Encapsulins—Bacterial Protein Nanocompartments: Structure, Properties, and Application
Abstract
1. Introduction
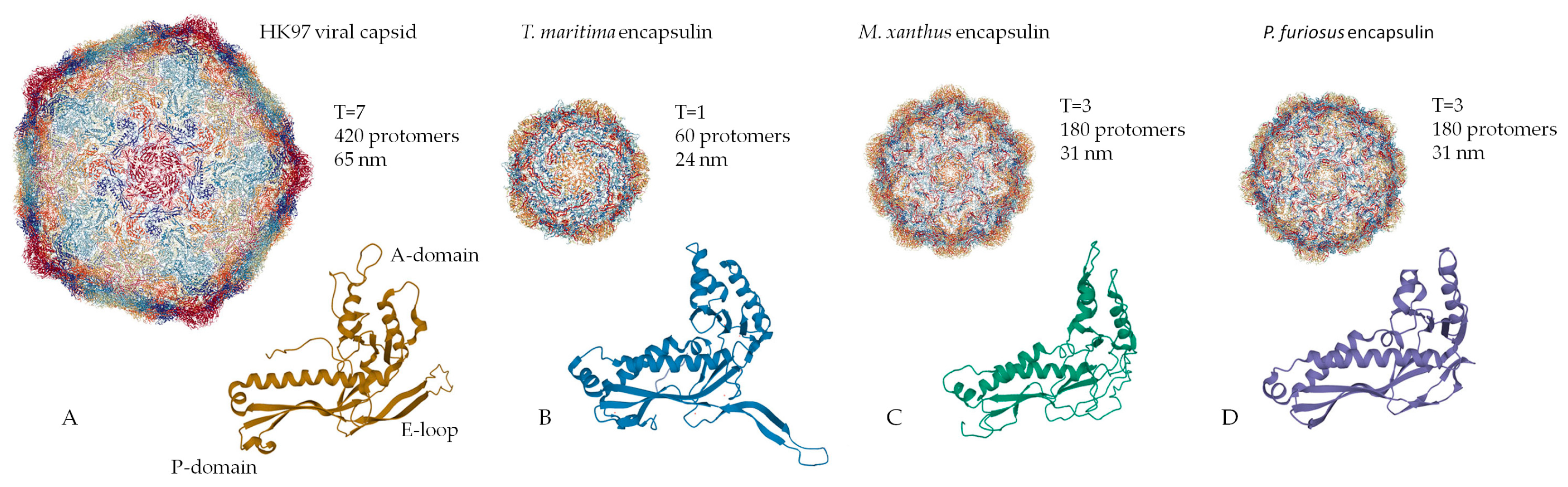
2. Structural Organization of the Shell
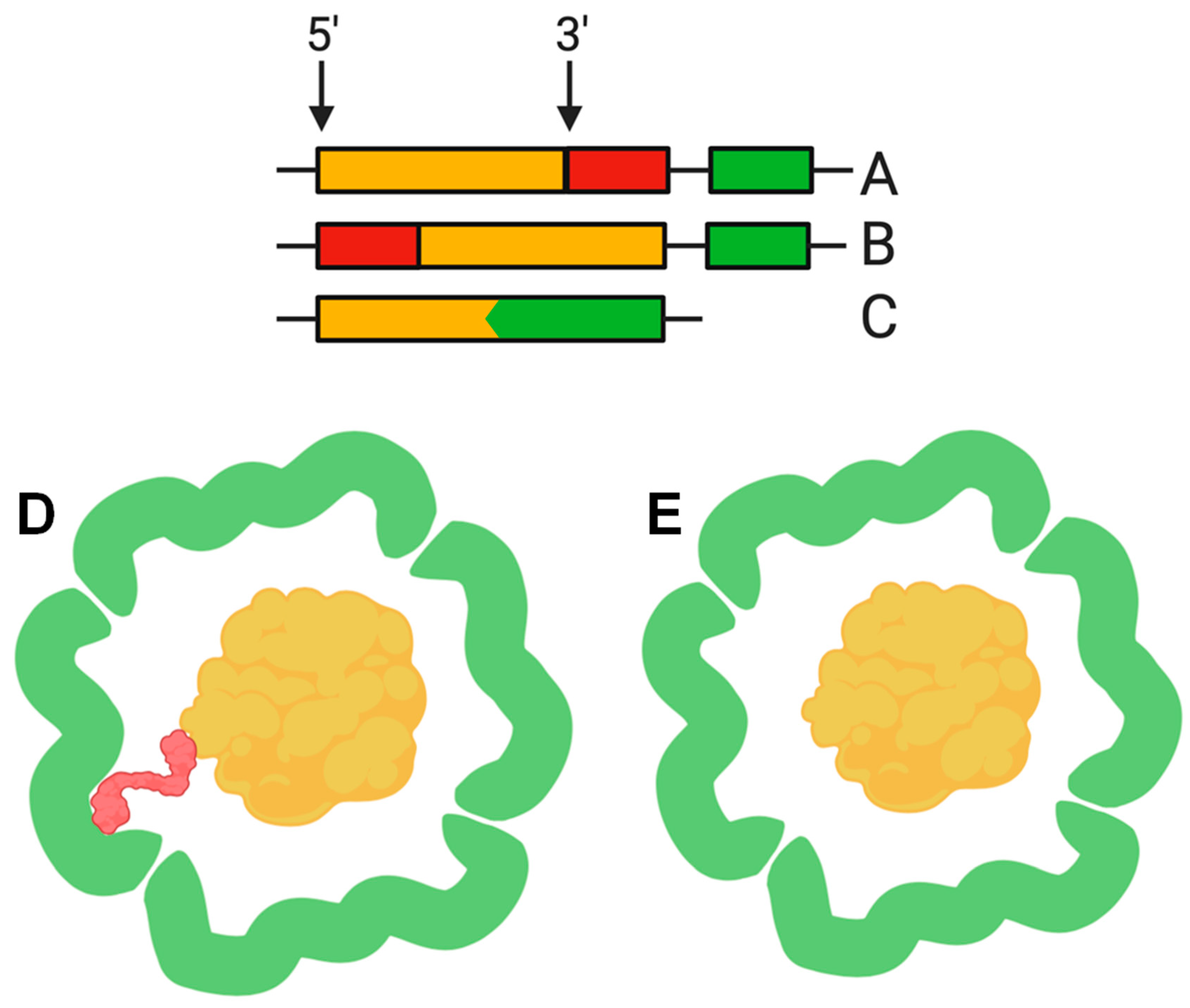
3. Cargo Proteins
4. Stability as the Main Property of Encapsulins
5. The Physiological Role
6. Nanocompartments and Nanotechnology
7. Nanocompartments as Cell and Tissue Markers
8. Conclusions
Funding
Conflicts of Interest
References
- Valdes-Stauber, N.; Scherer, S. Isolation and Characterization of Linocin M18, a Bacteriocin Produced by Brevibacterium Linens. Appl. Environ. Microbiol. 1994, 60, 3809–3814. [Google Scholar] [CrossRef] [PubMed]
- Sutter, M.; Boehringer, D.; Gutmann, S.; Günther, S.; Prangishvili, D.; Loessner, M.J.; Stetter, K.O.; Weber-Ban, E.; Ban, N. Structural Basis of Enzyme Encapsulation into a Bacterial Nanocompartment. Nat. Struct. Mol. Biol. 2008, 15, 939–947. [Google Scholar] [CrossRef] [PubMed]
- Giessen, T.W. Encapsulins: Microbial Nanocompartments with Applications in Biomedicine, Nanobiotechnology and Materials Science. Curr. Opin. Chem. Biol. 2016, 34, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Rosenkrands, I.; Rasmussen, P.B.; Carnio, M.; Jacobsen, S.; Theisen, M.; Andersen, P. Identification and Characterization of a 29-Kilodalton Protein from Mycobacterium Tuberculosis Culture Filtrate Recognized by Mouse Memory Effector Cells. Infect. Immun. 1998, 66, 2728–2735. [Google Scholar] [CrossRef] [PubMed]
- Hicks, P.M.; Rinker, K.D.; Baker, J.R.; Kelly, R.M. Homomultimeric Protease in the Hyperthermophilic Bacterium Thermotoga Maritima Has Structural and Amino Acid Sequence Homology to Bacteriocins in Mesophilic Bacteria. FEBS Lett. 1998, 440, 393–398. [Google Scholar] [CrossRef]
- Giessen, T.W.; Silver, P.A. Widespread Distribution of Encapsulin Nanocompartments Reveals Functional Diversity. Nat. Microbiol. 2017, 2, 17029. [Google Scholar] [CrossRef] [PubMed]
- Winter, N.; Triccas, J.A.; Rivoire, B.; Pessolani, M.C.V.; Eiglmeier, K.; Lim, E.-M.; Hunter, S.W.; Brennan, P.J.; Britton, W.J. Characterization of the Gene Encoding the Immunodominant 35 KDa Protein of Mycobacterium Leprae. Mol. Microbiol. 1995, 16, 865–876. [Google Scholar] [CrossRef]
- Triccas, J.A.; Roche, P.W.; Winter, N.; Feng, C.G.; Ruth Butlin, C.; Britton, W.J. A 35-Kilodalton Protein Is a Major Target of the Human Immune Response to Mycobacterium Leprae. Infect. Immun. 1996, 64, 5171–5177. [Google Scholar] [CrossRef]
- Kawamoto, S.; Watanabe, M.; Saito, N.; Hesketh, A.; Vachalova, K.; Matsubara, K.; Ochi, K. Molecular and Functional Analyses of the Gene (EshA) Encoding the 52-Kilodalton Protein of Streptomyces Coelicolor A3(2) Required for Antibiotic Production. J. Bacteriol. 2001, 183, 6009–6016. [Google Scholar] [CrossRef]
- Kwak, J.; McCue, L.A.; Trczianka, K.; Kendrick, K.E. Identification and Characterization of a Developmentally Regulated Protein, EshA, Required for Sporogenic Hyphal Branches in Streptomyces Griseus. J. Bacteriol. 2001, 183, 3004–3015. [Google Scholar] [CrossRef]
- Saito, N.; Matsubara, K.; Watanabe, M.; Kato, F.; Ochi, K. Genetic and Biochemical Characterization of EshA, a Protein That Forms Large Multimers and Affects Developmental Processes in Streptomyces Griseus. J. Biol. Chem. 2003, 278, 5902–5911. [Google Scholar] [CrossRef] [PubMed]
- Namba, K.; Hagiwara, K.; Tanaka, H.; Nakaishi, Y.; Chong, K.T.; Yamashita, E.; Armah, G.E.; Ono, Y.; Ishino, Y.; Omura, T.; et al. Expression and Molecular Characterization of Spherical Particles Derived from the Genome of the Hyperthermophilic Euryarchaeote Pyrococcus Furiosus. J. Biochem. 2005, 138, 193–199. [Google Scholar] [CrossRef] [PubMed]
- Akita, F.; Chong, K.T.; Tanaka, H.; Yamashita, E.; Miyazaki, N.; Nakaishi, Y.; Suzuki, M.; Namba, K.; Ono, Y.; Tsukihara, T.; et al. The Crystal Structure of a Virus-like Particle from the Hyperthermophilic Archaeon Pyrococcus Furiosus Provides Insight into the Evolution of Viruses. J. Mol. Biol. 2007, 368, 1469–1483. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.; Lee, J.; Min, J.; Kang, S. Developing Genetically Engineered Encapsulin Protein Cage Nanoparticles as a Targeted Delivery Nanoplatform. Biomacromolecules 2014, 15, 3794–3801. [Google Scholar] [CrossRef] [PubMed]
- Rurup, W.F.; Snijder, J.; Koay, M.S.T.; Heck, A.J.R.; Cornelissen, J.J.L.M. Self-Sorting of Foreign Proteins in a Bacterial Nanocompartment. J. Am. Chem. Soc. 2014, 136, 3828–3832. [Google Scholar] [CrossRef]
- Snijder, J.; Van De Waterbeemd, M.; Damoc, E.; Denisov, E.; Grinfeld, D.; Bennett, A.; Agbandje-Mckenna, M.; Makarov, A.; Heck, A.J.R. Defining the Stoichiometry and Cargo Load of Viral and Bacterial Nanoparticles by Orbitrap Mass Spectrometry. J. Am. Chem. Soc. 2014, 136, 7295–7299. [Google Scholar] [CrossRef]
- Tamura, A.; Fukutani, Y.; Takami, T.; Fujii, M.; Nakaguchi, Y.; Murakami, Y.; Noguchi, K.; Yohda, M.; Odaka, M. Packaging Guest Proteins into the Encapsulin Nanocompartment from Rhodococcus Erythropolis N771. Biotechnol. Bioeng. 2015, 112, 13–20. [Google Scholar] [CrossRef]
- Cassidy-Amstutz, C.; Oltrogge, L.; Going, C.C.; Lee, A.; Teng, P.; Quintanilla, D.; East-Seletsky, A.; Williams, E.R.; Savage, D.F. Identification of a Minimal Peptide Tag for in Vivo and in Vitro Loading of Encapsulin. Biochemistry 2016, 55, 3461–3468. [Google Scholar] [CrossRef]
- Choi, B.; Moon, H.; Hong, S.J.; Shin, C.; Do, Y.; Ryu, S.; Kang, S. Effective Delivery of Antigen-Encapsulin Nanoparticle Fusions to Dendritic Cells Leads to Antigen-Specific Cytotoxic T Cell Activation and Tumor Rejection. ACS Nano 2016, 10, 7339–7350. [Google Scholar] [CrossRef]
- Giessen, T.W.; Silver, P.A. Converting a Natural Protein Compartment into a Nanofactory for the Size-Constrained Synthesis of Antimicrobial Silver Nanoparticles. ACS Synth. Biol. 2016, 5, 1497–1504. [Google Scholar] [CrossRef]
- Sonotaki, S.; Takami, T.; Noguchi, K.; Odaka, M.; Yohda, M.; Murakami, Y. Successful PEGylation of Hollow Encapsulin Nanoparticles from: Rhodococcus Erythropolis N771 without Affecting Their Disassembly and Reassembly Properties. Biomater. Sci. 2017, 5, 1082–1089. [Google Scholar] [CrossRef] [PubMed]
- Rahmanpour, R.; Bugg, T.D.H. Assembly in Vitro of Rhodococcus Jostii RHA1 Encapsulin and Peroxidase DypB to Form a Nanocompartment. FEBS J. 2013, 280, 2097–2104. [Google Scholar] [CrossRef] [PubMed]
- Contreras, H.; Joens, M.S.; McMath, L.M.; Le, V.P.; Tullius, M.V.; Kimmey, J.M.; Bionghi, N.; Horwitz, M.A.; Fitzpatrick, J.A.J.; Goulding, C.W. Characterization of a Mycobacterium Tuberculosis Nanocompartment and Its Potential Cargo Proteins. J. Biol. Chem. 2014, 289, 18279–18289. [Google Scholar] [CrossRef] [PubMed]
- McHugh, C.A.; Fontana, J.; Nemecek, D.; Cheng, N.; Aksyuk, A.A.; Heymann, J.B.; Winkler, D.C.; Lam, A.S.; Wall, J.S.; Steven, A.C.; et al. A Virus Capsid-like Nanocompartment That Stores Iron and Protects Bacteria from Oxidative Stress. EMBO J. 2014, 33, 1896–1911. [Google Scholar] [CrossRef]
- He, D.; Hughes, S.; Vanden-Hehir, S.; Georgiev, A.; Altenbach, K.; Tarrant, E.; Mackay, C.L.; Waldron, K.J.; Clarke, D.J.; Marles-Wright, J. Structural Characterization of Encapsulated Ferritin Provides Insight into Iron Storage in Bacterial Nanocompartments. Elife 2016, 5, e18972. [Google Scholar] [CrossRef]
- Wikoff, W.R.; Liljas, L.; Duda, R.L.; Tsuruta, H.; Hendrix, R.W.; Johnson, J.E. Topologically Linked Protein Rings in the Bacteriophage HK97 Capsid. Science (80-) 2000, 289, 2129–2133. [Google Scholar] [CrossRef]
- Fokine, A.; Leiman, P.G.; Shneider, M.M.; Ahvazi, B.; Boeshans, K.M.; Steven, A.C.; Black, L.W.; Mesyanzhinov, V.V.; Rossmann, M.G. Structural and Functional Similarities between the Capsid Proteins of Bacteriophages T4 and HK97 Point to a Common Ancestry. Proc. Natl. Acad. Sci. USA 2005, 102, 7163–7168. [Google Scholar] [CrossRef]
- Baker, M.L.; Jiang, W.; Rixon, F.J.; Chiu, W. Common Ancestry of Herpesviruses and Tailed DNA Bacteriophages. J. Virol. 2005, 79, 14967–14970. [Google Scholar] [CrossRef]
- Pietilä, M.K.; Laurinmäki, P.; Russell, D.A.; Ko, C.C.; Jacobs-Sera, D.; Hendrix, R.W.; Bamford, D.H.; Butcher, S.J. Structure of the Archaeal Head-Tailed Virus HSTV-1 Completes the HK97 Fold Story. Proc. Natl. Acad. Sci. USA 2013, 110, 10604–10609. [Google Scholar] [CrossRef]
- CASPAR, D.L.; KLUG, A. Physical Principles in the Construction of Regular Viruses. Cold Spring Harb. Symp. Quant. Biol. 1962, 27, 1–24. [Google Scholar] [CrossRef]
- Giessen, T.W.; Orlando, B.J.; Verdegaal, A.A.; Chambers, M.G.; Gardener, J.; Bell, D.C.; Birrane, G.; Liao, M.; Silver, P.A. Large Protein Organelles Form a New Iron Sequestration System with High Storage Capacity. Elife 2019, 8, e46070. [Google Scholar] [CrossRef] [PubMed]
- Bamford, D.H.; Grimes, J.M.; Stuart, D.I. What Does Structure Tell Us about Virus Evolution? Curr. Opin. Struct. Biol. 2005, 15, 655–663. [Google Scholar] [CrossRef] [PubMed]
- Heinemann, J.; Maaty, W.S.; Gauss, G.H.; Akkaladevi, N.; Brumfield, S.K.; Rayaprolu, V.; Young, M.J.; Lawrence, C.M.; Bothner, B. Fossil Record of an Archaeal HK97-like Provirus. Virology 2011, 417, 362–368. [Google Scholar] [CrossRef] [PubMed]
- Williams, E.M.; Jung, S.M.; Coffman, J.L.; Lutz, S. Pore Engineering for Enhanced Mass Transport in Encapsulin Nanocompartments. ACS Synth. Biol. 2018, 7, 2514–2517. [Google Scholar] [CrossRef]
- Snijder, J.; Kononova, O.; Barbu, I.M.; Uetrecht, C.; Rurup, W.F.; Burnley, R.J.; Koay, M.S.T.; Cornelissen, J.J.L.M.; Roos, W.H.; Barsegov, V.; et al. Assembly and Mechanical Properties of the Cargo-Free and Cargo-Loaded Bacterial Nanocompartment Encapsulin. Biomacromolecules 2016, 17, 2522–2529. [Google Scholar] [CrossRef]
- Putri, R.M.; Allende-Ballestero, C.; Luque, D.; Klem, R.; Rousou, K.A.; Liu, A.; Traulsen, C.H.H.; Rurup, W.F.; Koay, M.S.T.; Castón, J.R.; et al. Structural Characterization of Native and Modified Encapsulins as Nanoplatforms for in Vitro Catalysis and Cellular Uptake. ACS Nano 2017, 11, 12796–12804. [Google Scholar] [CrossRef]
- Green, E.R.; Mecsas, J. Bacterial Secretion Systems: An Overview. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Wang, P.; Robert, L.; Pelletier, J.; Dang, W.L.; Taddei, F.; Wright, A.; Jun, S. Robust Growth of Escherichia Coli. Curr. Biol. 2010, 20, 1099–1103. [Google Scholar] [CrossRef]
- Imlay, J.A.; Chin, S.M.; Linn, S. Toxic DNA Damage by Hydrogen Peroxide through the Fenton Reaction in Vivo and in Vitro. Science (80-) 1988, 240, 640–642. [Google Scholar] [CrossRef]
- Andrews, S.C. Iron Storage in Bacteria. Adv. Microb. Physiol. 1998, 40, 281–351. [Google Scholar]
- Goulding, C.W.; Apostol, M.I.; Sawaya, M.R.; Phillips, M.; Parseghian, A.; Eisenberg, D. Regulation by Oligomerization in a Mycobacterial Folate Biosynthetic Enzyme. J. Mol. Biol. 2005, 349, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Sugano, Y.; Muramatsu, R.; Ichiyanagi, A.; Sato, T.; Shoda, M. DyP, a Unique Dye-Decolorizing Peroxidase, Represents a Novel Heme Peroxidase Family: ASP171 Replaces the Distal Histidine of Classical Peroxidases. J. Biol. Chem. 2007, 282, 36652–36658. [Google Scholar] [CrossRef] [PubMed]
- Pandey, R.; Rodriguez, G.M. A Ferritin Mutant of Mycobacterium Tuberculosis Is Highly Susceptible to Killing by Antibiotics and Is Unable to Establish a Chronic Infection in Mice. Infect. Immun. 2012, 80, 3650–3659. [Google Scholar] [CrossRef] [PubMed]
- Reddy, P.V.; Puri, R.V.; Khera, A.; Tyagi, A.K. Iron Storage Proteins Are Essential for the Survival and Pathogenesis of Mycobacterium Tuberculosis in THP-1 Macrophages and the Guinea Pig Model of Infection. J. Bacteriol. 2012, 194, 567–575. [Google Scholar] [CrossRef]
- Ahmad, M.; Roberts, J.N.; Hardiman, E.M.; Singh, R.; Eltis, L.D.; Bugg, T.D.H. Identification of DypB from Rhodococcus Jostii RHA1 as a Lignin Peroxidase. Biochemistry 2011, 50, 5096–5107. [Google Scholar] [CrossRef]
- Bobik, T.A.; Lehman, B.P.; Yeates, T.O. Bacterial Microcompartments: Widespread Prokaryotic Organelles for Isolation and Optimization of Metabolic Pathways. Mol. Microbiol. 2015, 98, 193–207. [Google Scholar] [CrossRef]
- Gong, J.; Chen, M.; Zheng, Y.; Wang, S.; Wang, Y. Polymeric Micelles Drug Delivery System in Oncology. J. Control. Release 2012, 159, 312–323. [Google Scholar] [CrossRef]
- Rösler, A.; Vandermeulen, G.W.M.; Klok, H.A. Advanced Drug Delivery Devices via Self-Assembly of Amphiphilic Block Copolymers. Adv. Drug Deliv. Rev. 2012, 64, 270–279. [Google Scholar] [CrossRef]
- Allen, T.M.; Cullis, P.R. Liposomal Drug Delivery Systems: From Concept to Clinical Applications. Adv. Drug Deliv. Rev. 2013, 65, 36–48. [Google Scholar] [CrossRef]
- Wang, A.Z.; Langer, R.; Farokhzad, O.C. Nanoparticle Delivery of Cancer Drugs. Annu. Rev. Med. 2012, 63, 185–198. [Google Scholar] [CrossRef]
- Haag, R.; Kratz, F. Polymer Therapeutics: Concepts and Applications. Angew. Chem. Int. Ed. 2006, 13, 1198–1215. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Nolte, R.J.M.; Cornelissen, J.J.L.M. Virus-Based Nanocarriers for Drug Delivery. Adv. Drug Deliv. Rev. 2012, 64, 811–825. [Google Scholar] [CrossRef] [PubMed]
- MaHam, A.; Tang, Z.; Wu, H.; Wang, J.; Lin, Y. Protein-Based Nanomedicine Platforms for Drug Delivery. Small 2009, 5, 1706–1721. [Google Scholar] [CrossRef] [PubMed]
- Brigger, I.; Dubernet, C.; Couvreur, P. Nanoparticles in Cancer Therapy and Diagnosis. Adv. Drug Deliv. Rev. 2002, 54, 631–651. [Google Scholar] [CrossRef]
- Toita, R.; Murata, M.; Tabata, S.; Abe, K.; Narahara, S.; Piao, J.S.; Kang, J.H.; Hashizume, M. Development of Human Hepatocellular Carcinoma Cell-Targeted Protein Cages. Bioconjug. Chem. 2012, 23, 1494–1501. [Google Scholar] [CrossRef]
- Toita, R.; Murata, M.; Abe, K.; Narahara, S.; Piao, J.S.; Kang, J.H.; Hashizume, M. A Nanocarrier Based on a Genetically Engineered Protein Cage to Deliver Doxorubicin to Human Hepatocellular Carcinoma Cells. Chem. Commun. 2013, 49, 7442–7444. [Google Scholar] [CrossRef]
- Putri, R.M.; Fredy, J.W.; Cornelissen, J.J.L.M.; Koay, M.S.T.; Katsonis, N. Labelling Bacterial Nanocages with Photo-Switchable Fluorophores. Chem. Phys. Chem. 2016, 1815–1818. [Google Scholar] [CrossRef]
- Klajn, R. Spiropyran-Based Dynamic Materials. Chem. Soc. Rev. 2014, 43, 148–184. [Google Scholar] [CrossRef]
- Fernández-Suárez, M.; Ting, A.Y. Fluorescent Probes for Super-Resolution Imaging in Living Cells. Nat. Rev. Mol. Cell Boil. 2008, 9, 929–943. [Google Scholar] [CrossRef]
- Fihey, A.; Perrier, A.; Browne, W.R.; Jacquemin, D. Multiphotochromic Molecular Systems. Chem. Soc. Rev. 2015, 44, 3719–3759. [Google Scholar] [CrossRef]
- Kortekaas, L.; Browne, W.R. The Evolution of Spiropyran: Fundamentals and Progress of an Extraordinarily Versatile Photochrome. Chem. Soc. Rev. 2019, 48, 3406–3424. [Google Scholar] [CrossRef] [PubMed]
- Moon, H.; Lee, J.; Kim, H.; Heo, S.; Min, J.; Kang, S. Genetically Engineering Encapsulin Protein Cage Nanoparticle as a SCC-7 Cell Targeting Optical Nanoprobe. Biomater. Res. 2014, 18, 21. [Google Scholar] [CrossRef] [PubMed]
- Jung, Y.; Kang, H.J.; Lee, J.M.; Jung, S.O.; Yun, W.S.; Chung, S.J.; Chung, B.H. Controlled Antibody Immobilization onto Immunoanalytical Platforms by Synthetic Peptide. Anal. Biochem. 2008, 374, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Lagoutte, P.; Mignon, C.; Stadthagen, G.; Potisopon, S.; Donnat, S.; Mast, J.; Lugari, A.; Werle, B. Simultaneous Surface Display and Cargo Loading of Encapsulin Nanocompartments and Their Use for Rational Vaccine Design. Vaccine 2018, 36, 3622–3628. [Google Scholar] [CrossRef]
- Cho, K.J.; Schepens, B.; Seok, J.H.; Kim, S.; Roose, K.; Lee, J.-H.; Gallardo, R.; Van Hamme, E.; Schymkowitz, J.; Rousseau, F.; et al. Structure of the Extracellular Domain of Matrix Protein 2 of Influenza A Virus in Complex with a Protective Monoclonal Antibody. J. Virol. 2015, 89, 3700–3711. [Google Scholar] [CrossRef]
- Maity, B.; Fujita, K.; Ueno, T. Use of the Confined Spaces of Apo-Ferritin and Virus Capsids as Nanoreactors for Catalytic Reactions. Curr. Opin. Chem. Boil. 2015, 25, 88–97. [Google Scholar] [CrossRef]
- Safont-Sempere, M.M.; Fernández, G.; Würthner, F. Self-Sorting Phenomena in Complex Supramolecular Systems. Chem. Rev. 2011, 111, 5784–5814. [Google Scholar] [CrossRef]
- Ai, H.W.; Henderson, J.N.; Remington, S.J.; Campbell, R.E. Directed Evolution of a Monomeric, Bright and Photostable Version of Clavularia Cyan Fluorescent Protein: Structural Characterization and Applications in Fluorescence Imaging. Biochem. J. 2006, 400, 531–540. [Google Scholar] [CrossRef]
- Künzle, M.; Mangler, J.; Lach, M.; Beck, T. Peptide-Directed Encapsulation of Inorganic Nanoparticles into Protein Containers. Nanoscale 2018, 10, 22917–22926. [Google Scholar] [CrossRef]
- Kim, T.; Momin, E.; Choi, J.; Yuan, K.; Zaidi, H.; Kim, J.; Park, M.; Lee, N.; McMahon, M.T.; Quinones-Hinojosa, A.; et al. Mesoporous Silica-Coated Hollow Manganese Oxide Nanoparticles as Positive T 1 Contrast Agents for Labeling and MRI Tracking of Adipose-Derived Mesenchymal Stem Cells. J. Am. Chem. Soc. 2011, 133, 2955–2961. [Google Scholar] [CrossRef]
- Di Corato, R.; Gazeau, F.; Le Visage, C.; Fayol, D.; Levitz, P.; Lux, F.; Letourneur, D.; Luciani, N.; Tillement, O.; Wilhelm, C. High-Resolution Cellular MRI: Gadolinium and Iron Oxide Nanoparticles for in-Depth Dual-Cell Imaging of Engineered Tissue Constructs. ACS Nano 2013, 7, 7500–7512. [Google Scholar] [CrossRef] [PubMed]
- Schrepfer, S.; Deuse, T.; Reichenspurner, H.; Fischbein, M.P.; Robbins, R.C.; Pelletier, M.P. Stem Cell Transplantation: The Lung Barrier. Transplant. Proc. 2007, 39, 573–576. [Google Scholar] [CrossRef] [PubMed]
- Bengel, F.M.; Schachinger, V.; Dimmeler, S. Cell-Based Therapies and Imaging in Cardiology. Eur. J. Nucl. Med. Mol. Imaging 2005, 32, 573–576. [Google Scholar] [CrossRef] [PubMed]
- Massoud, T.F.; Gambhir, S.S. Molecular Imaging in Living Subjects: Seeing Fundamental Biological Processes in a New Light. Genes Dev. 2003, 17, 545–580. [Google Scholar] [CrossRef]
- Gilad, A.A.; Winnard, P.T.; van Zijl, P.C.M.; Bulte, J.W.M. Developing MR Reporter Genes: Promises and Pitfalls. NMR Biomed. 2007, 20, 275–290. [Google Scholar] [CrossRef]
- Chasteen, N.D.; Harrison, P.M. Mineralization in Ferritin: An Efficient Means of Iron Storage. J. Struct. Biol. 1999, 126, 182–194. [Google Scholar] [CrossRef]
- Naumova, A.V.; Reinecke, H.; Yarnykh, V.; Deem, J.; Yuan, C.; Murry, C.E. Ferritin Overexpression for Noninvasive Magnetic Resonance Imaging-Based Tracking of Stem Cells Transplanted into the Heart. Mol. Imaging 2010, 9, 201–210. [Google Scholar] [CrossRef]
- Weissleder, R.; Simonova, M.; Bogdanova, A.; Bredow, S.; Enochs, W.S.; Bogdanov, A. MR Imaging and Scintigraphy of Gene Expression through Melanin Induction. Radiology 1997, 204, 425–429. [Google Scholar] [CrossRef]
- Sigmund, F.; Pettinger, S.; Kube, M.; Schneider, F.; Schifferer, M.; Schneider, S.; Efremova, M.V.; Pujol-Martí, J.; Aichler, M.; Walch, A.; et al. Iron-Sequestering Nanocompartments as Multiplexed Electron Microscopy Gene Reporters. ACS Nano 2019, 13, 8114–8123. [Google Scholar] [CrossRef]
- Sigmund, F.; Massner, C.; Erdmann, P.; Stelzl, A.; Rolbieski, H.; Desai, M.; Bricault, S.; Wörner, T.P.; Snijder, J.; Geerlof, A.; et al. Bacterial Encapsulins as Orthogonal Compartments for Mammalian Cell Engineering. Nat. Commun. 2018, 9, 1990. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gabashvili, A.N.; Chmelyuk, N.S.; Efremova, M.V.; Malinovskaya, J.A.; Semkina, A.S.; Abakumov, M.A. Encapsulins—Bacterial Protein Nanocompartments: Structure, Properties, and Application. Biomolecules 2020, 10, 966. https://doi.org/10.3390/biom10060966
Gabashvili AN, Chmelyuk NS, Efremova MV, Malinovskaya JA, Semkina AS, Abakumov MA. Encapsulins—Bacterial Protein Nanocompartments: Structure, Properties, and Application. Biomolecules. 2020; 10(6):966. https://doi.org/10.3390/biom10060966
Chicago/Turabian StyleGabashvili, Anna N., Nelly S. Chmelyuk, Maria V. Efremova, Julia A. Malinovskaya, Alevtina S. Semkina, and Maxim A. Abakumov. 2020. "Encapsulins—Bacterial Protein Nanocompartments: Structure, Properties, and Application" Biomolecules 10, no. 6: 966. https://doi.org/10.3390/biom10060966
APA StyleGabashvili, A. N., Chmelyuk, N. S., Efremova, M. V., Malinovskaya, J. A., Semkina, A. S., & Abakumov, M. A. (2020). Encapsulins—Bacterial Protein Nanocompartments: Structure, Properties, and Application. Biomolecules, 10(6), 966. https://doi.org/10.3390/biom10060966





