The First WHO International Standard for Harmonizing the Biological Activity of Bevacizumab
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials and Processing
2.2. Collaborative Study Participants
2.3. Collaborative Study Design
2.4. Assays Employed in the Study
2.5. Stability Studies
2.6. Statistical Analysis
potency of preparation (IU) × EC50 (ng)
Assumed mass content (ng)
3. Results
3.1. Evaluation of Bevacizumab Materials and Lyophilizing Formulations
3.2. Study Data Submitted by the Participants
3.3. Study Assay Validity
3.4. Potency Estimates Relative to the Candidate Standard 18/210
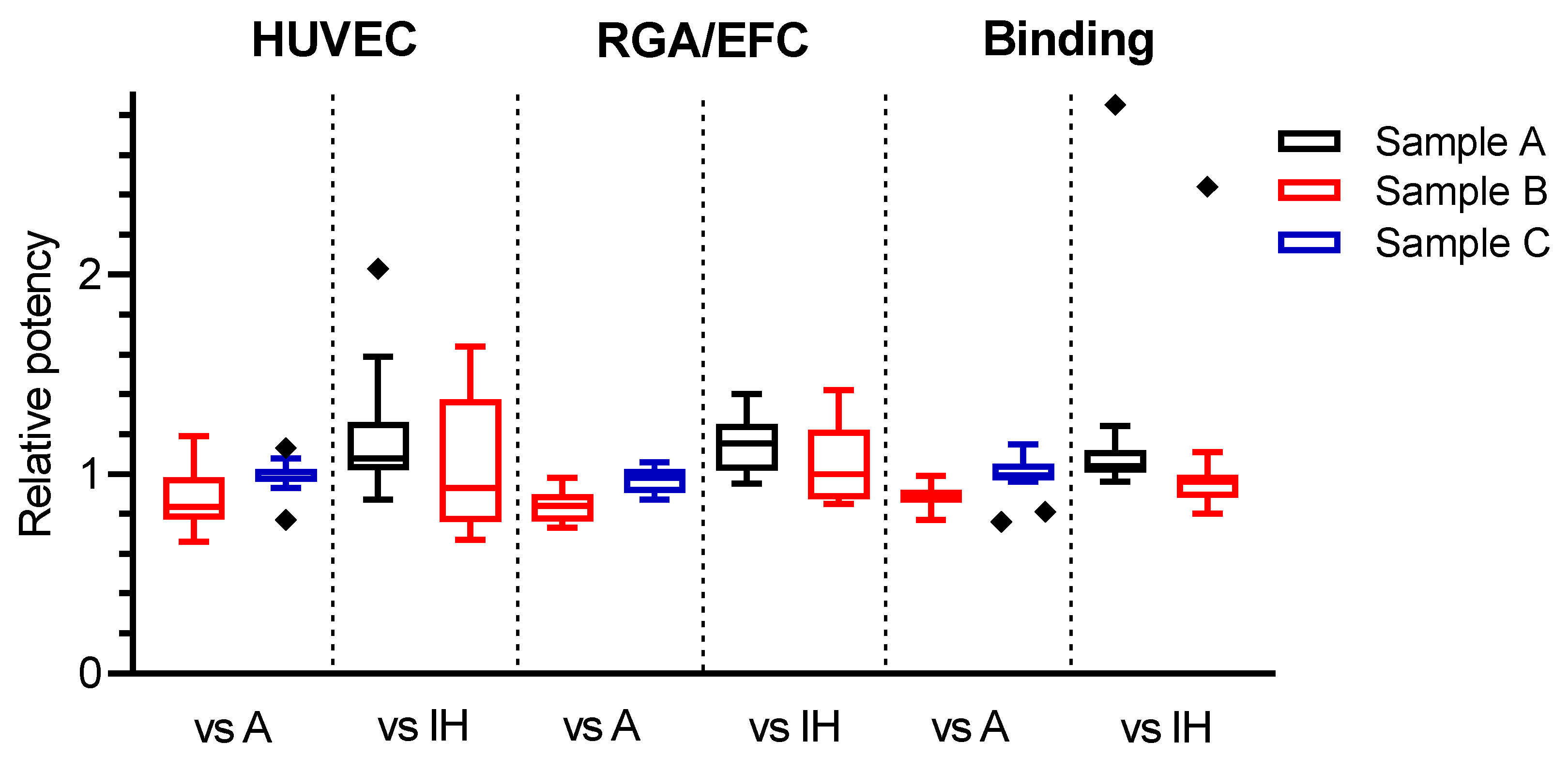
3.5. Improvement of Inter-Laboratory Variability by Use of the Candidate Standard 18/210
3.6. Estimates of EC50 Derived from Neutralisation Assays
3.7. Stability of the Candidate Preparations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Apte, R.S.; Chen, D.S.; Ferrara, N. VEGF in Signaling and Disease: Beyond Discovery and Development. Cell 2019, 176, 1248–1264. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferrara, N.; Adamis, A.P. Ten years of anti-vascular endothelial growth factor therapy. Nat. Rev. Drug Discov. 2016, 15, 385–403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Welti, J.; Loges, S.; Dimmeler, S.; Carmeliet, P. Recent molecular discoveries in angiogenesis and antiangiogenic therapies in cancer. J. Clin. Investig. 2013, 123, 3190–3200. [Google Scholar] [CrossRef] [Green Version]
- Hurwitz, H.; Fehrenbacher, L.; Novotny, W.; Cartwright, T.; Hainsworth, J.; Heim, W.; Berlin, J.; Baron, A.; Griffing, S.; Holmgren, E.; et al. Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N. Engl. J. Med. 2004, 350, 2335–2342. [Google Scholar] [CrossRef] [Green Version]
- Ferrara, N.; Hillan, K.J.; Gerber, H.P.; Novotny, W. Discovery and development of bevacizumab, an anti-VEGF antibody for treating cancer. Nat. Rev. Drug Discov. 2004, 3, 391–400. [Google Scholar] [CrossRef]
- US Food and Drug Administration—Approved Drugs: Avastin (Bevacizumab) Label, Letter and Review. Available online: https://www.accessdata.fda.gov/scripts/cder/daf/index.cfm?event=overview.process&ApplNo=125085 (accessed on 16 March 2021).
- Genentech Inc. Avastin Prescribing Information. 2018. Available online: https://www.accessdata.fda.gov/drugsatfda_docs/label/2018/125085s323lbl.pdf (accessed on 16 March 2021).
- European Medicines Agency EPAR—Avastin (Bevacizumab) 2006. Available online: https://www.ema.europa.eu/en/medicines/human/EPAR/avastin (accessed on 16 March 2021).
- Roche Registration GmbH. Summary of Product Characteristics: Avastin. 2018. Available online: https://www.ema.europa.eu/documents/product-information/avastin-epar-product-information_en.pdf (accessed on 16 March 2021).
- Garcia, J.; Hurwitz, H.I.; Sandler, A.B.; Miles, D.; Coleman, R.L.; Deurloo, R.; Chinot, O.L. Bevacizumab (Avastin®) in Cancer Treatment: A Review of 15 Years of Clinical Experience and Future Outlook. Cancer Treat. Rev. 2020, 86, 102017. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization Model List of Essential Medicines, 21st List. 2019. Available online: https://www.who.int/medicines/publications/essentialmedicines/en/ (accessed on 16 March 2021).
- Ziemssen, F.; Grisanti, S.; Bartz-Schmidt, K.U.; Spitzer, M.S. Off-label Use of Bevacizumab for the Treatment of Age-Related Macular Degeneration: What Is the Evidence? Drugs Aging 2009, 26, 295–320. [Google Scholar] [CrossRef]
- Ferrara, N. Vascular endothelial growth factor and age-related macular degeneration: From basic science to therapy. Nat. Med. 2010, 16, 1107–1111. [Google Scholar] [CrossRef] [PubMed]
- Virgili, G.; Parravano, M.; Evans, J.R.; Gordon, I.; Lucenteforte, E. Anti-vascular endothelial growth factor for diabetic macular oedema: A network meta-analysis. Cochrane Database Syst. Rev. 2018, 10, CD007419. [Google Scholar] [CrossRef] [Green Version]
- Ackermann, M.; Verleden, S.E.; Kuehnel, M.; Haverich, A.; Welte, T.; Laenger, F.; Vanstapel, A.; Werlein, C.; Stark, H.; Tzankov, A.; et al. Pulmonary Vascular Endothelialitis, Thrombosis, and Angiogenesis in COVID-19. N. Engl. J. Med. 2020, 383, 120–128. [Google Scholar] [CrossRef]
- El Bairi, K.; Trapani, D.; Petrillo, A.; Le Page, C.; Zbakh, H.; Daniele, B.; Belbaraka, R.; Curigliano, G.; Afqir, S. Repurposing anticancer drugs for the management of COVID-19. Eur. J. Cancer 2020, 141, 40–61. [Google Scholar] [CrossRef]
- Urguhart, L. Top Companies and Drugs by Sales in 2019. Nat. Rev. Drug Discov. 2020, 19, 228. [Google Scholar] [CrossRef]
- Thatcher, N.; Goldschmidt, J.H.; Thomas, M.; Schenker, M.; Pan, Z.; Paz-Ares, R.L.; Breder, V.; Ostoros, G.; Hanes, V. Efficacy and Safety of the Biosimilar ABP 215 Compared with Bevacizumab in Patients with Advanced Nonsquamous Non-small Cell Lung Cancer (MAPLE): A Randomized, Double-blind, Phase III Study. Clin. Cancer Res. 2019, 25, 2088–2095. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Thomas, M.; Thatcher, N.; Goldschmidt, J.; Ohe, Y.; McBride, H.J.; Hanes, V. Totality of Evidence in the Development of ABP 215, an Approved Bevacizumab Biosimilar. Immunotherapy 2019, 11, 1337–1351. [Google Scholar] [CrossRef] [PubMed]
- US Food and Drug Administration Approves First Biosimilar for the Treatment of Cancer—Mvasi, a Biosimilar to the Cancer Drug Avastin, Is Approved for Certain Colorectal, Lung, Brain, Kidney and Cervical Cancers. 2017. Available online: https://www.fda.gov/news-events/press-announcements/fda-approves-first-biosimilar-treatment-cancer (accessed on 16 March 2021).
- European Medicines Agency: Mvasi (Bevacizumab) 2018. Available online: https://www.ema.europa.eu/en/documents/overview/mvasi-epar-medicine-overview_en.pdf (accessed on 16 March 2021).
- Reinmuth, N.; Bryl, M.; Bondarenko, I.; Syrigos, K.; Vladimirov, V.; Zereu, M.; Bair, A.H.; Hilton, F.; Liau, K.; Kasahara, K. PF-06439535 (a Bevacizumab Biosimilar) Compared with Reference Bevacizumab (Avastin®), Both Plus Paclitaxel and Carboplatin, as First-Line Treatment for Advanced Non-Squamous Non-Small-Cell Lung Cancer: A Randomized, Double-Blind Study. BioDrugs 2019, 33, 555–570. [Google Scholar] [CrossRef] [Green Version]
- European Medicines Agency: Zirabev (Bevacizumab) 2019. Available online: https://www.ema.europa.eu/en/medicines/human/EPAR/zirabev (accessed on 16 March 2021).
- US Food and Drug Administration—Approved Biosimilar Products—Zirabev (Bevacizumab). Available online: https://www.fda.gov/drugs/biosimilars/biosimilar-product-information (accessed on 16 March 2021).
- Reck, M.; Luft, A.; Bondarenko, I.; Shevnia, S.; Trukhin, D.; Kovalenko, N.V.; Vacharadze, K.; Andrea, F.; Hontsa, A.; Choi, J.; et al. A phase III, randomized, double-blind, multicenter study to compare the efficacy, safety, pharmacokinetics, and immunogenicity between SB8 (proposed bevacizumab biosimilar) and reference bevacizumab in patients with metastatic or recurrent nonsquamous non-small cell lung cancer. Lung Cancer 2020, 146, 12–18. [Google Scholar] [PubMed]
- European Medicines Agency: Aybiotio (Bevacizumab) 2020. Available online: https://www.ema.europa.eu/en/medicines/human/EPAR/aybintio (accessed on 16 March 2021).
- European Medicines Agency: Equidacent (Bevacizumab) 2020. Available online: https://www.ema.europa.eu/en/medicines/human/EPAR/equidacent (accessed on 16 March 2021).
- European Medicines Agency: Onbevzi (Bevacizumab) 2020. Available online: https://www.ema.europa.eu/en/medicines/human/EPAR/onbevzi (accessed on 16 March 2021).
- Romera, A.; Peredpaya, S.; Shparyk, Y.; Bondarenko, I.; Mendonça, B.G.; Abdalla, K.C.; Roca, E.; Franke, F.; Melo, C.F.; Ramesh, A.; et al. Bevacizumab biosimilar BEVZ92 versus reference bevacizumab in combination with FOLFOX or FOLFIRI as first-line treatment for metastatic colorectal cancer: A multicentre, open-label, randomised controlled trial. Lancet Gastroenterol. Hepatol. 2018, 3, 845–855. [Google Scholar] [CrossRef]
- Eisenstein, M. Bring on the biosimilars. Nature 2019, 569, S2–S3. [Google Scholar] [CrossRef] [Green Version]
- Planinc, A.; Dejaegher, B.; Vander Heyden, Y.; Viaene, J.; Van Praet, S.; Rappez, F.; Van Antwerpen, P.; Delporte, C. Batch-to-batch N-glycosylation Study of Infliximab, Trastuzumab and Bevacizumab, and Stability Study of Bevacizumab. Eur. J. Hosp. Pharm. 2017, 24, 286–292. [Google Scholar] [CrossRef] [PubMed]
- Duivelshof, B.L.; Jiskoot, W.; Beck, A.; Veuthey, J.L.; Guillarme, D.; D’Atri, V. Glycosylation of biosimilars: Recent advances in analytical characterization and clinical implications. Anal. Chim. Acta 2019, 1089, 1–18. [Google Scholar] [CrossRef] [Green Version]
- Delobel, A. Glycosylation of Therapeutic Proteins: A Critical Quality Attribute. Methods Mol. Biol. 2021, 2271, 1–21. [Google Scholar] [PubMed]
- Montacir, O.; Montacir, H.; Eravci, M.; Springer, A.; Hinderlich, S.; Saadati, A.; Parr, M.K. Comparability study of Rituximab originator and follow-on biopharmaceutical. J. Pharm. Biomed. Anal. 2017, 140, 239–251. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Song, J.; Park, S.; Ham, S.; Paek, K.; Kang, M.; Chae, Y.; Seo, H.; Kim, H.C.; Flores, M. Drifts in ADCC-related quality attributes of Herceptin®: Impact on development of a trastuzumab biosimilar. MAbs 2017, 9, 704–714. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- World Health Organization Expert Committee on Biological Standardisation. Sixty-seventy Report. Proposed first WHO international standards for VEGF antagonists. WHO Tech. Rep. Series 2016, 1004, 52–53. [Google Scholar]
- Wadhwa, M.; Bird, C.; Dilger, P.; Rigsby, P.; Jia, H.; Gross, M.E.; participants of the study. Establishment of the first WHO International Standard for etanercept, a TNF receptor II Fc fusion protein: Report of an international collaborative study. J. Immunol. Methods 2017, 447, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Metcalfe, C.; Dougall, T.; Bird, C.; Rigsby, P.; Behr-Gross, M.E.; Wadhwa, M.; Study, P.O.T. The first World Health Organization International Standard for infliximab products: A step towards maintaining harmonized biological activity. Mabs 2019, 11, 13–25. [Google Scholar] [CrossRef] [Green Version]
- Wadhwa, M.; Bird, C.; Atkinson, E.; Cludts, I.; Rigsby, P. The First WHO International Standard for Adalimumab: Dual Role in Bioactivity and Therapeutic Drug Monitoring. Front. Immunol. 2021, 12, 636420. [Google Scholar] [CrossRef]
- World Health Organization Expert Committee on Biological Standardisation. Fifty-fifth Report. Recommendations for the preparation, characterization and establishment of international and other biological reference standards. WHO Tech. Rep. Series 2006, 932, 73–130. [Google Scholar]
- Peraza, M.A.; Rule, K.E.; Shiue, M.H.I.; Finch, G.L.; Thibault, S.; Brown, P.R.; Clarke, D.W.; Leach, M.W. Nonclinical assessments of the potential biosimilar PF-06439535 and bevacizumab. Regul. Toxicol. Pharmacol. 2018, 95, 236–243. [Google Scholar] [CrossRef]
- Wang, Y.; Fei, D.; Vanderlaan, M.; Song, A. Biological activity of bevacizumab, a humanized anti-VEGF antibody in vitro. Angiogenesis 2004, 7, 335–345. [Google Scholar] [CrossRef]
- Ebbers, H.C.; van Meer, P.J.; Moors, E.H.; Mantel-Teeuwisse, A.K.; Leufkens, H.G.; Schellekens, H. Measures of biosimilarity in monoclonal antibodies in oncology: The case of bevacizumab. Drug Discov. Today 2013, 18, 872–879. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jia, H.; Thelwell, C.; Dilger, P.; Bird, C.; Daniels, S.; Wadhwa, M. Endothelial cell functions impaired by interferon in vitro: Insights into the molecular mechanism of thrombotic microangiopathy associated with interferon therapy. Thromb. Res. 2018, 163, 105–116. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suehiro, J.; Kanki, Y.; Makihara, C.; Schadler, K.; Miura, M.; Manabe, Y.; Aburatani, H.; Kodama, T.; Minami, T. Genome-wide approaches reveal functional vascular endothelial growth factor (VEGF)-inducible nuclear factor of activated T cells (NFAT) c1 binding to angiogenesis-related genes in the endothelium. J. Biol. Chem. 2014, 289, 29044–29059. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Xu, G.L.; Gao, K.; Wilkinson, J.; Zhang, F.; Yu, L.; Liu, C.Y.; Yu, C.F.; Wang, W.B.; Li, M.; et al. Development of a robust reporter-based assay for the bioactivity determination of anti-VEGF therapeutic antibodies. J. Pharm. Biomed. Anal. 2016, 125, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Naqvi, T.; Lim, A.; Rouhani, R.; Singh, R.; Eglen, R.M. Beta galactosidase enzyme fragment complementation as a high-throughput screening protease technology. J. Biomol. Screen. 2004, 9, 398–408. [Google Scholar] [CrossRef]
- Lamerdin, J.; Daino-Laizure, H.; Saharia, A.; Charter, N.W. Accelerating Biologic and Biosimilar Drug Development: Ready-to-Use, Cell-Based Assays for Potency and Lot-Release Testing. BioProcess Inter. 2016, 14, 36–44. [Google Scholar]
- Kirkwood, T.B. Predicting the stability of biological standards and products. Biometrics 1977, 33, 736–742. [Google Scholar] [CrossRef]
- Ritz, C.; Baty, F.; Streibig, J.C.; Gerhard, D. Dose-Response Analysis Using R. PLoS ONE 2015, 10, e0146021. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2018; Available online: https://www.R-project.org/ (accessed on 16 March 2021).
- Jia, H.; Bagherzadeh, A.; Bicknell, R.; Duchen, M.R.; Liu, D.; Zachary, I. Vascular endothelial growth factor (VEGF)-D and VEGF-A differentially regulate KDR-mediated signalling and biological function in vascular endothelial cells. J. Biol. Chem. 2004, 279, 36148–36157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Damiano, V.; Caputo, R.; Garofalo, S.; Bianco, R.; Rosa, R.; Merola, G.; Gelardi, T.; Racioppi, L.; Fontanini, G.; De Placido, S.; et al. TLR9 agonist acts by different mechanisms synergizing with bevacizumab in sensitive and cetuximab-resistant colon cancer xenografts. Proc. Natl. Acad. Sci. USA 2007, 104, 12468–12473. [Google Scholar] [CrossRef] [Green Version]
- Prior, S.; Metcalfe, C.; Hufton, S.E.; Wadhwa, M.; Schneider, C.K.; Burns, C. Maintaining ‘standards’ for biosimilar monoclonal antibodies. Nat. Biotechnol. 2021, 39, 276–280. [Google Scholar] [CrossRef] [PubMed]
| INN | Brand Name | Manufacturer | Product Type | US Approval | EU Approval |
|---|---|---|---|---|---|
| Bevacizumab | Avastin® | Genentech/Roche | Full antibody | 2004 | 2005 |
| Ranibizumab | Lucentis® | Genentech/Novartis | Fab fragment | 2006 | 2007 |
| Aflibercept | Eylea® | Regeneron | VEGFR1/2-Fc fusion protein | 2011 | 2012 |
| Brolucizumab | Beovu® | Novartis | Single-chain antibody fragment | 2019 | 2020 |
| Participants | Laboratory Address | Country |
|---|---|---|
| Akiko Ishii-Watabe and Takuo Suzuki | National Institute of Health Sciences, Division of Biological Chemistry and Biologicals, 3-25-26, Tonomachi, Kawasaki-ku, Kawasaki, Kanagawa, 210-9501 | Japan |
| Chunping Deng | Bio-Thera Solutions Ltd., Bldg A6-5fl, 11 Kai-Yuan Blvd, Science City, Guangzhou, 510530 | China |
| Feng Zhang and Lan Wang | National Institutes for Food and Drug Control (NIFDC), Division of Monoclonal Antibodies, No. 31 Huatuo Road, Daxing District, Beijing, 102629 | China |
| Francesca Luciani and Agnese D’Angiò | ISS, Biologicals and Biotechnologicals Unit, National Centre for the Control and Evaluation of Medicines (CNCF), Istituto Superiore di Sanità, Viale Regina Elena 299, Rome, 161 | Italy |
| Guoping Wu and Christian Erickson | R&D Systems, Bio-Techne, Bioassay, 614 McKinley Place NE, Minneapolis, MN55413 | USA |
| He Chen and Jiemin Chen | Genor BioPharma, Building No. 3, 1690 Zhangheng Rd, Zhangjiang, Pudong District, Shanghai, 201203 | China |
| Hongyan Ye and Jiulin Wang | Qilu Pharmaceutical, No. 243 Gong Ye Bei Road, Licheng District, Jinan, 250000 | China |
| Jane Lamerdin and Ai Shih | Eurofins DiscoverX, 42501 Albrae Street, Fremont, CA94538 | USA |
| Jianying Fu and Chen Ma | Henlius Biopharmaceuticals, 1289 Yishan Road, Shanghai, 200030 | China |
| Jill Crouse-Zeineddini and Jolene Teraoka | Amgen Inc., One Amgen Center Dr., B30E Dropzone DZ-1B, Thousand Oaks, CA 91320 | USA |
| Jixiang Jiao and Karen Zhang | Shanghai Roche Pharmaceuticals Ltd., 1100 Long Dong Avenue, Pudong District, Shanghai, 201203 | China |
| Junxian Guo and Qingcheng Guo | Shanghai Biomabs Pharmaceuticals Co.,Ltd, NO. 301 Libing Road, Pilot Free Trade Zone, Shanghai, 201203 | China |
| Karin Blume and Kerstin Mårtensson | Svar Life Science, Lundavägen 151, Malmö, 21224 | Sweden |
| Keith Mortimer and Anita Carscadden | Therapeutic Goods Administration, TGA Laboratories, Biochemistry Section, 136 Narrabundah Lane, Symonston, Canberra ACT, 2609 | Australia |
| Kyumin Han and Joon Hyuk Lim | Samsung Bioepis, 107 Cheomdan-Daero, Yeonsu-gu, Incheon, 406-840 | Republic of Korea |
| Manuel Navarro and Daniela Lorenzo | mAbxience SAU, Carlos Villate 5148, Munro, Buenos Aires, 1605 | Argentina |
| Pankaj Kalita and Sanjay Bandyopadhyay | Zydus, Cadila Healthcare Ltd., Zydus Research Centre, Sarkhej Bavla N.H. No. 8A., Moraiya, Ahmedabad, 382213 | India |
| Parvathy Harikumar and Haiyan Jia | Cytokines and Growth Factors Section, Biotherapeutics Group, NIBSC, Blanche Lane, South Mimms, Potters Bar, Herts, EN6 3QG | UK |
| Shubrata Khedkar and Mitali Samaddar | United States Pharmacopeia–India (P) Ltd., Plot D6 and D8, IKP Knowledge Park, Genome valley, Shameerpet, R.R. Dist. Telangana, Hyderabad, 500078 | India |
| Nripendra Nath Mishra, Subhash Chand, Ratnesh K. Sharma and J. P. Prasad | National Institute of Biologicals, A-32, Sector-62, Noida, 201309 | India |
| Tina Kneeland and David Cirelli | Pfizer, Analytical Research and Development, 1 Burtt Rd, Andover, Massachusetts 01810 | USA |
| Valérie Ridoux and Jean-Claude Ourlin | ANSM, 635 rue de la garenne, Vendargues, 34740 | France |
| Yangdong Sun | Innoventbio, 168 Dongping Street, Industrial Park, Suzhou, 215123 | China |
| Yingchun LI and Tongjie Xu | CTTQ Pharma, No. 1099 Fuying Road, Jiangning Dist, Nanjing, 211100 | China |
| Yujie Zhang | Teruisi Pharmaceutical Inc., 3rd Floor, Building 5, 1366 Hongfeng Road, South Lake Tai Scientific Innovation Center, Huzhou, 313000 | China |
| Ampoule Code | Study Code | Protein (Predicted µg) | Mean Fill Weight (ng) | CV Fill Weight (%) | Mean Residual Moisture (%) | CV Residual Moisture (%) | Mean Headspace Oxygen (%) | CV Headspace Oxygen (%) |
|---|---|---|---|---|---|---|---|---|
| 18/210 | ~53 | 1.0083 (241) | 0.2193 | 0.56031 (12) | 13.59 | 0.15 (12) | 40.5 | |
| 18/214 | ~50 | 1.0100 (33) | 0.2187 | 0.08349 (12) | 11.21 | 0.34 (12) | 32.2 | |
| 18/216 | ~43 | 1.0093 (10) | 0.1450 | 0.11034 (6) | 8.49 | 0.35 (6) | 15.5 |
| Bioassay Type | Cell Line | Number of Participants | VEGF165 (U/mL) a | Assay Period (Hours) | Assay Readout | Readout Reagent b |
|---|---|---|---|---|---|---|
| Anti-proliferation | HUVEC | 13 | 10–50 | 48–99 | Absorbance (3) | CCK-8 |
| Fluorescence (8) | alamarBlue (6), CellTiter-Blue (1), resazurin dye (1) | |||||
| Luminescence (2) | CellTiter-Glo® | |||||
| Reporter gene | HEK293 | 12 | 3.75–75 | 3.5–18 | Luminescence | Bio-Glo™ luciferase (6), Bright-Glo™ luciferase (4), Steady-Glo® luciferase (1) |
| Enzyme-fragment complementation | HEK293 | 2 | 11–12 | 16–20 | Luminescence | PathHunter® bioassay detection kit |
| Assay Type | Number of Participants | IH Standard | Assay Description | Detection Reagent | Assay Readout | Readout Reagent |
|---|---|---|---|---|---|---|
| ELISA | Yes (or Avastin) | Bevacizumab binds to VEGF165 coated plate | Goat anti Human IgG-HRP | Absorbance | TMB substrate | |
| Competitive binding | 1 | Yes | Bevacizumab and VEGF165 complex is added to capture plate | Anti-biotinylated VEGF | Absorbance | SureBlue™ TMB substrate |
| Biolayer interferometry | 1 | Avastin | Bevacizumab binds to biotinylated VEGF165 captured onto streptavidin biosensor | Not relevant | Response binding rate (nm/s) | Not relevant |
| Lab Code | Potency Relative to Candidate (Sample A) | Potency Relative to In-House Reference | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample B | Sample C | Sample A | Sample B | |||||||||
| GM | GCV | N | GM | GCV | N | GM | GCV | N | GM | GCV | N | |
| 04 | 1.19 | 59.69 | 5 | 1.01 | 124.2 | 7 | 1.08 | 61.61 | 9 | 1.32 | 144.8 | 4 |
| 05 | 0.78 | 15.51 | 9 | 0.98 | 16.29 | 9 | 0.87 | 12.25 | 12 | 0.67 | 14.47 | 9 |
| 06 | 0.80 | 7.98 | 8 | 0.96 | 7.20 | 8 | 1.10 | 9.07 | 9 | 0.89 | 8.49 | 8 |
| 07a a | 0.79 | 5.66 | 8 | 1.01 | 5.05 | 8 | 1.02 | 7.51 | 7 | 0.79 | 7.14 | 9 |
| 07b a | 0.75 | 17.29 | 3 | 1.01 | 5.46 | 4 | 1.08 | n/a | 1 | 0.81 | 9.65 | 4 |
| 08 | 0.98 | 24.38 | 12 | 1.13 | 47.25 | 11 | 1.20 | 32.77 | 14 | 1.13 | 42.28 | 12 |
| 09 | 0.86 | 16.02 | 11 | 1.02 | 10.31 | 10 | - | - | - | - | - | - |
| 12 | 0.99 | 18.31 | 6 | 1.04 | 55.68 | 4 | - | - | - | - | - | - |
| 14 | 0.98 | 26.27 | 7 | 0.96 | 40.82 | 8 | 1.59 | 12.26 | 7 | 1.54 | 26.14 | 5 |
| 21 | 0.93 | 10.62 | 8 | 0.99 | 14.39 | 8 | 1.04 | 8.08 | 12 | 0.97 | 8.62 | 8 |
| 22 | 1.03 | 33.38 | 8 | 1.08 | 18.93 | 9 | - | - | - | - | - | - |
| 23 | 0.81 | 3.98 | 12 | 1.01 | 5.47 | 12 | 2.03 | 8.82 | 12 | 1.64 | 9.32 | 12 |
| 25a | 0.70 | 31.26 | 5 | 0.93 | 3.22 | 6 | 0.95 | 7.14 | 9 | 0.67 | 42.71 | 5 |
| 25b | 0.66 | n/a | 1 | 0.77 | 3.17 | 6 | 1.26 | 0.82 | 3 | n/a | n/a | n/a |
| Assay Type | Lab Code | Potency Relative to Candidate (Sample A) | Potency Relative to In-House Reference | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample B | Sample C | Sample A | Sample B | ||||||||||
| GM | GCV | N | GM | GCV | N | GM | GCV | N | GM | GCV | N | ||
| RGA | 01 | 0.89 | 11.86 | 11 | 1.04 | 10.93 | 8 | 1.40 | n/a | 1 | 1.42 | n/a | 1 |
| RGA | 02 | 0.77 | 20.85 | 9 | 0.91 | 19.84 | 9 | 1.14 | 9.78 | 9 | 0.88 | 17.53 | 9 |
| RGA | 04 | 0.86 | 10.00 | 9 | 1.02 | 10.30 | 9 | 1.04 | 8.89 | 12 | 0.88 | 6.70 | 9 |
| RGA | 05 | 0.80 | 7.14 | 9 | 1.01 | 7.98 | 9 | 1.01 | 13.48 | 12 | 0.86 | 12.46 | 9 |
| RGA | 08 | 0.95 | 12.66 | 6 | 0.90 | n/a | 2 | 1.38 | 14.85 | 5 | 1.26 | 22.26 | 5 |
| RGA | 10 | 0.83 | 9.84 | 9 | 1.06 | 16.08 | 9 | 1.06 | 13.30 | 9 | 0.85 | 16.52 | 7 |
| RGA | 13 | 0.74 | 29.12 | 8 | 0.94 | 52.00 | 7 | 1.22 | 28.29 | 6 | 1.01 | 27.61 | 6 |
| RGA | 15 | 0.98 | 30.94 | 3 | 0.90 | n/a | 2 | 1.26 | 35.18 | 3 | 1.25 | 32.38 | 3 |
| RGA | 16 | 0.73 | n/a | 1 | - | - | - | 0.95 | n/a | 1 | 1.04 | n/a | 1 |
| RGA | 17 | 0.89 | 8.92 | 9 | 1.03 | 7.41 | 8 | 1.01 | 11.31 | 12 | 0.87 | 6.88 | 9 |
| RGA | 18 | 0.82 | 7.39 | 6 | 0.98 | 9.54 | 6 | n/a | n/a | n/a | n/a | n/a | n/a |
| RGA | 19 | 0.85 | 5.74 | 9 | 1.01 | 10.15 | 8 | 1.17 | 6.83 | 9 | 0.99 | 5.76 | 9 |
| EFC | 21 | 0.92 | 8.81 | 7 | 0.87 | 12.58 | 7 | 1.17 | 17.21 | 10 | 1.14 | 26.03 | 7 |
| EFC | 24 | 0.73 | 6.58 | 9 | 0.94 | 18.94 | 9 | - | - | - | - | - | - |
| Lab Code | Potency Relative to Candidate (Sample A) | Potency Relative to In-House Reference | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample B | Sample C | Sample A | Sample B | |||||||||
| GM | GCV | N | GM | GCV | N | GM | GCV | N | GM | GCV | N | |
| 02 | 0.88 | 6.07 | 9 | 1.01 | 9.39 | 9 | 1.01 | 6.38 | 9 | 0.90 | 6.73 | 9 |
| 03 a | 0.90 | 2.73 | 4 | 1.00 | 2.98 | 4 | 1.24 | 4.92 | 4 | 1.11 | 6.96 | 4 |
| 04 | 0.99 | 11.02 | 9 | 1.15 | 16.42 | 9 | 1.01 | 15.73 | 12 | 0.97 | 17.03 | 9 |
| 05 | 0.88 | 7.02 | 9 | 0.99 | 12.83 | 9 | 0.96 | 10.35 | 9 | 0.84 | 12.36 | 9 |
| 10 | 0.86 | 6.05 | 9 | 0.99 | 6.02 | 9 | 1.00 | 7.41 | 9 | 0.86 | 9.24 | 9 |
| 11 | 0.90 | n/a | 1 | 0.81 | n/a | 1 | - | - | - | - | - | - |
| 13 | 0.90 | 9.33 | 9 | 1.06 | 14.60 | 9 | 1.02 | 13.19 | 9 | 0.93 | 21.33 | 9 |
| 15 | 0.77 | 27.87 | 3 | 1.06 | 19.44 | 3 | 1.04 | 21.95 | 3 | 0.80 | 23.46 | 3 |
| 16 | 0.92 | n/a | 1 | 0.76 | n/a | 1 | 1.01 | 5.17 | 3 | 0.98 | n/a | 2 |
| 18 | 0.92 | 11.72 | 9 | 1.02 | 2.62 | 9 | 1.10 | 4.39 | 9 | 1.01 | 11.27 | 9 |
| 19 | 0.85 | 10.24 | 8 | 0.96 | 19.15 | 8 | 2.85 | 20.75 | 8 | 2.44 | 29.40 | 8 |
| 20 | 0.86 | 6.96 | 9 | 0.97 | 4.66 | 9 | 1.10 | 8.08 | 11 | 0.97 | 14.93 | 8 |
| 21 | 0.92 | 4.81 | 9 | 1.05 | 4.26 | 9 | 1.04 | 4.91 | 9 | 0.96 | 6.11 | 9 |
| 23 | 0.83 | 2.10 | 9 | 0.99 | 4.28 | 9 | 1.14 | 7.09 | 9 | 0.95 | 7.26 | 9 |
| Method | Sample | Potencies Relative to Sample A | Potencies Relative to IH Reference | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| GM | LCL | UCL | GCV | N | GM | LCL | UCL | GCV | N | ||
| Neutralisation (All) a | A | - | - | - | - | - | 1.16 | 1.05 | 1.28 | 21.87 | 18 |
| B | 0.85 | 0.82 | 0.89 | 11.35 | 25 | 0.98 | 0.87 | 1.10 | 27.98 | 19 | |
| C | 1.00 | 0.97 | 1.02 | 5.90 | 23 | 1.14 | 1.03 | 1.26 | 21.94 | 18 | |
| D | 0.86 | 0.81 | 0.91 | 12.59 | 17 | 0.95 | 0.80 | 1.13 | 30.29 | 12 | |
| Neutralisation (HUVEC) a | A | - | - | - | - | - | 1.18 | 0.93 | 1.50 | 32.91 | 8 |
| B | 0.86 | 0.79 | 0.93 | 13.61 | 12 | 0.96 | 0.75 | 1.24 | 39.14 | 9 | |
| C | 1.01 | 0.98 | 1.04 | 5.39 | 12 | 1.16 | 0.94 | 1.42 | 30.90 | 9 | |
| D | 0.84 | 0.79 | 0.90 | 9.86 | 10 | 0.95 | 0.72 | 1.25 | 38.60 | 8 | |
| Neutralisation (RGA/EFC) a | A | - | - | - | - | - | 1.14 | 1.06 | 1.23 | 10.73 | 10 |
| B | 0.85 | 0.80 | 0.89 | 9.38 | 13 | 0.99 | 0.88 | 1.10 | 16.89 | 10 | |
| C | 0.98 | 0.94 | 1.02 | 6.37 | 11 | 1.13 | 1.04 | 1.22 | 10.96 | 9 | |
| D | 0.88 | 0.77 | 1.01 | 16.22 | 7 | 0.96 | 0.83 | 1.10 | 9.37 | 4 | |
| Binding b | A | - | - | - | - | - | 1.06 | 1.01 | 1.11 | 7.50 | 11 |
| B | 0.88 | 0.85 | 0.91 | 6.26 | 12 | 0.93 | 0.88 | 0.99 | 9.61 | 11 | |
| C | 1.02 | 0.99 | 1.05 | 5.08 | 12 | 1.09 | 1.03 | 1.14 | 7.75 | 11 | |
| D | 0.81 | 0.78 | 0.84 | 5.16 | 8 | 0.86 | 0.79 | 0.95 | 11.80 | 8 | |
| Sample | GM | LCL | UCL | GCV | N |
|---|---|---|---|---|---|
| A | 132.2 | 63.9 | 273.6 | 99.9 | 6 |
| B | 152.8 | 78.2 | 298.5 | 89.3 | 6 |
| C | 141.5 | 61.5 | 325.5 | 121.2 | 6 |
| IH | 194.6 | 117.6 | 322.1 | 50.1 | 5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jia, H.; Harikumar, P.; Atkinson, E.; Rigsby, P.; Wadhwa, M. The First WHO International Standard for Harmonizing the Biological Activity of Bevacizumab. Biomolecules 2021, 11, 1610. https://doi.org/10.3390/biom11111610
Jia H, Harikumar P, Atkinson E, Rigsby P, Wadhwa M. The First WHO International Standard for Harmonizing the Biological Activity of Bevacizumab. Biomolecules. 2021; 11(11):1610. https://doi.org/10.3390/biom11111610
Chicago/Turabian StyleJia, Haiyan, Parvathy Harikumar, Eleanor Atkinson, Peter Rigsby, and Meenu Wadhwa. 2021. "The First WHO International Standard for Harmonizing the Biological Activity of Bevacizumab" Biomolecules 11, no. 11: 1610. https://doi.org/10.3390/biom11111610
APA StyleJia, H., Harikumar, P., Atkinson, E., Rigsby, P., & Wadhwa, M. (2021). The First WHO International Standard for Harmonizing the Biological Activity of Bevacizumab. Biomolecules, 11(11), 1610. https://doi.org/10.3390/biom11111610





