The Role of LSD1 and LSD2 in Cancers of the Gastrointestinal System: An Update
Abstract
1. Introduction
2. Gastric Cancer
2.1. LSD1 in Gastric Cancer
2.1.1. LSD1 and LincRNAFEZF1-AS1
2.1.2. LSD1 and lncRNA HOXA11-AS
2.1.3. LSD1 and Long Noncoding RNA FOXD2-AS1
2.1.4. LSD1 and LINC00673
2.2. LSD2 in Gastric Cancer
3. Liver Cancer
3.1. LSD1 in Liver Cancer
LSD1 Inhibition as a Treatment Strategy for Liver Cancer
3.2. LSD2 in Liver Cancer
4. Pancreatic Cancer
5. Colorectal Cancer
5.1. LSD1 in Colorectal Cancer
LSD1-Associated Noncoding RNAs Involved in Colorectal Cancer Progression
5.2. LSD2 in Colorectal Cancer
6. Therapeutic Implications of LSD1 and LSD2 in Various Cancers
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Conflicts of Interest
References
- Cancer. 2022. Available online: https://www.who.int/news-room/fact-sheets/detail/cancer (accessed on 15 January 2022).
- Global Cancer Observatory. International Agency for Research on Cancer. 2022. Available online: https://gco.iarc.fr/ (accessed on 15 January 2022).
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Hanahan, D. Hallmarks of Cancer: New Dimensions. Cancer Discov. 2022, 12, 31–46. [Google Scholar] [CrossRef] [PubMed]
- Nirmaladevi, R.; Ilango, S.; Paital, B.; Jayachandran, P.; Padma, P.R. Epigenetic alterations in cancer. Front. Biosci. 2020, 25, 1058–1109. [Google Scholar] [CrossRef] [PubMed]
- Burg, J.M.; Link, J.E.; Morgan, B.; Heller, F.J.; Hargrove, A.E.; McCafferty, D.G. KDM1 class flavin-dependent protein lysine demethylases. Biopolymers 2015, 104, 213–246. [Google Scholar] [CrossRef] [PubMed]
- Maiques-Diaz, A.; Somervaille, T.C. LSD1: Biologic roles and therapeutic targeting. Epigenomics 2016, 8, 1103–1116. [Google Scholar] [CrossRef] [PubMed]
- Perillo, B.; Tramontano, A.; Pezone, A.; Migliaccio, A. LSD1: More than demethylation of histone lysine residues. Exp. Mol. Med. 2020, 52, 1936–1947. [Google Scholar] [CrossRef] [PubMed]
- Niwa, H.; Sato, S.; Hashimoto, T.; Matsuno, K.; Umehara, T. Crystal Structure of LSD1 in Complex with 4-[5-(Piperidin-4-ylmethoxy)-2-(p-tolyl)pyridin-3-yl]benzonitrile. Molecules 2018, 23, 1538. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Vasilatos, S.N.; Qin, Y.; Katz, T.A.; Cao, C.; Wu, H.; Tasdemir, N.; Levine, K.M.; Oesterreich, S.; Davidson, N.E.; et al. Functional characterization of lysine-specific demethylase 2 (LSD2/KDM1B) in breast cancer progression. Oncotarget 2017, 8, 81737–81753. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Yang, H. Upregulation of the long noncoding RNA ADPGK-AS1 promotes carcinogenesis and predicts poor prognosis in gastric cancer. Biochem. Biophys. Res. Commun. 2019, 513, 127–134. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Yang, H.; Dong, Z.; Fang, J.; Wang, P.; Zhu, T.; Gong, W.; Fang, R.; Shi, Y.G.; Li, Z.; et al. Structural insight into substrate recognition by histone demethylase LSD2/KDM1b. Cell Res. 2013, 23, 306–309. [Google Scholar] [CrossRef]
- Han, D.; Huang, M.; Wang, T.; Li, Z.; Chen, Y.; Liu, C.; Lei, Z.; Chu, X. Lysine methylation of transcription factors in cancer. Cell Death Dis. 2019, 10, 290. [Google Scholar] [CrossRef] [PubMed]
- Marayati, B.F.; Tucker, J.F.; De La Cerda, D.A.; Hou, T.-C.; Chen, R.; Sugiyama, T.; Pease, J.B.; Zhang, K. The Catalytic-Dependent and -Independent Roles of Lsd1 and Lsd2 Lysine Demethylases in Heterochromatin Formation in Schizosaccharomyces pombe. Cells 2020, 9, 955. [Google Scholar] [CrossRef] [PubMed]
- Wojcieszyńska, D.; Hupert-Kocurek, K.; Guzik, U. Flavin-Dependent Enzymes in Cancer Prevention. Int. J. Mol. Sci. 2012, 13, 16751–16768. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Sengupta, R.; Espejo, A.; Lee, M.G.; Dorsey, J.A.; Richter, M.; Opravil, S.; Shiekhattar, R.; Bedford, M.T.; Jenuwein, T.; et al. p53 is regulated by the lysine demethylase LSD1. Nature 2007, 449, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.-S.; Suzuki, T.; Dohmae, N.; Hayami, S.; Unoki, M.; Yoshimatsu, M.; Toyokawa, G.; Takawa, M.; Chen, T.; Kurash, J.K.; et al. Demethylation of RB Regulator MYPT1 by Histone Demethylase LSD1 Promotes Cell Cycle Progression in Cancer Cells. Cancer Res. 2010, 71, 655–660. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Hoang, N.; Leng, F.; Saxena, L.; Lee, L.; Alejo, S.; Qi, D.; Khal, A.; Sun, H.; Lu, F.; et al. LSD1 demethylase and the methyl-binding protein PHF20L1 prevent SET7 methyltransferase–dependent proteolysis of the stem-cell protein SOX2. J. Biol. Chem. 2018, 293, 3663–3674. [Google Scholar] [CrossRef]
- Fujikuni, N.; Tanabe, K.; Hattori, M.; Yamamoto, Y.; Tazawa, H.; Toyota, K.; Tokumoto, N.; Hotta, R.; Yanagawa, S.; Saeki, Y.; et al. Distal Gastrectomy for Symptomatic Stage IV Gastric Cancer Contributes to Prognosis with Acceptable Safety Compared to Gastrojejunostomy. Cancers 2022, 14, 388. [Google Scholar] [CrossRef]
- Salati, M.; Spallanzani, A. New Horizons for Personalised Treatment in Gastroesophageal Cancer. J. Clin. Med. 2022, 11, 311. [Google Scholar] [CrossRef]
- Liu, Y.-W.; Xia, R.; Lu, K.; Xie, M.; Yang, F.; Sun, M.; De, W.; Wang, C.; Ji, G. LincRNAFEZF1-AS1 represses p21 expression to promote gastric cancer proliferation through LSD1-Mediated H3K4me2 demethylation. Mol. Cancer 2017, 16, 39. [Google Scholar] [CrossRef]
- Majello, B.; Gorini, F.; Saccà, C.D.; Amente, S. Expanding the Role of the Histone Lysine-Specific Demethylase LSD1 in Cancer. Cancers 2019, 11, 324. [Google Scholar] [CrossRef]
- Collette, J.; Le Bourhis, X.; Adriaenssens, E. Regulation of Human Breast Cancer by the Long Non-Coding RNA H19. Int. J. Mol. Sci. 2017, 18, 2319. [Google Scholar] [CrossRef] [PubMed]
- Huang, M.; Hou, J.; Wang, Y.; Xie, M.; Wei, C.; Nie, F.; Wang, Z.; Sun, M. Long Noncoding RNA LINC00673 Is Activated by SP1 and Exerts Oncogenic Properties by Interacting with LSD1 and EZH2 in Gastric Cancer. Mol. Ther. 2017, 25, 1014–1026. [Google Scholar] [CrossRef] [PubMed]
- Takeiwa, T.; Ikeda, K.; Mitobe, Y.; Horie-Inoue, K.; Inoue, S. Long Noncoding RNAs Involved in the Endocrine Therapy Resistance of Breast Cancer. Cancers 2020, 12, 1424. [Google Scholar] [CrossRef]
- Ocker, M.; Al Bitar, S.; Monteiro, A.C.; Gali-Muhtasib, H.; Schneider-Stock, R. Epigenetic Regulation of p21cip1/waf1 in Human Cancer. Cancers 2019, 11, 1343. [Google Scholar] [CrossRef] [PubMed]
- Shin, T.-J.; Lee, K.-H.; Cho, J.-Y. Epigenetic Mechanisms of LncRNAs Binding to Protein in Carcinogenesis. Cancers 2020, 12, 2925. [Google Scholar] [CrossRef] [PubMed]
- Etani, T.; Naiki, T.; Naiki-Ito, A.; Suzuki, T.; Iida, K.; Nozaki, S.; Kato, H.; Nagayasu, Y.; Suzuki, S.; Kawai, N.; et al. NCL1, A Highly Selective Lysine-Specific Demethylase 1 Inhibitor, Suppresses Castration-Resistant Prostate Cancer Growth via Regulation of Apoptosis and Autophagy. J. Clin. Med. 2019, 8, 442. [Google Scholar] [CrossRef] [PubMed]
- Bure, I.; Nemtsova, M. Methylation and Noncoding RNAs in Gastric Cancer: Everything Is Connected. Int. J. Mol. Sci. 2021, 22, 5683. [Google Scholar] [CrossRef]
- Sun, M.; Nie, F.; Wang, Y.; Zhang, Z.; Hou, J.; He, D.; Xie, M.; Xu, L.; De, W.; Wang, Z.; et al. LncRNA HOXA11-AS Promotes Proliferation and Invasion of Gastric Cancer by Scaffolding the Chromatin Modification Factors PRC2, LSD1, and DNMT1. Cancer Res. 2016, 76, 6299–6310. [Google Scholar] [CrossRef] [PubMed]
- Abdullah, O.; Alhosin, M. HAUSP Is a Key Epigenetic Regulator of the Chromatin Effector Proteins. Genes 2022, 13, 42. [Google Scholar] [CrossRef]
- Morlando, M.; Fatica, A. Alteration of Epigenetic Regulation by Long Noncoding RNAs in Cancer. Int. J. Mol. Sci. 2018, 19, 570. [Google Scholar] [CrossRef]
- Uchida, S.; Kauppinen, S. Long Non-Coding RNAs in Liver Cancer and Nonalcoholic Steatohepatitis. Non-Coding RNA 2020, 6, 34. [Google Scholar] [CrossRef]
- Han, T.-S.; Hur, K.; Cho, H.-S.; Ban, H.S. Epigenetic Associations between lncRNA/circRNA and miRNA in Hepatocellular Carcinoma. Cancers 2020, 12, 2622. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.-P.; Wang, W.-Y.; Ma, P.; Shuai, Y.; Zhao, K.; Wang, Y.-F.; Li, W.; Xia, R.; Chen, W.-M.; Zhang, E.-B.; et al. Upregulation of the long noncoding RNA FOXD2-AS1 promotes carcinogenesis by epigenetically silencing EphB3 through EZH2 and LSD1, and predicts poor prognosis in gastric cancer. Oncogene 2018, 37, 5020–5036. [Google Scholar] [CrossRef]
- Schwarzmueller, L.; Bril, O.; Vermeulen, L.; Léveillé, N. Emerging Role and Therapeutic Potential of lncRNAs in Colorectal Cancer. Cancers 2020, 12, 3843. [Google Scholar] [CrossRef]
- Siddaiah, R.; Oji-Mmuo, C.; Montes, D.; Fuentes, N.; Spear, D.; Donnelly, A.; Silveyra, P. MicroRNA Signatures Associated with Bronchopulmonary Dysplasia Severity in Tracheal Aspirates of Preterm Infants. Biomedicines 2021, 9, 257. [Google Scholar] [CrossRef] [PubMed]
- Barth, D.; Juracek, J.; Slaby, O.; Pichler, M.; Calin, G. lncRNA and Mechanisms of Drug Resistance in Cancers of the Genitourinary System. Cancers 2020, 12, 2148. [Google Scholar] [CrossRef] [PubMed]
- Gianì, F.; Vella, V.; Tumino, D.; Malandrino, P.; Frasca, F. The Possible Role of Cancer Stem Cells in the Resistance to Kinase Inhibitors of Advanced Thyroid Cancer. Cancers 2020, 12, 2249. [Google Scholar] [CrossRef] [PubMed]
- Su, S.-C.; Lin, C.-W.; Ju, P.-C.; Chang, L.-C.; Chuang, C.-Y.; Liu, Y.-F.; Hsieh, M.-J.; Yang, S.-F. Association of LINC00673 Genetic Variants with Progression of Oral Cancer. J. Pers. Med. 2021, 11, 468. [Google Scholar] [CrossRef] [PubMed]
- Jing, R.; Liu, S.; Jiang, Y.; Zong, W.; Ju, S.; Cui, M. Determination of serum RP11-731F5.2 as a noninvasive biomarker for gastric cancer diagnosis and prognosis. Pathol. Res. Pract. 2020, 216, 153261. [Google Scholar] [CrossRef]
- Jiang, H.-Y.; Wang, Z.-J. ADPGK-AS1 promotes the progression of colorectal cancer via sponging miR-525 to upregulate FUT1. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 2380–2386. [Google Scholar] [CrossRef]
- Hu, Z.Q.; Li, H.C.; Teng, F.; Chang, Q.M.; Wu, X.B.; Feng, J.F.; Zhang, Z.P. Long noncoding RNA MAFG-AS1 facilitates the progression of hepatocellular carcinoma via targeting miR-3196/OTX1 axis. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 12131–12143. [Google Scholar] [CrossRef] [PubMed]
- Cai, S.; Wang, J.; Zeng, W.; Cheng, X.; Liu, L.; Li, W. Lysine-specific histone demethylase 1B (LSD2/KDM1B) represses p53 expression to promote proliferation and inhibit apoptosis in colorectal cancer through LSD2-mediated H3K4me2 demethylation. Aging 2020, 12, 14990–15001. [Google Scholar] [CrossRef]
- Llovet, J.M.; Kelley, R.K.; Villanueva, A.; Singal, A.G.; Pikarsky, E.; Roayaie, S.; Lencioni, R.; Koike, K.; Zucman-Rossi, J.; Finn, R.S. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 2021, 7, 6. [Google Scholar] [CrossRef]
- Caines, A.; Selim, R.; Salgia, R. The Changing Global Epidemiology of Hepatocellular Carcinoma. Clin. Liver Dis. 2020, 24, 535–547. [Google Scholar] [CrossRef]
- Marcon, P.D.S.; Tovo, C.V.; Kliemann, D.A.; Fisch, P.; De Mattos, A.A. Incidence of hepatocellular carcinoma in patients with chronic liver disease due to hepatitis B or C and coinfected with the human immunodeficiency virus: A retrospective cohort study. World J. Gastroenterol. 2018, 24, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Ochiai, Y.; Kawamura, Y.; Kobayashi, M.; Shindoh, J.; Kobayashi, Y.; Okubo, S.; Muraishi, N.; Kajiwara, A.; Iritani, S.; Fujiyama, S.; et al. Effects of alcohol consumption on multiple hepatocarcinogenesis in patients with fatty liver disease. Hepatol. Res. 2021, 51, 62–68. [Google Scholar] [CrossRef] [PubMed]
- Ayuso, C.; Rimola, J.; Vilana, R.; Burrel, M.; Darnell, A.; García-Criado, Á.; Bianchi, L.; Belmonte, E.; Caparroz, C.; Barrufet, M.; et al. Diagnosis and staging of hepatocellular carcinoma (HCC): Current guidelines. Eur. J. Radiol. 2018, 101, 72–81. [Google Scholar] [CrossRef]
- Vogel, A.; Cervantes, A.; Chau, I.; Daniele, B.; Llovet, J.M.; Meyer, T.; Nault, J.-C.; Neumann, U.; Ricke, J.; Sangro, B.; et al. Hepatocellular carcinoma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 2018, 29, iv238–iv255. [Google Scholar] [CrossRef]
- Lee, G.C.; Ferrone, C.R.; Vagefi, P.A.; Uppot, R.N.; Tanabe, K.K.; Lillemoe, K.D.; Blaszkowsky, L.S.; Qadan, M. Surgical resection versus ablation for early-stage hepatocellular carcinoma: A retrospective cohort analysis. Am. J. Surg. 2019, 218, 157–163. [Google Scholar] [CrossRef] [PubMed]
- Mazzarelli, C.; Cannon, M.D.; Hudson, M.; Heaton, N.; Sarker, D.; Kane, P.; Quaglia, A.; Suddle, A. Hepatocellular Carcinoma as a Complication of Vascular Disease of the Liver After Fontan Procedure. Hepatology 2018, 69, 911–913. [Google Scholar] [CrossRef]
- Rimassa, L.; Personeni, N.; Czauderna, C.; Foerster, F.; Galle, P. Systemic treatment of HCC in special populations. J. Hepatol. 2021, 74, 931–943. [Google Scholar] [CrossRef] [PubMed]
- Tsuchiya, H.; Shiota, G. Clinical and Biological Implications of Cancer Stem Cells in Hepatocellular Carcinoma. Yonago Acta Med. 2021, 64, 1–11. [Google Scholar] [CrossRef]
- Hayami, S.; Kelly, J.D.; Cho, H.-S.; Yoshimatsu, M.; Unoki, M.; Tsunoda, T.; Field, H.I.; Neal, D.E.; Yamaue, H.; Ponder, B.A.; et al. Overexpression of LSD1 contributes to human carcinogenesis through chromatin regulation in various cancers. Int. J. Cancer 2011, 128, 574–586. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Wang, Y.; Yang, X.H.; Kang, T.; Zhao, Y.; Wang, C.; Evers, B.M.; Zhou, B.P. The Deubiquitinase USP28 Stabilizes LSD1 and Confers Stem-Cell-like Traits to Breast Cancer Cells. Cell Rep. 2013, 5, 224–236. [Google Scholar] [CrossRef]
- Adamo, A.; Barrero, M.; Belmonte, J.C.I. LSD1 and pluripotency: A new player in the network. Cell Cycle 2011, 10, 3215–3216. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Whyte, W.A.; Bilodeau, S.; Orlando, D.A.; Hoke, H.A.; Frampton, G.M.; Foster, C.T.; Cowley, S.M.; Young, R.A. Author Correction: Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature 2018, 562, E24. [Google Scholar] [CrossRef] [PubMed]
- Thambyrajah, R.; Mazan, M.; Patel, R.; Moignard, V.R.; Stefańska, M.; Marinopoulou, E.; Li, Y.; Lancrin, C.; Clapes, T.; Möröy, T.; et al. GFI1 proteins orchestrate the emergence of haematopoietic stem cells through recruitment of LSD1. Nat. Cell Biol. 2016, 18, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Choi, H.-J.; Park, J.-H.; Park, M.; Won, H.-Y.; Joo, H.-S.; Lee, C.H.; Lee, J.-Y.; Kong, G. UTX inhibits EMT-induced breast CSC properties by epigenetic repression of EMT genes in cooperation with LSD1 and HDAC1. EMBO Rep. 2015, 16, 1288–1298. [Google Scholar] [CrossRef] [PubMed]
- Mohammad, H.P.; Smitheman, K.N.; Kamat, C.D.; Soong, D.; Federowicz, K.E.; Van Aller, G.S.; Schneck, J.L.; Carson, J.D.; Liu, Y.; Butticello, M.; et al. A DNA Hypomethylation Signature Predicts Antitumor Activity of LSD1 Inhibitors in SCLC. Cancer Cell 2015, 28, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Lei, Z.-J.; Wang, J.; Xiao, H.-L.; Guo, Y.; Wang, T.; Li, Q.; Liu, L.; Luo, X.; Fan, L.-L.; Lin, L.; et al. Lysine-specific demethylase 1 promotes the stemness and chemoresistance of Lgr5+ liver cancer initiating cells by suppressing negative regulators of β-catenin signaling. Oncogene 2015, 34, 3188–3198. [Google Scholar] [CrossRef]
- Zhao, Z.-K.; Dong, P.; Gu, J.; Chen, L.; Zhuang, M.; Lu, W.-J.; Wang, D.-R.; Liu, Y.-B. Overexpression of LSD1 in hepatocellular carcinoma: A latent target for the diagnosis and therapy of hepatoma. Tumor Biol. 2013, 34, 173–180. [Google Scholar] [CrossRef]
- Kim, S.; Bolatkan, A.; Kaneko, S.; Ikawa, N.; Asada, K.; Komatsu, M.; Hayami, S.; Ojima, H.; Abe, N.; Yamaue, H.; et al. Deregulation of the Histone Lysine-Specific Demethylase 1 Is Involved in Human Hepatocellular Carcinoma. Biomolecules 2019, 9, 810. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-Y.; Park, J.-H.; Choi, H.-J.; Won, H.-Y.; Joo, H.-S.; Shin, D.-H.; Park, M.K.; Han, B.; Kim, K.P.; Lee, T.J.; et al. LSD1 demethylates HIF1α to inhibit hydroxylation and ubiquitin-mediated degradation in tumor angiogenesis. Oncogene 2017, 36, 5512–5521. [Google Scholar] [CrossRef]
- Mould, D.; McGonagle, A.E.; Wiseman, D.; Williams, E.L.; Jordan, A.M. Reversible Inhibitors of LSD1 as Therapeutic Agents in Acute Myeloid Leukemia: Clinical Significance and Progress to Date. Med. Res. Rev. 2015, 35, 586–618. [Google Scholar] [CrossRef]
- Fang, Y.; Liao, G.; Yu, B. LSD1/KDM1A inhibitors in clinical trials: Advances and prospects. J. Hematol. Oncol. 2019, 12, 129. [Google Scholar] [CrossRef]
- Ma, L.; Xu, A.; Kang, L.; Cong, R.; Fan, Z.; Zhu, X.; Huo, N.; Liu, W.; Xue, C.; Ji, Q.; et al. LSD1 -Demethylated LINC01134 Confers Oxaliplatin Resistance Through SP1 -Induced p62 Transcription in HCC. Hepatology 2021, 74, 3213–3234. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Lv, R.; Wu, T.; Zhang, X.; Sun, Y.; Yan, J.; Zhang, Z.; Zhao, D.; Cheng, M. Design, synthesis, and biological evaluation of coumarin analogs as novel LSD1 inhibitors. Arch. Pharm. 2021, 355, e2100311. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Huang, H.; Zhang, Z.; Yan, J.; Wu, T.; Yin, W.; Sun, Y.; Wang, X.; Gu, Y.; Zhao, D.; et al. Design, synthesis and biological evaluation of novel benzofuran derivatives as potent LSD1 inhibitors. Eur. J. Med. Chem. 2021, 220, 113501. [Google Scholar] [CrossRef]
- Ding, L.; Wei, F.; Wang, N.; Sun, Y.; Wang, Q.; Fan, X.; Qi, L.; Wang, S. Tertiary sulphonamide derivatives as dual acting small molecules that inhibit LSD1 and suppress tubulin polymerisation against liver cancer. J. Enzym. Inhib. Med. Chem. 2021, 36, 1562–1571. [Google Scholar] [CrossRef] [PubMed]
- Bayo, J.; Fiore, E.J.; Dominguez, L.M.; Real, A.; Malvicini, M.; Rizzo, M.; Atorrasagasti, C.; García, M.G.; Argemi, J.; Martinez, E.D.; et al. A comprehensive study of epigenetic alterations in hepatocellular carcinoma identifies potential therapeutic targets. J. Hepatol. 2019, 71, 78–90. [Google Scholar] [CrossRef]
- Huang, M.; Chen, C.; Geng, J.; Han, D.; Wang, T.; Xie, T.; Wang, L.; Wang, Y.; Wang, C.; Lei, Z.; et al. Targeting KDM1A attenuates Wnt/β-catenin signaling pathway to eliminate sorafenib-resistant stem-like cells in hepatocellular carcinoma. Cancer Lett. 2017, 398, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Lian, Y.; Li, Z.; Fan, Y.; Huang, Q.; Chen, J.; Liu, W.; Xu, H. The lncRNA-HOXA-AS2/EZH2/LSD1 oncogene complex promotes cell proliferation in pancreatic cancer. Am. J. Transl. Res. 2017, 9, 5496–5506. [Google Scholar]
- Lian, Y.; Xiao, C.; Yan, C.; Chen, D.; Huang, Q.; Fan, Y.; Xu, H. Knockdown of pseudogene derived from lncRNA DUXAP10 inhibits cell proliferation, migration, invasion, and promotes apoptosis in pancreatic cancer. J. Cell Biochem. 2018, 119, 3671–3682. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Sun, L.; Luo, Y.; He, S. Knockdown of KDM1B inhibits cell proliferation and induces apoptosis of pancreatic cancer cells. Pathol. Res. Pract. 2019, 215, 1054–1060. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Zhao, J.; Ding, J.; Wang, Z.; Du, J.; Wu, C. Knocking down LSD1 inhibits the stemness features of colorectal cancer stem cells. Braz. J. Med. Biol. Res. 2020, 53, e9230. [Google Scholar] [CrossRef]
- Chen, J.; Ding, J.; Wang, Z.; Zhu, J.; Wang, X.; Du, J. Identification of downstream metastasis-associated target genes regulated by LSD1 in colon cancer cells. Oncotarget 2017, 8, 19609–19630. [Google Scholar] [CrossRef]
- Miller, S.A.; Policastro, R.A.; Sriramkumar, S.; Lai, T.; Huntington, T.D.; Ladaika, C.A.; Kim, D.; Hao, C.; Zentner, G.E.; O’Hagan, H.M. LSD1 and aberrant DNA methylation mediate persistence of enteroendocrine progenitors that support BRAF mutant colorectal cancer. Cancer Res. 2021, 81, 3791–3805. [Google Scholar] [CrossRef]
- Ramírez-Ramírez, R.; Gutiérrez-Angulo, M.; Peregrina-Sandoval, J.; Moreno-Ortiz, J.M.; Franco-Topete, R.A.; Cerda-Camacho, F.D.J.; Ayala-Madrigal, M.D.L.L. Somatic deletion of KDM1A/LSD1 gene is associated to advanced colorectal cancer stages. J. Clin. Pathol. 2020, 73, 107–111. [Google Scholar] [CrossRef]
- Carvalho, S.; Freitas, M.; Antunes, L.; Monteiro-Reis, S.; Vieira-Coimbra, M.; Tavares, A.; Paulino, S.; Videira, J.F.; Jerónimo, C.; Henrique, R. Prognostic value of histone marks H3K27me3 and H3K9me3 and modifying enzymes EZH2, SETDB1 and LSD-1 in colorectal cancer. J. Cancer Res. Clin. Oncol. 2018, 144, 2127–2137. [Google Scholar] [CrossRef]
- Miller, S.A.; Policastro, R.A.; Savant, S.S.; Sriramkumar, S.; Ding, N.; Lu, X.; Mohammad, H.P.; Cao, S.; Kalin, J.H.; Cole, P.A.; et al. Lysine-Specific Demethylase 1 Mediates AKT Activity and Promotes Epithelial-to-Mesenchymal Transition in PIK3CA-Mutant Colorectal Cancer. Mol. Cancer Res. 2020, 18, 264–277. [Google Scholar] [CrossRef]
- Hong, X.; Huang, H.; Qiu, X.; Ding, Z.; Feng, X.; Zhu, Y.; Zhuo, H.; Hou, J.; Zhao, J.; Cai, W.; et al. Targeting posttranslational modifications of RIOK1 inhibits the progression of colorectal and gastric cancers. eLife 2018, 7, e29511. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Zhang, Q.; Fang, Y.; Wang, Y. The deubiquitinase USP38 affects cellular functions through interacting with LSD1. Biol. Res. 2018, 51, 53. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Zhang, C.; Feng, Y.; Quan, M.; Cui, Y.; Xuan, Y. Tenascin-C predicts poor outcomes for patients with colorectal cancer and drives cancer stemness via Hedgehog signaling pathway. Cancer Cell Int. 2020, 20, 122. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.-S.; Liu, H.-Y.; Zhou, Z.; Sun, H.-L.; Liu, M.-Y. TSPAN8 promotes colorectal cancer cell growth and migration in LSD1-dependent manner. Life Sci. 2020, 241, 117114. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Qian, X.; Lin, Y.; Tao, L.; Zuo, Z.; Zhang, H.; Yang, S.; Cen, X.; Zhao, Y. Lipidomic profiling reveals lipid regulation by a novel LSD1 inhibitor treatment. Oncol. Rep. 2021, 46, 233. [Google Scholar] [CrossRef] [PubMed]
- Lian, Y.; Xu, Y.; Xiao, C.; Xia, R.; Gong, H.; Yang, P.; Chen, T.; Wu, D.; Cai, Z.; Zhang, J.; et al. The pseudogene derived from long non-coding RNA DUXAP10 promotes colorectal cancer cell growth through epigenetically silencing of p21 and PTEN. Sci. Rep. 2017, 7, 7312. [Google Scholar] [CrossRef] [PubMed]
- Gong, A.; Huang, Z.; Ge, H.; Cai, Y.; Yang, C. The carcinogenic complex lncRNA DUXAP8/EZH2/LSD1 accelerates the proliferation, migration and invasion of colorectal cancer. J. Buon. 2019, 24, 1830–1836. [Google Scholar] [PubMed]
- Wu, X.; Zhu, H.; Xie, Y.; Gu, X.; Zhang, L.; Huang, L. Knockdown of ZEB2-AS1 inhibits cell invasion and induces apoptosis in colorectal cancer. J. BUON 2020, 25, 194–201. [Google Scholar]
- Ding, J.; Xie, M.; Lian, Y.; Zhu, Y.; Peng, P.; Wang, J.; Wang, L.; Wang, K. Long noncoding RNA HOXA-AS2 represses P21 and KLF2 expression transcription by binding with EZH2, LSD1 in colorectal cancer. Oncogenesis 2017, 6, e288. [Google Scholar] [CrossRef]
- Ding, X.; Zhang, J.; Feng, Z.; Tang, Q.; Zhou, X. MiR-137-3p Inhibits Colorectal Cancer Cell Migration by Regulating a KDM1A-Dependent Epithelial–Mesenchymal Transition. Am. J. Dig. Dis. 2020, 66, 2272–2282. [Google Scholar] [CrossRef]
- Wojtala, M.; Rybaczek, D.; Wielgus, E.; Sobalska-Kwapis, M.; Strapagiel, D.; Balcerczyk, A. The Role of Lysine-Specific Demethylase 1 (LSD1) in Shaping the Endothelial Inflammatory Response. Cell. Physiol. Biochem. 2021, 55, 569–589. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.-J.; Lei, P.-M.; Wong, S.-Y.; Ma, D.-L.; Leung, C.-H. Pharmacological Inhibition of LSD1 for Cancer Treatment. Molecules 2018, 23, 3194. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.C.; Yu, B.; Jiang, G.Z.; Feng, X.J.; He, P.X.; Chu, X.Y.; Zhao, W.; Liu, H.M. Irreversible LSD1 Inhibitors: Application of Tranylcypromine and Its Derivatives in Cancer Treatment. Curr. Top. Med. Chem. 2016, 16, 2179–2188. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.-C.; Yu, B.; Chen, Z.-S.; Liu, Y.; Liu, H.-M. TCPs: Privileged scaffolds for identifying potent LSD1 inhibitors for cancer therapy. Epigenomics 2016, 8, 651–666. [Google Scholar] [CrossRef] [PubMed]
- Drugs Approved for Pancreatic Cancer—National Cancer Institute. 2022. Available online: https://www.cancer.gov/about-cancer/treatment/drugs/pancreatic (accessed on 20 February 2022).
- Drugs Approved for Colon and Rectal Cancer—National Cancer Institute. 2022. Available online: https://www.cancer.gov/about-cancer/treatment/drugs/colorectal (accessed on 10 March 2022).
- Kalin, J.H.; Wu, M.; Gomez, A.V.; Song, Y.; Das, J.; Hayward, D.; Adejola, N.; Wu, M.; Panova, I.; Chung, H.J.; et al. Targeting the CoREST complex with dual histone deacetylase and demethylase inhibitors. Nat. Commun. 2018, 9, 53. [Google Scholar] [CrossRef] [PubMed]
- Lan, H.; Tan, M.; Zhang, Q.; Yang, F.; Wang, S.; Li, H.; Xiong, X.; Sun, Y. LSD1 destabilizes FBXW7 and abrogates FBXW7 functions independent of its demethylase activity. Proc. Natl. Acad. Sci. USA 2019, 116, 12311–12320. [Google Scholar] [CrossRef]
- Hong, S.H.; Lee, J.; Jang, E.B.; Park, S.Y.; Moon, H.-S.; Yoon, Y.E. Epigenetic Approaches to the Treatment of Renal Cell Cancer. Korean J. Urol. Oncol. 2020, 18, 78–90. [Google Scholar] [CrossRef]
- Kharkar, V.J.; Ast, A.; Gupta, S.; Sharma, S. LSD2/KDM1B/AOF1 and Human Cancer Pathways: A Literature Review. Cancer Stud. Ther. J. 2016, 1, 1–5. [Google Scholar] [CrossRef]
- Takeuchi, M.; Fuse, Y.; Watanabe, M.; Andrea, C.-S.; Takeuchi, M.; Nakajima, H.; Ohashi, K.; Kaneko, H.; Kobayashi-Osaki, M.; Yamamoto, M.; et al. LSD1/KDM1A promotes hematopoietic commitment of hemangioblasts through downregulation of Etv2. Proc. Natl. Acad. Sci. USA 2015, 112, 13922–13927. [Google Scholar] [CrossRef]
- Nicosia, L.; Boffo, F.L.; Ceccacci, E.; Conforti, F.; Pallavicini, I.; Bedin, F.; Ravasio, R.; Massignani, E.; Somervaille, T.C.P.; Minucci, S.; et al. Pharmacological inhibition of LSD1 triggers myeloid differentiation by targeting GSE1 oncogenic functions in AML. Oncogene 2022, 41, 878–894. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.-C.; Ma, J.; Wang, Z.; Bailing, J.; Jiang, B.; Zhou, W.; Shi, X.; Wang, X.; Zhao, W.; Liu, H.-M. A Systematic Review of Histone Lysine-Specific Demethylase 1 and Its Inhibitors. Med. Res. Rev. 2015, 35, 1032–1071. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, X.; Wu, T.; Yin, W.; Yan, J.; Sun, Y.; Zhao, D. Therapeutic potential of targeting LSD1/ KDM1A in cancers. Pharmacol. Res. 2021, 175, 105958. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Doyle, K.; Mosbruger, T.L.; Butterfield, A.; Weston, A.; Ast, A.; Kaadige, M.; Verma, A.; Sharma, S. Reversible LSD1 inhibition with HCI-2509 induces the p53 gene expression signature and disrupts the MYCN signature in high-risk neuroblastoma cells. Oncotarget 2018, 9, 9907–9924. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Tang, T.; Pang, J.; Xu, J.; Yang, X.; Wang, L.; Huang, Y.; Huang, Z.; Liu, G.; Tong, D.; et al. LSD1 Promotes Bladder Cancer Progression by Upregulating LEF1 and Enhancing EMT. Front. Oncol. 2020, 10, 1234. [Google Scholar] [CrossRef] [PubMed]
- Sehrawat, A.; Gao, L.; Wang, Y.; Bankhead, A.; McWeeney, S.K.; King, C.J.; Schwartzman, J.; Urrutia, J.; Bisson, W.H.; Coleman, D.J.; et al. LSD1 activates a lethal prostate cancer gene network independently of its demethylase function. Proc. Natl. Acad. Sci. USA 2018, 115, E4179–E4188. [Google Scholar] [CrossRef]
- Augert, A.; Eastwood, E.; Ibrahim, A.H.; Wu, N.; Grunblatt, E.; Basom, R.; Liggitt, D.; Eaton, K.D.; Martins, R.; Poirier, J.T.; et al. Targeting NOTCH activation in small cell lung cancer through LSD1 inhibition. Sci. Signal. 2019, 12, eaau2922. [Google Scholar] [CrossRef] [PubMed]
- Qin, Y.; Vasilatos, S.N.; Chen, L.; Wu, H.; Cao, Z.; Fu, Y.; Huang, M.; Vlad, A.M.; Lu, B.; Oesterreich, S.; et al. Inhibition of histone lysine-specific demethylase 1 elicits breast tumor immunity and enhances antitumor efficacy of immune checkpoint blockade. Oncogene 2019, 38, 390–405. [Google Scholar] [CrossRef] [PubMed]
- Lynch, J.T.; Harris, W.J.; Somervaille, T.C.P. LSD1 inhibition: A therapeutic strategy in cancer? Expert Opin. Ther. Targets 2012, 16, 1239–1249. [Google Scholar] [CrossRef] [PubMed]
- Katz, T.A.; Vasilatos, S.N.; Harrington, E.; Oesterreich, S.; Davidson, N.E.; Huang, Y. Inhibition of histone demethylase, LSD2 (KDM1B), attenuates DNA methylation and increases sensitivity to DNMT inhibitor-induced apoptosis in breast cancer cells. Breast Cancer Res. Treat. 2014, 146, 99–108. [Google Scholar] [CrossRef]
- Tang, D.; He, J.; Dai, Y.; Geng, X.; Leng, Q.; Jiang, H.; Sun, R.; Xu, S. Targeting KDM1B-dependent miR-215-AR-AGR2-axis promotes sensitivity to enzalutamide-resistant prostate cancer. Cancer Gene Ther. 2021, 1–15. [Google Scholar] [CrossRef]
- Venkata, P.P.; Chen, Y.; Alejo, S.; He, Y.; Palacios, B.E.; Loeffel, I.; Liu, J.; Pratap, U.P.; Gray, G.; Pillai, S.M.A.; et al. KDM1A inhibition augments the efficacy of rapamycin for the treatment of endometrial cancer. Cancer Lett. 2022, 524, 219–231. [Google Scholar] [CrossRef] [PubMed]

| LSD | Model | Main Findings | Reference |
|---|---|---|---|
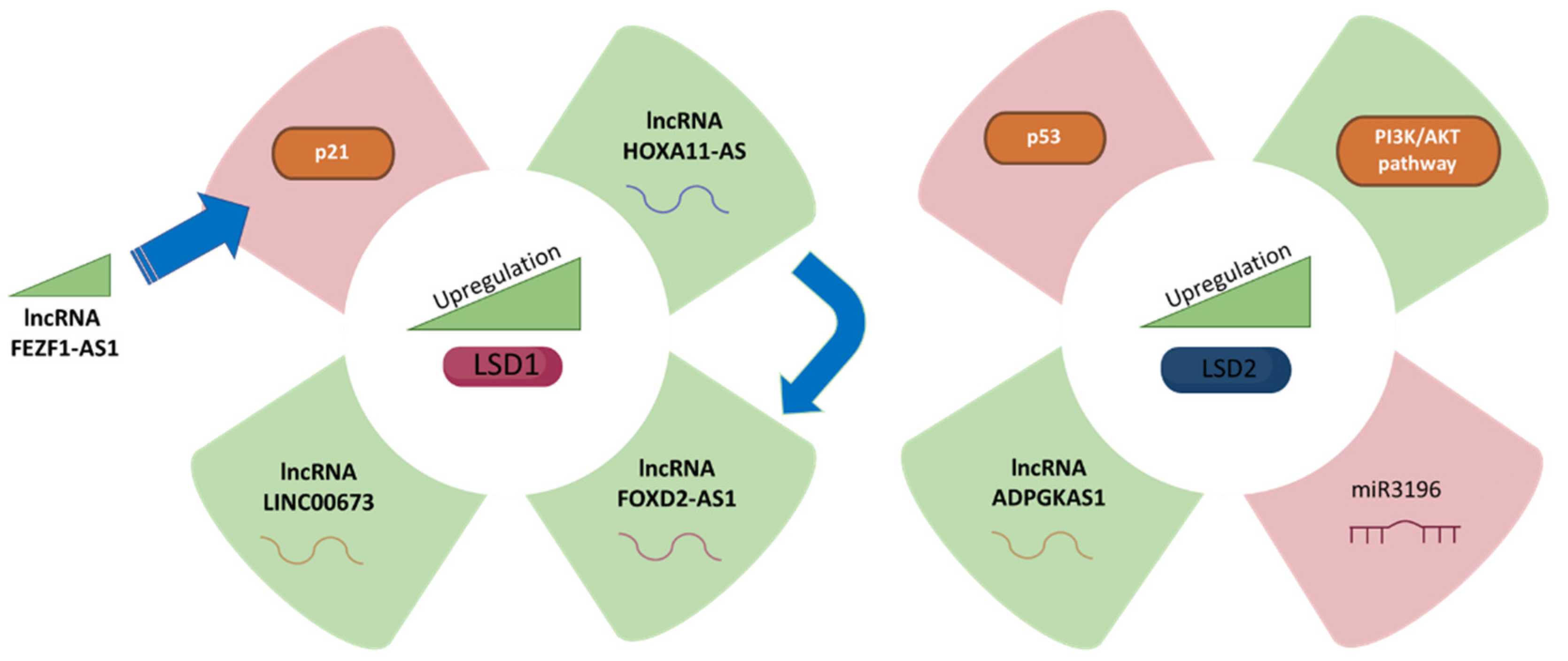
| LSD1 | The GC cell lines (MGC-803, SGC-7901 and AGS) | LSD1 hyperactivates gastric cancer cells with the assistance of LincRNAFEZF1-AS1 to repress p21 expression. LSD1 promotes GC cell proliferation. | [21] |
| LSD1 | (BGC823, SGC7901, MGC803, AGS) and normal gastric mucosa GES1 cell lines. | HOXA11-AS potentially binds LSD1 in GC cells and functions as a scaffold to regulate PRSS8 and KLF2 at transcriptional level. | [30] |
| LSD1 | The GC cell lines (GES1, BGC823, AGS, MKN45, SGC7901 and MGC803) | FOXD2-AS1 is dysregulated GC. FOXD2-AS1 promotes GC tumorigenesis partly through EZH2- and LSD1-mediated EphB3 downregulation. | [35] |
| LSD1 | The GC cell lines (GES1, BGC823, AGS, SGC7901 and MGC803) | LINC00673 works as a scaffold for LSD1 in GC. LINC00673 binds directly to LSD1 in GC cells. LINC00673 and LSD1 are involved in regulating CADM4, LATS2, and KLF2. | [24] |
| LSD2 | The GC cell lines (BGC-823, MGC-803, AGS and SGC-7901) | Cell proliferation due to inhibition of ADPGKAS1 is mostly restored after overexpression of LSD2. | [11] |
| LSD2 | AGS and MKN45 gastric cancer cell lines. | The PI3K/AKT/mTOR pathway is an important intracellular signalling pathway involved in GC prognosis. ADPGKAS1 activation induces the PI3K/AKT/mTOR signalling pathway to promote GC cell emergence and progression by regulating LSD2/KDM1B in GCs via miR3196. | [11,43] |
| LSD | Model | Main Findings | Reference |
|---|---|---|---|
| LSD1 | Clinical data from 188 primary HCC patients. Primary HCC cells, Male nude mice injected with tumour suspension for HCC tumour model. | LSD1 expression is positively associated with LGR5 expression and poor survival in HCC patients. Induction of LSD1 overexpression expands the pool of LGR5+ cells (cancer initiating cells), and drug resistance in HCC cells. Depletion of LSD1 attenuates the self-renewal of CSCs and their drug resistance. LSD1 reduces the H3K4me1/2 methylation at the promoters of several repressors of β-catenin signalling to enhance β-catenin activity in CSCs and enhances tumour formation | [62] |
| LSD1 | Data from 303 HCC patients, Crispr/Cas9 for LSD1-KO SNU-423 and SNU-475 cell lines | LSD1 expression is associated with poorer outcome for overall and disease-free survival for HCC. LSD1 knockout results in reduced S-phase population and it is involved in retinoic acid (RA) pathway. | [64] |
| LSD1 | Human HCC cell lines (Huh7, HCC-LM3, HepG2, MHCC97H and Hep3B), the normal liver cell line (LO2), and OXA-resistant liver cancer cell lines, Clinical data from 153 HCC patients, xenograft HepG2 mice models | LSD1 demethylates LINC01134 for its up-regulation and subsequently confers resistance against oxaliplatin in HCC cells. LSD1 knockdown results in deregulation of LINC01134. LSD1 expression is correlated with LINC01134 in HCC patients. | [68] |
| LSD1 | Liver cancer Bel-7402 cells, xenograft mice models using Bel-7402 cells. | Inhibition of LSD1 leads to attenuated migration of liver cancer cells and show potential antitumor activity in vivo. | [71] |
| LSD1 | HCC TCGA dataset, HCC cell lines HuH7, Hep3B, HepG2, SK-Hep1, PLC/PRF/5 and FOCUS. | Higher LSD1 expression is associated with poor survival in HCC. LSD1 inhibitor exhibited poor effect on HCC cell survival. | [72] |
| LSD1 | HCC cell lines (PLC/PRF/5 and Huh7), PLC and Huh7 sorafenib-resistant cell lines, mice transfected with sh-RNA (LSD1 and LSD2) containing stable clones of HCC cell lines | LSD1 is critical for the induction of a stem-like population and inhibiting its activity attenuates stemness in sorafenib-resistant HCC cells. LSD1 inhibitors derepress the transcription of Wnt antagonists and down-regulate β -catenin signalling activity in sorafenib-resistant cells and in vivo. | [73] |
| LSD2 | HCC tissue samples, n = 365. | Higher expression of LSD2 is associated with a worse prognosis. | [72] |
| LSD2 | Sorafenib resistant Huh7 Cell line. | No change in the expression levels of LSD2. Depletion of LSD2 did not affect sensitivity to sorafenib. | [73] |
| LSD | Model | Main Findings | Reference |
|---|---|---|---|
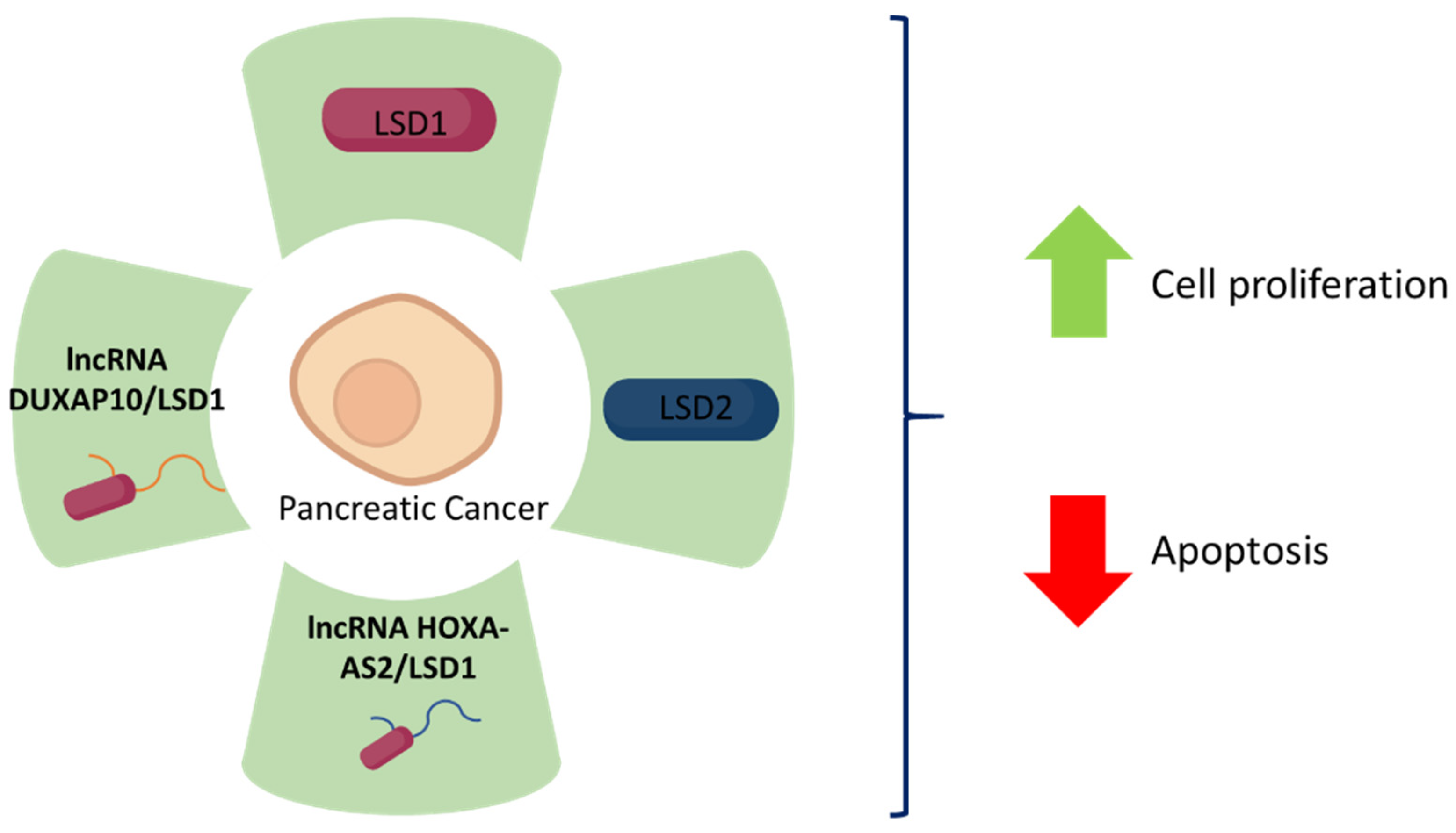
| LSD1 | Gene Expression Omnibus data sets GSE15471. Human pancreatic cancer and matched paracancerous tissue samples, n = 28. BxPC-3 and PANC-1 human pancreatic cancer cell lines. Xenografts using BxPC-3 cells in BALB/C mice, n = 12. | lncRNA HOXA-AS2 promotes pancreatic cancer cell proliferation and reduces apoptosis. HOXA-AS2 binds to LSD1 in lncRNA-HOXA-AS2/EZH2/LSD1 complex to exert its oncogenic functions. | [74] |
| LSD1 | Gene Expression Omnibus data sets GSE15471, GSE15932, GSE16515. Human pancreatic cancer tissue samples, n = 48. AsPC-1, BxPC-3, and PANC-1 human pancreatic cancer cell lines. Xenografts using BxPC-3 cells in BALB/c mice, n = 10. | lncRNA DUXAP10 promotes pancreatic cancer cells, reduces apoptosis, and is associated with poor prognosis. DUXAP10 correlates and binds to LSD1. | [75] |
| LSD2 | Human pancreatic cancer and matched paracancerous tissue samples, n = 20. BxPC-3, CFPAC-1, PANC-1, SW1990 human pancreatic cancer cell lines, and HPDE6-C7 normal human pancreatic duct epithelial cell line. | LSD2 is highly expressed in pancreatic cancer. LSD2 promotes pancreatic cancer cell proliferation and reduces apoptosis. LSD2 knockdown upregulates phosphorylation of ERK1/2, Smad2, p53, cleaved PARP, cleaved Caspase-3, cleaved Caspase-7, eIF2a and Survivin. | [76] |
| LSD | Model | Main Findings | Reference |
|---|---|---|---|
| LSD1 | SW620 human CRC cell line. Xenografts using SW620 cells in BALB/c mice, n = 24. | LSD1 is upregulated in cells presenting cancer stem cell marker CD133. LSD1 KD impairs the stemness of CD133+ cells, decreasing cell viability, colony formation rate, migration, and invasion. | [77] |
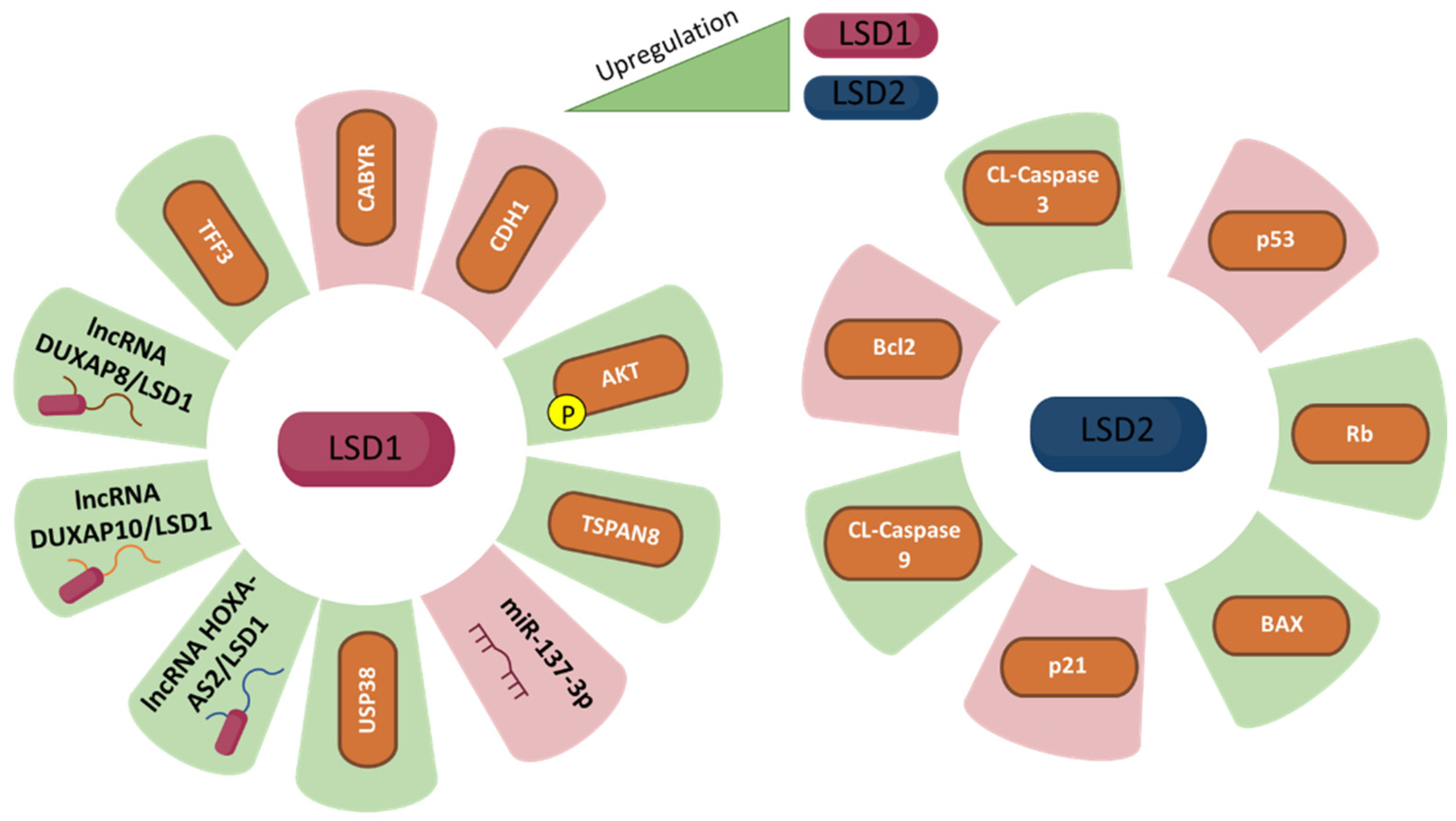
| LSD1 | SW620 and HT-29 human CRC cell lines. | CABYR, FOX2, TLE4 and CDH1 are 4 key LSD-1 target genes associated with proliferation, metastasis, and invasion. LSD1 downregulates CABYR expression by decreasing H3K4me1/2 and downregulates CDH1 by decreasing H3K4me2. LSD1 knockdown affects most frequently p53 pathway. LSD1 affects IG-1/mTOR pathway. | [78] |
| LSD1 | Gene Expression Omnibus data set GSE167262. HT-29, LSD-174T, NCI-H508 human CRC cell lines. Normal human organoids derived from the ascending colon. Colon cancer organoides derived from patient-derived xenografts models 519858 162-T and 817829 284-R. | BRAF mutation is associated with poorly differentiated enteroendocrine cells. LSD1 is upregulated in early secretory cells and early enteroendocrine cells. LSD1 KD results in loss of secretory cells. LSD1 KD reduces TFF3 expression. LSD1 KD leads to loss of pS473-AKT and abrogates tumour growth and metastasis. | [79] |
| LSD1 | CRC tissue samples, n =50. | Loss of LSD1 is associated with metastasis and higher TNM stages. | [80] |
| LSD1 | CRC tissue samples, n = 207. | LSD1 is associated with lower TNM stages. LSD1 low expression is associated with lower disease-specific and disease-free survival. | [81] |
| LSD1 | HT-29, SW480, HCT116, LoVo, and RKO human CRC cell lines. AGS human gastric cancer cell line. | LSD1 is upregulated in PIK3CA mutant CRC compared to PIK3CA wt CRC. LSD1 increases pS473-AKT by scaffolding the CoREST complex. LSD1 enhances EMT-associated gene programmes in PIK3CA mutant cells. LSD1 regulates protein stability of Snail by regulating AKT. LSD1 is required for EGF induced migration mediated by AKT-GSK3β-Snail pathway. | [82] |
| LSD1 | HCT116 human CRC cell line. | RIOK1 promotes CRC cell proliferation and migration through PI3K/AKT pathway. LSD1 demethylates RIOK1 stabilizing it. | [83] |
| LSD1 | HCT116 and SW48 human CRC cell lines. HEK293T human embryonic kidney cell line. | USP38 binds and deubiquitinase LSD1, enhancing the activity of the signalling pathways activated by LSD1. | [84] |
| LSD1 | Tissue microarray containing 100 human CRC cases. HT-29 and HCT116 human CRC cell lines. | TNC expression is associated with poor clinical outcomes, proliferation, and migration. TNC is positively correlated with the LSD1 protein in CRC. TNC KD decreases LSD1 expression. | [85] |
| LSD1 | SW620, SW480, DLD-1, HTC116, and HT-29 human CRC cell lines. NCM460 normal human colon epithelial cell line. HEK293T human embryonic kidney cell line. | LSD1 and TSPAN8 are overexpressed in CRC. LSD1 upregulates TSPAN8 expression by reducing H3K9me2 occupancy on TSPAN8 promoter. TSPAN8 enhances tumorigenicity and EMT in CRC cells in an LSD-1 dependent manner. LSD1 and TSPAN8 KD results in an upregulation of E-cadherin and ZO-1, and a downregulation of n-cadherin, Vimentin, Slug and Snail. | [86] |
| LSD1 | HCT116 human CRC cell line. HeLa human cervical cancer cell line. | LSD1 inhibition modifies the lipidome of cancer cells, specially it dysregulates sphingolipids and glycolipids. LSD1 inhibition enhances bioactive lipids such as ceramides and sphingomyelin which are involved in signalling pathways such as apoptosis. | [87] |
| LSD1 | CRC tissue samples, n = 58. DLD-1, HCT116, SW480, and SW620 human CRC cell lines. | lncRNA DUXAP10 is positively associated with CRC cell proliferation, tumour size, advanced TNM stages and lymph node metastasis. LSD1 interacts with DUXAP10 and decreases p21 and PTEN expression. | [88] |
| LSD1 | Human CRC and matched paracancerous tissue samples. DLD-1 and SW480 human colorectal cancer cell lines. | lncRNA DUXAP8 is positively associated with CRC cell proliferation, tumour size, and advanced TNM stages. LSD1 interacts with DUXAP8 and promotes CRC cell proliferation. | [89] |
| LSD1 | Human CRC and matched paracancerous tissue samples, n = 60. HT-29, HCT116, SW480 and DLD-1 human colorectal cancer tissue samples. NCM460 human colon epithelial normal cells. | lncRNA ZEB2-AS1 is associated with increased proliferation rate, tumour size, higher TNM stage and lymph node metastasis. LSD1 interacts with ZEB-AS1 promoting cell proliferation. | [90] |
| LSD1 | Human CRC and matched paracancerous tissue samples, n = 69. HCT116, DLD-1, SW480, SW620, HT-29, and LoVo human CRC cell lines. | lncRNA HOXA-AS2 is associated with increased proliferation rate, tumour size, higher TNM stage and lymph node metastasis. LSD1 interacts with HOXA-AS2 silencing p21 and KLF2 transcription and promotes cell proliferation. | [91] |
| LSD1 | Human CRC and matched paracancerous tissue samples, n = 98. LoVo and HCT116 human CRC cell lines. | miR-137-3p is negatively associated with the invasiveness of CRC cells. LSD1 is regulated by miR-137-3p and is involved in CRC cell proliferation, adhesion, and invasion. LSD1 and miR-137-3p respond to hypoxia. | [92] |
| LSD2 | LoVo, HCT116, SW1116, andCaco2 human CRC cell lines. Xenografts assays using LoVo cells in BALB/c mice, n = 6. | LSD2 is upregulated in CRC. LSD2 reduces Bcl-2 and increases cleaved caspase 3, cleaved caspase 9 and BAX levels. LSD2 downregulates p53 expression and p21 and drives the cell cycle through p53-p21-Rb. | [44] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malagraba, G.; Yarmohammadi, M.; Javed, A.; Barceló, C.; Rubio-Tomás, T. The Role of LSD1 and LSD2 in Cancers of the Gastrointestinal System: An Update. Biomolecules 2022, 12, 462. https://doi.org/10.3390/biom12030462
Malagraba G, Yarmohammadi M, Javed A, Barceló C, Rubio-Tomás T. The Role of LSD1 and LSD2 in Cancers of the Gastrointestinal System: An Update. Biomolecules. 2022; 12(3):462. https://doi.org/10.3390/biom12030462
Chicago/Turabian StyleMalagraba, Gianluca, Mahdieh Yarmohammadi, Aadil Javed, Carles Barceló, and Teresa Rubio-Tomás. 2022. "The Role of LSD1 and LSD2 in Cancers of the Gastrointestinal System: An Update" Biomolecules 12, no. 3: 462. https://doi.org/10.3390/biom12030462
APA StyleMalagraba, G., Yarmohammadi, M., Javed, A., Barceló, C., & Rubio-Tomás, T. (2022). The Role of LSD1 and LSD2 in Cancers of the Gastrointestinal System: An Update. Biomolecules, 12(3), 462. https://doi.org/10.3390/biom12030462









