125–132. [Google Scholar] [CrossRef]
- Mccormick, F.; Clark, B.F.C.; La Cour, T.F.M.; Kjeldgaard, M.; Norskov-Lauritsen, L.; Nyborg, J. A Model for the Tertiary Structure of P21, the Product of the Ras Oncogene. Science 1985, 230, 78–82. [Google Scholar] [CrossRef]
- Liao, J.; Wu, C.X.; Burlak, C.; Zhang, S.; Sahm, H.; Wang, M.; Zhang, Z.Y.; Vogel, K.W.; Federici, M.; Riddle, S.M.; et al. Parkinson Disease-Associated Mutation R1441H in LRRK2 Prolongs the “Active State” of Its GTPase Domain. Proc. Natl. Acad. Sci. USA 2014, 111, 4055–4060. [Google Scholar] [CrossRef]
- Deyaert, E.; Wauters, L.; Guaitoli, G.; Konijnenberg, A.; Leemans, M.; Terheyden, S.; Petrovic, A.; Gallardo, R.; Nederveen-Schippers, L.M.; Athanasopoulos, P.S.; et al. A Homologue of the Parkinson’s Disease-Associated Protein LRRK2 Undergoes a Monomer-Dimer Transition during GTP Turnover. Nat. Commun. 2017, 8, 1008. [Google Scholar] [CrossRef]
- Mills, R.D.; Mulhern, T.D.; Cheng, H.C.; Culvenor, J.G. Analysis of LRRK2 Accessory Repeat Domains: Prediction of Repeat Length, Number and Sites of Parkinson’s Disease Mutations. Biochem. Soc. Trans. 2012, 40, 1086–1089. [Google Scholar] [CrossRef]
- Rosenbusch, K.E.; Kortholt, A. Activation Mechanism of LRRK2 and Its Cellular Functions in Parkinson’s Disease. Parkinsons. Dis. 2016, 2016, 7351985. [Google Scholar] [CrossRef] [PubMed]
- Piccoli, G.; Onofri, F.; Cirnaru, M.D.; Kaiser, C.J.O.; Jagtap, P.; Kastenmüller, A.; Pischedda, F.; Marte, A.; von Zweydorf, F.; Vogt, A.; et al. Leucine-Rich Repeat Kinase 2 Binds to Neuronal Vesicles through Protein Interactions Mediated by Its C-Terminal WD40 Domain. Mol. Cell. Biol. 2014, 34, 2147–2161. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.J.; Weis, W.I. Structure of the Armadillo Repeat Domain of Plakophilin 1. J. Mol. Biol. 2005, 346, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Cardona, F.; Tormos-Pérez, M.; Pérez-Tur, J. Structural and Functional in Silico Analysis of LRRK2 Missense Substitutions. Mol. Biol. Rep. 2014, 41, 2529–2542. [Google Scholar] [CrossRef] [PubMed]
- Michaely, P.; Tomchick, D.R.; Machius, M.; Anderson, R.G.W. Crystal Structure of a 12 ANK Repeat Stack from Human AnkyrinR. EMBO J. 2002, 21, 6387–6396. [Google Scholar] [CrossRef] [PubMed]
- Mosavi, L.K.; Cammett, T.J.; Desrosiers, D.C.; Peng, Z. The Ankyrin Repeat as Molecular Architecture for Protein Recognition. Protein Sci. 2004, 13, 1435–1448. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Mahajan, A.; Tsai, M.D. Ankyrin Repeat: A Unique Motif Mediating Protein-Protein Interactions. Biochemistry 2006, 45, 15168–15178. [Google Scholar] [CrossRef]
- Berger, Z.; Smith, K.A.; Lavoie, M.J. Membrane Localization of LRRK2 Is Associated with Increased Formation of the Highly Active Lrrk2 Dimer and Changes in Its Phosphorylation. Biochemistry 2010, 49, 5511–5523. [Google Scholar] [CrossRef]
- Manschwetus, J.T.; Wallbott, M.; Fachinger, A.; Obergruber, C.; Pautz, S.; Bertinetti, D.; Schmidt, S.H.; Herberg, F.W. Binding of the Human 14-3-3 Isoforms to Distinct Sites in the Leucine-Rich Repeat Kinase 2. Front. Neurosci. 2020, 14, 302. [Google Scholar] [CrossRef]
- McGrath, E.; Waschbüsch, D.; Baker, B.M.; Khan, A.R. LRRK2 Binds to the Rab32 Subfamily in a GTP-Dependent Manner via Its Armadillo Domain. Small GTPases 2021, 12, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Antoniou, N.; Vlachakis, D.; Memou, A.; Leandrou, E.; Valkimadi, P.E.; Melachroinou, K.; Re, D.B.; Przedborski, S.; Dauer, W.T.; Stefanis, L.; et al. A Motif within the Armadillo Repeat of Parkinson’s-Linked LRRK2 Interacts with FADD to Hijack the Extrinsic Death Pathway. Sci. Rep. 2018, 8, 3455. [Google Scholar] [CrossRef] [PubMed]
- Ding, X.; Goldberg, M.S. Regulation of LRRK2 Stability by the E3 Ubiquitin Ligase CHIP. PLoS ONE 2009, 4, e5949. [Google Scholar] [CrossRef]
- Zhu, H.; Tonelli, F.; Alessi, D.R.; Sun, J. Structural Basis of Human LRRK2 Membrane Recruitment and Activation. bioRxiv 2022. [Google Scholar] [CrossRef]
- Vides, E.G.; Adhikari, A.; Chiang, C.Y.; Lis, P.; Purlyte, E.; Limouse, C.; Shumate, J.L.; Spinola-Lasso, E.; Dhekne, H.S.; Alessi, D.R.; et al. A Feed-Forward Pathway Drives LRRK2 Kinase Membrane Recruitment and Activation. Elife 2022, 11, e79771. [Google Scholar] [CrossRef] [PubMed]
- Taymans, J.M.; Vancraenenbroeck, R.; Ollikainen, P.; Beilina, A.; Lobbestael, E.; de Maeyer, M.; Baekelandt, V.; Cookson, M.R. LRRK2 Kinase Activity Is Dependent on LRRK2 Gtp Binding Capacity but Independent of LRRK2 GTP Binding. PLoS ONE 2011, 6, e23207. [Google Scholar] [CrossRef] [PubMed]
- Ito, G.; Okai, T.; Fujino, G.; Takeda, K.; Ichijo, H.; Katada, T.; Iwatsubo, T. GTP Binding Is Essential to the Protein Kinase Activity of LRRK2, a Causative Gene Product for Familial Parkinson’s Disease. Biochemistry 2007, 46, 1380–1388. [Google Scholar] [CrossRef]
- Weng, J.H.; Aoto, P.C.; Lorenz, R.; Wu, J.; Schmidt, S.H.; Manschwetus, J.T.; Kaila-Sharma, P.; Silletti, S.; Mathea, S.; Chatterjee, D.; et al. LRRK2 Dynamics Analysis Identifies Allosteric Control of the Crosstalk between Its Catalytic Domains. PLoS Biol. 2022, 20, e3001427. [Google Scholar] [CrossRef] [PubMed]
- Kett, L.R.; Boassa, D.; Ho, C.C.Y.; Rideout, H.J.; Hu, J.; Terada, M.; Ellisman, M.; Dauer, W.T. LRRK2 Parkinson Disease Mutations Enhance Its Microtubule Association. Hum. Mol. Genet. 2012, 21, 890–899. [Google Scholar] [CrossRef]
- Gilsbach, B.K.; Kortholt, A. Structural Biology of the LRRK2 GTPase and Kinase Domains: Implications for Regulation. Front. Mol. Neurosci. 2014, 7, 32. [Google Scholar] [CrossRef]
- Muda, K.; Bertinetti, D.; Gesellchen, F.; Hermann, J.S.; Von Zweydorf, F.; Geerlof, A.; Jacob, A.; Ueffing, M.; Gloeckner, C.J.; Herberg, F.W. Parkinson-Related LRRK2 Mutation R1441C/G/H Impairs PKA Phosphorylation of LRRK2 and Disrupts Its Interaction with 14-3-3. Proc. Natl. Acad. Sci. USA 2014, 111, E34–E43. [Google Scholar] [CrossRef] [PubMed]
- Fraser, K.B.; Moehle, M.S.; Daher, J.P.L.; Webber, P.J.; Williams, J.Y.; Stewart, C.A.; Yacoubian, T.A.; Cowell, R.M.; Dokland, T.; Ye, T.; et al. LRRK2 Secretion in Exosomes Is Regulated by 14-3-3. Hum. Mol. Genet. 2013, 22, 4988–5000. [Google Scholar] [CrossRef]
- Zhao, J.; Molitor, T.P.; Langston, J.W.; Nichols, R.J. LRRK2 Dephosphorylation Increases Its Ubiquitination. Biochem. J. 2015, 469, 107–120. [Google Scholar] [CrossRef]
- Rudi, K.; Ho, F.Y.; Gilsbach, B.K.; Pots, H.; Wittinghofer, A.; Kortholt, A.; Klare, J.P. Conformational Heterogeneity of the Roc Domains in C. Tepidum Roc-COR and Implications for Human LRRK2 Parkinson Mutations. Biosci. Rep. 2015, 35, e00254. [Google Scholar] [CrossRef] [PubMed]
- Nixon-Abell, J.; Berwick, D.C.; Grannó, S.; Spain, V.A.; Blackstone, C.; Harvey, K. Protective LRRK2 R1398H Variant Enhances GTPase and Wnt Signaling Activity. Front. Mol. Neurosci. 2016, 9, 18. [Google Scholar] [CrossRef] [PubMed]
- Ho, D.H.; Jang, J.; Joe, E.H.; Son, I.; Seo, H.; Seol, W. G2385R and I2020T Mutations Increase LRRK2 GTPase Activity. Biomed. Res. Int. 2016, 2016, 7917128. [Google Scholar] [CrossRef] [PubMed]
- Di Fonzo, A.; Wu-Chou, Y.H.; Lu, C.S.; Van Doeselaar, M.; Simons, E.J.; Rohé, C.F.; Chang, H.C.; Chen, R.S.; Weng, Y.H.; Vanacore, N.; et al. A Common Missense Variant in the LRRK2 Gene, Gly2385Arg, Associated with Parkinson’s Disease Risk in Taiwan. Neurogenetics 2006, 7, 133–138. [Google Scholar] [CrossRef]
- Rudenko, I.N.; Kaganovich, A.; Hauser, D.N.; Beylina, A.; Chia, R.; Ding, J.; Maric, D.; Jaffe, H.; Cookson, M.R. The G2385R Variant of Leucine-Rich Repeat Kinase 2 Associated with Parkinson’s Disease Is a Partial Loss-of-Function Mutation. Biochem. J. 2012, 446, 99–111. [Google Scholar] [CrossRef]
- Carrion, M.D.P.; Marsicano, S.; Daniele, F.; Marte, A.; Pischedda, F.; Di Cairano, E.; Piovesana, E.; Von Zweydorf, F.; Kremmer, E.; Gloeckner, C.J.; et al. The LRRK2 G2385R Variant Is a Partial Loss-of-Function Mutation That Affects Synaptic Vesicle Trafficking through Altered Protein Interactions. Sci. Rep. 2017, 7, 5377. [Google Scholar] [CrossRef]
- Ogata, J.; Hirao, K.; Nishioka, K.; Hayashida, A.; Li, Y.; Yoshino, H.; Shimizu, S.; Hattori, N.; Imai, Y. A Novel Lrrk2 Variant p.G2294r in the Wd40 Domain Identified in Familial Parkinson’s Disease Affects Lrrk2 Protein Levels. Int. J. Mol. Sci. 2021, 22, 3708. [Google Scholar] [CrossRef]
- Liu, Z.; Lee, J.; Krummey, S.; Lu, W.; Cai, H.; Lenardo, M.J. The Kinase LRRK2 Is a Regulator of the Transcription Factor NFAT That Modulates the Severity of Inflammatory Bowel Disease. Nat. Immunol. 2011, 12, 1063–1070. [Google Scholar] [CrossRef] [PubMed]
- Perez Carrion, M.; Pischedda, F.; Biosa, A.; Russo, I.; Straniero, L.; Civiero, L.; Guida, M.; Gloeckner, C.J.; Ticozzi, N.; Tiloca, C.; et al. The LRRK2 Variant E193K Prevents Mitochondrial Fission upon MPP+ Treatment by Altering LRRK2 Binding to DRP1. Front. Mol. Neurosci. 2018, 11, 64. [Google Scholar] [CrossRef] [PubMed]
- Marku, A.; Carrion, M.D.P.; Pischedda, F.; Marte, A.; Casiraghi, Z.; Marciani, P.; von Zweydorf, F.; Gloeckner, C.J.; Onofri, F.; Perego, C.; et al. The LRRK2 N-Terminal Domain Influences Vesicle Trafficking: Impact of the E193K Variant. Sci. Rep. 2020, 10, 3799. [Google Scholar] [CrossRef]
- Berg, D.; Schweitzer, K.; Leitner, P.; Zimprich, A.; Lichtner, P.; Belcredi, P.; Brussel, T.; Schulte, C.; Maass, S.; Nagele, T. Type and Frequency of Mutations in the LRRK2 Gene in Familial and Sporadic Parkinson’s Disease*. Brain 2005, 128, 3000–3011. [Google Scholar] [CrossRef] [PubMed]
- Skipper, L.; Shen, H.; Chua, E.; Bonnard, C.; Kolatkar, P.; Tan, L.C.S.; Jamora, R.D.; Puvan, K.; Puong, K.Y.; Zhao, Y.; et al. Analysis of LRRK2 Functional Domains in Nondominant Parkinson Disease. Neurology 2005, 65, 1319–1321. [Google Scholar] [CrossRef]
- Vancraenenbroeck, R.; Lobbestael, E.; Weeks, S.D.; Strelkov, S.V.; Baekelandt, V.; Taymans, J.M.; De Maeyer, M. Expression, Purification and Preliminary Biochemical and Structural Characterization of the Leucine Rich Repeat Namesake Domain of Leucine Rich Repeat Kinase 2. Biochim. Biophys. Acta Proteins Proteom. 2012, 1824, 450–460. [Google Scholar] [CrossRef]
- De Wit, J.; Hong, W.; Luo, L.; Ghosh, A. Role of Leucine-Rich Repeat Proteins in the Development and Function of Neural Circuits. Annu. Rev. Cell Dev. Biol. 2011, 27, 697–729. [Google Scholar] [CrossRef]
- Helton, L.G.; Soliman, A.; Von Zweydorf, F.; Kentros, M.; Manschwetus, J.T.; Hall, S.; Gilsbach, B.; Ho, F.Y.; Athanasopoulos, P.S.; Singh, R.K.; et al. Allosteric Inhibition of Parkinson’s-Linked LRRK2 by Constrained Peptides. ACS Chem. Biol. 2021, 16, 2326–2338. [Google Scholar] [CrossRef]
- Soliman, A.; Cankara, F.N.; Kortholt, A. Allosteric Inhibition of LRRK2, Where Are We Now. Biochem. Soc. Trans. 2020, 48, 2185–2194. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).