Self-Attention-Based Models for the Extraction of Molecular Interactions from Biological Texts
Abstract
:1. Why Text Mining?
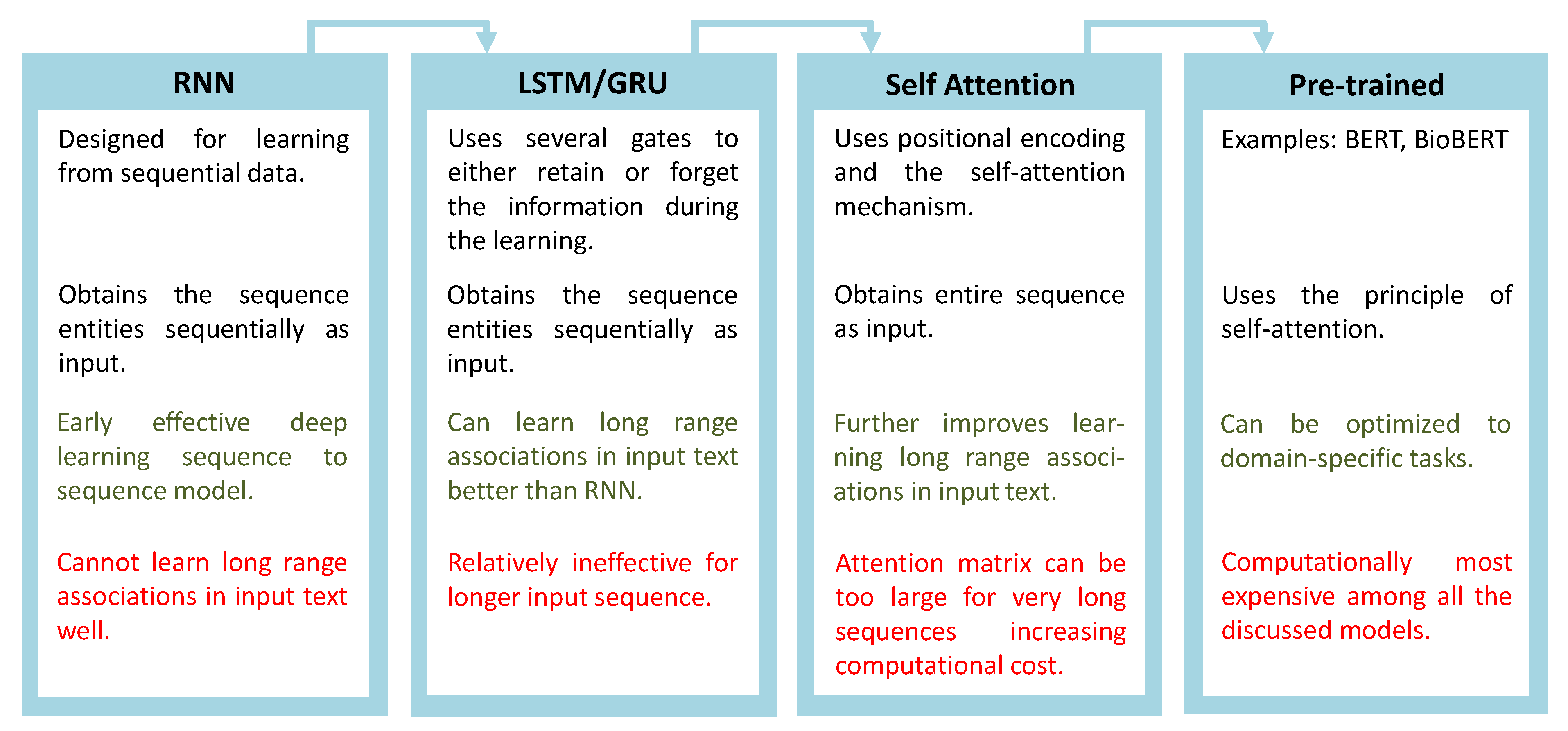
2. Evolution of Deep Learning Models for NLP
2.1. Self-Attention and Its Advantages
- The encoder compresses all input information into a vector of fixed length, which is passed to the decoder, causing significant information loss [18];
- Such models are unable to model the alignment between input and output vectors. “Intuitively, in sequence-to-sequence tasks, each output token is expected to be more influenced by some specific parts of the input sequence. However, decoder lacks any mechanism to selectively focus on relevant input tokens while generating each output token” [18].
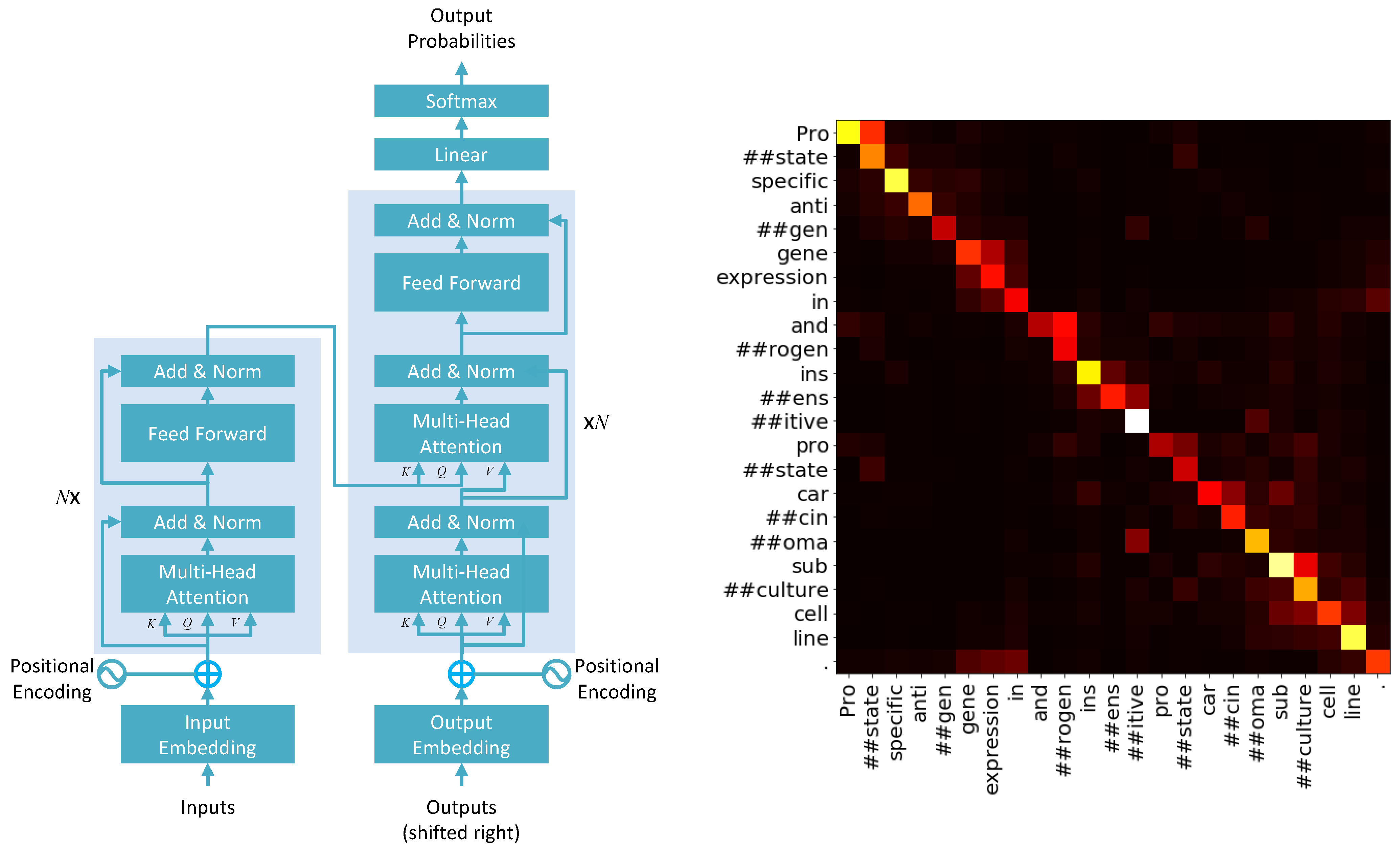
- Positional encoding: Given an input sentence in a transformer model, the model first creates a vectorized representation of the sentence S, such that each word in the sentence is represented by a vector of a user-defined dimension. The vectorized version of the sentence S is then integrated with positional encoding. Recall that, unlike sequence-to-sequence models such as RNNs and LSTMs, which would feed the sequence elements (words in a sentence) as the input sequentially, self-attention-based models feed the entire sequence (sentence) as the input at a time. This requires a mechanism that can account for the sequential structure of the input sequence/sentence. This is achieved through positional encoding. The formal expression for positional encoding is given by a pair of equations:In Equations (1) and (2), the expression pos is used to denote the position of a word in a sentence and d denotes the dimension of user-defined dimensions for the word embeddings, that is each word is essentially perceived by the model as a d-dimensional vector. The index i runs over the dimensions of these word embeddings and take values in the range . Note that Equations (1) and (2) propose two different functions over the vector, depending on whether one is calculating an odd index or even index of the word-embedding vector. The dependence of the positional encoding functions on , given that these functions are periodic functions by design, ensures that several frequencies are captured over several dimensions of the word-embedding vectors. “The wavelengths form a geometric progression from to 10,000 × ” [5]. Intuitively, proximal words in a sentence are likely to have a similar P value in a lower frequency, but can still be differentiated in the higher frequencies. For far apart words in a sentence, the case is just the opposite. Equations (1) and (2) also ensure the robustness and uniformity of the positional encoding function P, over all sentences, independent of their length [5];
- The self-attention mechanism: Once the positional encoding is integrated with the word embedding of an input sentence S, the resultant vector W is fed into the mechanism of self-attention. There is a popular analogy used by many data scientists to explain the concepts of Query (Q), Key (K), and Value (V), which are central to the idea of self-attention. When we search for a particular video on YouTube, we submit a query to the search engine, which then maps our query to a set of keys (video title and descriptions), associated with existing videos in the database. The algorithm then presents to us the best possible values as the search result we see. For a self-attention mechanism [5],A dot product between Q and K in the form can measure the attention between pairwise words in a sentence, to generate attention weights. The attention weights are used to generate a weighted mean over the word vectors in V, to obtain relevant information from the input as per the given task. As these vectors are learned through the training procedure of the model, the framework can help the model retrieve relevant information from an input, for a given task. The equation governing the process is given by [5]:In practice, however, a multiheaded attention mechanism is used. The idea of multiple heads is again often compared to the use of different filters in CNNs, where each filter learns latent features from the input. Similarly, in multiheaded attention, different heads learn different latent features from the input. The information from all heads is later integrated by a concatenation operation. To account for multiple heads, Equation (3) is violated of course, and the dimension of the positionally encoded word vector W is distributed over the multiple heads. Equation (4) is also adjusted accordingly by replacing the denominator of by , where is the dimension of the keys considering multiple heads. Several other concepts such as layer normalization and masking are also used in transformer models, which we will not discuss in detail here. A representation of the transformer architecture and attention map over a sentence is provided in Figure 3 [5].
2.2. Pretrained Models
3. Applications of Self-Attention-Based Models in BLM
3.1. Commonly Used Datasets
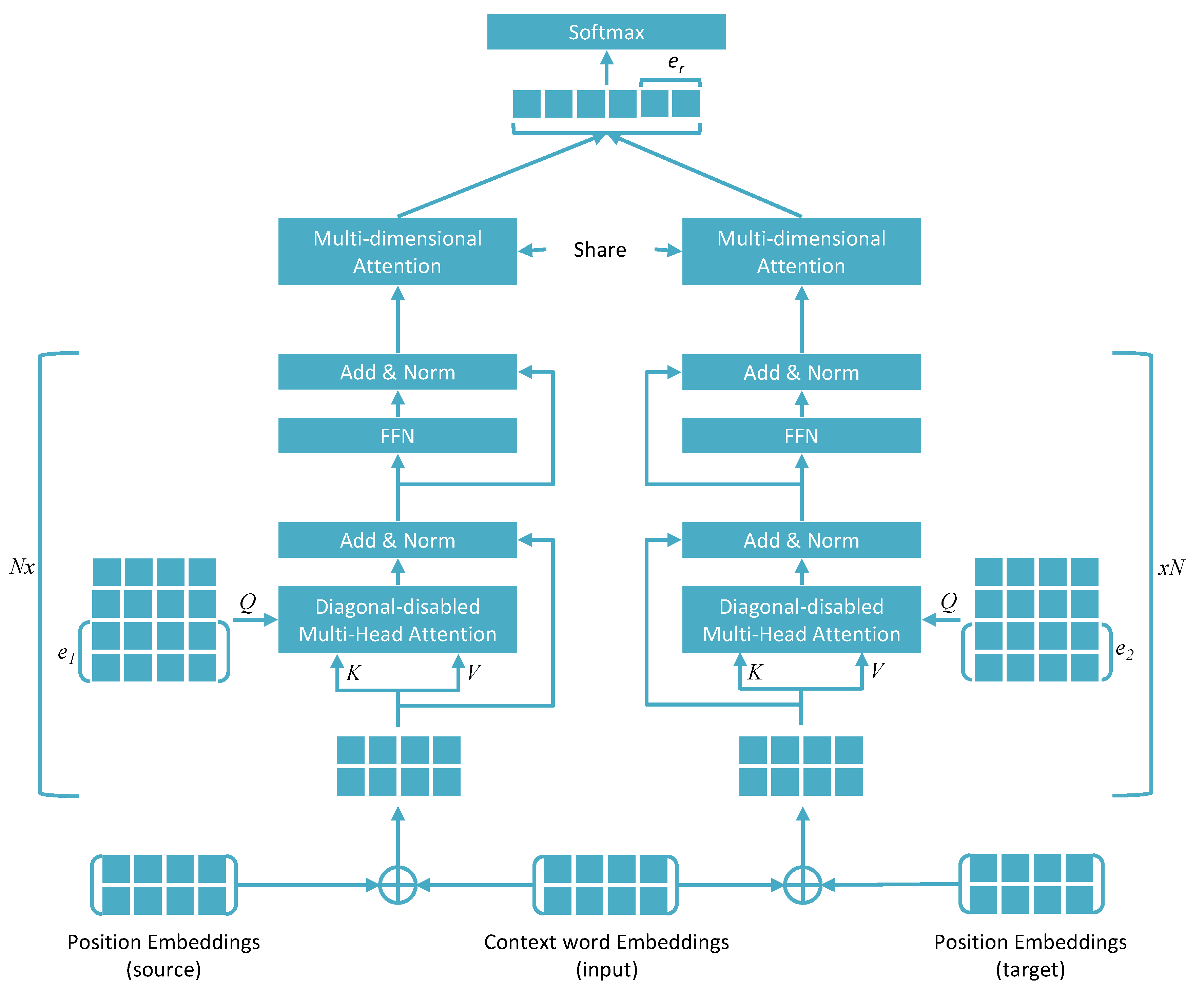
3.2. Architectural Comparison of Some Recent Attention-Based Models
- Identifying documents describing mutations affecting PPI;
- Extracting relevant PPI through RE.
3.3. Performance Comparison among the Discussed Models
4. Discussion
- Reducing the number of inappropriate instances; the sentence distance between a protein pair was assumed to be less than three;
- Selecting the words among a protein pair and three expansion words on both sides as the context word sequence with respect to the protein pair;
- Removing protein names from the input string;
- Replacing numeric entries by predefined strings;
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Kotu, V.; Deshpande, B. Chapter 9–Text Mining. In Data Science, 2nd ed.; Kotu, V., Deshpande, B., Eds.; Morgan Kaufmann: Burlington, MA, USA, 2019; pp. 281–305. ISBN 9780128147610. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Nastou, A.L.; Lyon, K.C.; Kirsch, D.; Pyysalo, R.; Doncheva, S.; Legeay, N.T.; Fang, M.; Bork, T.; Jensen, P.L.; et al. The STRING database in 2021: Customizable protein—Protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef] [PubMed]
- Serhan, C.; Gupta, S.; Perretti, M.; Godson, C.; Brennan, E.; Li, Y.; Soehnlein, O.; Shimizu, T.; Werz, O.; Chiurchiù, V.; et al. The Atlas of Inflammation Resolution (AIR). Mol. Asp. Med. 2020, 74, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Su, C.; Lu, Z.; Wang, F. Recent advances in biomedical literature mining. Briefings Bioinform. 2021, 22, 1–19. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.M.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, L.; Polosukhin, I. Attention is All you Need. In Proceedings of the 31st Conference on Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 6000–6010. [Google Scholar]
- Zhang, Y.; Lin, H.; Yang, Z.; Wang, J.; Sun, Y.; Xu, B.; Zhao, Z. Neural network-based approaches for biomedical relation classification: A review. J. Biomed. Inform. 2019, 99, 103294. [Google Scholar] [CrossRef]
- Papanikolaou, N.; Pavlopoulos, G.; Theodosiou, T.; Iliopoulos, I. Protein-protein interaction predictions using text mining methods. Methods 2015, 74, 47–53. [Google Scholar] [CrossRef]
- Bahdanau, D.; Cho, K.; Bengio, Y. Neural Machine Translation by Jointly Learning to Align and Translate. arXiv 2015, arXiv:1409.0473. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale. arXiv 2021, arXiv:2010.11929. [Google Scholar]
- Tran, D.T.; Iosifidis, A.; Kanniainen, J.; Gabbouj, M. Temporal Attention-Augmented Bilinear Network for Financial Time-Series Data Analysis. IEEE Trans. Neural Netw. Learn. Syst. 2019, 30, 1407–1418. [Google Scholar] [CrossRef] [Green Version]
- Choi, J.; Lee, B.; Zhang, B. Multi-focus Attention Network for Efficient Deep Reinforcement Learning. arXiv 2017. Available online: https://arxiv.org/ftp/arxiv/papers/1712/1712.04603.pdf (accessed on 2 October 2021).
- Luong, T.; Pham, H.; Manning, C.D. Effective Approaches to Attention-based Neural Machine Translation. In Proceedings of the 2015 Conference on Empirical Methods in Natural Language Processing, Lisbon, Portugal, 7–11 September.
- Hermann, K.M.; Kočiský, T.; Grefenstette, E.; Espeholt, L.; Kay, W.; Suleyman, M.; Blunsom, P. Teaching machines to read and comprehend. In Proceedings of the 28th International Conference on Neural Information Processing Systems (NIPS’15), Montreal, QC, Canada, 7–12 December 2015; MIT Press: Cambridge, MA, USA, 2015; Volume 1, pp. 1693–1701. [Google Scholar]
- Yang, Z.; Yang, D.; Dyer, C.; He, X.; Smola, A.; Hovy, E. Hierarchical Attention Networks for Document Classification. In Proceedings of the 2016 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, San Diego, CA, USA, 12–17 June 2016; pp. 1480–1489. [Google Scholar]
- Devlin, J.; Ming-Wei, C.; Kenton, L.; Kristina, T. BERT: Pretraining of Deep Bidirectional Transformers for Language Understanding. NAACL-HLT 2019, 1, 4171–4186. [Google Scholar]
- Ambartsoumian, A.; Popowich, F. Self-Attention: A Better Building Block for Sentiment Analysis Neural Network Classifiers. In Proceedings of the 9th Workshop on Computational Approaches to Subjectivity, Sentiment and Social Media Analysis, Brussels, Belgium, 31 October 2018; Association for Computational Linguistics: Stroudsburg, PA, USA; pp. 130–139. [Google Scholar]
- Cho, K.; van Merrienboer, B.; Gülçehre, C.; Bahdanau, D.; Bougares, F.; Schwenk, H.; Bengio, Y. Learning Phrase Representations using RNN Encoder-Decoder for Statistical Machine Translation. In Proceedings of the 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP), Doha, Qatar, 25–29 October 2014; pp. 1724–1734. [Google Scholar]
- Chaudhari, S.; Polatkan, G.; Ramanath, R.; Mithal, V. An Attentive Survey of Attention Models. arXiv 2019, arXiv:1904.02874. [Google Scholar]
- Radford, A.; Narasimhan, K. Improving Language Understanding by Generative Pre-Training. 2018. Available online: https://openai.com/blog/language-unsupervised/ (accessed on 2 October 2021).
- Dehghani, M.; Azarbonyad, H.; Kamps, J.; de Rijke, M. Learning to Transform, Combine, and Reason in Open-Domain Question Answering. In Proceedings of the Twelfth ACM International Conference on Web Search and Data Mining (WSDM’19), Melbourne, Australia, 11–15 February 2019; Association for Computing Machinery: New York, NY, USA; pp. 681–689. [Google Scholar] [CrossRef]
- Zhu, Y.; Kiros, R.; Zemel, R.; Salakhutdinov, R.; Urtasun, R.; Torralba, A.; Fidler, S. Aligning Books and Movies: Towards Story-Like Visual Explanations by Watching Movies and Reading Books. In Proceedings of the IEEE International Conference on Computer Vision (ICCV), Las Condes, Chile, 11–18 December 2015; pp. 19–27. [Google Scholar] [CrossRef] [Green Version]
- Wu, Y.; Schuster, M.; Chen, Z.; Le, Q.V.; Norouzi, M.; Macherey, W.; Krikun, M.; Cao, Y.; Gao, Q.; Macherey, K.; et al. Google’s Neural Machine Translation System: Bridging the Gap between Human and Machine Translation. arXiv 2016, arXiv:1609.08144. [Google Scholar]
- Wang, A.; Singh, A.; Michael, J.; Hill, F.; Levy, O.; Bowman, S.R. GLUE: A Multi-Task Benchmark and Analysis Platform for Natural Language Understanding. In Proceedings of the 2018 EMNLP Workshop BlackboxNLP: Analyzing and Interpreting Neural Networks for NLP, Brussels, Belgium, 1 November 2018; pp. 353–355. [Google Scholar]
- Rajpurkar, P.; Zhang, J.; Lopyrev, K.; Liang, P. SQuAD: 100,000+ Questions for Machine Comprehension of Text. In Proceedings of the 2016 Conference on Empirical Methods in Natural Language Processing, Austin, TX, USA, 1–5 November 2016; 2016; pp. 1264–2383. [Google Scholar]
- Zellers, R.; Bisk, Y.; Schwartz, R.; Choi, Y. SWAG: A Large-Scale Adversarial Dataset for Grounded Commonsense Inference. In Proceedings of the 2018 Conference on Empirical Methods in Natural Language Processing, Brussels, Belgium, 31 October–4 November 2018; pp. 93–104. [Google Scholar]
- Lan, Z.; Chen, M.; Goodman, S.; Gimpel, K.; Sharma, P.; Soricut, R. ALBERT: A Lite BERT for Self-supervised Learning of Language Representations. In Proceedings of the International Conference on Learning Representations 2020, Addis Ababa, Ethiopia, 26–30 April 2020. [Google Scholar]
- Nguyen, D.Q.; Vu, T.; Nguyen, A. BERTweet: A pretrained language model for English Tweets. In Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing: System Demonstrations, Virtual Event, 16–20 November 2020; Association for Computational Linguistics: Stroudsburg, PA, USA, 2020; pp. 9–14. [Google Scholar]
- Martin, L.; Muller, B.; Ortiz, S.J.P.; Dupont, Y.; Romary, L.; De la Clergerie, E.; Seddah, D.; Sagot, B. CamemBERT: A Tasty French Language Model. In Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics, Virtual Event, 5–10 July 2020; pp. 7203–7219. [Google Scholar]
- Liu, Y.; Ott, M.; Goyal, N.; Du, J.; Joshi, M.; Chen, D.; Levy, O.; Lewis, M.; Zettlemoyer, L.; Stoyanov, V. RoBERTa: A Robustly Optimized BERT Pretraining Approach. arXiv 2019, arXiv:1907.11692. [Google Scholar]
- Lee, J.; Yoon, W.; Kim, S.; Kim, D.; Kim, S.; So, C.H.; Kang, J. BioBERT: A pretrained biomedical language representation model for biomedical text mining. Bioinformatics 2020, 36, 1234–1240. [Google Scholar] [CrossRef]
- Beltagy, I.; Lo, K.; Cohan, A. SciBERT: A Pretrained Language Model for Scientific Text. In Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing and the 9th International Joint Conference on Natural Language Processing (EMNLP-IJCNLP), China, Hong Kong, 3–7 December 2019; pp. 3615–3620. [Google Scholar]
- Oughtred, R.; Rust, J.M.; Chang, C.S.; Breitkreutz, B.; Stark, C.; Willems, A.; Boucher, L.; Leung, G.; Kolas, N.; Zhang, F.; et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci. Publ. Protein Soc. 2020, 30, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Hermjakob, H.; Montecchi-Palazzi, L.; Lewington, C.; Mudali, S.K.; Kerrien, S.; Orchard, S.; Vingron, M.; Roechert, B.; Roepstorff, P.; Valencia, A.; et al. IntAct: An open source molecular interaction database. Nucleic Acids Res. 2004, 32, D452–D455. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wishart, D.; Knox, C.; Guo, A.; Shrivastava, S.; Hassanali, M.; Stothard, P.; Chang, Z.; Woolsey, J. DrugBank: A comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 2006, 34, D668–D672. [Google Scholar] [CrossRef] [PubMed]
- Taboureau, O.; Nielsen, S.K.; Audouze, K.; Weinhold, N.; Edsgärd, D.; Roque, F.S.; Kouskoumvekaki, I.; Bora, A.; Curpan, R.; Jensen, T.; et al. ChemProt: A disease chemical biology database. Nucleic Acids Res. 2011, 39, D367–D372. [Google Scholar] [CrossRef] [Green Version]
- Zanzoni, A.; Montecchi-Palazzi, L.; Quondam, M.; Ausiello, G.; Helmer-Citterich, M.; Cesareni, G. MINT: A Molecular INTeraction database. FEBS Lett. 2007, 513, 135–140. [Google Scholar] [CrossRef] [Green Version]
- Bader, G.D.; Donaldson, I.; Wolting, C.; Ouellette, B.; Pawson, T.; Hogue, C. BIND: The Biomolecular Interaction Network Database. Nucleic Acids Res. 2001, 31, 248–250. [Google Scholar] [CrossRef]
- Han, H.; Shim, H.; Shin, D.; Shim, J.; Ko, Y.; Shin, J.; Kim, H.; Cho, A.; Kim, E.; Lee, T.; et al. TRRUST: A reference database of human transcriptional regulatory interactions. Sci. Rep. 2015, 11432. [Google Scholar] [CrossRef] [PubMed]
- Elangovan, A.; Davis, M.J.; Verspoor, K. Assigning function to protein-protein interactions: A weakly supervised BioBERT based approach using PubMed abstracts. arXiv 2020, arXiv:2008.08727. [Google Scholar]
- Giles, O.; Karlsson, A.; Masiala, S.; White, S.; Cesareni, G.; Perfetto, L.; Mullen, J.; Hughes, M.; Harl, L.; Malone, J. Optimising biomedical relationship extraction with BioBERT. bioRxiv 2020. Available online: https://www.biorxiv.org/content/10.1101/2020.09.01.277277v1.full (accessed on 2 October 2021).
- Su, P.; Vijay-Shanker, K. Investigation of BERT Model on Biomedical Relation Extraction Based on Revised Fine-tuning Mechanism. In Proceedings of the IEEE International Conference on Bioinformatics and Biomedicine, Houston, TX, USA, 8–11 December 2020; pp. 2522–2529. [Google Scholar]
- Su, P.; Peng, Y.; Vijay-Shanker, K. Improving BERT Model Using Contrastive Learning for Biomedical Relation Extraction. arXiv 2021, arXiv:2104.13913. [Google Scholar]
- Wang, Y.; Zhang, S.; Zhang, Y.; Wang, J.; Lin, H. Extracting Protein-Protein Interactions Affected by Mutations via Auxiliary Task and Domain Pretrained Model. In Proceedings of the 2020 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Seoul, Korea, 16–19 December 2020; pp. 495–498. [Google Scholar]
- Zhou, H.; Liu, Z.; Ning, S.; Lang, C.; Lin, Y.; Du, L. Knowledge-aware attention network for protein-protein interaction extraction. J. Biomed. Inform. 2019, 96, 103234. [Google Scholar] [CrossRef] [Green Version]
- Fujita, K.A.; Ostaszewski, M.; Matsuoka, Y.; Ghosh, S.; Glaab, E.; Trefois, C.; Crespo, I.; Perumal, T.M.; Jurkowski, W.; Antony, P.M.; et al. Integrating Pathways of Parkinson’s Disease in a Molecular Interaction Map. Mol. Neurobiol. 2013, 49, 88–102. [Google Scholar] [CrossRef] [Green Version]
- Liu, Y.; Hildebr, M.; Joblin, M.; Ringsqu, I.M.; Raissouni, R.; Tresp, V. Neural Multi-Hop Reasoning with Logical Rules on Biomedical Knowledge Graphs. In Proceedings of the Semantic Web 18th International Conference, ESWC 2021; Virtual Event, 6–10 June 2021, Lecture Notes in Computer Science Book Series (LNCS); Springer: Berlin/Heidelberg, Germany, 2021; Volume 4825, pp. 375–391. [Google Scholar]

| Abbreviation | Full Name | Description |
|---|---|---|
| BLM | Biological Literature Mining | Mining information from biological literature/publications |
| NLP | Natural Language Processing | Ability of a computer program to understand human language |
| RE | Relationship Extraction | Extracting related entities and the relationship type from biological texts |
| NER | Named Entity Recognition | NLP-based approaches to identify context-specific entity names from text |
| CNN | Convolutional Neural Network | A type of neural network popularly used in computer vision |
| RNN | Recurrent Neural Network | One of the neural network models designed to handle sequential data |
| LSTM | Long Short-Term Memory | A successor of RNN useful for handling sequential data |
| GRUs | Gated Recurrent Units | A successor of RNN useful for handling sequential data |
| BERT | Bidirectional Encoder Representations from Transformer | A pretrained neural network popularly used for NLP tasks |
| KAN | Knowledge-aware Attention Network | A self-attention-based network for RE problems |
| PPI | Protein–Protein Interaction | Interactions among proteins, a popular problem in RE |
| DDI | Drug–Drug Interaction | Interactions among drugs, a popular problem in RE |
| ChemProt | Chemical–Protein Interaction | Interactions among chemicals and proteins, a popular problem in RE |
| Work | Datasets | Model | Tasks Performed | Performance |
|---|---|---|---|---|
| Elangovan et al. (2020) [39] | Processed version of the IntAct dataset with seven types of interactions | Ensemble of fine-tuned BioBERT models; no external knowledge used | Typed and untyped RE with relationship types such as phosphorylation, acetylation, etc. | Typed PPI: 0.540; untyped PPI: 0.717; metric: F1 score |
| Giles et al. (2020) [40] | Manually curated from the MEDLINE database | Fine-tuned BioBERT model; used STRING database knowledge during dataset curation | Classification problem with classes coincidental mention, positive, negative, incorrect entity recognition, and unclear | Curated data and BioBERT: 0.889; metric: F1 score |
| Su et al. (2020) [41] | Processed versions of the BioGRID, DrugBank, and IntAct datasets | Fine-tuned the BERT model integrated with LSTM and additive attention; no external knowledge used | Classification tasks on PPI (binary), DDI (multiclass), and ChemProt (multiclass) | PPI: 0.828; DDI: 0.807; ChemProt: 0.768; metric: F1 score |
| Su et al. (2021) [42] | Processed versions of the BioGRID, DrugBank, and IntAct datasets | Contrastive learning model; no external knowledge used in dataset curation or as a part of the model | Classification tasks on PPI (binary), DDI (multiclass), and ChemProt (multiclass) | PPI: 0.827; DDI: 0.829; ChemProt: 0.787; metric: F1 score |
| Wang et al. (2020) [43] | Processed versions of the BioCreative VI PPI dataset | A multitasking architecture based on BERT, BioBERT, BiLSTM, and text CNN; no external knowledge used | Document triage classification, NER (auxiliary tasks), and PPI RE (main task). | NER task: 0.936, PPI RE (exact match evaluation): 0.431; metric: F1 score |
| Zhou et al. (2019) [44] | Processed versions of the BioCreative VI PPI dataset | KAN; TransE used to integrate prior knowledge from the BioGRID and IntAct datasets on triplets to the model | PPI-RE classification task from BioCreative VI | PPI RE (exact match evaluation): 0.382 PPI RE: (HomoloGene evaluation): 0.404; metric: F1 score |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Srivastava, P.; Bej, S.; Yordanova, K.; Wolkenhauer, O. Self-Attention-Based Models for the Extraction of Molecular Interactions from Biological Texts. Biomolecules 2021, 11, 1591. https://doi.org/10.3390/biom11111591
Srivastava P, Bej S, Yordanova K, Wolkenhauer O. Self-Attention-Based Models for the Extraction of Molecular Interactions from Biological Texts. Biomolecules. 2021; 11(11):1591. https://doi.org/10.3390/biom11111591
Chicago/Turabian StyleSrivastava, Prashant, Saptarshi Bej, Kristina Yordanova, and Olaf Wolkenhauer. 2021. "Self-Attention-Based Models for the Extraction of Molecular Interactions from Biological Texts" Biomolecules 11, no. 11: 1591. https://doi.org/10.3390/biom11111591
APA StyleSrivastava, P., Bej, S., Yordanova, K., & Wolkenhauer, O. (2021). Self-Attention-Based Models for the Extraction of Molecular Interactions from Biological Texts. Biomolecules, 11(11), 1591. https://doi.org/10.3390/biom11111591







