Deoxyribonucleases and Their Applications in Biomedicine
Abstract
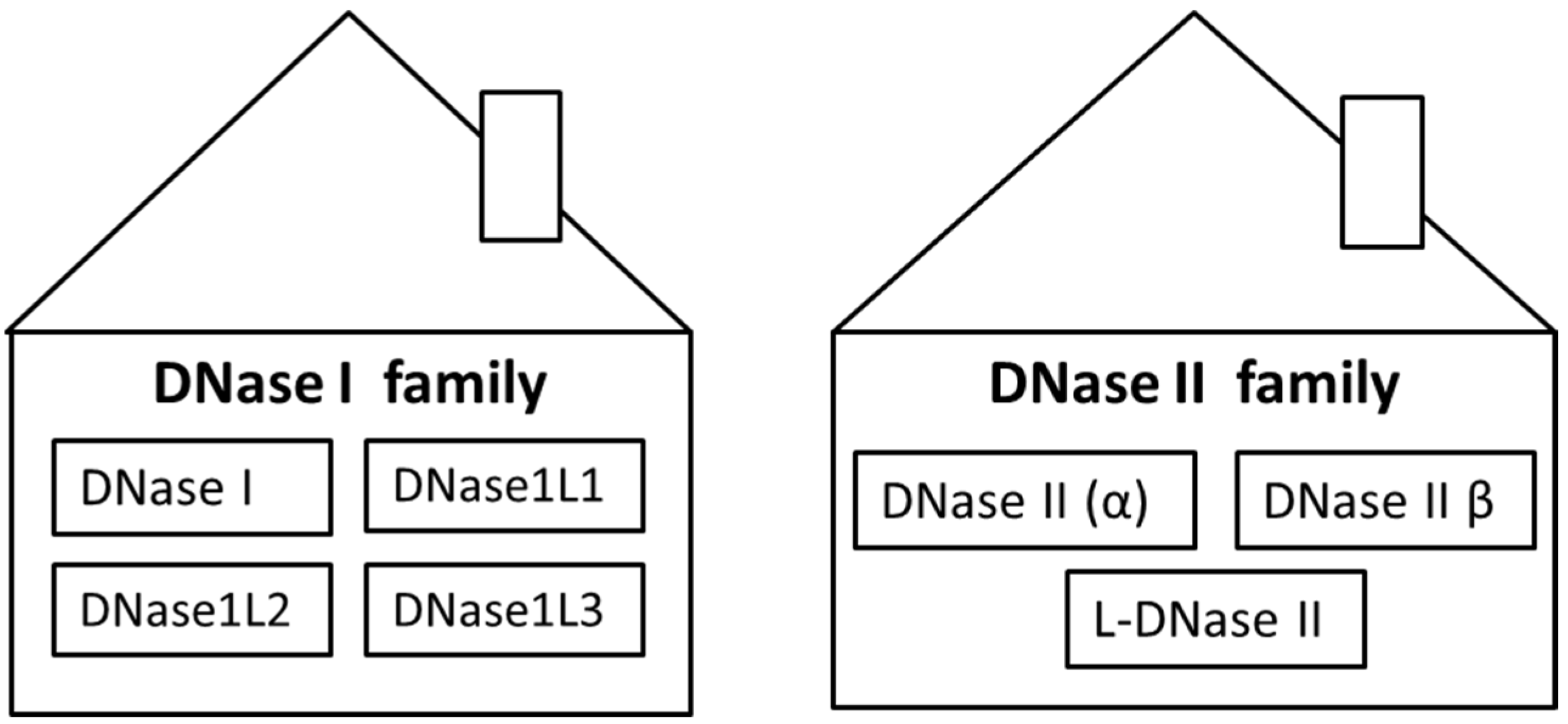
1. Introduction
2. DNase I
3. DNase1L1, DNase1L2 and DNase1L3
4. DNase II Family
5. DNase II α
6. DNase II β
7. L-DNase II
8. TREX1 and TREX2
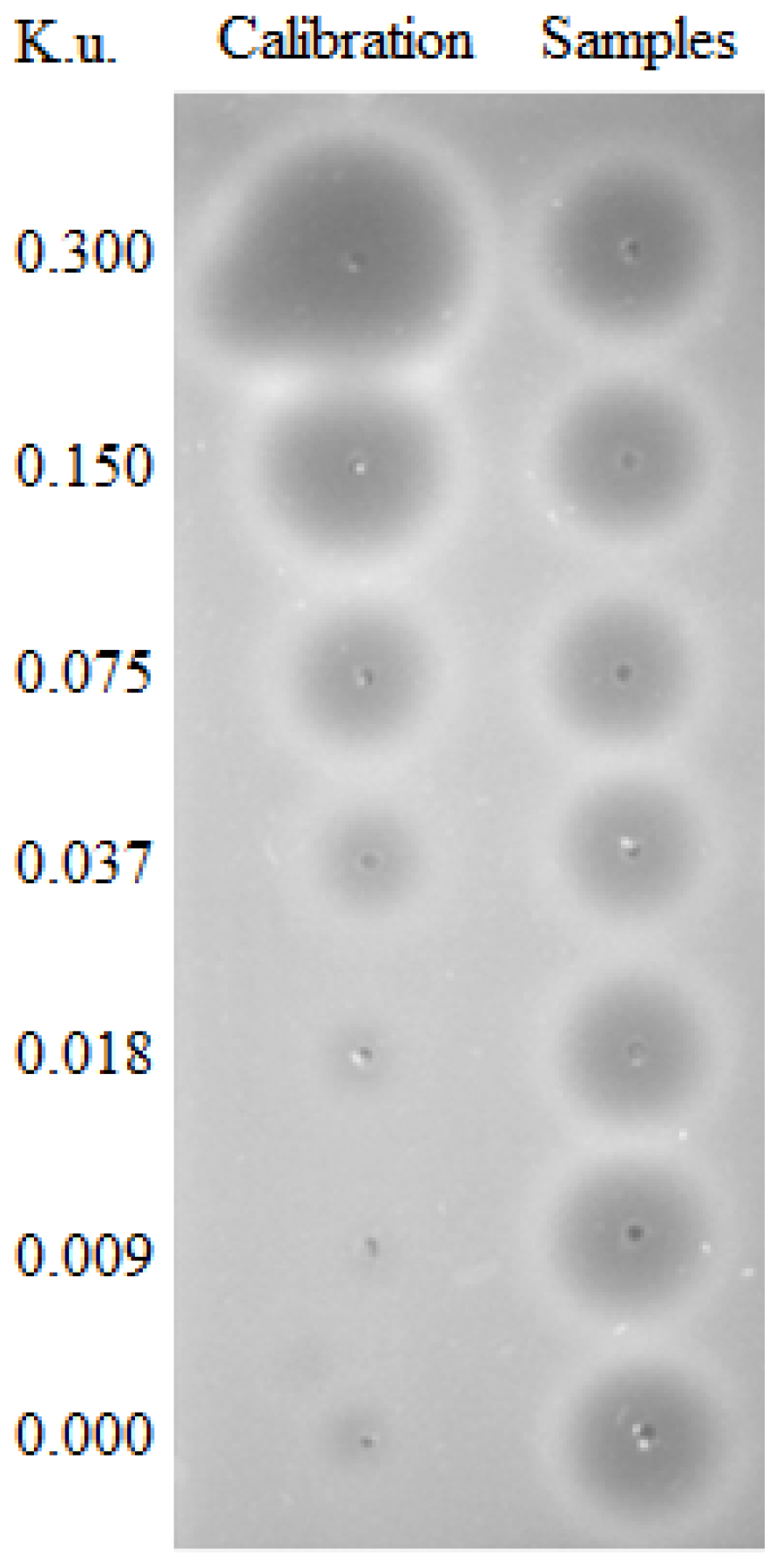
9. Methods of the Measurement of DNase Activity
10. DNase Activity as a Biomarker
11. Application of DNase and DNase Treatment
12. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Mandel, P.; Metais, P. Les acides nucléiques du plasma sanguin chez l’homme. C. R. Seances Soc. Biol. Fil. 1948, 142, 241–243. [Google Scholar] [PubMed]
- Magna, M.; Pisetsky, D.S. The Alarmin Properties of DNA and DNA-associated Nuclear Proteins. Clin. Ther. 2016, 38, 1029–1041. [Google Scholar] [CrossRef] [PubMed]
- Han, D.S.C.; Ni, M.; Chan, R.W.Y.; Chan, V.W.H.; Lui, K.O.; Chiu, R.W.K.; Lo, Y.M.D. The Biology of Cell-free DNA Fragmentation and the Roles of DNASE1, DNASE1L3, and DFFB. Am. J. Hum. Genet. 2020, 106, 202–214. [Google Scholar] [CrossRef] [PubMed]
- Shpak, M.; Kugelman, J.R.; Varela-Ramirez, A.; Aguilera, R.J. The phylogeny and evolution of deoxyribonuclease II: An enzyme essential for lysosomal DNA degradation. Mol. Phylogenet. Evol. 2008, 47, 841–854. [Google Scholar] [CrossRef][Green Version]
- Junowicz, E.; Spencer, J.H. Studies on bovine pancreatic deoxyribonuclease A. II. The effect of different bivalent metals on the specificity of degradation of DNA. Biochim. Biophys. Acta 1973, 312, 85–102. [Google Scholar] [CrossRef]
- Suck, D. DNA recognition by DNase I. J. Mol. Recognit. 1994, 7, 65–70. [Google Scholar] [CrossRef]
- Kishi, K.; Yasuda, T.; Takeshita, H. DNase I: Structure, function, and use in medicine and forensic science. Leg. Med. 2001, 3, 69–83. [Google Scholar] [CrossRef]
- Kolarevic, A.; Pavlovic, A.; Djordjevic, A.; Lazarevic, J.; Savic, S.; Kocic, G.; Anderluh, M.; Smelcerovic, A. Rutin as Deoxyribonuclease I Inhibitor. Chem. Biodivers. 2019, 16, e1900069. [Google Scholar] [CrossRef]
- Yasuda, T.; Awazu, S.; Sato, W.; Iida, R.; Tanaka, Y.; Kishi, K. Human genetically polymorphic deoxyribonuclease: Purification, characterization, and multiplicity of urine deoxyribonuclease I. J. Biochem. 1990, 108, 393–398. [Google Scholar] [CrossRef]
- Takeshita, H.; Mogi, K.; Yasuda, T.; Nakajima, T.; Nakashima, Y.; Mori, S.; Hoshino, T.; Kishi, K. Mammalian deoxyribonucleases I are classified into three types: Pancreas, parotid, and pancreas-parotid (mixed), based on differences in their tissue concentrations. Biochem. Biophys. Res. Commun. 2000, 269, 481–484. [Google Scholar] [CrossRef]
- Fujihara, J.; Yasuda, T.; Ueki, M.; Iida, R.; Takeshita, H. Comparative biochemical properties of vertebrate deoxyribonuclease I. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2012, 163, 263–273. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Nadano, D.; Iida, R.; Takeshita, H.; Lane, S.A.; Callen, D.F.; Kishi, K. Chromosomal assignment of the human deoxyribonuclease I gene, DNASE 1 (DNL1), to band 16p13.3 using the polymerase chain reaction. Cytogenet. Cell Genet. 1995, 70, 221–223. [Google Scholar] [CrossRef]
- Yasuda, T.; Kishi, K.; Yanagawa, Y.; Yoshida, A. Structure of the human deoxyribonuclease I (DNase I) gene: Identification of the nucleotide substitution that generates its classical genetic polymorphism. Ann. Hum. Genet. 1995, 59, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Iida, R.; Yasuda, T.; Aoyama, M.; Tsubota, E.; Kobayashi, M.; Yuasa, I.; Matsuki, T.; Kishi, K. The fifth allele of the human deoxyribonuclease I (DNase I) polymorphism. Electrophoresis 1997, 18, 1936–1939. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Takeshita, H.; Iida, R.; Kogure, S.; Kishi, K. A new allele, DNASE1*6, of human deoxyribonuclease I polymorphism encodes an Arg to Cys substitution responsible for its instability. Biochem. Biophys. Res. Commun. 1999, 260, 280–283. [Google Scholar] [CrossRef] [PubMed]
- Ueki, M.; Kimura-Kataoka, K.; Fujihara, J.; Iida, R.; Kawai, Y.; Kusaka, A.; Sasaki, T.; Takeshita, H.; Yasuda, T. Evaluation of the functional effects of genetic variants‒missense and nonsense SNPs, indels and copy number variations‒in the gene encoding human deoxyribonuclease I potentially implicated in autoimmunity. Sci. Rep. 2019, 9, 13660. [Google Scholar] [CrossRef]
- Jones, S.J.; Worrall, A.F.; Connolly, B.A. Site-directed mutagenesis of the catalytic residues of bovine pancreatic deoxyribonuclease I. J. Mol. Biol. 1996, 264, 1154–1163. [Google Scholar] [CrossRef]
- Pan, C.Q.; Ulmer, J.S.; Herzka, A.; Lazarus, R.A. Mutational analysis of human DNase I at the DNA binding interface: Implications for DNA recognition, catalysis, and metal ion dependence. Protein Sci. 1998, 7, 628–636. [Google Scholar] [CrossRef]
- Oefner, C.; Suck, D. Crystallographic refinement and structure of DNase I at 2 A resolution. J. Mol. Biol. 1986, 192, 605–632. [Google Scholar] [CrossRef]
- Mannherz, H.G. Crystallization of actin in complex with actin-binding proteins. J. Biol. Chem. 1992, 267, 11661–11664. [Google Scholar]
- Ulmer, J.S.; Herzka, A.; Toy, K.J.; Baker, D.L.; Dodge, A.H.; Sinicropi, D.; Shak, S.; Lazarus, R.A. Engineering actin-resistant human DNase I for treatment of cystic fibrosis. Proc. Natl. Acad. Sci. USA 1996, 93, 8225–8229. [Google Scholar] [CrossRef]
- Fujihara, J.; Yasuda, T.; Kunito, T.; Fujii, Y.; Takatsuka, H.; Moritani, T.; Takeshita, H. Two N-linked glycosylation sites (Asn18 and Asn106) are both required for full enzymatic activity, thermal stability, and resistance to proteolysis in mammalian deoxyribonuclease I. Biosci. Biotechnol. Biochem. 2008, 72, 3197–3205. [Google Scholar] [CrossRef] [PubMed]
- Martínez Valle, F.; Balada, E.; Ordi-Ros, J.; Vilardell-Tarres, M. DNase 1 and systemic lupus erythematosus. Autoimmun. Rev. 2008, 7, 359–363. [Google Scholar] [CrossRef] [PubMed]
- Leffler, J.; Ciacma, K.; Gullstrand, B.; Bengtsson, A.A.; Martin, M.; Blom, A.M. A subset of patients with systemic lupus erythematosus fails to degrade DNA from multiple clinically relevant sources. Arthritis Res. Ther. 2015, 17, 205. [Google Scholar] [CrossRef]
- Cheng, T.H.T.; Lui, K.O.; Laura Peng, X.; Cheng, S.H.; Jiang, P.; Chan, K.C.A.; Chiu, R.W.K.; Lo, Y.M.D. DNase1 Does Not Appear to Play a Major Role in the Fragmentation of Plasma DNA in a Knockout Mouse Model. Clin. Chem. 2018, 64, 106–408. [Google Scholar] [CrossRef] [PubMed]
- Napirei, M.; Karsunky, H.; Zevnik, B.; Stephan, H.; Mannherz, H.G.; Möröy, T. Features of systemic lupus erythematosus in Dnase1-deficient mice. Nat. Genet. 2000, 25, 177–181. [Google Scholar] [CrossRef] [PubMed]
- Zykova, S.N.; Tveita, A.A.; Rekvig, O.P. Renal Dnase1 enzyme activity and protein expression is selectively shut down in murine and human membranoproliferative lupus nephritis. PLoS ONE 2010, 5, e12096. [Google Scholar] [CrossRef]
- Jiménez-Alcázar, M.; Rangaswamy, C.; Panda, R.; Bitterling, J.; Simsek, Y.J.; Long, A.T.; Bilyy, R.; Krenn, V.; Renné, C.; Renné, T.; et al. Host DNases prevent vascular occlusion by neutrophil extracellular traps. Science 2017, 358, 1202–1206. [Google Scholar] [CrossRef]
- Basnakian, A.G.; Apostolov, E.O.; Yin, X.; Napirei, M.; Mannherz, H.G.; Shah, S. V Cisplatin nephrotoxicity is mediated by deoxyribonuclease I. J. Am. Soc. Nephrol. 2005, 16, 697–702. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, A.M.; Rodin, D.; Nomura, H.; Morton, C.C.; Weremowicz, S.; Schneider, M.C. Identification, localization, and expression of two novel human genes similar to deoxyribonuclease I. Genomics 1997, 42, 507–513. [Google Scholar] [CrossRef]
- Parrish, J.E.; Ciccodicola, A.; Wehhert, M.; Cox, G.F.; Chen, E.; Nelson, D.L. A muscle-specific DNase I-like gene in human Xq28. Hum. Mol. Genet. 1995, 4, 1557–1564. [Google Scholar]
- Shiokawa, D.; Tanuma, S. Characterization of human DNase I family endonucleases and activation of DNase gamma during apoptosis. Biochemistry 2001, 40, 143–152. [Google Scholar] [CrossRef] [PubMed]
- Baron, W.F.; Pan, C.Q.; Spencer, S.A.; Ryan, A.M.; Lazarus, R.A.; Baker, K.P. Cloning and characterization of an actin-resistant DNase I-like endonuclease secreted by macrophages. Gene 1998, 215, 291–301. [Google Scholar] [CrossRef]
- Pergolizzi, R.; Appierto, V.; Bosetti, A.; DeBellis, G.L.; Rovida, E.; Biunno, I. Cloning of a gene encoding a DNase I-like endonuclease in the human Xq28 region. Gene 1996, 168, 267–270. [Google Scholar] [CrossRef]
- Fischer, H.; Buchberger, M.; Napirei, M.; Tschachler, E.; Eckhart, L. Inactivation of DNase1L2 and DNase2 in keratinocytes suppresses DNA degradation during epidermal cornification and results in constitutive parakeratosis. Sci. Rep. 2017, 7, 6433. [Google Scholar] [CrossRef]
- Eckhart, L.; Fischer, H.; Barken, K.B.; Tolker-Nielsen, T.; Tschachler, E. DNase1L2 suppresses biofilm formation by Pseudomonas aeruginosa and Staphylococcus aureus. Br. J. Dermatol. 2007, 156, 1342–1345. [Google Scholar] [CrossRef]
- Los, M.; Neubüser, D.; Coy, J.F.; Mozoluk, M.; Poustka, A.; Schulze-Osthoff, K. Functional characterization of DNase X, a novel endonuclease expressed in muscle cells. Biochemistry 2000, 39, 7365–7373. [Google Scholar] [CrossRef]
- Shiokawa, D.; Tanuma, S. Differential DNases are selectively used in neuronal apoptosis depending on the differentiation state. Cell Death Differ. 2004, 11, 1112–1120. [Google Scholar] [CrossRef]
- Shiokawa, D.; Kobayashi, T.; Tanuma, S. Involvement of DNase gamma in apoptosis associated with myogenic differentiation of C2C12 cells. J. Biol. Chem. 2002, 277, 31031–31037. [Google Scholar] [CrossRef]
- Sun, Y.; Ouyang, B.; Xie, Q.Q.; Wang, L.; Zhu, S.; Jia, Y. Serum Deoxyribonuclease 1-like 3 is a potential biomarker for diagnosis of ankylosing spondylitis. Clin. Chim. Acta 2020, 503, 197–202. [Google Scholar] [CrossRef]
- Serpas, L.; Chan, R.W.Y.; Jiang, P.; Ni, M.; Sun, K.; Rashidfarrokhi, A.; Soni, C.; Sisirak, V.; Lee, W.S.; Cheng, S.H.; et al. Dnase1l3 deletion causes aberrations in length and end-motif frequencies in plasma DNA. Proc. Natl. Acad. Sci. USA 2019, 116, 641–649. [Google Scholar] [CrossRef] [PubMed]
- Weisenburger, T.; von Neubeck, B.; Schneider, A.; Ebert, N.; Schreyer, D.; Acs, A.; Winkler, T.H. Epistatic interactions between mutations of deoxyribonuclease 1-Like 3 and the inhibitory Fc gamma receptor IIB result in very early and Massive Autoantibodies against double-stranded DNA. Front. Immunol. 2018, 9, 1551. [Google Scholar] [CrossRef]
- Baker, K.P.; Baron, W.F.; Henzel, W.J.; Spencer, S.A. Molecular cloning and characterization of human and murine DNase II. Gene 1998, 215, 281–289. [Google Scholar] [CrossRef]
- Harosh, I.; Binninger, D.M.; Harris, P.V.; Mezzina, M.; Boyd, J.B. Mechanism of action of deoxyribonuclease II from human lymphoblasts. Eur. J. Biochem. 1991, 202, 479–484. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Nadano, D.; Awazu, S.; Kishi, K. Human urine deoxyribonuclease II (DNase II) isoenzymes: A novel immunoaffinity purification, biochemical multiplicity, genetic heterogeneity and broad distribution among tissues and body fluids. Biochim. Biophys. Acta 1992, 1119, 185–193. [Google Scholar] [CrossRef]
- Ohkouchi, S.; Shibata, M.; Sasaki, M.; Koike, M.; Safig, P.; Peters, C.; Nagata, S.; Uchiyama, Y. Biogenesis and proteolytic processing of lysosomal DNase II. PLoS ONE 2013, 8, e59148. [Google Scholar] [CrossRef]
- Lyon, C.J.; Aguilera, R.J. Purification and characterization of the immunoglobulin switch sequence-specific endonuclease (Endo-SR) from bovine spleen. Mol. Immunol. 1997, 34, 209–219. [Google Scholar] [CrossRef]
- Baranovskii, A.G.; Buneva, V.N.; Nevinsky, G.A. Human deoxyribonucleases. Biochem. Biokhimiia 2004, 69, 587–601. [Google Scholar] [CrossRef]
- Yasuda, T.; Takeshita, H.; Nakazato, E.; Nakajima, T.; Hosomi, O.; Nakashima, Y.; Kishi, K. Activity Measurement for Deoxyribonucleases I and II with Picogram Sensitivity Based on DNA/SYBR Green I Fluorescence. Anal. Biochem. 1998, 255, 274–276. [Google Scholar] [CrossRef]
- Yasuda, T.; Takeshita, H.; Iida, R.; Nakajima, T.; Hosomi, O.; Nakashima, Y.; Mogi, K.; Kishi, K. Chromosomal localization of a human deoxyribonuclease II gene (DNASE2) to 19p13.2-p13.1 using both the polymerase chain reaction and fluorescence in situ hybridization analysis. Biochem. Biophys. Res. Commun. 1998, 244, 815–818. [Google Scholar] [CrossRef]
- Yasuda, T.; Takeshita, H.; Iida, R.; Tsutsumi, S.; Nakajima, T.; Hosomi, O.; Nakashima, Y.; Mori, S.; Kishi, K. Structure and organization of the human deoxyribonuclease II (DNase II) gene. Ann. Hum. Genet. 1998, 62, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Nadano, D.; Sawazaki, K.; Kishi, K. Genetic polymorphism of human deoxyribonuclease II (DNase II): Low activity levels in urine and leukocytes are due to an autosomal recessive allele. Ann. Hum. Genet. 1992, 56, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Krieser, R.J.; Eastman, A. The cloning and expression of human deoxyribonuclease II. A possible role in apoptosis. J. Biol. Chem. 1998, 273, 30909–30914. [Google Scholar] [CrossRef] [PubMed]
- MacLea, K.S.; Krieser, R.J.; Eastman, A. Revised structure of the active form of human deoxyribonuclease IIalpha. Biochem. Biophys. Res. Commun. 2002, 292, 415–421. [Google Scholar]
- MacLea, K.S.; Krieser, R.J.; Eastman, A. Structural requirements of human DNase II alpha for formation of the active enzyme: The role of the signal peptide, N-glycosylation, and disulphide bridging. Biochem. J. 2003, 371, 867–876. [Google Scholar] [CrossRef]
- Gottlieb, R.A.; Nordberg, J.; Skowronski, E.; Babior, B.M. Apoptosis induced in Jurkat cells by several agents is preceded by intracellular acidification. Proc. Natl. Acad. Sci. USA 1996, 93, 654–658. [Google Scholar] [CrossRef]
- Kawane, K.; Fukuyama, H.; Yoshida, H.; Nagase, H.; Ohsawa, Y.; Uchiyama, Y.; Okada, K.; Iida, T.; Nagata, S. Impaired thymic development in mouse embryos deficient in apoptotic DNA degradation. Nat. Immunol. 2003, 4, 138–144. [Google Scholar] [CrossRef]
- Howell, D.P.G.; Krieser, R.J.; Eastman, A.; Barry, M.A. Deoxyribonuclease II is a lysosomal barrier to transfection. Mol. Ther. 2003, 8, 957–963. [Google Scholar] [CrossRef]
- Chan, M.P.; Onji, M.; Fukui, R.; Kawane, K.; Shibata, T.; Saitoh, S.; Ohto, U.; Shimizu, T.; Barber, G.N.; Miyake, K. DNase II-dependent DNA digestion is required for DNA sensing by TLR9. Nat. Commun. 2015, 6, 5853. [Google Scholar] [CrossRef]
- Pawaria, S.; Moody, K.; Busto, P.; Nündel, K.; Choi, C.H.; Ghayur, T.; Marshak-Rothstein, A. Cutting Edge: DNase II deficiency prevents activation of autoreactive B cells by double-stranded DNA endogenous ligands. J. Immunol. 2015, 194, 1403–1407. [Google Scholar] [CrossRef]
- Kawane, K.; Fukuyama, H.; Kondoh, G.; Takeda, J.; Ohsawa, Y.; Uchiyama, Y.; Nagata, S. Requirement of DNase II for definitive erythropoiesis in the mouse fetal liver. Science 2001, 292, 1546–1549. [Google Scholar] [CrossRef] [PubMed]
- Krieser, R.J.; MacLea, K.S.; Longnecker, D.S.; Fields, J.L.; Fiering, S.; Eastman, A. Deoxyribonuclease IIalpha is required during the phagocytic phase of apoptosis and its loss causes perinatal lethality. Cell Death Differ. 2002, 9, 956–962. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kawane, K.; Ohtani, M.; Miwa, K.; Kizawa, T.; Kanbara, Y.; Yoshioka, Y.; Yoshikawa, H.; Nagata, S. Chronic polyarthritis caused by mammalian DNA that escapes from degradation in macrophages. Nature 2006, 443, 998–1002. [Google Scholar] [CrossRef]
- Shin, H.D.; Park, B.L.; Cheong, H.S.; Lee, H.-S.; Jun, J.-B.; Bae, S.-C. DNase II polymorphisms associated with risk of renal disorder among systemic lupus erythematosus patients. J. Hum. Genet. 2005, 50, 107–111. [Google Scholar] [CrossRef] [PubMed]
- Fischer, H.; Scherz, J.; Szabo, S.; Mildner, M.; Benarafa, C.; Torriglia, A.; Tschachler, E.; Eckhart, L. Dnase 2 is the main DNA-degrading enzyme of the stratum corneum. PLoS ONE 2011, 6, e17581. [Google Scholar] [CrossRef] [PubMed]
- Fischer, H.; Fumicz, J.; Rossiter, H.; Napirei, M.; Buchberger, M.; Tschachler, E.; Eckhart, L. Holocrine Secretion of Sebum Is a Unique DNase2-Dependent Mode of Programmed Cell Death. J. Investig. Dermatol. 2017, 137, 587–594. [Google Scholar] [CrossRef]
- Fischer, H.; Szabo, S.; Scherz, J.; Jaeger, K.; Rossiter, H.; Buchberger, M.; Ghannadan, M.; Hermann, M.; Theussl, H.C.; Tobin, D.J.; et al. Essential role of the keratinocyte-specific endonuclease DNase1L2 in the removal of nuclear DNA from hair and nails. J. Investig. Dermatol. 2011, 131, 1208–1215. [Google Scholar] [CrossRef]
- Manils, J.; Fischer, H.; Climent, J.; Casas, E.; García-Martínez, C.; Bas, J.; Sukseree, S.; Vavouri, T.; Ciruela, F.; De Anta, J.M.; et al. Double deficiency of Trex2 and DNase1L2 nucleases leads to accumulation of DNA in lingual cornifying keratinocytes without activating inflammatory responses. Sci. Rep. 2017, 7, 11902. [Google Scholar] [CrossRef]
- Shiokawa, D.; Tanuma, S. DLAD, a novel mammalian divalent cation-independent endonuclease with homology to DNase II. Nucleic Acids Res. 1999, 27, 4083–4089. [Google Scholar] [CrossRef]
- Krieser, R.J.; MacLea, K.S.; Park, J.P.; Eastman, A. The cloning, genomic structure, localization, and expression of human deoxyribonuclease IIbeta. Gene 2001, 269, 205–216. [Google Scholar] [CrossRef]
- Tanuma, S.; Shiokawa, D. Cloning of a cDNA Encoding a Rat DNase II-like Acid DNase. Biochem. Biophys. Res. Commun. 1999, 265, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Nishimoto, S.; Kawane, K.; Watanabe-Fukunaga, R.; Fukuyama, H.; Ohsawa, Y.; Uchiyama, Y.; Hashida, N.; Ohguro, N.; Tano, Y.; Morimoto, T.; et al. Nuclear cataract caused by a lack of DNA degradation in the mouse eye lens. Nature 2003, 424, 1071–1074. [Google Scholar] [CrossRef] [PubMed]
- Torriglia, A.; Perani, P.; Brossas, J.Y.; Chaudun, E.; Treton, J.; Courtois, Y.; Counis, M.F. L-DNase II, a Molecule That Links Proteases and Endonucleases in Apoptosis, Derives from the Ubiquitous Serpin Leukocyte Elastase Inhibitor. Mol. Cell. Biol. 1998, 18, 3612–3619. [Google Scholar] [CrossRef]
- Potempa, J.; Dubin, A.; Watorek, W.; Travis, J. An elastase inhibitor from equine leukocyte cytosol belongs to the serpin superfamily. Further characterization and amino acid sequence of the reactive center. J. Biol. Chem. 1988, 263, 7364–7369. [Google Scholar] [PubMed]
- Altairac, S.; Wright, S.C.; Courtois, Y.; Torriglia, A. L-DNase II activation by the 24 kDa apoptotic protease (AP24) in TNFalpha-induced apoptosis. Cell Death Differ. 2003, 10, 1109–1111. [Google Scholar] [CrossRef] [PubMed]
- Chahory, S.; Padron, L.; Daniel, C.; Keller, N.; Torriglia, A. Involvement of the LEI/L—DNase II Pathway in Light–Induced Retinal Degeneration. Investig. Ophthalmol. Vis. Sci. 2006, 47, 2045. [Google Scholar]
- Torriglia, A.; Jaadane, I.; Lebon, C. Mechanisms of cell death in neurodegenerative and retinal diseases: Common pathway? Curr. Opin. Neurol. 2016, 29, 55–60. [Google Scholar] [CrossRef]
- Gorrini, C.; Donzelli, M.; Torriglia, A.; Supino, R.; Brison, O.; Bernardi, R.; Negri, C.; Denegri, M.; Counis, M.F.; Ranzani, G.N.; et al. Effect of apoptogenic stimuli on colon carcinoma cell lines with a different c-myc expression level. Int. J. Mol. Med. 2003, 11, 737–742. [Google Scholar] [CrossRef]
- Stetson, D.B.; Ko, J.S.; Heidmann, T.; Medzhitov, R. Trex1 Prevents Cell-Intrinsic Initiation of Autoimmunity. Cell 2008, 134, 587–598. [Google Scholar] [CrossRef]
- Chowdhury, D.; Beresford, P.J.; Zhu, P.; Zhang, D.; Sung, J.S.; Demple, B.; Perrino, F.W.; Lieberman, J. The Exonuclease TREX1 Is in the SET Complex and Acts in Concert with NM23-H1 to Degrade DNA during Granzyme A-Mediated Cell Death. Mol. Cell 2006, 23, 133–142. [Google Scholar] [CrossRef]
- Lee-Kirsch, M.A.; Gong, M.; Chowdhury, D.; Senenko, L.; Engel, K.; Lee, Y.A.; De Silva, U.; Bailey, S.L.; Witte, T.; Vyse, T.J.; et al. Mutations in the gene encoding the 3′–5′ DNA exonuclease TREX1 are associated with systemic lupus erythematosus. Nat. Genet. 2007, 39, 1065–1067. [Google Scholar] [CrossRef] [PubMed]
- Lazzaretto, B.; Fadeel, B. Intra- and Extracellular Degradation of Neutrophil Extracellular Traps by Macrophages and Dendritic Cells. J. Immunol. 2019, 203, 2276–2290. [Google Scholar] [CrossRef] [PubMed]
- Mazur, D.J.; Perrino, F.W. Structure and Expression of the TREX1 and TREX2 3′–5′ Exonuclease Genes. J. Biol. Chem. 2001, 276, 14718–14727. [Google Scholar] [CrossRef] [PubMed]
- Jani, D.; Lutz, S.; Laskey, R.A.; Stewart, M.; Wickramasinghe, V.O. Functional and Structural Characterization of the Mammalian TREX-2 Complex That Links Transcription With Nuclear Messenger RNA Export—PubMed. Nucleic Acids Res. 2012, 40, 4562–4573. [Google Scholar] [CrossRef] [PubMed]
- KUNITZ, M. Crystalline desoxyribonuclease; isolation and general properties; spectrophotometric method for the measurement of desoxyribonuclease activity. J. Gen. Physiol. 1950, 33, 349–362. [Google Scholar] [CrossRef] [PubMed]
- Nadano, D.; Yasuda, T.; Kishi, K. Measurement of deoxyribonuclease I activity in human tissues and body fluids by a single radial enzyme-diffusion method. Clin. Chem. 1993, 39, 448–452. [Google Scholar] [CrossRef]
- Horney, D.L.; Webster, D.A. Deoxyribonuclease: A sensitive assay using radial diffusion in agarose containing methyl green-DNA complex. Biochim. Biophys. Acta 1971, 247, 54–61. [Google Scholar] [CrossRef]
- Choi, S.J.; Szoka, F.C. Fluorometric Determination of Deoxyribonuclease I Activity with PicoGreen. Anal. Biochem. 2000, 281, 95–97. [Google Scholar] [CrossRef]
- Cherepanova, A.; Tamkovich, S.; Pyshnyi, D.; Kharkova, M.; Vlassov, V.; Laktionov, P. Immunochemical assay for deoxyribonuclease activity in body fluids. J. Immunol. Methods 2007, 325, 96–103. [Google Scholar] [CrossRef]
- Nakajima, T.; Takagi, R.; Tajima, Y.; Makita, C.; Kominato, Y.; Kuribara, J.; Ohshima, S.; Tada, H.; Tsurugaya, H.; Kobayashi, Y.; et al. Development of a sensitive enzyme-linked immunosorbent assay for measurement of DNase I in human serum. Clin. Chim. Acta 2009, 403, 219–222. [Google Scholar] [CrossRef]
- Fujihara, J.; Tabuchi, M.; Inoue, T.; Yasuda, T.; Fujita, Y.; Takeshita, H. Rapid measurement of deoxyribonuclease I activity with the use of microchip electrophoresis based on DNA degradation. Anal. Biochem. 2011, 413, 78–79. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Min, D.H. A simple fluorometric assay for DNA exonuclease activity based on graphene oxide. Analyst 2012, 137, 2024–2026. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Ying, J.Y. Homogeneous Immunochemical Assay on the Lateral Flow Strip for Measurement of DNase I Activity. Anal. Chem. 2015, 87, 10193–10198. [Google Scholar] [CrossRef]
- Vogel, B.; Frantz, S. Determination of DNase activity by degradation of ethidium bromide–DNA complexes using a fluorescence plate reader. Anal. Biochem. 2015, 471, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Funakoshi, A.; Wakasugi, H.; Ibayashi, H. Clinical investigation of serum deoxyribonuclease: II. Clinical studies of serum deoxyribonuclease activity in pancreatic disease. Gastroenterol. Jpn. 1979, 14, 436–440. [Google Scholar] [CrossRef] [PubMed]
- Economidou-Karaoglou, A.; Lans, M.; Taper, H.S.; Michaux, J.L.; Roberfroid, M. variations in serum alkaline dnase activity. A new means for therapeutic monitoring of malignant lymphomas. Cancer 1988, 61, 1838–1843. [Google Scholar] [CrossRef]
- Spandidos, D.A.; Ramandanis, G.; Garas, J.; Kottaridis, S.D. Serum deoxyribonucleases in patients with breast cancer. Eur. J. Cancer 1980, 16, 1615–1619. [Google Scholar] [CrossRef]
- Scully, C.; Spandidos, D.A.; Ward Booth, P.; McGregor, I.A.; Boyle, P. Serum alkaline deoxyribonuclease in oral cancer and premalignant lesions. Biomedicine 1981, 35, 179–180. [Google Scholar]
- Kawai, Y.; Yoshida, M.; Arakawa, K.; Kumamoto, T.; Morikawa, N.; Masamura, K.; Tada, H.; Ito, S.; Hoshizaki, H.; Oshima, S.; et al. Diagnostic use of serum deoxyribonuclease I activity as a novel early-phase marker in acute myocardial infarction. Circulation 2004, 109, 2398–2400. [Google Scholar] [CrossRef]
- Fujibayashi, K.; Kawai, Y.; Kitayama, M.; Akao, H.; Ishida, R.; Motoyama, A.; Wakasa, M.; Arakawa, K.; Ueki, M.; Kajinami, K.; et al. Serum deoxyribonuclease I activity can be a useful diagnostic marker for the early diagnosis of unstable angina pectoris or non-ST-segment elevation myocardial infarction. J. Cardiol. 2012, 59, 258–265. [Google Scholar] [CrossRef]
- Morikawa, N.; Kawai, Y.; Arakawa, K.; Kumamoto, T.; Miyamori, I.; Akao, H.; Kitayama, M.; Kajinami, K.; Lee, J.D.; Takeshita, H.; et al. Serum deoxyribonuclease I activity can be used as a novel marker of transient myocardial ischaemia: Results in vasospastic angina pectoris induced by provocation test. Eur. Heart J. 2007, 28, 2992–2997. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Iida, R.; Kawai, Y.; Nakajima, T.; Kominato, Y.; Fujihara, J.; Takeshita, H. Serum deoxyribonuclease I can be used as a useful marker for diagnosis of death due to ischemic heart disease. Leg. Med. 2009, 11, S213–S215. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Kawai, Y.; Ueki, M.; Kishi, K. Clinical applications of DNase I, a genetic marker already used for forensic identification. Leg. Med. 2005, 7, 274–277. [Google Scholar] [CrossRef] [PubMed]
- Creeth, J.M.; Gulland, J.M.; Jordan, D.O. Deoxypentose nucleic acids. Part III. Viscosity and streaming birefringence of solutions of the sodium salt of the deoxypentose nucleic acid of calf thymus. J. Chem. Soc. 1947, 1141–1145. [Google Scholar] [CrossRef] [PubMed]
- Elkin, I.; Weight, A.K.; Klibanov, A.M. Markedly lowering the viscosity of aqueous solutions of DNA by additives. Int. J. Pharm. 2015, 494, 66–72. [Google Scholar] [CrossRef] [PubMed]
- Matthews, L.W.; Spector, S.; Lemm, J.; Potter, J.L. Studied on pulmonary secretions. I. The over-all chemical composition of pulmonary secretions from patients with cystic fibrosis, bronchiectasis and laryngectomy. Am. Rev. Respir. Dis. 1963, 88, 199–204. [Google Scholar]
- Lethem, M.I.; James, S.L.; Marriott, C.; Burke, J.F. The origin of DNA associated with mucus glycoproteins in cystic fibrosis sputum. Eur. Respir. J. 1990, 3, 19–23. [Google Scholar]
- Yoo, D.; Floyd, M.; Winn, M.; Moskowitz, S.M.; Rada, B. NET formation induced by Pseudomonas aeruginosa cystic fibrosis isolates measured as release of myeloperoxidase-DNA and neutrophil elastase-DNA complexes. Immunol. Lett. 2014, 160, 186–194. [Google Scholar] [CrossRef]
- Armstrong, J.; White, J.C. Liquefaction of viscous purulent exudates by deoxyribonuclease. Lancet 1950, 256, 739–742. [Google Scholar] [CrossRef]
- Shak, S.; Capon, D.J.; Hellmiss, R.; Marsters, S.A.; Baker, C.L. Recombinant human DNase I reduces the viscosity of cystic fibrosis sputum. Proc. Natl. Acad. Sci. USA 1990, 87, 9188–9192. [Google Scholar] [CrossRef]
- Yang, C.; Chilvers, M.; Montgomery, M.; Nolan, S.J. Dornase alfa for cystic fibrosis. Cochrane Database Syst. Rev. 2016, 4, CD001127. [Google Scholar] [CrossRef] [PubMed]
- Cantin, A.M. DNase I Acutely Increases Cystic Fibrosis Sputum Elastase Activity and its Potential to Induce Lung Hemorrhage in Mice. Am. J. Respir. Crit. Care Med. 1998, 157, 464–469. [Google Scholar] [CrossRef] [PubMed]
- Sawicki, G.S.; Chou, W.; Raimundo, K.; Trzaskoma, B.; Konstan, M.W. Randomized trial of efficacy and safety of dornase alfa delivered by eRapid nebulizer in cystic fibrosis patients. J. Cyst. Fibros. 2015, 14, 777–783. [Google Scholar] [CrossRef] [PubMed]
- Mainz, J.G.; Schien, C.; Schiller, I.; Schädlich, K.; Koitschev, A.; Koitschev, C.; Riethmüller, J.; Graepler-Mainka, U.; Wiedemann, B.; Beck, J.F. Sinonasal inhalation of dornase alfa administered by vibrating aerosol to cystic fibrosis patients: A double-blind placebo-controlled cross-over trial. J. Cyst. Fibros. 2014, 13, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Amin, R.; Subbarao, P.; Lou, W.; Jabar, A.; Balkovec, S.; Jensen, R.; Kerrigan, S.; Gustafsson, P.; Ratjen, F. The effect of dornase alfa on ventilation inhomogeneity in patients with cystic fibrosis. Eur. Respir. J. 2011, 37, 806–812. [Google Scholar] [CrossRef] [PubMed]
- Henry, R.L.; Gibson, P.G.; Carty, K.; Cai, Y.; Francis, J.L. Airway inflammation after treatment with aerosolized deoxyribonuclease in cystic fibrosis. Pediatr. Pulmonol. 1998, 26, 97–100. [Google Scholar] [CrossRef]
- Patel, A.; Harrison, E.; Durward, A.; Murdoch, I.A. Intratracheal recombinant human deoxyribonuclease in acute life-threatening asthma refractory to conventional treatment. Br. J. Anaesth. 2000, 84, 505–507. [Google Scholar] [CrossRef]
- Durward, A.; Forte, V.; Shemie, S.D. Resolution of mucus plugging and atelectasis after intratracheal rhDNase therapy in a mechanically ventilated child with refractory status asthmaticus. Crit. Care Med. 2000, 28, 560–562. [Google Scholar] [CrossRef]
- Silverman, R.A.; Foley, F.; Dalipi, R.; Kline, M.; Lesser, M. The use of rhDNAse in severely ill, non-intubated adult asthmatics refractory to bronchodilators: A pilot study. Respir. Med. 2012, 106, 1096–1102. [Google Scholar] [CrossRef]
- Macanovic, M.; Sinicropi, D.; Shak, S.; Baughman, S.; Thiru, S.; Lachmann, P.J. The treatment of systemic lupus erythematosus (SLE) in NZB/W F1 hybrid mice; studies with recombinant murine DNase and with dexamethasone. Clin. Exp. Immunol. 1996, 106, 243–252. [Google Scholar] [CrossRef]
- Verthelyi, D.; Dybdal, N.; Elias, K.A.; Klinman, D.M. DNAse treatment does not improve the survival of lupus prone (NZB x NZW)F1 mice. Lupus 1998, 7, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Davis, J.C.; Manzi, S.; Yarboro, C.; Rairie, J.; Mcinnes, I.; Averthelyi, D.; Sinicropi, D.; Hale, V.G.; Balow, J.; Austin, H.; et al. Recombinant human Dnase I (rhDNase) in patients with lupus nephritis. Lupus 1999, 8, 68–76. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, S.; Yamamoto, T.; Tsuruta, J.; Tanaka, J.; Kambara, T.; Hiraoka, T.; Miyauchi, Y. Serine protease-induced enhancement of blood-borne metastasis of rat ascites tumour cells and its prevention with deoxyribonuclease. Br. J. Cancer 1990, 62, 607–613. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, S.; Yamamoto, T.; Tanaka, H.; Kambara, T.; Hiraoka, T.; Miyauchi, Y. Deoxyribonuclease treatment prevents blood-borne liver metastasis of cutaneously transplanted tumour cells in mice. Br. J. Cancer 1993, 67, 66–70. [Google Scholar] [CrossRef]
- Tokita, K.; Sugihara, S.; Hiraoka, T.; Miyauchi, Y.; Kambara, T.; Yamamoto, T. Effects of serine protease and deoxyribonuclease on intravascular tumor cell arrest in rat blood-borne lung metastasis. Invasion Metastasis 1995, 15, 46–59. [Google Scholar]
- Tetz, V.; Tetz, G. Effect of deoxyribonuclease I treatment for dementia in end-stage Alzheimer’s disease: A case report. J. Med. Case Rep. 2016, 10, 131. [Google Scholar] [CrossRef]
- Gao, X.; Hao, S.; Yan, H.; Ding, W.; Li, K.; Li, J. Neutrophil extracellular traps contribute to the intestine damage in endotoxemic rats. J. Surg. Res. 2015, 195, 211–218. [Google Scholar] [CrossRef]
- Mai, S.H.C.; Khan, M.; Dwivedi, D.J.; Ross, C.A.; Zhou, J.; Gould, T.J.; Gross, P.L.; Weitz, J.I.; Fox-Robichaud, A.E.; Liaw, P.C. Delayed but not Early Treatment with DNase Reduces Organ Damage and Improves Outcome in a Murine Model of Sepsis. Shock 2015, 44, 166–172. [Google Scholar] [CrossRef]
- Meng, W.; Paunel-Görgülü, A.; Flohé, S.; Hoffmann, A.; Witte, I.; MacKenzie, C.; Baldus, S.E.; Windolf, J.; Lögters, T.T. Depletion of neutrophil extracellular traps in vivo results in hypersusceptibility to polymicrobial sepsis in mice. Crit. Care 2012, 16, R137. [Google Scholar] [CrossRef]
- Lauková, L.; Konečná, B.; Bábíčková, J.; Wagnerová, A.; Melišková, V.; Vlková, B.; Celec, P. Exogenous deoxyribonuclease has a protective effect in a mouse model of sepsis. Biomed. Pharmacother. 2017, 93, 8–16. [Google Scholar] [CrossRef]
- Peer, V.; Abu Hamad, R.; Berman, S.; Efrati, S. Renoprotective Effects of DNAse-I Treatment in a Rat Model of Ischemia/Reperfusion-Induced Acute Kidney Injury. Am. J. Nephrol. 2016, 43, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Vokálová, L.; Lauková, L.; Čonka, J.; Melišková, V.; Borbélyová, V.; Bábíčková, J.; Tóthová, L.; Hodosy, J.; Vlková, B.; Celec, P. Deoxyribonuclease partially ameliorates thioacetamide-induced hepatorenal injury. Am. J. Physiol. Gastrointest. Liver Physiol. 2017, 312, G457–G463. [Google Scholar] [CrossRef] [PubMed]
- Albadawi, H.; Oklu, R.; Raacke Malley, R.E.; O’Keefe, R.M.; Uong, T.P.; Cormier, N.R.; Watkins, M.T.; Oklu, R.; Albadawi, H.; Jones, J.E.; et al. Effect of DNase I treatment and neutrophil depletion on acute limb ischemia-reperfusion injury in mice. J. Vasc. Surg. 2016, 64, 484–493. [Google Scholar] [CrossRef] [PubMed]
- Bobek, V.; Majewski, A.; Kolostova, K.; Rzechonek, A.; Lischke, R.; Schutzner, J.; Kacprzak, G. Intrapleural administration of DNase alone for pleural empyema. Int. J. Clin. Exp. Med. 2015, 8, 22011–22015. [Google Scholar] [PubMed]
- Simpson, G.; Roomes, D.; Heron, M. Effects of streptokinase and deoxyribonuclease on viscosity of human surgical and empyema pus. Chest 2000, 117, 1728–1733. [Google Scholar] [CrossRef]
- Pedersen, H.L.; Horvei, K.D.; Thiyagarajan, D.; Seredkina, N.; Rekvig, O.P. Murine and Human Lupus Nephritis: Pathogenic Mechanisms and Theoretical Strategies for Therapy. Semin. Nephrol. 2015, 35, 427–438. [Google Scholar] [CrossRef]
- Alcázar-Leyva, S.; Cerón, E.; Masso, F.; Montaño, L.F.; Gorocica, P.; Alvarado-Vásquez, N. Incubation with DNase I inhibits tumor cell proliferation. Med. Sci. Monit. 2009, 15, CR51–CR55. [Google Scholar]
- Linardou, H.; Epenetos, A.A.; Deonarain, M.P. A recombinant cytotoxic chimera based on mammalian deoxyribonuclease-I. Int. J. Cancer 2000, 86, 561–569. [Google Scholar] [CrossRef]
- Golonka, R.M.; Yeoh, B.S.; Petrick, J.L.; Weinstein, S.J.; Albanes, D.; Gewirtz, A.T.; McGlynn, K.A.; Vijay-Kumar, M. Deoxyribonuclease I Activity, Cell-Free DNA, and Risk of Liver Cancer in a Prospective Cohort. JNCI Cancer Spectr. 2018, 2, pky083. [Google Scholar] [CrossRef]
- Whitchurch, C.B.; Tolker-Nielsen, T.; Ragas, P.C.; Mattick, J.S.; Costerton, J.W.; Stewart, P.S.; Greenberg, E.P.; Kolter, R.; Sutherland, I.W.; Muto, Y.; et al. Extracellular DNA required for bacterial biofilm formation. Science 2002, 295, 1487. [Google Scholar] [CrossRef]
- Tetz, V.V.; Tetz, G.V. Effect of Extracellular DNA Destruction by DNase I on Characteristics of Forming Biofilms. DNA Cell Biol. 2010, 29, 399–405. [Google Scholar] [CrossRef]
- Izano, E.A.; Shah, S.M.; Kaplan, J.B. Intercellular adhesion and biocide resistance in nontypeable Haemophilus influenzae biofilms. Microb. Pathog. 2009, 46, 207–213. [Google Scholar] [CrossRef] [PubMed]
- Seper, A.; Fengler, V.H.I.; Roier, S.; Wolinski, H.; Kohlwein, S.D.; Bishop, A.L.; Camilli, A.; Reidl, J.; Schild, S. Extracellular nucleases and extracellular DNA play important roles in Vibrio cholerae biofilm formation. Mol. Microbiol. 2011, 82, 1015–1037. [Google Scholar] [CrossRef] [PubMed]
- Harmsen, M.; Lappann, M.; Knøchel, S.; Molin, S. Role of extracellular DNA during biofilm formation by listeria monocytogenes. Appl. Environ. Microbiol. 2010, 76, 2271–2279. [Google Scholar] [CrossRef] [PubMed]
- Hall-Stoodley, L.; Nistico, L.; Sambanthamoorthy, K.; Dice, B.; Nguyen, D.; Mershon, W.J.; Johnson, C.; Hu, F.Z.; Stoodley, P.; Ehrlich, G.D.; et al. Characterization of biofilm matrix, degradation by DNase treatment and evidence of capsule downregulation in Streptococcus pneumoniae clinical isolates. BMC Microbiol. 2008, 8, 173. [Google Scholar] [CrossRef]
- Cagliani, J.; Yang, W.L.; Brenner, M.; Wang, P. Deoxyribonuclease Reduces Tissue Injury and Improves Survival After Hemorrhagic Shock. J. Surg. Res. 2020, 249, 104–113. [Google Scholar] [CrossRef]
- Mun, C.; Gulati, S.; Tibrewal, S.; Chen, Y.F.; An, S.; Surenkhuu, B.; Raju, I.; Buwick, M.; Ahn, A.; Kwon, J.E.; et al. A phase I/II placebo-controlled randomized pilot clinical trial of recombinant deoxyribonuclease (DNase) eye drops use in patients with dry eye disease. Transl. Vis. Sci. Technol. 2019, 8, 10. [Google Scholar] [CrossRef]
- McIlroy, D.J.; Minahan, K.; Keely, S.; Lott, N.; Hansbro, P.; Smith, D.W.; Balogh, Z.J. Reduced deoxyribonuclease enzyme activity in response to high postinjury mitochondrial DNA concentration provides a therapeutic target for Systemic Inflammatory Response Syndrome. J. Trauma Acute Care Surg. 2018, 85, 354–358. [Google Scholar] [CrossRef]
- Chen, Q.; Ye, L.; Jin, Y.; Zhang, N.; Lou, T.; Qiu, Z.; Jin, Y.; Cheng, B.; Fang, X. Circulating nucleosomes as a predictor of sepsis and organ dysfunction in critically ill patients. Int. J. Infect. Dis. 2012, 16, e558–e564. [Google Scholar] [CrossRef]
- Zeerleder, S.; Zwart, B.; Wuillemin, W.A.; Aarden, L.A.; Groeneveld, A.B.J.; Caliezi, C.; van Nieuwenhuijze, A.E.M.; van Mierlo, G.J.; Eerenberg, A.J.M.; Lämmle, B.; et al. Elevated nucleosome levels in systemic inflammation and sepsis. Crit. Care Med. 2003, 31, 1947–1951. [Google Scholar] [CrossRef]
- Dwivedi, D.J.; Toltl, L.J.; Swystun, L.L.; Pogue, J.; Liaw, K.L.; Weitz, J.I.; Cook, D.J.; Fox-Robichaud, A.E.; Liaw, P.C.; Canadian Critical Care Translational Biology Group. Prognostic utility and characterization of cell-free DNA in patients with severe sepsis. Crit. Care 2012, 16, R151. [Google Scholar] [CrossRef] [PubMed]


| Family | DNase I Family | DNase II Family | |||||
|---|---|---|---|---|---|---|---|
| Name | DNase I | DNase1L3 | DNase1L1 | DNase1L2 | DNase II α | DNase II β | L-DNase II |
| Molecular mass (kDa) | 38 | 33 | 34 | 33 | 43 | 40 | 27 |
| Optimal pH | 6.5–8 | 6.5–8 | 6.5–8 | 5.6 | 4.8–5.2 | 4.8–5.2 | 4.8–5.2 |
| Activation by Mg2+ and Ca2+ | + | + | + | + | - | - | - |
| Inhibition by EDTA/EGTA | + | + | + | + | - | - | - |
| Inhibition by G-actin | + | - | - | - | - | - | - |
| Productive organs | pancreas | spleen, liver | muscles, myocardium | brain, lungs, placenta, skin | all tissues | salivary glands | spleen |
| Name of coding gene | DNASE1 | DNASE1L3 | DNASE1L1 | DNASE1L2 | DNASE2 | DNASE2B | SERPINB1 |
| Localization of human gene | 16p13.3 | 3p14.3 | Xq28 | 16p13.3 | 19p13.2 | 1p22.3 | 6q25.2 |
| Number of exons | 17 | 8 | 10 | 7 | 6 | 6 | 9 |
| Disease | Organism | Administration | Dose | Effect | Reference |
|---|---|---|---|---|---|
| Cystic fibrosis | humans | inhalation | 2.5 mg once or twice daily | positive | [110] |
| humans | inhalation | 2.5 mg/day, 1 month | positive | [116] | |
| mice and humans | inhalation (humans) at the nares (mice) | 2.5 mg/day (humans) 50 µL of the sputum (mice) | negative | [112] | |
| humans | inhalation | 2.5 mg/day in 2-week periods | positive | [113] | |
| humans | inhalation | 2.5 mg/day, 28 days | positive | [114] | |
| humans | inhalation | 2.5 mg/day, 2-times, 4 weeks, with a 4 week pause | positive | [115] | |
| Asthma | humans | intratracheal | 2.5 mg | positive | [117] |
| humans | intratracheal | 10 mg, 2 times 8 h apart | positive | [118] | |
| humans | inhalation | 2.5, 5.0 or 7.5 mg | no | [119] | |
| Systemic lupus erythematosus | mice | intraperitoneal | 150 µg/day, 3 months | positive | [120] |
| mice | intraperitoneal | 0–15 µg/g, 1–6 months | no | [121] | |
| humans | single intravenous 10 subcutaneous | 25 µg/kg, 125 µg/kg | no | [122] | |
| Cancer | rats | intravenous | 1.5 U | positive | [123] |
| mice | intravenous | 0.1 U | positive | [124] | |
| rats | intravenous | 1 U | positive | [125] | |
| Alzheimer disease | humans | oral | 120 mg/day, 2 months | positive | [126] |
| rats | intravenous | 10 mg/kg | positive | [127] | |
| Sepsis | mice | intraperitoneal | 20 mg/kg (every 2, 4 or 6 h) | positive | [128] |
| mice | intraperitoneal | 5 mg/kg (7 times) | positive | [129] | |
| mice | intravenous | 10 mg/kg | positive | [130] | |
| Acute kidney injury | rats | intraperitoneal | 0.1 mg/kg | positive | [131] |
| Acute liver injury | rats | intravenous | 10 mg/kg | positive | [132] |
| Ischemic-reperfusion syndrome | mice | intraperitoneal intravenous | 50 µg, 2 times 10 µg, 1 time | positive | [133] |
| Empyema thoracis | humans | intrapleural | 2.5 mg (1–2 times) | positive | [134] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lauková, L.; Konečná, B.; Janovičová, Ľ.; Vlková, B.; Celec, P. Deoxyribonucleases and Their Applications in Biomedicine. Biomolecules 2020, 10, 1036. https://doi.org/10.3390/biom10071036
Lauková L, Konečná B, Janovičová Ľ, Vlková B, Celec P. Deoxyribonucleases and Their Applications in Biomedicine. Biomolecules. 2020; 10(7):1036. https://doi.org/10.3390/biom10071036
Chicago/Turabian StyleLauková, Lucia, Barbora Konečná, Ľubica Janovičová, Barbora Vlková, and Peter Celec. 2020. "Deoxyribonucleases and Their Applications in Biomedicine" Biomolecules 10, no. 7: 1036. https://doi.org/10.3390/biom10071036
APA StyleLauková, L., Konečná, B., Janovičová, Ľ., Vlková, B., & Celec, P. (2020). Deoxyribonucleases and Their Applications in Biomedicine. Biomolecules, 10(7), 1036. https://doi.org/10.3390/biom10071036






