COVID-19: Unmasking Emerging SARS-CoV-2 Variants, Vaccines and Therapeutic Strategies
Abstract
:1. Introduction
2. The Pathophysiology of COVID-19 in Immune-Dysregulated Patients
2.1. Multiple Sclerosis (MS)
2.2. Rheumatoid Arthritis (RA)
2.3. Systemic Lupus Erythematosus (SLE)
2.4. Cancer
2.5. Inflammatory Bowel Disease (IBD)
3. Vaccine Platforms
3.1. Protein-Based Vaccines (PV)
3.2. Nucleic Acid Vaccines
3.2.1. mRNA Vaccines
3.2.2. DNA Vaccines
3.3. Viral Vector-Based Vaccines
3.3.1. Non-Replicating Viral Vector Vaccines
3.3.2. Replicating Viral Vector Vaccines
3.4. Inactivated Vaccines (IVs)
3.5. Live-Attenuated Vaccines (LAV)
4. Pharmacological Therapies
4.1. Remdesivir
4.2. Dexamethasone
4.3. Favipiravir
4.4. Opaganib
4.5. Tocilizumab
4.6. Chloroquine (CQ) and Hydroxychloroquine (HCQ)
4.7. Baricitinib
4.8. SARS-CoV-2 Monoclonal Antibodies
4.9. Convalescent Plasma Therapy (CPT)
4.10. Additional Novel Therapies
4.10.1. Inhaled Nanobodies
4.10.2. Mesenchymal Stem Cells Therapy
| Manufacturer | Name | Target | Mechanism of Action | Phase | RoA |
|---|---|---|---|---|---|
| Gilead Sciences Inc | Veklury (Remdesivir) | Viral RNA polymerase | Inhibitor of viral replication | Approved | IV |
| None (generic) | Dexamethasone | Glucocorticoid receptor agonist | Alters the body’s normal immune system responses | Approved | Oral |
| Fujifilm Toyama Chemical | Favipiravir | Viral RNA polymerase | Inhibitor of viral replication | Approved (as generics) | Oral |
| Eli Lilly | Olumiant (Baricitinib) | JAK1/2 inhibitor | Decreases immune system activation | EUA | Oral |
| Regeneron/Sanofi | Casirivimab and Imdevimab (REGN-COV2) | Viral epitopes | Binds to virus and neutralizes its ability for infection | EUA | IV |
| Eli Lilly | Bamlanivimab (LY-CoV555) and Etesevimab (LY-CoV016) | Viral epitopes | Binds to virus and neutralizes its ability for infection | EUA | IV |
| GSK/ Vir Biotech | Sotrovimab | Viral epitopes | Binds to virus and neutralizes its ability for infection | EUA | IV |
| Roche/Chugai | Tocilizumab (Actemra) | IL-6 | Decreases immune system activation | Phase 3 | IV and SC |
| Sanofi | Chloroquine/Hydroxychloroquine | Endosomal vesicles | Antiviral activity through pH change | Phase 3 | Oral |
| Humanigen [178] | Lenzilumab | GM-CSF | Neutralizes circulating GM-CSF | Phase 3 | IV |
| RedHill [145] | Opaganib | Sphingosine kinase-2 (SK2) | SK2 inhibitor | Phase 3 | Oral |
| EUSA Pharma [179] | Siltuximab | IL-6 | Decreases immune system activation | Phase 3 | IV |
| Merck [180] | MK-4482 | Viral RNA polymerase | Inhibitor of viral replication | Phase 3 | Oral |
| Synairgen [181] | SNG001 | IFN-beta-1a | Delivery of FN-beta inhibits viral replication | Phase 3 | IN |
| GSK/Vir [168] | GSK4182136 | Viral epitopes | Binds to virus and neutralizes its ability for infection | Phase 3 | IV, IM |
| PharmaMar [182] | Plitidepsin (Aplidin) | eEF1A | eEF1A inhibitor | Phase 2 | IV |
| Pfizer [183] | PF-07321332 | 3CL protease | 3CL protease inhibitor | Phase 1 | Oral |
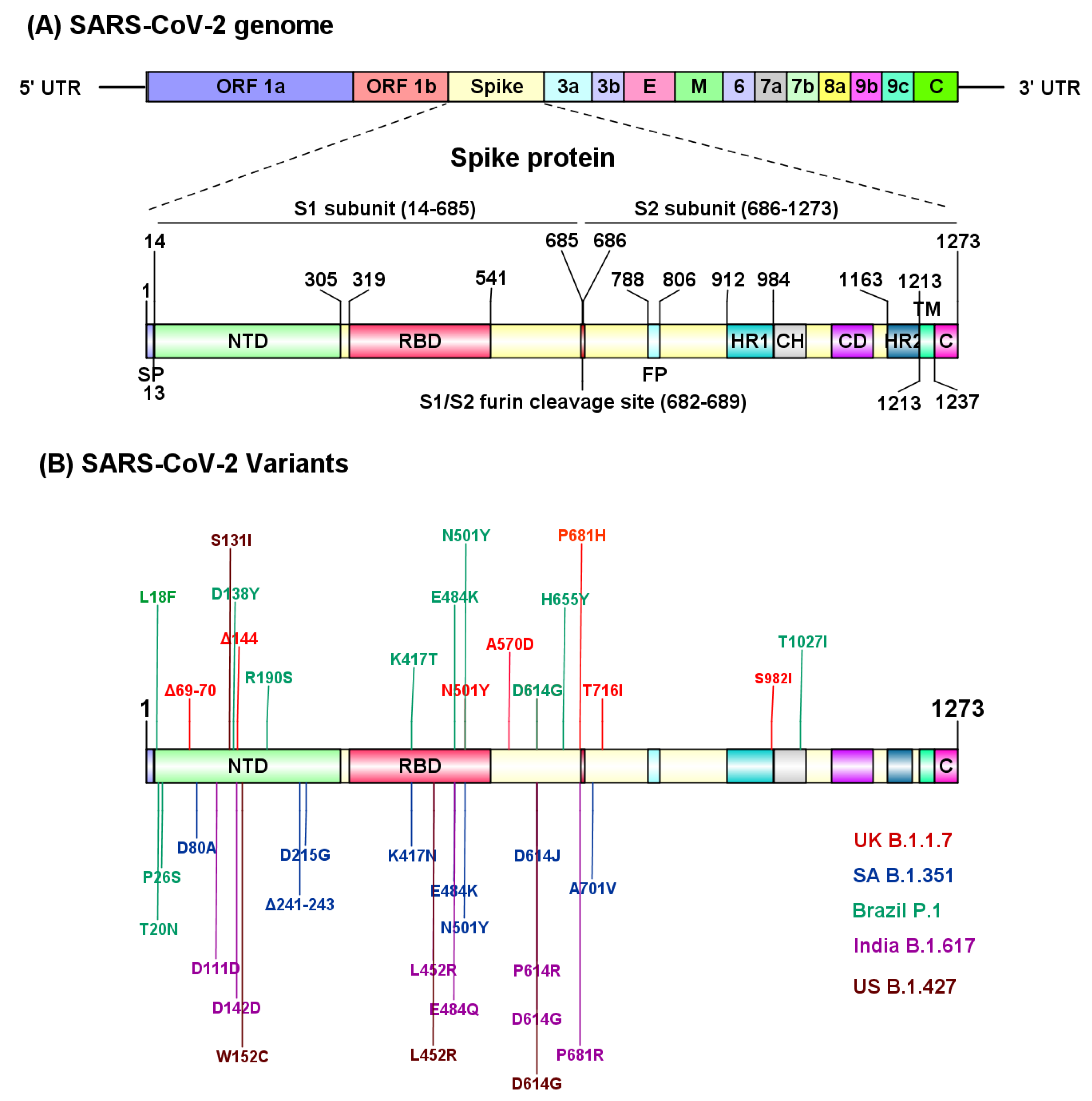
5. SARS-CoV-2 Variants
5.1. The B.1.1.7 Lineage (Alpha Variant)
5.2. The B.1.351 Lineage (Beta Variant)
5.3. P.1 Variant (Gamma Variant)
5.4. The B.1.617.2 (Delta Variant)
5.5. The CAL.20C Variant
5.6. Other Variants of Interest (VOI)
| Name | Country of Origin | Mutations in Spike Protein | Effect on Monoclonal Antibody Treatment Regimens and Neutralization of Convalescent Sera | Effect on Vaccine Efficacy |
|---|---|---|---|---|
| B.1.1.7 (Alpha) † | United Kingdom | N501Y *, A570D, D614G, P681H *, T716I, S982A, Δ69/70 *, Δ144 * |
| Vaccine efficacy slightly lower or unchanged, largely preserved neutralizing titers
|
| B.1.351 (Beta) † | South Africa | D80A, D215G, Δ241/242/243, K417N *, E484K *, N501Y *, D614G, A701V |
| Reduced efficacy for some vaccines, completely abolished for others. |
| P.1 (Gamma) † | Japan/Brazil | L18F, T20N, P26S, D138Y, R190S, K417T *, E484K *, N501Y *, D614G, H655Y, T1027I |
| |
| B.1.617.2 (Delta) † | India | L452R *, E484Q *, D614GD111D, G142D, P614R, P681R * | ||
| CAL.20C | California, USA | S13I *, W152C *, L452R *, D614G |
| No evidence yet |
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- WHO. Coronavirus Disease (Covid-19) Pandemic. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019 (accessed on 28 May 2021).
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar]
- Tay, M.Z.; Poh, C.M.; Renia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of covid-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.M.; Wang, W.; Song, Z.G.; Hu, Y.; Tao, Z.W.; Tian, J.H.; Pei, Y.Y.; et al. A new coronavirus associated with human respiratory disease in china. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early transmission dynamics in wuhan, china, of novel coronavirus-infected pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A novel coronavirus from patients with pneumonia in china, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [Green Version]
- Chen, L.; Liu, W.; Zhang, Q.; Xu, K.; Ye, G.; Wu, W.; Sun, Z.; Liu, F.; Wu, K.; Zhong, B.; et al. Rna based mngs approach identifies a novel human coronavirus from two individual pneumonia cases in 2019 wuhan outbreak. Emerg. Microbes Infect. 2020, 9, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Hillyer, C.; Du, L. Neutralizing antibodies against sars-cov-2 and other human coronaviruses: (trends in immunology 41, 355-359; 2020). Trends Immunol. 2020, 41, 545. [Google Scholar] [CrossRef] [PubMed]
- Walls, A.C.; Park, Y.J.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, function, and antigenicity of the sars-cov-2 spike glycoprotein. Cell 2020, 181, 281–292. [Google Scholar] [CrossRef]
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Genome composition and divergence of the novel coronavirus (2019-ncov) originating in china. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blanco-Melo, D.; Nilsson-Payant, B.E.; Liu, W.C.; Uhl, S.; Hoagland, D.; Moller, R.; Jordan, T.X.; Oishi, K.; Panis, M.; Sachs, D.; et al. Imbalanced host response to sars-cov-2 drives development of covid-19. Cell 2020, 181, 1036–1045. [Google Scholar] [CrossRef] [PubMed]
- Matsuyama, S.; Nao, N.; Shirato, K.; Kawase, M.; Saito, S.; Takayama, I.; Nagata, N.; Sekizuka, T.; Katoh, H.; Kato, F.; et al. Enhanced isolation of sars-cov-2 by tmprss2-expressing cells. Proc. Natl. Acad. Sci. USA 2020, 117, 7001–7003. [Google Scholar] [CrossRef] [Green Version]
- Guo, Y.R.; Cao, Q.D.; Hong, Z.S.; Tan, Y.Y.; Chen, S.D.; Jin, H.J.; Tan, K.S.; Wang, D.Y.; Yan, Y. The origin, transmission and clinical therapies on coronavirus disease 2019 (covid-19) outbreak—An update on the status. Mil. Med. Res. 2020, 7, 11. [Google Scholar] [CrossRef] [Green Version]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of sars-cov-2 by full-length human ace2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [Green Version]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First case of 2019 novel coronavirus in the united states. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Puelles, V.G.; Lutgehetmann, M.; Lindenmeyer, M.T.; Sperhake, J.P.; Wong, M.N.; Allweiss, L.; Chilla, S.; Heinemann, A.; Wanner, N.; Liu, S.; et al. Multiorgan and renal tropism of sars-cov-2. N. Engl. J. Med. 2020, 383, 590–592. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Du, R.H.; Li, B.; Zheng, X.S.; Yang, X.L.; Hu, B.; Wang, Y.Y.; Xiao, G.F.; Yan, B.; Shi, Z.L.; et al. Molecular and serological investigation of 2019-ncov infected patients: Implication of multiple shedding routes. Emerg. Microbes Infect. 2020, 9, 386–389. [Google Scholar] [CrossRef] [Green Version]
- Zhou, J.; Li, C.; Liu, X.; Chiu, M.C.; Zhao, X.; Wang, D.; Wei, Y.; Lee, A.; Zhang, A.J.; Chu, H.; et al. Infection of bat and human intestinal organoids by sars-cov-2. Nat. Med. 2020, 26, 1077–1083. [Google Scholar] [CrossRef] [PubMed]
- Pung, R.; Chiew, C.J.; Young, B.E.; Chin, S.; Chen, M.I.; Clapham, H.E.; Cook, A.R.; Maurer-Stroh, S.; Toh, M.; Poh, C.; et al. Investigation of three clusters of covid-19 in singapore: Implications for surveillance and response measures. Lancet 2020, 395, 1039–1046. [Google Scholar] [CrossRef]
- Guan, W.J.; Ni, Z.Y.; Hu, Y.; Liang, W.H.; Ou, C.Q.; He, J.X.; Liu, L.; Shan, H.; Lei, C.L.; Hui, D.S.C.; et al. Clinical characteristics of coronavirus disease 2019 in china. N. Engl. J. Med. 2020, 382, 1708–1720. [Google Scholar] [CrossRef]
- Lauer, S.A.; Grantz, K.H.; Bi, Q.; Jones, F.K.; Zheng, Q.; Meredith, H.R.; Azman, A.S.; Reich, N.G.; Lessler, J. The incubation period of coronavirus disease 2019 (covid-19) from publicly reported confirmed cases: Estimation and application. Ann. Intern. Med. 2020, 172, 577–582. [Google Scholar] [CrossRef] [Green Version]
- Bai, Y.; Yao, L.; Wei, T.; Tian, F.; Jin, D.Y.; Chen, L.; Wang, M. Presumed asymptomatic carrier transmission of covid-19. JAMA 2020, 323, 1406–1407. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rothe, C.; Schunk, M.; Sothmann, P.; Bretzel, G.; Froeschl, G.; Wallrauch, C.; Zimmer, T.; Thiel, V.; Janke, C.; Guggemos, W.; et al. Transmission of 2019-ncov infection from an asymptomatic contact in germany. N. Engl. J. Med. 2020, 382, 970–971. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thompson, B.T.; Chambers, R.C.; Liu, K.D. Acute respiratory distress syndrome. N. Engl. J. Med. 2017, 377, 562–572. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in wuhan, china. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in wuhan, china. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef]
- Bhatraju, P.K.; Ghassemieh, B.J.; Nichols, M.; Kim, R.; Jerome, K.R.; Nalla, A.K.; Greninger, A.L.; Pipavath, S.; Wurfel, M.M.; Evans, L.; et al. Covid-19 in critically ill patients in the seattle region—Case series. N. Engl. J. Med. 2020, 382, 2012–2022. [Google Scholar] [CrossRef]
- Liang, W.; Liang, H.; Ou, L.; Chen, B.; Chen, A.; Li, C.; Li, Y.; Guan, W.; Sang, L.; Lu, J.; et al. Development and validation of a clinical risk score to predict the occurrence of critical illness in hospitalized patients with covid-19. JAMA Intern. Med. 2020, 180, 1081–1089. [Google Scholar] [CrossRef]
- Liu, W.; Xie, Y.; Ma, J.; Luo, X.; Nie, P.; Zuo, Z.; Lahrmann, U.; Zhao, Q.; Zheng, Y.; Zhao, Y.; et al. Ibs: An illustrator for the presentation and visualization of biological sequences. Bioinformatics 2015, 31, 3359–3361. [Google Scholar] [CrossRef] [Green Version]
- Abraham, C.; Cho, J.H. Inflammatory bowel disease. N. Engl. J. Med. 2009, 361, 2066–2078. [Google Scholar] [CrossRef]
- Conti, P.; Ronconi, G.; Caraffa, A.; Gallenga, C.E.; Ross, R.; Frydas, I.; Kritas, S.K. Induction of pro-inflammatory cytokines (il-1 and il-6) and lung inflammation by coronavirus-19 (covi-19 or sars-cov-2): Anti-inflammatory strategies. J. Biol. Regul. Homeost. Agents 2020, 34, 327–331. [Google Scholar]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; Xiang, J.; Wang, Y.; Song, B.; Gu, X.; et al. Clinical course and risk factors for mortality of adult inpatients with covid-19 in wuhan, china: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- DeLuca, J.; Chiaravalloti, N.D.; Sandroff, B.M. Treatment and management of cognitive dysfunction in patients with multiple sclerosis. Nat. Rev. Neurol. 2020, 16, 319–332. [Google Scholar] [CrossRef]
- Williamson, E.J.; Walker, A.J.; Bhaskaran, K.; Bacon, S.; Bates, C.; Morton, C.E.; Curtis, H.J.; Mehrkar, A.; Evans, D.; Inglesby, P.; et al. Factors associated with covid-19-related death using opensafely. Nature 2020, 584, 430–436. [Google Scholar] [CrossRef]
- Crescenzo, F.; Marastoni, D.; Bovo, C.; Calabrese, M. Frequency and severity of covid-19 in multiple sclerosis: A short single-site report from northern italy. Mult. Scler. Relat. Disord. 2020, 44, 102372. [Google Scholar] [CrossRef] [PubMed]
- Chou, S.H.; Beghi, E.; Helbok, R.; Moro, E.; Sampson, J.; Altamirano, V.; Mainali, S.; Bassetti, C.; Suarez, J.I.; McNett, M.; et al. Global incidence of neurological manifestations among patients hospitalized with covid-19-a report for the gcs-neurocovid consortium and the energy consortium. JAMA Netw Open 2021, 4, e2112131. [Google Scholar] [CrossRef] [PubMed]
- Fan, M.; Qiu, W.; Bu, B.; Xu, Y.; Yang, H.; Huang, D.; Lau, A.Y.; Guo, J.; Zhang, M.N.; Zhang, X.; et al. Risk of covid-19 infection in ms and neuromyelitis optica spectrum disorders. Neurol. Neuroimmunol. Neuroinflamm. 2020, 7, e787. [Google Scholar] [CrossRef] [PubMed]
- Louapre, C.; Collongues, N.; Stankoff, B.; Giannesini, C.; Papeix, C.; Bensa, C.; Deschamps, R.; Creange, A.; Wahab, A.; Pelletier, J.; et al. Clinical characteristics and outcomes in patients with coronavirus disease 2019 and multiple sclerosis. JAMA Neurol. 2020, 77, 1079–1088. [Google Scholar] [CrossRef] [PubMed]
- Aletaha, D.; Funovits, J.; Smolen, J.S. Physical disability in rheumatoid arthritis is associated with cartilage damage rather than bone destruction. Ann. Rheum. Dis. 2011, 70, 733–739. [Google Scholar] [CrossRef] [Green Version]
- Sparks, J.A. Rheumatoid arthritis. Ann. Intern. Med. 2019, 170, ITC1–ITC16. [Google Scholar] [CrossRef] [PubMed]
- Ji, W.; Huh, K.; Kang, M.; Hong, J.; Bae, G.H.; Lee, R.; Na, Y.; Choi, H.; Gong, S.Y.; Choi, Y.H.; et al. Effect of underlying comorbidities on the infection and severity of covid-19 in korea: A nationwide case-control study. J. Korean Med. Sci. 2020, 35, e237. [Google Scholar] [CrossRef] [PubMed]
- Schett, G.; Manger, B.; Simon, D.; Caporali, R. Covid-19 revisiting inflammatory pathways of arthritis. Nat. Rev. Rheumatol. 2020, 16, 465–470. [Google Scholar] [CrossRef] [PubMed]
- Monti, S.; Balduzzi, S.; Delvino, P.; Bellis, E.; Quadrelli, V.S.; Montecucco, C. Clinical course of covid-19 in a series of patients with chronic arthritis treated with immunosuppressive targeted therapies. Ann. Rheum. Dis. 2020, 79, 667–668. [Google Scholar] [CrossRef] [Green Version]
- Tsokos, G.C. Systemic lupus erythematosus. N. Engl. J. Med. 2011, 365, 2110–2121. [Google Scholar] [CrossRef] [Green Version]
- McNab, F.; Mayer-Barber, K.; Sher, A.; Wack, A.; O’Garra, A. Type i interferons in infectious disease. Nat. Rev. Immunol. 2015, 15, 87–103. [Google Scholar] [CrossRef]
- Horisberger, A.; Moi, L.; Ribi, C.; Comte, D. Impact of covid-19 pandemic on sle: Beyond the risk of infection. Lupus Sci. Med. 2020, 7, e000408. [Google Scholar] [CrossRef]
- Favalli, E.G.; Gerosa, M.; Murgo, A.; Caporali, R. Are patients with systemic lupus erythematosus at increased risk for covid-19? Ann. Rheum. Dis. 2021, 80, e25. [Google Scholar] [CrossRef]
- Holubar, J.; Le Quintrec, M.; Letaief, H.; Faillie, J.L.; Pers, Y.M.; Jorgensen, C. Monitoring of patients with systemic lupus erythematosus during the covid-19 outbreak. Ann. Rheum. Dis. 2020. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Ruiz, R.; Paredes, J.L.; Niewold, T.B. Covid-19 in patients with systemic lupus erythematosus: Lessons learned from the inflammatory disease. Transl. Res. 2021, 232, 13–36. [Google Scholar] [CrossRef]
- Gendebien, Z.; von Frenckell, C.; Ribbens, C.; Andre, B.; Thys, M.; Gangolf, M.; Seidel, L.; Malaise, M.G.; Malaise, O. Systematic analysis of covid-19 infection and symptoms in a systemic lupus erythematosus population: Correlation with disease characteristics, hydroxychloroquine use and immunosuppressive treatments. Ann. Rheum. Dis. 2020. [CrossRef] [PubMed]
- Xia, Y.; Jin, R.; Zhao, J.; Li, W.; Shen, H. Risk of covid-19 for patients with cancer. Lancet Oncol. 2020, 21, e180. [Google Scholar] [CrossRef]
- Liang, W.; Guan, W.; Chen, R.; Wang, W.; Li, J.; Xu, K.; Li, C.; Ai, Q.; Lu, W.; Liang, H.; et al. Cancer patients in sars-cov-2 infection: A nationwide analysis in china. Lancet Oncol. 2020, 21, 335–337. [Google Scholar] [CrossRef]
- Dai, M.; Liu, D.; Liu, M.; Zhou, F.; Li, G.; Chen, Z.; Zhang, Z.; You, H.; Wu, M.; Zheng, Q.; et al. Patients with cancer appear more vulnerable to sars-cov-2: A multicenter study during the covid-19 outbreak. Cancer Discov. 2020, 10, 783–791. [Google Scholar] [PubMed]
- Kuderer, N.M.; Choueiri, T.K.; Shah, D.P.; Shyr, Y.; Rubinstein, S.M.; Rivera, D.R.; Shete, S.; Hsu, C.Y.; Desai, A.; de Lima Lopes, G., Jr.; et al. Clinical impact of covid-19 on patients with cancer (ccc19): A cohort study. Lancet 2020, 395, 1907–1918. [Google Scholar] [CrossRef]
- Kamboj, M.; Sepkowitz, K.A. Nosocomial infections in patients with cancer. Lancet Oncol. 2009, 10, 589–597. [Google Scholar] [CrossRef]
- Longbottom, E.R.; Torrance, H.D.; Owen, H.C.; Fragkou, P.C.; Hinds, C.J.; Pearse, R.M.; O’Dwyer, M.J. Features of postoperative immune suppression are reversible with interferon gamma and independent of interleukin-6 pathways. Ann. Surg. 2016, 264, 370–377. [Google Scholar] [CrossRef]
- Bersanelli, M. Controversies about covid-19 and anticancer treatment with immune checkpoint inhibitors. Immunotherapy 2020, 12, 269–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, Z.; Shi, L.; Wang, Y.; Zhang, J.; Huang, L.; Zhang, C.; Liu, S.; Zhao, P.; Liu, H.; Zhu, L.; et al. Pathological findings of covid-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 2020, 8, 420–422. [Google Scholar] [CrossRef]
- Chan, J.F.; Yuan, S.; Kok, K.H.; To, K.K.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.; Poon, R.W.; et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: A study of a family cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef] [Green Version]
- Lui, R.N.; Wong, S.H.; Sanchez-Luna, S.A.; Pellino, G.; Bollipo, S.; Wong, M.-Y.; Chiu, P.W.Y.; Sung, J.J.Y. Overview of guidance for endoscopy during the coronavirus disease 2019 pandemic. J. Gastroenterol. Hepatol. 2020, 35, 749–759. [Google Scholar] [CrossRef]
- Lin, L.; Jiang, X.; Zhang, Z.; Huang, S.; Zhang, Z.; Fang, Z.; Gu, Z.; Gao, L.; Shi, H.; Mai, L.; et al. Gastrointestinal symptoms of 95 cases with sars-cov-2 infection. Gut 2020, 69, 997–1001. [Google Scholar] [CrossRef] [PubMed]
- Ranjan, K. Intestinal immune homeostasis and inflammatory bowel disease: A perspective on intracellular response mechanisms. Gastrointest. Disord. 2020, 2, 246–266. [Google Scholar] [CrossRef]
- Garg, M.; Royce, S.G.; Tikellis, C.; Shallue, C.; Batu, D.; Velkoska, E.; Burrell, L.M.; Patel, S.K.; Beswick, L.; Jackson, A.; et al. Imbalance of the renin-angiotensin system may contribute to inflammation and fibrosis in ibd: A novel therapeutic target? Gut 2020, 69, 841–851. [Google Scholar] [CrossRef] [PubMed]
- An, P.; Ji, M.; Ren, H.; Su, J.; Kang, J.; Yin, A.; Zhou, Q.; Shen, L.; Zhao, L.; Jiang, X. Protection of 318 inflammatory bowel disease patients from the outbreak and rapid spread of covid-19 infection in Wuhan, China. SSRN J. 2020. [Google Scholar] [CrossRef]
- Higgins, P.D.R.; Ng, S.; Danese, S.; Rao, K. The risk of sars-cov-2 in immunosuppressed ibd patients. Crohns Colitis 360 2020, 2, otaa026. [Google Scholar] [CrossRef] [PubMed]
- Harmer, D.; Gilbert, M.; Borman, R.; Clark, K.L. Quantitative mrna expression profiling of ace 2, a novel homologue of angiotensin converting enzyme. FEBS Lett. 2002, 532, 107–110. [Google Scholar] [CrossRef] [Green Version]
- Tursi, A.; Angarano, G.; Monno, L.; Saracino, A.; Signorile, F.; Ricciardi, A.; Papa, A. Covid-19 infection in crohn’s disease under treatment with adalimumab. Gut 2020, 69, 1364–1365. [Google Scholar] [CrossRef]
- Amanat, F.; Krammer, F. Sars-cov-2 vaccines: Status report. Immunity 2020, 52, 583–589. [Google Scholar] [CrossRef]
- Grifoni, A.; Weiskopf, D.; Ramirez, S.I.; Mateus, J.; Dan, J.M.; Moderbacher, C.R.; Rawlings, S.A.; Sutherland, A.; Premkumar, L.; Jadi, R.S.; et al. Targets of t cell responses to sars-cov-2 coronavirus in humans with covid-19 disease and unexposed individuals. Cell 2020, 181, 1489–1501. [Google Scholar] [CrossRef]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA 2004, 101, 6641–6646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- WHO. Draft Landscape and Tracker of Covid-19 Candidate Vaccines. 2021. Available online: https://www.who.int/publications/m/item/draft-landscape-of-covid-19-candidate-vaccines (accessed on 26 May 2021).
- Merlin, M.; Gecchele, E.; Capaldi, S.; Pezzotti, M.; Avesani, L. Comparative evaluation of recombinant protein production in different biofactories: The green perspective. BioMed Res. Int. 2014, 2014, 136419. [Google Scholar] [CrossRef] [Green Version]
- Keech, C.; Albert, G.; Cho, I.; Robertson, A.; Reed, P.; Neal, S.; Plested, J.S.; Zhu, M.; Cloney-Clark, S.; Zhou, H.; et al. Phase 1-2 trial of a sars-cov-2 recombinant spike protein nanoparticle vaccine. N. Engl. J. Med. 2020, 383, 2320–2332. [Google Scholar] [CrossRef] [PubMed]
- Novavax. Novavax Confirms High Levels of Efficacy against Original and Variant Covid-19 Strains in United Kingdom and South Africa Trials. CISION 2021. Available online: https://www.prnewswire.com/news-releases/novavax-confirms-high-levels-of-efficacy-against-original-and-variant-covid-19-strains-in-united-kingdom-and-south-africa-trials-301246019.html (accessed on 20 May 2021).
- Folegatti, P.M.; Ewer, K.J.; Aley, P.K.; Angus, B.; Becker, S.; Belij-Rammerstorfer, S.; Bellamy, D.; Bibi, S.; Bittaye, M.; Clutterbuck, E.A.; et al. Safety and immunogenicity of the chadox1 ncov-19 vaccine against sars-cov-2: A preliminary report of a phase 1/2, single-blind, randomised controlled trial. Lancet 2020, 396, 467–478. [Google Scholar] [CrossRef]
- Jackson, L.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; McCullough, M.P.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; et al. An mrna vaccine against sars-cov-2—Preliminary report. N. Engl. J. Med. 2020, 383, 1920–1931. [Google Scholar] [CrossRef] [PubMed]
- Walsh, E.E.; Frenck, R.; Falsey, A.R.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Mulligan, M.J.; Bailey, R.; et al. Rna-based covid-19 vaccine bnt162b2 selected for a pivotal efficacy study. medRxiv 2020. [Google Scholar] [CrossRef]
- Yan, Z.P.; Yang, M.; Lai, C.L. Covid-19 vaccines: A review of the safety and efficacy of current clinical trials. Pharmaceuticals 2021, 14, 406. [Google Scholar] [CrossRef] [PubMed]
- Machhi, J.; Shahjin, F.; Das, S.; Patel, M.; Abdelmoaty, M.M.; Cohen, J.D.; Singh, P.A.; Baldi, A.; Bajwa, N.; Kumar, R.; et al. Nanocarrier vaccines for sars-cov-2. Adv. Drug Deliv. Rev. 2021, 171, 215–239. [Google Scholar] [CrossRef]
- Biospace. Covaxx’s Covid-19 Vaccine, Ub-612, Induced Neutralizing Antibodies in 100% of Participants during Phase 1 Clinical Trial. 2021. Available online: https://www.biospace.com/article/releases/covaxx-s-covid-19-vaccine-ub-612-induced-neutralizing-antibodies-in-100-percent-of-participants-during-phase-1-clinical-trial/ (accessed on 20 May 2021).
- King, A. Protein-Based Covid-19 Vaccines Could Overshadow Rivals. 2020. Available online: https://www.chemistryworld.com/news/protein-based-covid-19-vaccines-could-overshadow-rivals/4012450.article (accessed on 20 May 2021).
- Li, Y.R.T.; Smoot, J.; Liu, C.; Watkins, S.; Zhou, O. A comprehensive review of the global efforts on covid-19 vaccine development. ACS Cent. Sci. 2021, 7, 512–533. [Google Scholar] [CrossRef] [PubMed]
- Bonam, S.R.; Partidos, C.D.; Halmuthur, S.K.M.; Muller, S. An overview of novel adjuvants designed for improving vaccine efficacy. Trends Pharm. Sci. 2017, 38, 771–793. [Google Scholar] [CrossRef]
- Amanat, F.; Stadlbauer, D.; Strohmeier, S.; Nguyen, T.H.O.; Chromikova, V.; McMahon, M.; Jiang, K.; Arunkumar, G.A.; Jurczyszak, D.; Polanco, J.; et al. A serological assay to detect sars-cov-2 seroconversion in humans. Nat. Med. 2020, 26, 1033–1036. [Google Scholar] [CrossRef]
- Hotez, P.J.; Corry, D.B.; Bottazzi, M.E. Covid-19 vaccine design: The janus face of immune enhancement. Nat. Rev. Immunol. 2020, 20, 347–348. [Google Scholar] [CrossRef] [PubMed]
- Krammer, F. Sars-cov-2 vaccines in development. Nature 2020, 586, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Syomin, B.V.; Ilyin, Y.V. Virus-like particles as an instrument of vaccine production. Mol. Biol. 2019, 53, 367–379. [Google Scholar] [CrossRef] [PubMed]
- Kushnir, N.; Streatfield, S.J.; Yusibov, V. Virus-like particles as a highly efficient vaccine platform: Diversity of targets and production systems and advances in clinical development. Vaccine 2012, 31, 58–83. [Google Scholar] [CrossRef]
- Corbett, K.S.; Edwards, D.K.; Leist, S.R.; Abiona, O.M.; Boyoglu-Barnum, S.; Gillespie, R.A.; Himansu, S.; Schafer, A.; Ziwawo, C.T.; DiPiazza, A.T.; et al. Sars-cov-2 mrna vaccine design enabled by prototype pathogen preparedness. Nature 2020, 586, 567–571. [Google Scholar] [CrossRef]
- Ulmer, J.B.; Geall, A.J. Recent innovations in mrna vaccines. Curr. Opin. Immunol. 2016, 41, 18–22. [Google Scholar] [CrossRef] [PubMed]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Perez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and efficacy of the bnt162b2 mrna covid-19 vaccine. N. Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and safety of the mrna-1273 sars-cov-2 vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef]
- Cohen, J. ‘Absolutely Remarkable’: No One Who Got Moderna’s Vaccine in Trial Developed Severe Covid-19. 30 November 2020. Available online: https://www.sciencemag.org/news/2020/11/absolutely-remarkable-no-one-who-got-modernas-vaccine-trial-developed-severe-covid-19 (accessed on 20 May 2021).
- Kamar, N.; Abravanel, F.; Marion, O.; Couat, C.; Izopet, J.; Del Bello, A. Three doses of an mrna covid-19 vaccine in solid-organ transplant recipients. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef]
- Callaway, E. Mix-and-match covid vaccines trigger potent immune response. Nature 2021, 593, 491. [Google Scholar] [CrossRef]
- Kremsner, P.; Mann, P.; Bosch, J.; Fendel, R.; Gabor, J.J.; Kreidenweiss, A.; Kroidl, A.; Leroux-Roels, I.; Leroux-Roels, G.; Schindler, C.; et al. Phase 1 assessment of the safety and immunogenicity of an mrna- lipid nanoparticle vaccine candidate against sars-cov-2 in human volunteers. medRxiv 2020. [Google Scholar] [CrossRef]
- Curevac’s Covid-19 Vaccine Candidate, Cvncov, Suitable for Standard Fridge Temperature Logistics. 2020. Available online: https://www.curevac.com/en/2020/11/12/curevacs-covid-19-vaccine-candidate-cvncov-suitable-for-standard-fridge-temperature-logistics/ (accessed on 26 May 2021).
- Dolgin, E. Curevac covid vaccine let-down spotlights mrna design challenges. Nature 2021, 594, 483. [Google Scholar] [CrossRef] [PubMed]
- Silveira, M.M.; Oliveira, T.L.; Schuch, R.A.; McBride, A.J.A.; Dellagostin, O.A.; Hartwig, D.D. DNA vaccines against leptospirosis: A literature review. Vaccine 2017, 35, 5559–5567. [Google Scholar] [CrossRef]
- Smith, T.R.F.; Patel, A.; Ramos, S.; Elwood, D.; Zhu, X.; Yan, J.; Gary, E.N.; Walker, S.N.; Schultheis, K.; Purwar, M.; et al. Immunogenicity of a DNA vaccine candidate for covid-19. Nat. Commun. 2020, 11, 2601. [Google Scholar] [CrossRef]
- Izda, V.; Jeffries, M.A.; Sawalha, A.H. Covid-19: A review of therapeutic strategies and vaccine candidates. Clin. Immunol. 2021, 222, 108634. [Google Scholar] [CrossRef]
- Zhu, F.C.; Guan, X.H.; Li, Y.H.; Huang, J.Y.; Jiang, T.; Hou, L.H.; Li, J.X.; Yang, B.F.; Wang, L.; Wang, W.J.; et al. Immunogenicity and safety of a recombinant adenovirus type-5-vectored covid-19 vaccine in healthy adults aged 18 years or older: A randomised, double-blind, placebo-controlled, phase 2 trial. Lancet 2020, 396, 479–488. [Google Scholar] [CrossRef]
- van Doremalen, N.; Lambe, T.; Spencer, A.; Belij-Rammerstorfer, S.; Purushotham, J.N.; Port, J.R.; Avanzato, V.A.; Bushmaker, T.; Flaxman, A.; Ulaszewska, M.; et al. Chadox1 ncov-19 vaccine prevents sars-cov-2 pneumonia in rhesus macaques. Nature 2020, 586, 578–582. [Google Scholar] [CrossRef]
- Ewer, K.J.; Barrett, J.R.; Belij-Rammerstorfer, S.; Sharpe, H.; Makinson, R.; Morter, R.; Flaxman, A.; Wright, D.; Bellamy, D.; Bittaye, M.; et al. T cell and antibody responses induced by a single dose of chadox1 ncov-19 (azd1222) vaccine in a phase 1/2 clinical trial. Nat. Med. 2021, 27, 270–278. [Google Scholar] [CrossRef] [PubMed]
- Voysey, M.; Clemens, S.A.C.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E.; et al. Safety and efficacy of the chadox1 ncov-19 vaccine (azd1222) against sars-cov-2: An interim analysis of four randomised controlled trials in brazil, south africa, and the uk. Lancet 2021, 397, 99–111. [Google Scholar] [CrossRef]
- Ohnson & Johnson Announces Single-Shot Janssen Covid-19 Vaccine Candidate Met Primary Endpoints in Interim Analysis of Its Phase 3 Ensemble Trial. 2021. Available online: https://www.janssen.com/johnson-johnson-announces-single-shot-janssen-covid-19-vaccine-candidate-met-primary-endpoints (accessed on 26 May 2021).
- Kowarz, E.L.K.; Reis, J.; Bracharz, S.; Kochanek, S.; Marschalek, R. Vaccine-induced covid-19 mimicry” syndrome:Splice reactions within the sars-cov-2 spike open reading frame result in spike protein variants that may cause thromboembolic events in patients immunized with vector-based vaccines. Res. Sq. 2021. [Google Scholar] [CrossRef]
- Logunov, D.Y.; Dolzhikova, I.V.; Shcheblyakov, D.V.; Tukhvatulin, A.I.; Zubkova, O.V.; Dzharullaeva, A.S.; Kovyrshina, A.V.; Lubenets, N.L.; Grousova, D.M.; Erokhova, A.S.; et al. Safety and efficacy of an rad26 and rad5 vector-based heterologous prime-boost covid-19 vaccine: An interim analysis of a randomised controlled phase 3 trial in russia. Lancet 2021, 397, 671–681. [Google Scholar] [CrossRef]
- CNBC. Russia Authorizes use of ‘Sputnik Light,’ a One-Shot Covid Vaccine It Says is 79% Effective. CNBC. 6 May 2021. Available online: https://www.cnbc.com/2021/05/06/sputnik-light-russia-authorizes-use-of-one-shot-covid-vaccine.html (accessed on 26 May 2021).
- Sun, W.; Leist, S.R.; McCroskery, S.; Liu, Y.; Slamanig, S.; Oliva, J.; Amanat, F.; Schafer, A.; Dinnon, K.H., 3rd; Garcia-Sastre, A.; et al. Newcastle disease virus (ndv) expressing the spike protein of sars-cov-2 as a live virus vaccine candidate. EBioMedicine 2020, 62, 103132. [Google Scholar] [CrossRef] [PubMed]
- Case, J.B.; Rothlauf, P.W.; Chen, R.E.; Kafai, N.M.; Fox, J.M.; Smith, B.K.; Shrihari, S.; McCune, B.T.; Harvey, I.B.; Keeler, S.P.; et al. Replication-competent vesicular stomatitis virus vaccine vector protects against sars-cov-2-mediated pathogenesis in mice. Cell Host Microbe 2020, 28, 465–474. [Google Scholar] [CrossRef] [PubMed]
- Gao, Q.; Bao, L.; Mao, H.; Wang, L.; Xu, K.; Yang, M.; Li, Y.; Zhu, L.; Wang, N.; Lv, Z.; et al. Development of an inactivated vaccine candidate for sars-cov-2. Science 2020, 369, 77–81. [Google Scholar] [CrossRef]
- Xia, S.; Duan, K.; Zhang, Y.; Zhao, D.; Zhang, H.; Xie, Z.; Li, X.; Peng, C.; Zhang, Y.; Zhang, W.; et al. Effect of an inactivated vaccine against sars-cov-2 on safety and immunogenicity outcomes: Interim analysis of 2 randomized clinical trials. JAMA 2020, 324, 951–960. [Google Scholar] [CrossRef] [PubMed]
- Mallapaty, S. China covid vaccine reports mixed results—what does that mean for the pandemic? Nature 2021. [Google Scholar] [CrossRef]
- Biotech, B. Covaxin—ndia’s First Indigenous Covid-19 Vaccine. 2021. Available online: https://www.bharatbiotech.com/covaxin.html (accessed on 26 May 2021).
- Broadbent, A.J.; Santos, C.P.; Anafu, A.; Wimmer, E.; Mueller, S.; Subbarao, K. Evaluation of the attenuation, immunogenicity, and efficacy of a live virus vaccine generated by codon-pair bias de-optimization of the 2009 pandemic h1n1 influenza virus, in ferrets. Vaccine 2016, 34, 563–570. [Google Scholar] [CrossRef] [Green Version]
- Codagenix. Covi-Vac for Sars-Cov-2 (Covid-19). Available online: https://codagenix.com/vaccine-programs/covid-19/ (accessed on 26 May 2021).
- Meissa Vaccines, Attenublock for Optimized Immunity. Available online: https://www.meissavaccines.com/technolog (accessed on 27 May 2021).
- Wang, Q.; Wu, J.; Wang, H.; Gao, Y.; Liu, Q.; Mu, A.; Ji, W.; Yan, L.; Zhu, Y.; Zhu, C.; et al. Structural basis for rna replication by the sars-cov-2 polymerase. Cell 2020, 182, 417–428. [Google Scholar] [CrossRef]
- Gordon, C.J.; Tchesnokov, E.P.; Woolner, E.; Perry, J.K.; Feng, J.Y.; Porter, D.P.; Gotte, M. Remdesivir is a direct-acting antiviral that inhibits rna-dependent rna polymerase from severe acute respiratory syndrome coronavirus 2 with high potency. J. Biol. Chem. 2020, 295, 6785–6797. [Google Scholar] [CrossRef] [Green Version]
- Choy, K.T.; Wong, A.Y.; Kaewpreedee, P.; Sia, S.F.; Chen, D.; Hui, K.P.Y.; Chu, D.K.W.; Chan, M.C.W.; Cheung, P.P.; Huang, X.; et al. Remdesivir, lopinavir, emetine, and homoharringtonine inhibit sars-cov-2 replication in vitro. Antivir. Res. 2020, 178, 104786. [Google Scholar] [CrossRef]
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-ncov) in vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef]
- Pizzorno, A.; Padey, B.; Julien, T.; Trouillet-Assant, S.; Traversier, A.; Errazuriz-Cerda, E.; Fouret, J.; Dubois, J.; Gaymard, A.; Lescure, F.X.; et al. Characterization and treatment of sars-cov-2 in nasal and bronchial human airway epithelia. Cell Rep. Med. 2020, 1, 100059. [Google Scholar] [CrossRef]
- Bafna, K.; White, K.; Harish, B.; Rosales, R.; Ramelot, T.A.; Acton, T.B.; Moreno, E.; Kehrer, T.; Miorin, L.; Royer, C.A.; et al. Hepatitis c virus drugs that inhibit sars-cov-2 papain-like protease synergize with remdesivir to suppress viral replication in cell culture. Cell Rep. 2021, 35, 109133. [Google Scholar] [CrossRef]
- Williamson, B.N.; Feldmann, F.; Schwarz, B.; Meade-White, K.; Porter, D.P.; Schulz, J.; van Doremalen, N.; Leighton, I.; Yinda, C.K.; Perez-Perez, L.; et al. Clinical benefit of remdesivir in rhesus macaques infected with sars-cov-2. Nature 2020, 585, 273–276. [Google Scholar] [CrossRef]
- Beigel, J.H.; Tomashek, K.M.; Dodd, L.E.; Mehta, A.K.; Zingman, B.S.; Kalil, A.C.; Hohmann, E.; Chu, H.Y.; Luetkemeyer, A.; Kline, S.; et al. Remdesivir for the treatment of covid-19—final report. N. Engl. J. Med. 2020, 383, 1813–1826. [Google Scholar] [CrossRef]
- Robinson, R.; Prakash, V.; Al Tamimi, R.; Albast, N.; Al-Bast, B.; Wieland, E.; Garcia, C. Impact of remdesivir on 28 day mortality in hospitalized patients with covid-19: February 2021 meta-analysis. medRxiv 2021. [Google Scholar] [CrossRef]
- Consortium, W.H.O.S.T.; Pan, H.; Peto, R.; Henao-Restrepo, A.M.; Preziosi, M.P.; Sathiyamoorthy, V.; Abdool Karim, Q.; Alejandria, M.M.; Hernandez Garcia, C.; Kieny, M.P.; et al. Repurposed antiviral drugs for covid-19—Interim who solidarity trial results. N. Engl. J. Med. 2021, 384, 497–511. [Google Scholar] [CrossRef] [PubMed]
- NIH. Therapeutic Management of Adults with Covid-19. 2021. Available online: https://www.covid19treatmentguidelines.nih.gov/management/therapeutic-management/ (accessed on 22 May 2021).
- Summary, P.C. Pubchem: Dexamethasone. Available online: https://pubchem.ncbi.nlm.nih.gov/compound/Dexamethasone (accessed on 22 May 2021).
- Ahmed, M.H.; Hassan, A. Dexamethasone for the treatment of coronavirus disease (covid-19): A review. SN Compr. Clin. Med. 2020, 1–10. [Google Scholar] [CrossRef]
- Stockman, L.J.; Bellamy, R.; Garner, P. Sars: Systematic review of treatment effects. PLoS Med. 2006, 3, e343. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Villar, J.; Confalonieri, M.; Pastores, S.M.; Meduri, G.U. Rationale for prolonged corticosteroid treatment in the acute respiratory distress syndrome caused by coronavirus disease 2019. Crit. Care Explor. 2020, 2, e0111. [Google Scholar] [CrossRef] [Green Version]
- Villar, J.; Ferrando, C.; Martinez, D.; Ambros, A.; Munoz, T.; Soler, J.A.; Aguilar, G.; Alba, F.; Gonzalez-Higueras, E.; Conesa, L.A.; et al. Dexamethasone treatment for the acute respiratory distress syndrome: A multicentre, randomised controlled trial. Lancet Respir. Med. 2020, 8, 267–276. [Google Scholar] [CrossRef]
- Group, R.C.; Horby, P.; Lim, W.S.; Emberson, J.R.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Brightling, C.; Ustianowski, A.; et al. Dexamethasone in hospitalized patients with covid-19. N. Engl. J. Med. 2021, 384, 693–704. [Google Scholar]
- The WHO Rapid Evidence Appraisal for COVID-19 Therapies (REACT) Working Group; Sterne, J.A.C.; Murthy, S.; Diaz, J.V.; Slutsky, A.S.; Villar, J.; Angus, D.C.; Annane, D.; Azevedo, L.C.P.; Berwanger, O.; et al. Association between administration of systemic corticosteroids and mortality among critically ill patients with covid-19: A meta-analysis. JAMA 2020, 324, 1330–1341. [Google Scholar]
- NIH COVID-19 Treatment Guidelines, Corticosteroids. Available online: https://www.covid19treatmentguidelines.nih.gov/immunomodulators/corticosteroids/ (accessed on 22 May 2021).
- Delang, L.; Abdelnabi, R.; Neyts, J. Favipiravir as a potential countermeasure against neglected and emerging rna viruses. Antivir. Res. 2018, 153, 85–94. [Google Scholar] [CrossRef]
- Sissoko, D.; Laouenan, C.; Folkesson, E.; M’Lebing, A.B.; Beavogui, A.H.; Baize, S.; Camara, A.M.; Maes, P.; Shepherd, S.; Danel, C.; et al. Experimental treatment with favipiravir for ebola virus disease (the jiki trial): A historically controlled, single-arm proof-of-concept trial in guinea. PLoS Med. 2016, 13, e1001967. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Furuta, Y.; Komeno, T.; Nakamura, T. Favipiravir (t-705), a broad spectrum inhibitor of viral rna polymerase. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2017, 93, 449–463. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cai, Q.; Yang, M.; Liu, D.; Chen, J.; Shu, D.; Xia, J.; Liao, X.; Gu, Y.; Cai, Q.; Yang, Y.; et al. Experimental treatment with favipiravir for covid-19: An open-label control study. Engineering 2020, 6, 1192–1198. [Google Scholar] [CrossRef]
- Hassanipour, S.; Arab-Zozani, M.; Amani, B.; Heidarzad, F.; Fathalipour, M.; Martinez-de-Hoyo, R. The efficacy and safety of favipiravir in treatment of covid-19: A systematic review and meta-analysis of clinical trials. medRxiv 2021. [Google Scholar] [CrossRef]
- Redhill Biopharma. 2020. Available online: https://www.redhillbio.com/RedHill/Templates/showpage.asp?DBID=1&LNGID=1&TMID=178&FID=2432&PID=0&IID=19319 (accessed on 20 May 2021).
- Villiger, P.M.; Adler, S.; Kuchen, S.; Wermelinger, F.; Dan, D.; Fiege, V.; Butikofer, L.; Seitz, M.; Reichenbach, S. Tocilizumab for induction and maintenance of remission in giant cell arteritis: A phase 2, randomised, double-blind, placebo-controlled trial. Lancet 2016, 387, 1921–1927. [Google Scholar] [CrossRef]
- Galeotti, C.; Boucheron, A.; Guillaume, S.; Kone-Paut, I. Sustained remission of multicentric castleman disease in children treated with tocilizumab, an anti-interleukin-6 receptor antibody. Mol. Cancer 2012, 11, 1623–1626. [Google Scholar] [CrossRef] [Green Version]
- Le, R.Q.; Li, L.; Yuan, W.; Shord, S.S.; Nie, L.; Habtemariam, B.A.; Przepiorka, D.; Farrell, A.T.; Pazdur, R. Fda approval summary: Tocilizumab for treatment of chimeric antigen receptor t cell-induced severe or life-threatening cytokine release syndrome. Oncologist 2018, 23, 943–947. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Investigators, R.-C.; Gordon, A.C.; Mouncey, P.R.; Al-Beidh, F.; Rowan, K.M.; Nichol, A.D.; Arabi, Y.M.; Annane, D.; Beane, A.; van Bentum-Puijk, W.; et al. Interleukin-6 receptor antagonists in critically ill patients with covid-19. N. Engl. J. Med. 2021, 384, 1491–1502. [Google Scholar] [CrossRef] [PubMed]
- Horby, P.W.; Pessoa-Amorim, G.; Peto, L.; Brightling, C.E.; Sarkar, R.; Thomas, K.; Jeebun, V.; Ashish, A.; Tully, R.; Chadwick, D.; et al. Tocilizumab in patients admitted to hospital with covid-19 (recovery): Preliminary results of a randomised, controlled, open-label, platform trial. medRxiv 2021. [Google Scholar] [CrossRef]
- NIH COVID-19 Treatment Guidelines, Tocilizumab. Available online: https://www.covid19treatmentguidelines.nih.gov/immunomodulators/interleukin-6-inhibitors/ (accessed on 26 May 2021).
- Savarino, A.; Boelaert, J.R.; Cassone, A.; Majori, G.; Cauda, R. Effects of chloroquine on viral infections: An old drug against today’s diseases? Lancet Infect. Dis. 2003, 3, 722–727. [Google Scholar] [CrossRef]
- Colson, P.; Rolain, J.M.; Raoult, D. Chloroquine for the 2019 novel coronavirus sars-cov-2. Int. J. Antimicrob. Agents 2020, 55, 105923. [Google Scholar] [CrossRef]
- Keyaerts, E.; Vijgen, L.; Maes, P.; Neyts, J.; Van Ranst, M. In vitro inhibition of severe acute respiratory syndrome coronavirus by chloroquine. Biochem. Biophys. Res. Commun. 2004, 323, 264–268. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Cao, R.; Xu, M.; Wang, X.; Zhang, H.; Hu, H.; Li, Y.; Hu, Z.; Zhong, W.; Wang, M. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting sars-cov-2 infection in vitro. Cell Discov. 2020, 6, 16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Funnell, S.G.P.; Dowling, W.E.; Munoz-Fontela, C.; Gsell, P.S.; Ingber, D.E.; Hamilton, G.A.; Delang, L.; Rocha-Pereira, J.; Kaptein, S.; Dallmeier, K.H.; et al. Emerging preclinical evidence does not support broad use of hydroxychloroquine in covid-19 patients. Nat. Commun 2020, 11, 4253. [Google Scholar] [CrossRef] [PubMed]
- Horby, P.; Mafham, M.; Linsell, L.; Bell, J.L.; Staplin, N.; Emberson, J.R.; Wiselka, M.; Ustianowski, A.; Elmahi, E.; Prudon, B.; et al. Effect of hydroxychloroquine in hospitalized patients with covid-19: Preliminary results from a multi-centre, randomized, controlled trial. medRxiv 2020. [Google Scholar] [CrossRef]
- Geleris, J.; Sun, Y.; Platt, J.; Zucker, J.; Baldwin, M.; Hripcsak, G.; Labella, A.; Manson, D.K.; Kubin, C.; Barr, R.G.; et al. Observational study of hydroxychloroquine in hospitalized patients with covid-19. N. Engl. J. Med. 2020, 382, 2411–2418. [Google Scholar] [CrossRef]
- Molina, J.M.; Delaugerre, C.; Le Goff, J.; Mela-Lima, B.; Ponscarme, D.; Goldwirt, L.; de Castro, N. No evidence of rapid antiviral clearance or clinical benefit with the combination of hydroxychloroquine and azithromycin in patients with severe covid-19 infection. Med. Mal. Infect. 2020, 50, 384. [Google Scholar] [CrossRef] [PubMed]
- NIH COVID-19 Treatment Guidelines, Chloroquine/Hydroxychloroquine. Available online: https://www.covid19treatmentguidelines.nih.gov/antiviral-therapy/chloroquine-or-hydroxychloroquine-with-or-without-azithromycin/ (accessed on 20 May 2021).
- FDA. Olumiant (Baricitinib). Available online: https://www.fda.gov/drugs/drug-approvals-and-databases/drug-trials-snapshots-olumiant (accessed on 26 May 2021).
- McInnes, I.B.; Byers, N.L.; Higgs, R.E.; Lee, J.; Macias, W.L.; Na, S.; Ortmann, R.A.; Rocha, G.; Rooney, T.P.; Wehrman, T.; et al. Comparison of baricitinib, upadacitinib, and tofacitinib mediated regulation of cytokine signaling in human leukocyte subpopulations. Arthritis Res. 2019, 21, 183. [Google Scholar] [CrossRef] [Green Version]
- Stebbing, J.; Krishnan, V.; de Bono, S.; Ottaviani, S.; Casalini, G.; Richardson, P.J.; Monteil, V.; Lauschke, V.M.; Mirazimi, A.; Youhanna, S.; et al. Mechanism of baricitinib supports artificial intelligence-predicted testing in covid-19 patients. EMBO Mol. Med. 2020, 12, e12697. [Google Scholar] [CrossRef]
- Kalil, A.C.; Patterson, T.F.; Mehta, A.K.; Tomashek, K.M.; Wolfe, C.R.; Ghazaryan, V.; Marconi, V.C.; Ruiz-Palacios, G.M.; Hsieh, L.; Kline, S.; et al. Baricitinib plus remdesivir for hospitalized adults with covid-19. N. Engl. J. Med. 2021, 384, 795–807. [Google Scholar] [CrossRef] [PubMed]
- Marovich, M.; Mascola, J.R.; Cohen, M.S. Monoclonal antibodies for prevention and treatment of covid-19. JAMA 2020, 324, 131–132. [Google Scholar] [CrossRef] [PubMed]
- FDA. Coronavirus (Covid-19) Update: Fda Authorizes Monoclonal Antibodies for Treatment of Covid-19. Available online: https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-authorizes-monoclonal-antibodies-treatment-covid-19 (accessed on 26 May 2021).
- Gottlieb, R.L.; Nirula, A.; Chen, P.; Boscia, J.; Heller, B.; Morris, J.; Huhn, G.; Cardona, J.; Mocherla, B.; Stosor, V.; et al. Effect of bamlanivimab as monotherapy or in combination with etesevimab on viral load in patients with mild to moderate covid-19: A randomized clinical trial. JAMA 2021, 325, 632–644. [Google Scholar] [CrossRef]
- Gsk and Vir Biotechnology Announce Sotrovimab (vir-7831) Receives Emergency Use Authorization from the US FDA for Treatment of Mild-to-Moderate Covid-19 in High-Risk Adults and Pediatric Patients. Available online: https://www.globenewswire.com/news-release/2021/05/26/2236926/0/en/GSK-and-Vir-Biotechnology-Announce-Sotrovimab-VIR-7831-Receives-Emergency-Use-Authorization-from-the-US-FDA-for-Treatment-of-Mild-to-Moderate-COVID-19-in-High-Risk-Adults-and-Pedia.html (accessed on 24 May 2021).
- Maiztegui, J.I.; Fernandez, N.J.; de Damilano, A.J. Efficacy of immune plasma in treatment of argentine haemorrhagic fever and association between treatment and a late neurological syndrome. Lancet 1979, 2, 1216–1217. [Google Scholar] [CrossRef]
- Chen, L.; Xiong, J.; Bao, L.; Shi, Y. Convalescent plasma as a potential therapy for covid-19. Lancet Infect. Dis. 2020, 20, 398–400. [Google Scholar] [CrossRef]
- Horby, P.W.; Estcourt, L.; Peto, L.; Emberson, J.R.; Staplin, N.; Spata, E.; Pessoa-Amorim, G.; Campbell, M.; Roddick, A.; Brunskill, N.E.; et al. Convalescent plasma in patients admitted to hospital with covid-19 (recovery): A randomised, controlled, open-label, platform trial. medRxiv 2021. [Google Scholar] [CrossRef]
- Simonovich, V.A.; Burgos Pratx, L.D.; Scibona, P.; Beruto, M.V.; Vallone, M.G.; Vazquez, C.; Savoy, N.; Giunta, D.H.; Perez, L.G.; Sanchez, M.D.L.; et al. A randomized trial of convalescent plasma in covid-19 severe pneumonia. N. Engl. J. Med. 2021, 384, 619–629. [Google Scholar] [CrossRef]
- FDA Updates Emergency Use Authorization for Covid-19 Convalescent Plasma to Reflect New Data. Available online: https://www.fda.gov/news-events/fda-brief/fda-brief-fda-updates-emergency-use-authorization-covid-19-convalescent-plasma-reflect-new-data (accessed on 26 May 2021).
- Martinez-Delgado, G. Inhaled nanobodies against covid-19. Nat. Rev. Immunol. 2020, 20, 593. [Google Scholar] [CrossRef]
- Nambulli, S.; Xiang, Y.; Tilston-Lunel, N.L.; Rennick, L.J.; Sang, Z.; Klimstra, W.B.; Reed, D.S.; Crossland, N.A.; Shi, Y.; Duprex, W.P. Inhalable nanobody (pin-21) prevents and treats sars-cov-2 infections in syrian hamsters at ultra-low doses. bioRxiv 2021. [Google Scholar] [CrossRef]
- Shetty, A.K. Mesenchymal stem cell infusion shows promise for combating coronavirus (covid-19)—Induced pneumonia. Aging Dis. 2020, 11, 462–464. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leng, Z.; Zhu, R.; Hou, W.; Feng, Y.; Yang, Y.; Han, Q.; Shan, G.; Meng, F.; Du, D.; Wang, S.; et al. Transplantation of ace2(-) mesenchymal stem cells improves the outcome of patients with covid-19 pneumonia. Aging Dis. 2020, 11, 216–228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Humanigen. Humanigen Reports Positive Phase 3 Topline Results Demonstrating That Lenzilumab™ Improves Survival without Need for Mechanical Ventilation in Hospitalized Patients with Covid-19. 2021. Available online: https://www.biospace.com/article/releases/humanigen-reports-positive-phase-3-topline-results-demonstrating-that-lenzilumab-improves-survival-without-need-for-mechanical-ventilation-in-hospitalized-patients-with-covid-19/ (accessed on 26 May 2021).
- Eusa Pharma Announces FDA Approval of Phase 3 Clinical Trial for Siltuximab in Hospitalized Patients with Covid-19 Associated Acute Respiratory Distress Syndrome. 2020. Available online: https://eusapharma.com/news/eusa-pharma-announces-fda-approval-of-phase-3-clinical-trial-for-siltuximab-in-hospitalized-patients-with-covid-19-associated-acute-respiratory-distress-syndrome (accessed on 20 May 2021).
- Merck and Ridgeback Biotherapeutics Provide Update on Progress of Clinical Development Program for Molnupiravir, An Investigational Oral Therapeutic for the Treatment of Mild-To-Moderate Covid-19. 2021. Available online: https://www.merck.com/news/merck-and-ridgeback-biotherapeutics-provide-update-on-progress-of-clinical-development-program-for-molnupiravir-an-investigational-oral-therapeutic-for-the-treatment-of-mild-to-moderate-covid-19/ (accessed on 20 May 2021).
- Sng001 Reduces Viral Load. Available online: https://www.synairgen.com/covid-19/ (accessed on 20 May 2021).
- Proof of Concept Study to Evaluate the Safety Profile of Plitidepsin in Patients with Covid-19 (Aplicov-pc). Available online: https://clinicaltrials.gov/ct2/show/NCT04382066 (accessed on 20 May 2021).
- Pfizer Initiates Phase 1 Study of Novel Oral Antiviral Therapeutic Agent against Sars-Cov-2. 2021. Available online: https://www.pfizer.com/news/press-release/press-release-detail/pfizer-initiates-phase-1-study-novel-oral-antiviral (accessed on 20 May 2021).
- Graham, R.L.; Becker, M.M.; Eckerle, L.D.; Bolles, M.; Denison, M.R.; Baric, R.S. A live, impaired-fidelity coronavirus vaccine protects in an aged, immunocompromised mouse model of lethal disease. Nat. Med. 2012, 18, 1820–1826. [Google Scholar] [CrossRef] [Green Version]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking changes in sars-cov-2 spike: Evidence that d614g increases infectivity of the covid-19 virus. Cell 2020, 182, 812–827. [Google Scholar] [CrossRef]
- Plante, J.A.; Liu, Y.; Liu, J.; Xia, H.; Johnson, B.A.; Lokugamage, K.G.; Zhang, X.; Muruato, A.E.; Zou, J.; Fontes-Garfias, C.R.; et al. Spike mutation d614g alters sars-cov-2 fitness. Nature 2021, 592, 116–121. [Google Scholar] [CrossRef]
- Zhou, B.; Thao, T.T.N.; Hoffmann, D.; Taddeo, A.; Ebert, N.; Labroussaa, F.; Pohlmann, A.; King, J.; Steiner, S.; Kelly, J.N.; et al. Sars-cov-2 spike d614g change enhances replication and transmission. Nature 2021, 592, 122–127. [Google Scholar] [CrossRef]
- Anderson, E.J.; Rouphael, N.G.; Widge, A.T.; Jackson, L.A.; Roberts, P.C.; Makhene, M.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; Pruijssers, A.J.; et al. Safety and immunogenicity of sars-cov-2 mrna-1273 vaccine in older adults. N. Engl. J. Med. 2020, 383, 2427–2438. [Google Scholar] [CrossRef]
- Chand, G.B.; Banerjee, A.; Azad, G.K. Identification of novel mutations in rna-dependent rna polymerases of sars-cov-2 and their implications on its protein structure. PeerJ 2020, 8, e9492. [Google Scholar] [CrossRef] [PubMed]
- Davies, N.G.; Abbott, S.; Barnard, R.C.; Jarvis, C.I.; Kucharski, A.J.; Munday, J.D.; Pearson, C.A.B.; Russell, T.W.; Tully, D.C.; Washburne, A.D.; et al. Estimated transmissibility and impact of sars-cov-2 lineage b.1.1.7 in england. Science 2021, 372. [Google Scholar] [CrossRef] [PubMed]
- Leung, K.; Shum, M.H.; Leung, G.M.; Lam, T.T.; Wu, J.T. Early transmissibility assessment of the n501y mutant strains of sars-cov-2 in the united kingdom, october to november 2020. Euro Surveill. 2021, 26, 2002106. [Google Scholar] [CrossRef] [PubMed]
- Volz, E.; Mishra, S.; Chand, M.; Barrett, J.C.; Johnson, R.; Geidelberg, L.; Hinsley, W.R.; Laydon, D.J.; Dabrera, G.; O’Toole, A.; et al. Assessing transmissibility of sars-cov-2 lineage b.1.1.7 in england. Nature 2021, 593, 266–269. [Google Scholar] [CrossRef]
- Kemp, S.A.; Collier, D.A.; Datir, R.; Ferreira, I.; Gayed, S.; Jahun, A.; Hosmillo, M.; Rees-Spear, C.; Mlcochova, P.; Lumb, I.U.; et al. Neutralising antibodies in spike mediated sars-cov-2 adaptation. medRxiv 2020. [Google Scholar] [CrossRef]
- McCarthy, K.R.; Rennick, L.J.; Nambulli, S.; Robinson-McCarthy, L.R.; Bain, W.G.; Haidar, G.; Duprex, W.P. Recurrent deletions in the sars-cov-2 spike glycoprotein drive antibody escape. Science 2021, 371, 1139–1142. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, J.; Plante, K.S.; Plante, J.A.; Xie, X.; Zhang, X.; Ku, Z.; An, Z.; Scharton, D.; Schindewolf, C.; et al. The n501y spike substitution enhances sars-cov-2 transmission. bioRxiv 2021. [Google Scholar] [CrossRef]
- Starr, T.N.; Greaney, A.J.; Addetia, A.; Hannon, W.W.; Choudhary, M.C.; Dingens, A.S.; Li, J.Z.; Bloom, J.D. Prospective mapping of viral mutations that escape antibodies used to treat covid-19. Science 2021, 371, 850–854. [Google Scholar] [CrossRef]
- Peacock, T.P.; Goldhill, D.H.; Zhou, J.; Baillon, L.; Frise, R.; Swann, O.C.; Kugathasan, R.; Penn, R.; Brown, J.C.; Sanchez-David, R.Y.; et al. The furin cleavage site in the sars-cov-2 spike protein is required for transmission in ferrets. Nat. Microbiol. 2021. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Pohlmann, S. A multibasic cleavage site in the spike protein of sars-cov-2 is essential for infection of human lung cells. Mol. Cell 2020, 78, 779–784. [Google Scholar] [CrossRef] [PubMed]
- Collier, D.A.; De Marco, A.; Ferreira, I.; Meng, B.; Datir, R.P.; Walls, A.C.; Kemp, S.A.; Bassi, J.; Pinto, D.; Silacci-Fregni, C.; et al. Sensitivity of sars-cov-2 b.1.1.7 to mrna vaccine-elicited antibodies. Nature 2021, 593, 136–141. [Google Scholar] [CrossRef]
- Xie, X.; Liu, Y.; Liu, J.; Zhang, X.; Zou, J.; Fontes-Garfias, C.R.; Xia, H.; Swanson, K.A.; Cutler, M.; Cooper, D.; et al. Neutralization of sars-cov-2 spike 69/70 deletion, e484k and n501y variants by bnt162b2 vaccine-elicited sera. Nat. Med. 2021, 27, 620–621. [Google Scholar] [CrossRef]
- Wang, P.; Nair, M.S.; Liu, L.; Iketani, S.; Luo, Y.; Guo, Y.; Wang, M.; Yu, J.; Zhang, B.; Kwong, P.D.; et al. Increased resistance of sars-cov-2 variants b.1.351 and b.1.1.7 to antibody neutralization. bioRxiv 2021. [Google Scholar] [CrossRef]
- Wu, K.; Werner, A.P.; Moliva, J.I.; Koch, M.; Choi, A.; Stewart-Jones, G.B.E.; Bennett, H.; Boyoglu-Barnum, S.; Shi, W.; Graham, B.S.; et al. Mrna-1273 vaccine induces neutralizing antibodies against spike mutants from global sars-cov-2 variants. bioRxiv 2021. [Google Scholar] [CrossRef]
- Ikegame, S.; Siddiquey, M.; Hung, C.T.; Haas, G.; Brambilla, L.; Oguntuyo, K.; Kowdle, S.; Vilardo, A.; Edelstein, A.; Perandones, C.; et al. Neutralizing activity of sputnik v vaccine sera against sars-cov-2 variants. Res. Sq. 2021. [Google Scholar] [CrossRef]
- NIH. Covid-19 Treatment Guidelines Panel. Coronavirus Disease 2019 (Covid-19) Treatment Guidelines. National Institutes of Health. Available online: https://www.Covid19treatmentguidelines.Nih.Gov/ (accessed on 29 May 2021).
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; Doolabh, D.; Pillay, S.; San, E.J.; Msomi, N.; et al. Detection of a sars-cov-2 variant of concern in south africa. Nature 2021, 592, 438–443. [Google Scholar] [CrossRef]
- Gard Nelson, O.B.; Spilman, P.; Niazi, K.; Rabizadeh, S.; Soon-Shiong, P. Molecular dynamic simulation reveals e484k mutation enhances spike rbd-ace2 affinity and the combination of e484k, k417n and n501y mutations (501y.V2 variant) induces conformational change greater than n501y mutant alone, potentially resulting in an escape mutant. BioRxiv 2021. [Google Scholar] [CrossRef]
- Wang, G.L.; Wang, Z.Y.; Duan, L.J.; Meng, Q.C.; Jiang, M.D.; Cao, J.; Yao, L.; Zhu, K.L.; Cao, W.C.; Ma, M.J. Susceptibility of circulating sars-cov-2 variants to neutralization. N. Engl. J. Med. 2021, 384, 2354–2356. [Google Scholar] [CrossRef] [PubMed]
- Shinde, V.; Bhikha, S.; Hoosain, Z.; Archary, M.; Bhorat, Q.; Fairlie, L.; Lalloo, U.; Masilela, M.S.L.; Moodley, D.; Hanley, S.; et al. Efficacy of nvx-cov2373 covid-19 vaccine against the b.1.351 variant. N. Engl. J. Med. 2021, 384, 1899–1909. [Google Scholar] [CrossRef] [PubMed]
- Madhi, S.A.; Baillie, V.; Cutland, C.L.; Voysey, M.; Koen, A.L.; Fairlie, L.; Padayachee, S.D.; Dheda, K.; Barnabas, S.L.; Bhorat, Q.E.; et al. Efficacy of the chadox1 ncov-19 covid-19 vaccine against the b.1.351 variant. N. Engl. J. Med. 2021, 384, 1885–1898. [Google Scholar] [CrossRef]
- Abu-Raddad, L.J.; Chemaitelly, H.; Butt, A.A.; National Study Group for COVID-19 Vaccination. Effectiveness of the bnt162b2 covid-19 vaccine against the b.1.1.7 and b.1.351 variants. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef] [PubMed]
- Dagan, N.; Barda, N.; Kepten, E.; Miron, O.; Perchik, S.; Katz, M.A.; Hernan, M.A.; Lipsitch, M.; Reis, B.; Balicer, R.D. Bnt162b2 mrna covid-19 vaccine in a nationwide mass vaccination setting. N. Engl. J. Med. 2021, 384, 1412–1423. [Google Scholar] [CrossRef] [PubMed]
- Thompson, M.G.; Burgess, J.L.; Naleway, A.L.; Tyner, H.L.; Yoon, S.K.; Meece, J.; Olsho, L.E.W.; Caban-Martinez, A.J.; Fowlkes, A.; Lutrick, K.; et al. Interim estimates of vaccine effectiveness of bnt162b2 and mrna-1273 covid-19 vaccines in preventing sars-cov-2 infection among health care personnel, first responders, and other essential and frontline workers—Eight u.S. Locations, december 2020-march 2021. MMWR Morb. Mortal Wkly. Rep. 2021, 70, 495–500. [Google Scholar]
- Wibmer, C.K.; Ayres, F.; Hermanus, T.; Madzivhandila, M.; Kgagudi, P.; Oosthuysen, B.; Lambson, B.E.; de Oliveira, T.; Vermeulen, M.; van der Berg, K.; et al. Sars-cov-2 501y.V2 escapes neutralization by south african covid-19 donor plasma. bioRxiv 2021. [Google Scholar] [CrossRef]
- Sabino, E.C.; Buss, L.F.; Carvalho, M.P.S.; Prete, C.A., Jr.; Crispim, M.A.E.; Fraiji, N.A.; Pereira, R.H.M.; Parag, K.V.; da Silva Peixoto, P.; Kraemer, M.U.G.; et al. Resurgence of covid-19 in manaus, brazil, despite high seroprevalence. Lancet 2021, 397, 452–455. [Google Scholar] [CrossRef]
- Faria, N.R.; Mellan, T.A.; Whittaker, C.; Claro, I.M.; Candido, D.D.S.; Mishra, S.; Crispim, M.A.E.; Sales, F.C.S.; Hawryluk, I.; McCrone, J.T.; et al. Genomics and epidemiology of the p.1 sars-cov-2 lineage in manaus, brazil. Science 2021, 372, 815–821. [Google Scholar] [CrossRef]
- Naveca, F.G.; Nascimento, V.; de Souza, V.C.; Corado, A.L.; Nascimento, F.; Silva, G.; Costa, A.; Duarte, D.; Pessoa, K.; Mejia, M.; et al. Covid-19 in amazonas, brazil, was driven by the persistence of endemic lineages and p.1 emergence. Nat. Med. 2021. [Google Scholar] [CrossRef]
- Abdool Karim, S.S.; de Oliveira, T. New sars-cov-2 variants—clinical, public health, and vaccine implications. N. Engl. J. Med. 2021, 384, 1866–1868. [Google Scholar] [CrossRef]
- de Souza, W.M.; Amorim, M.R.; Sesti-Costa, R.; Coimbra, L.D.; de Toledo-Teixeira, D.A.; Parise, P.L.; Barbosa, P.P.; Bispo-dos-Santos, K.; Mofatto, L.S.; Simeoni, C.L.; et al. Levels of sars-cov-2 lineage p.1 neutralization by antibodies elicited after natural infection and vaccination. Lancet 2021. [Google Scholar] [CrossRef]
- Moutinho, S. Chinese covid-19 vaccine maintains protection in variant-plagued brazil. Science 2021. [Google Scholar] [CrossRef]
- Deng, X.; Garcia-Knight, M.A.; Khalid, M.M.; Servellita, V.; Wang, C.; Morris, M.K.; Sotomayor-Gonzalez, A.; Glasner, D.R.; Reyes, K.R.; Gliwa, A.S.; et al. Transmission, infectivity, and neutralization of a spike l452r sars-cov-2 variant. Cell 2021, 184, 3426–3437. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Davis, B.D.; Chen, S.S.; Sincuir Martinez, J.M.; Plummer, J.T.; Vail, E. Emergence of a novel sars-cov-2 variant in southern california. JAMA 2021, 325, 1324–1326. [Google Scholar] [CrossRef]
- Rosa-Aquino, C.D.a.P. What we know about the dangerous covid b.1.617.2 (delta) variant. N. Y. Intell. 2021. [Google Scholar]
- Mahase, E. Delta variant: What is happening with transmission, hospital admissions, and restrictions? BMJ 2021, 373, n1513. [Google Scholar] [CrossRef] [PubMed]
- Wall, E.C.; Wu, M.; Harvey, R.; Kelly, G.; Warchal, S.; Sawyer, C.; Daniels, R.; Hobson, P.; Hatipoglu, E.; Ngai, Y.; et al. Neutralising antibody activity against sars-cov-2 vocs b.1.617.2 and b.1.351 by bnt162b2 vaccination. Lancet 2021, 397, 2331–2333. [Google Scholar] [CrossRef]
- Yeung, J. There Are at Least 200 Known Cases of the Delta Plus Coronavirus Variant Worldwide. Here’s What We Know. CNN: 2021. Available online: https://www.cnn.com/2021/06/25/health/delta-plus-variant-explainer-intl-hnk-scn/index.html (accessed on 29 June 2021).
- Sars-Cov-2 Variants of Concern and Variants under Investigation in England. 2021. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/997418/Variants_of_Concern_VOC_Technical_Briefing_17.pdf (accessed on 19 May 2021).
- Lacobucci, G. Covid-19: Single vaccine dose is 33% effective against variant from india, data show. BMJ 2021, 373, n1346. [Google Scholar] [CrossRef]
- England, P.H. Effectiveness of Covid-19 Vaccines on Hospitalisation Disease with the Delta Variant. 2021. Available online: https://media.tghn.org/articles/Effectiveness_of_COVID-19_vaccines_against_hospital_admission_with_the_Delta_B._G6gnnqJ.pdf (accessed on 20 May 2021).
- Takuya Tada, H.Z.; Dcosta, B.M.; Samanovic, M.I.; Mulligan, M.J.; Landau, N.R. The spike proteins of sars-cov-2 b.1.617 and b.1.618 variants identified in india provide partial resistance to vaccine-elicited and therapeutic monoclonal antibodies. bioRxiv 2021. [Google Scholar] [CrossRef]
- McCallum, M.; Bassi, J.; Marco, A.; Chen, A.; Walls, A.C.; Iulio, J.D.; Tortorici, M.A.; Navarro, M.J.; Silacci-Fregni, C.; Saliba, C.; et al. Sars-cov-2 immune evasion by variant b.1.427/b.1.429. bioRxiv 2021. [Google Scholar] [CrossRef]
- Annavajhala, M.K.; Mohri, H.; Zucker, J.E.; Sheng, Z.; Wang, P.; Gomez-Simmonds, A.; Ho, D.D.; Uhlemann, A.C. A novel sars-cov-2 variant of concern, b.1.526, identified in new york. medRxiv 2021. [Google Scholar] [CrossRef]
- Bugembe, D.L.; Kayiwa, J.; Phan, M.V.T.; Tushabe, P.; Balinandi, S.; Dhaala, B.; Lexow, J.; Mwebesa, H.; Aceng, J.; Kyobe, H.; et al. Main routes of entry and genomic diversity of sars-cov-2, uganda. Emerg Infect. Dis. 2020, 26, 2411–2415. [Google Scholar] [CrossRef]
- Lopez Bernal, J.; Andrews, N.; Gower, C.; Gallagher, E.; Simmons, R.; Thelwall, S.; Stowe, J.; Tessier, E.; Groves, N.; Dabrera, G.; et al. Effectiveness of covid-19 vaccines against the b.1.617.2 variant. BioRxiv 2021. [Google Scholar] [CrossRef]
- Mahase, E. Covid-19: Novavax vaccine efficacy is 86% against uk variant and 60% against south african variant. BMJ 2021, 372, n296. [Google Scholar] [CrossRef] [PubMed]
- Emary, K.R.W.; Golubchik, T.; Aley, P.K.; Ariani, C.V.; Angus, B.; Bibi, S.; Blane, B.; Bonsall, D.; Cicconi, P.; Charlton, S.; et al. Efficacy of chadox1 ncov-19 (azd1222) vaccine against sars-cov-2 variant of concern 202012/01 (b.1.1.7): An exploratory analysis of a randomised controlled trial. Lancet 2021, 397, 1351–1362. [Google Scholar] [CrossRef]
- Jansen. Emergency Use Authorization (eua) for an Unapproved Product Review Memorandum. FDA, Ed. 2021. Available online: https://www.fda.gov/media/146338/download (accessed on 20 May 2021).
- Sadoff, J.; Gray, G.; Vandebosch, A.; Cardenas, V.; Shukarev, G.; Grinsztejn, B.; Goepfert, P.A.; Truyers, C.; Fennema, H.; Spiessens, B.; et al. Safety and efficacy of single-dose ad26.Cov2.S vaccine against covid-19. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef] [PubMed]
- Yadav, P.D.; Sapkal, G.N.; Abraham, P.; Ella, R.; Deshpande, G.; Patil, D.Y.; Nyayanit, D.A.; Gupta, N.; Sahay, R.R.; Shete, A.M.; et al. Neutralization of variant under investigation b.1.617 with sera of bbv152 vaccinees. Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef] [PubMed]

| Type | Manufacturer | Name | Phase | RoA | Trial Registration |
|---|---|---|---|---|---|
| PV | Novavax | NVX-CoV2373 | Phase 3 | IM | NCT04611802 |
| PV | Anhui Zhifei Longcom Biopharmaceutical | SARS-CoV-2 vaccine | Phase 3 | IM | NCT04466085 |
| PV | Center for Genetic Engineering and Biotechnology (CIGB) | CIGB-66 | Phase 3 | IM | RPCEC00000359 |
| PV | Federal Budgetary Research Institution State Research Center of Virology and Biotechnology “Vector” | EpiVacCorona | EUA (Russia) | IM | NCT04780035 |
| PV | Instituto Finlay de Vacunas | FINLAY-FR-2 | Phase 3 | IM | RPCEC00000354 |
| PV | Sanofi Pasteur + GSK | VAT00002 | Phase 3 | IM | PACTR20201152310190 |
| VLP | VBI Vaccines Inc. | VBI-2902a | Phase 1/2 | IM | NCT04773665 |
| VLP | The Scientific and Technological Research Council of Turkey | SARS-CoV-2 VLP Vaccine | Phase 1 | SC | NCT04818281 |
| VLP | Radboud University | ABNCoV2 | Phase 1 | IM | NCT04839146 |
| Type | Manufacturer | Name | Phase | RoA | Trial Registration |
|---|---|---|---|---|---|
| RNA | Pfizer-BioNTech + Fosun Pharma | BNT162b2 (Comirnaty) | Approved (US, EU, Canada, Israel) | IM | NCT04760132 |
| RNA | Moderna | mRNA -1273 | Approved in Switzerland. EUA (US, EU, UK, Canada, Israel) | IM | NCT04760132 |
| RNA | CureVac AG | CVnCoV Vaccine | Phase 3 | IM | NCT04674189 |
| RNA | Walvax Biotechnology | ARCoV | Phase 3 | IM | NCT04847102 |
| DNA | Zydus Cadila | nCov vaccine | Phase 3 | ID | CTRI/2020/07/026352 |
| DNA | Inovio Pharmaceuticals | INO-4800 | Phase 2/3 | ID | NCT04642638 |
| DNA | AnGes + Takara Bio + Osaka Univ | AG0301 | Phase 2/3 | IM | NCT04655625 |
| Type | Manufacturer | Name | Phase | RoA | Trial Registration |
|---|---|---|---|---|---|
| VVnra | AstraZeneca + University of Oxford | ChAdOx1-S (Covishield) | Approved (UK, India, Argentina, México) | IM | NCT04760132 |
| VVnra | CanSino Biological | Recombinant coronavirus vaccine (Ad5 vector) | EUA (Mexico) | IM | NCT04526990 |
| VVnra | Gamaleya Research Institute | Gam-COVID-Vac | EUA (Russia, Argentina, Bolivia, UAE) | IM | NCT04530396 |
| VVnra | Janssen Pharmaceutical | Ad26.COV2. S | EUA (US, Canada) | IM | NCT04505722 |
| VVrb | Beijing Wantai Biological Pharmacy | DelNS1-2019-nCoV-RBD-OPT1 | Phase 2 | IN | ChiCTR2000039715 |
| VVrb | Israel Institute for Biological Research | rVSV-SARS-CoV-2-S Vaccine | Phase 1/2 | IM | NCT04608305 |
| Type | Manufacturer | Name | Phase | RoA | Trial Registration |
|---|---|---|---|---|---|
| IV | Sinovac | CoronaVac | Approved (China, Indonesia) | IM | NCT04756830 |
| IV | Sinopharm | SARS-CoV-2 vaccine | Phase 3 | IM | ChiCTR2000034780 |
| IV | Sinopharm | BBIBP-CorV | Approved (China, Bahrain, UAE) | IM | NCT04863638 |
| IV | Institute of Medical Biology + Chinese Academy of Medical Sciences | SARS-CoV-2 vaccine | Phase 3 | IM | NCT04659239 |
| IV | Research Institute for Biological Safety Problem (Kazakhstan) | QazCovid-in® | Phase 3 | IM | NCT04691908 |
| IV | Bharat Biotech | COVAXIN® | EUA (India) | IM | NCT04641481; CTRI/2020/11/028976 |
| IV | Beijing Minhai Biotechnology | Inactivated SARS-CoV-2 vaccine | Phase 3 | IM | NCT04852705 |
| IV | Valneva, National Institute for Health Research, United Kingdom | VLA2001 | Phase 3 | IM | NCT04864561 |
| LAV | Codagenix/Serum Institute of India | COVI-VAC | Phase 1 | IN | NCT04619628 |
| LAV | Meissa Vaccines | MV-014-212 | Phase 1 | IN | NCT04798001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raman, R.; Patel, K.J.; Ranjan, K. COVID-19: Unmasking Emerging SARS-CoV-2 Variants, Vaccines and Therapeutic Strategies. Biomolecules 2021, 11, 993. https://doi.org/10.3390/biom11070993
Raman R, Patel KJ, Ranjan K. COVID-19: Unmasking Emerging SARS-CoV-2 Variants, Vaccines and Therapeutic Strategies. Biomolecules. 2021; 11(7):993. https://doi.org/10.3390/biom11070993
Chicago/Turabian StyleRaman, Renuka, Krishna J. Patel, and Kishu Ranjan. 2021. "COVID-19: Unmasking Emerging SARS-CoV-2 Variants, Vaccines and Therapeutic Strategies" Biomolecules 11, no. 7: 993. https://doi.org/10.3390/biom11070993
APA StyleRaman, R., Patel, K. J., & Ranjan, K. (2021). COVID-19: Unmasking Emerging SARS-CoV-2 Variants, Vaccines and Therapeutic Strategies. Biomolecules, 11(7), 993. https://doi.org/10.3390/biom11070993





