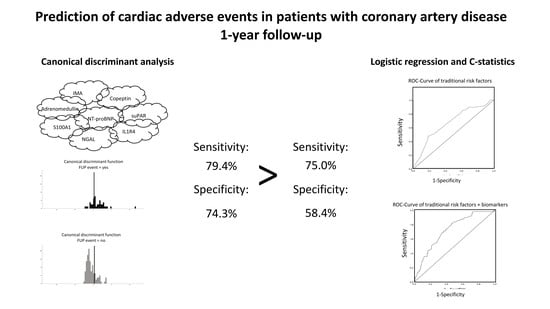
Multimarker Approach to Identify Patients with Coronary Artery Disease at High Risk for Subsequent Cardiac Adverse Events: The Multi-Biomarker Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Patient Population, Inclusion and Exclusion Criteria
2.3. Endpoints of the Study
2.4. Clinical Data Collection
2.5. Laboratory Procedures
2.6. Statistics
3. Results
3.1. Clinical Events during the 1-Year FUP
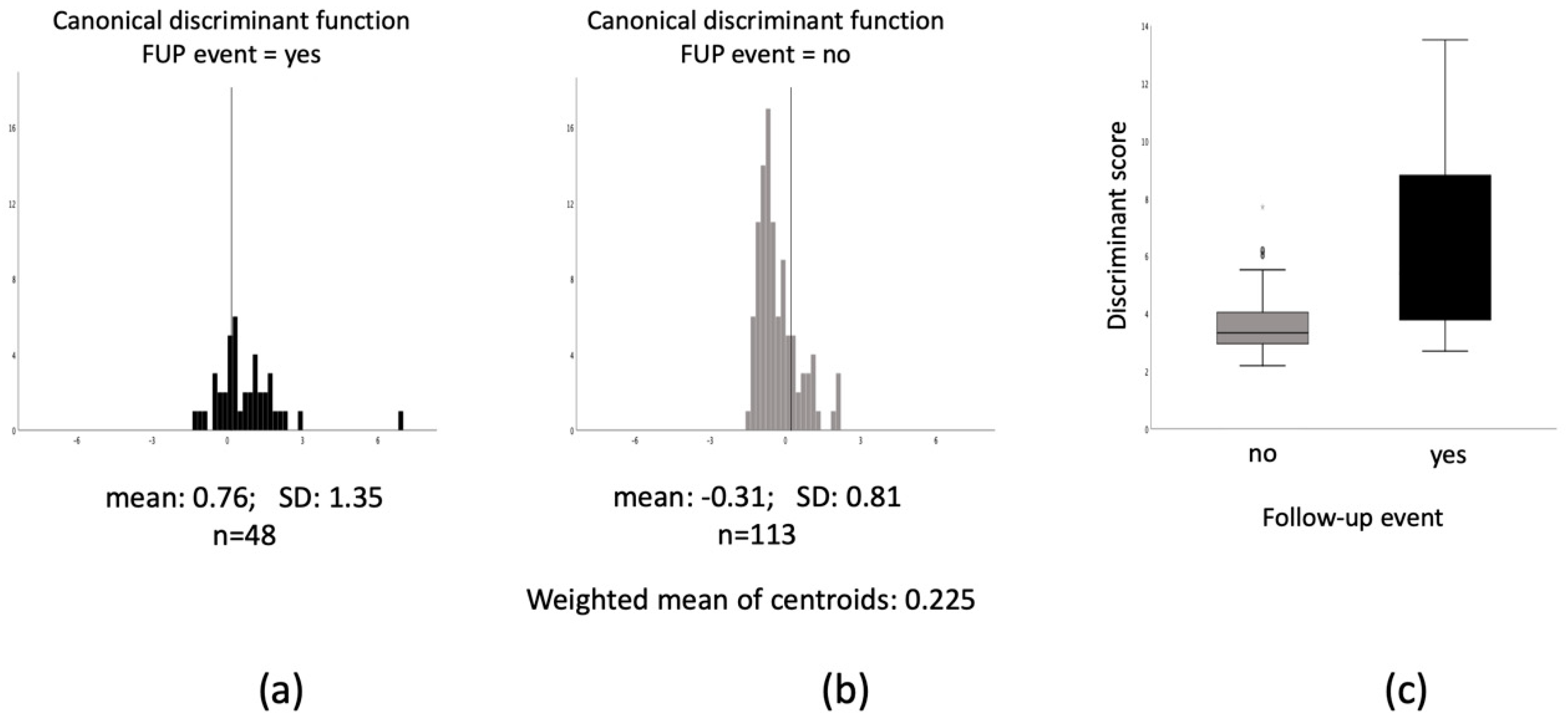
3.2. Multimarker Approach for Prediction of Adverse Events
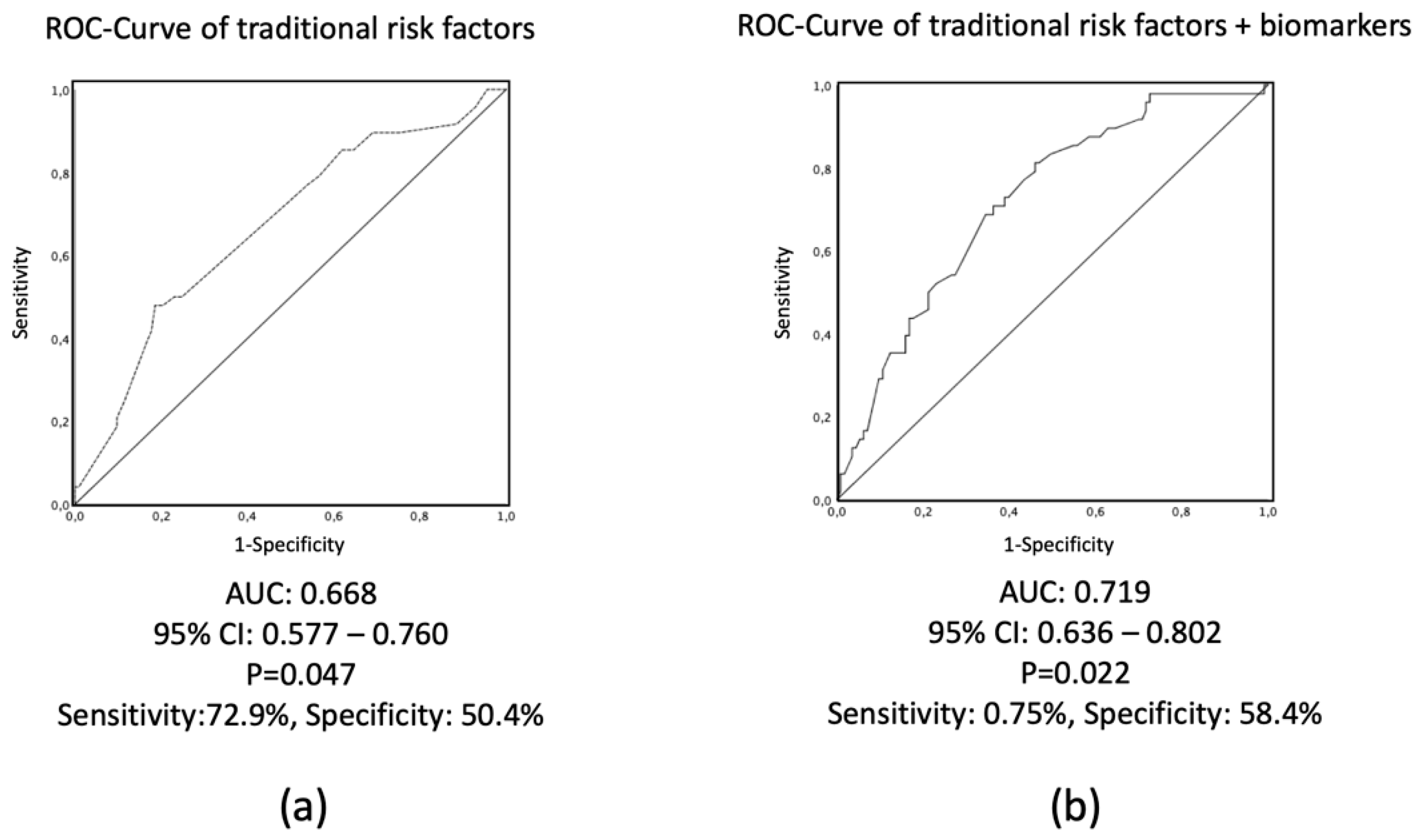
3.3. Risk Prediction Including Clinical Variables by Using Logistic Regression and C-Statistics
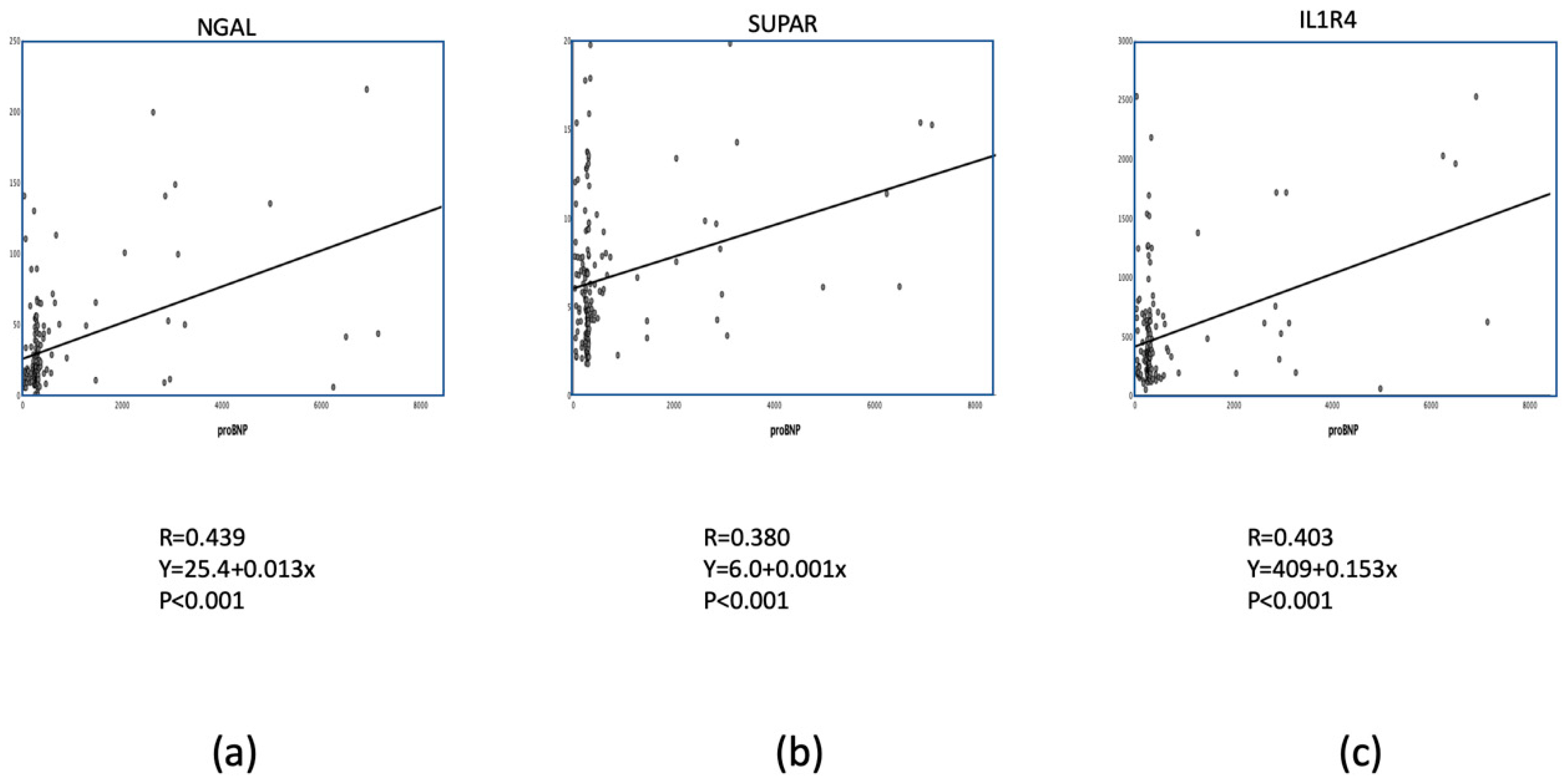
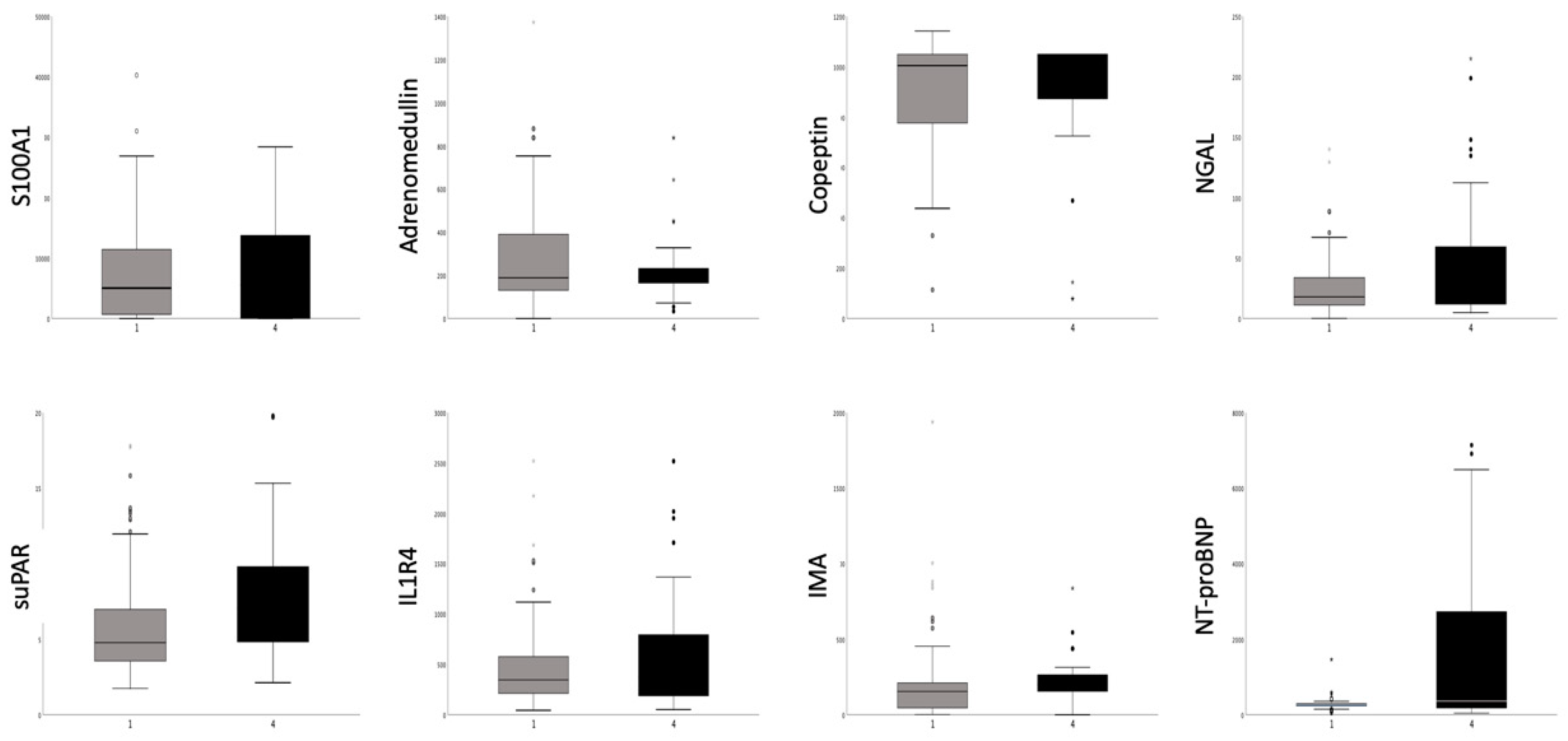
3.4. Diagnostic Value of Selected Biomarkers in Recent AMI
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Morrow, D.A. Cardiovascular risk prediction in patients with stable and unstable coronary heart disease. Circulation 2010, 121, 2681–2691. [Google Scholar] [CrossRef]
- Schnabel, R.B.; Sullivan, L.M.; Levy, D.; Pencina, M.J.; Massaro, J.M.; D’Agostino, R.B., Sr.; Newton-Cheh, C.; Yamamoto, J.F.; Magnani, J.W.; Tadros, T.M.; et al. Development of a risk score for atrial fibrillation (Framingham Heart Study): A community-based cohort study. Lancet 2009, 373, 739–745. [Google Scholar] [CrossRef]
- Goliasch, G.; Kleber, M.E.; Richter, B.; Plischke, M.; Hoke, M.; Haschemi, A.; Marculescu, R.; Endler, G.; Grammer, T.B.; Pilz, S.; et al. Routinely available biomarkers improve prediction of long-term mortality in stable coronary artery disease: The Vienna and Ludwigshafen Coronary Artery Disease (VILCAD) risk score. Eur. Heart J. 2012, 33, 2282–2289. [Google Scholar] [CrossRef] [PubMed]
- Conroy, R.M.; Pyörälä, K.; Fitzgerald, A.P.; Sans, S.; Menotti, A.; De Backer, G.; De Bacquer, D.; Ducimetière, P.; Jousilahti, P.; Keil, U.; et al. SCORE project group. Estimation of ten-year risk of fatal cardiovascular disease in Europe: The SCORE project. Eur. Heart J. 2003, 24, 987–1003. [Google Scholar] [CrossRef]
- Meeuwsen, J.A.L.; Wesseling, M.; Hoefer, I.E.; de Jager, S.C.A. Prognostic value of circulating inflammatory cells in patients with stable and acute coronary artery disease. Front. Cardiovasc. Med. 2017, 4, 44. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.H. Contemporary challenges in translating biomarker evidence into clinical practice. J. Am. Coll. Cardiol. 2010, 55, 2077–2079. [Google Scholar] [CrossRef][Green Version]
- Reichlin, T.; Hochholzer, W.; Stelzig, C.; Laule, K.; Freidank, H.; Morgenthaler, N.G.; Bergmann, A.; Potocki, M.; Noveanu, M.; Breidthardt, T.; et al. Incremental value of copeptin for rapid rule out of acute myocardial infarction. J. Am. Coll. Cardiol. 2009, 54, 60–68. [Google Scholar] [CrossRef]
- Nakada, Y.; Kawakami, R.; Matsui, M.; Ueda, T.; Nakano, T.; Takitsume, A.; Nakagawa, H.; Nishida, T.; Onoue, K.; Soeda, T.; et al. Prognostic value of urinary neutrophil gelatinase-associated lipocalin on the first day of admission for adverse events in patients with acute decompensated heart failure. J. Am. Heart Assoc. 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Sehestedt, T.; Lyngbaek, S.; Eugen-Olsen, J.; Jeppesen, J.; Andersen, O.; Hansen, T.W.; Linneberg, A.; Jørgensen, T.; Haugaard, S.B.; Olsen, M.H.; et al. Soluble urokinase plasminogen activator receptor is associated with subclinical organ damage and cardiovascular events. Atherosclerosis 2011, 216, 237–243. [Google Scholar] [CrossRef]
- Wang, T.J. Multiple biomarkers for predicting cardiovascular events: Lessons learned. J. Am. Coll. Cardiol. 2010, 55, 2092–2095. [Google Scholar] [CrossRef][Green Version]
- Mori, Y.; Kosaki, A.; Kishimoto, N.; Kimura, T.; Iida, K.; Fukui, M.; Nakajima, F.; Nagahara, M.; Urakami, M.; Iwasaka, T.; et al. Increased plasma S100A12 (EN-RAGE) levels in hemodialysis patients with atherosclerosis. Am. J. Nephrol. 2009, 29, 18–24. [Google Scholar] [CrossRef] [PubMed]
- Shiotsu, Y.; Mori, Y.; Nishimura, M.; Sakoda, C.; Tokoro, T.; Hatta, T.; Maki, N.; Iida, K.; Iwamoto, N.; Ono, T.; et al. Plasma S100A12 level is associated with cardiovascular disease in hemodialysis patients. Clin. J. Am. Soc. Nephrol. 2011, 6, 718–723. [Google Scholar] [CrossRef]
- Khan, S.Q.; O’Brien, R.J.; Struck, J.; Quinn, P.; Morgenthaler, N.; Squire, I.; Davies, J.; Bergmann, A.; Ng, L.L. Prognostic value of midregional pro-adrenomedullin in patients with acute myocardial infarction: The LAMP (Leicester Acute Myocardial Infarction Peptide) study. J. Am. Coll. Cardiol. 2007, 49, 1525–1532. [Google Scholar] [CrossRef]
- Wild, P.S.; Schnabel, R.B.; Lubos, E.; Zeller, T.; Sinning, C.R.; Keller, T.; Tzikas, S.; Lackner, K.J.; Peetz, D.; Rupprecht, H.J.; et al. Midregional proadrenomedullin for prediction of cardiovascular events in coronary artery disease: Results from the AtheroGene study. Clin. Chem. 2012, 58, 226–236. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Keller, T.; Tzikas, S.; Zeller, T.; Czyz, E.; Lillpopp, L.; Ojeda, F.M.; Roth, A.; Bickel, C.; Baldus, S.; Sinning, C.R.; et al. Copeptin improves early diagnosis of acute myocardial infarction. J. Am. Coll. Cardiol. 2010, 55, 2096–2106. [Google Scholar] [CrossRef] [PubMed]
- Yndestad, A.; Landro, L.; Ueland, T.; Dahl, C.P.; Flo, T.H.; Vinge, L.E.; Espevik, T.; Frøland, S.S.; Husberg, C.; Christensen, G.; et al. Increased systemic and myocardial expression of neutrophil gelatinase-associated lipocalin in clinical and experimental heart failure. Eur. Heart J. 2009, 30, 1229–1236. [Google Scholar] [CrossRef] [PubMed]
- Eugen-Olsen, J.; Andersen, O.; Linneberg, A.; Ladelund, S.; Hansen, T.W.; Langkilde, A.; Petersen, J.; Pielak, T.; Møller, L.N.; Jeppesen, J.; et al. Circulating soluble urokinase plasminogen activator receptor predicts cancer, cardiovascular disease, diabetes and mortality in the general population. J. Intern. Med. 2010, 268, 296–308. [Google Scholar] [CrossRef]
- Koller, L.; Stojkovic, S.; Richter, B.; Sulzgruber, P.; Potolidis, C.; Liebhart, F.; Mörtl, D.; Berger, R.; Goliasch, G.; Wojta, J.; et al. Soluble urokinase-type plasminogen activator receptor improves risk prediction in patients with chronic heart failure. JACC Heart Fail. 2017, 5, 268–277. [Google Scholar] [CrossRef]
- Dieplinger, B.; Mueller, T. Soluble ST2 in heart failure. ClinChimActa 2015, 443, 57–70. [Google Scholar] [CrossRef]
- Sabatine, M.S.; Morrow, D.A.; Higgins, L.J.; MacGillivray, C.; Guo, W.; Bode, C.; Rifai, N.; Cannon, C.P.; Gerszten, R.E.; Lee, R.T.; et al. Complementary roles for biomarkers of biomechanical strain ST2 and N-terminal prohormone B-type natriuretic peptide in patients with ST-elevation myocardial infarction. Circulation 2008, 117, 1936–1944. [Google Scholar] [CrossRef] [PubMed]
- Dhillon, O.S.; Narayan, H.K.; Quinn, P.A.; Squire, I.B.; Davies, J.E.; Ng, L.L. Interleukin 33 and ST2 in non-ST-elevation myocardial infarction: Comparison with Global Registry of Acute Coronary Events Risk Scoring and NT-proBNP. Am. Heart J. 2011, 161, 1163–1170. [Google Scholar] [CrossRef] [PubMed]
- Eggers, K.M.; Armstrong, P.W.; Califf, R.M.; Simoons, M.L.; Venge, P.; Wallentin, L.; James, S.K. ST2 and mortality in non-ST-segment elevation acute coronary syndrome. Am. Heart J. 2010, 159, 788–794. [Google Scholar] [CrossRef]
- Dieplinger, B.; Egger, M.; Haltmayer, M.; Kleber, M.E.; Scharnagl, H.; Silbernagel, G.; de Boer, R.A.; Maerz, W.; Mueller, T. Increased soluble ST2 predicts long-term mortality in patients with stable coronary artery disease: Results from the Ludwigshafen risk and cardiovascular health study. Clin. Chem. 2014, 60, 530–540. [Google Scholar] [CrossRef] [PubMed]
- Turedi, S.; Gunduz, A.; Mentese, A.; Dasdibi, B.; Karahan, S.C.; Sahin, A.; Tuten, G.; Kopuz, M.; Alver, A. Investigation of the possibility of using ischemia-modified albumin as a novel and early prognostic marker in cardiac arrest patients after cardiopulmonary resuscitation. Resuscitation 2009, 80, 994–999. [Google Scholar] [CrossRef] [PubMed]
- Kanko, M.; Yavuz, S.; Duman, C.; Hosten, T.; Oner, E.; Berki, T. Ischemia-modified albumin use as a prognostic factor in coronary bypass surgery. J. Cardiothorac. Surg. 2012, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.C.; Greenland, P.; Rossouw, J.E.; Manson, J.E.; Cochrane, B.B.; Lasser, N.L.; Limacher, M.C.; Lloyd-Jones, D.M.; Margolis, K.L.; Robinson, J.G. Multimarker prediction of coronary heart disease risk. J. Am. Coll. Cardiol. 2010, 55, 2080–2091. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.J.; Gona, P.; Larson, M.G.; Tofler, G.H.; Levy, D.; Newton-Cheh, C.; Jacques, P.F.; Rifai, N.; Selhub, J.; Robins, S.J.; et al. Multiple biomarkers for the prediction of first major cardiovascular events and death. N. Engl. J. Med. 2006, 355, 2631–2639. [Google Scholar] [CrossRef] [PubMed]
- Uno, H.; Cai, T.; Pencina, M.J.v.; D’Agostino, R.B.; Wei, L.J. On the C-statistics for evaluating overall adequacy of risk prediction procedures with censored survival data. Stat. Med. 2011, 30, 1105–1117. [Google Scholar] [CrossRef]
- Baranowska-Bik, A.; Kochanowski, J.; Uchman, D.; Litwiniuk, A.; Kalisz, M.; Martynska, L.; Wolinska-Witort, E.; Baranowska, B.; Bik, W. Association of copeptin and cortisol in newly diagnosed multiple sclerosis patients. J. Neuroimmunol. 2015, 282, 21–24. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Puerta, J.A.; Ortiz-Reyes, B.; Urrego, T.; Vanegas-García, A.L.; Muñoz, C.H.; González, L.A.; Cervera, R.; Vásquez, G. Urinary neutrophil gelatinase-associated lipocalin and monocyte chemoattractant protein 1 as biomarkers for lupus nephritis in Colombian SLE patients. Lupus 2018, 27, 637–646. [Google Scholar] [CrossRef]
| Clinical and Laboratory Parameter | All Patients n = 161 | Group AE n = 48 | Group Non-AE n = 113 | p Value between Groups |
|---|---|---|---|---|
| Male gender n (%) | 126 (78.3%) | 35 (72.9%) | 91 (80.5%) | 0.301 |
| Age (y; mean ± SD) | 67.0 ± 11.9 | 70.4 ± 11.1 | 65.6 ± 12.0 | 0.017 |
| Diabetes mellitus n (%) | 41 (25.5%) | 18 (37.5%) | 23 (20.4%) | 0.030 |
| Hypertension n (%) | 129 (80.1%) | 41 (85.4%) | 88 (77.9%) | 0.388 |
| Hyperlipidemia n (%) | 117 (72.7%) | 37 (77.1%) | 80 (70.8%) | 0.447 |
| Smoking n (%) | 58 (36.0%) | 26 (55.3%) | 32 (28.3%) | 0.011 |
| Previous MI n (%) | 83 (51.6%) | 29 (60.4%) | 54 (47.8%) | 0.295 |
| Previous PCI n (%) | 85 (52.8%) | 22 (45.8%) | 63 (55.8%) | 0.301 |
| Previous CABG n (%) | 31 (19.3%) | 17 (35.4%) | 14 (12.4%) | 0.002 |
| PAD n (%) | 20 (12.4%) | 9 (18.8%) | 11 (9.7%) | 0.123 |
| Carotid artery disease n (%) | 28 (17.4%) | 9 (18.8%) | 19 (17.0%) | 0.822 |
| Aspirin (%) | 161 (100%) | 48 (100%) | 113 (100%) | 1 |
| Clopidogrel n (%) | 132 (82.0%) | 38 (79.2%) | 94 (83.2%) | 0.408 |
| Beta-blocker n (%) | 152 (94.4%) | 43 (89.6%) | 109 (96.4%) | 0.398 |
| ACE-inhibitor/ARB n (%) | 146 (90.7%) | 42 (87.5%) | 104 (92.0%) | 0.780 |
| NT-proBNP (median; Q) (pg/mL) | 280 (232; 335) | 282 (243; 507) | 279 (232; 309) | 0.224 |
| Troponin T (ng/L) (median; Q) (ng/mL) | 0.01 (0.01; 0.41) | 0.01 (0.01; 0.04) | 0.01 (0.01; 0.08) | 0.313 |
| Creatine kinase (U/L) (median; Q) | 116 (66; 188) | 88 (67; 177) | 118 (66; 106) | 0.541 |
| Baseline WMSI (mean ± SD) | 1.31 ± 0.46 | 1.22 ± 0.45 | 1.34 ± 0.47 | 0.115 |
| Follow-up WMSI (mean ± SD) | 1.31 ± 0.51 | 1.28 ± 0.51 | 1.32 ± 0.51 | 0.651 |
| Selected biomarkers (Median, 25% and 75% Quartiles) | ||||
| S100A1(ng/mL) | 5504 (0; 11858) | 5311 (0; 14462) | 5674 (0; 11327) | 0.724 |
| Adrenomedullin (ng/mL) | 189 (146; 328) | 178 (144; 279) | 194 (150; 343) | 0.187 |
| Copeptin (pg/mL) | 1049 (817; 1049) | 938 (731; 1049) | 1049 (844; 1049) | 0.058 |
| NGAL (pg/mL) | 18.5 (11.0; 39.0) | 40.3 (16.5; 57.7) | 16.2 (9.9; 27.6) | <0.05 |
| suPAR (ng/mL) | 5.37 (3.70; 8.51) | 9.18 (3.58; 11.99) | 4.94 (3.70; 6.95) | <0.05 |
| IL1R4 (pg/mL) | 347 (211; 616) | 542 (222; 769) | 318 (193; 548) | <0.05 |
| IMA (ng/mL) | 167 (53; 219) | 156 (34; 211) | 178 (76; 221) | 0.120 |
| Clinical and Laboratory Parameter | Subgroup CAD n = 111 | Subgroup AMI n = 50 | p Value between the Groups |
|---|---|---|---|
| Male gender n (%) | 89 (80.2%) | 37 (74.0%) | 0.412 |
| Age (y; mean ± SD) | 68.4 ± 11.5 | 64.0 ± 12.3 | 0.036 |
| Diabetes mellitus n (%) | 34 (30.6%) | 7 (14.0%) | 0.031 |
| Hypertension n (%) | 92 (82.9%) | 37 (74.0%) | 0.205 |
| Hyperlipidemia n (%) | 77 (69.4%) | 40 (80.0%) | 0.185 |
| Smoking n (%) | 34 (30.6%) | 24 (48.0%) | 0.167 |
| Previous MI n (%) | 71 (64.0%) | 12 (24.0%) | <0.001 |
| Previous PCI n (%) | 74 (66.7%) | 11 (22.0%) | <0.001 |
| Previous CABG n (%) | 25 (22.5%) | 6 (12.0%) | 0.135 |
| PAD n (%) | 15 (13.5%) | 5 (10.0%) | 0.614 |
| Carotid artery disease n (%) | 23 (20.7%) | 5 (10%) | 0.117 |
| Aspirin n (%) | 111 (100%) | 50 (100%) | 1 |
| Clopidogrel n (%) | 92 (82.9%) | 40 (80.0%) | 0.666 |
| Beta-blocker n (%) | 102 (91.9%) | 50 (100%) | 0.863 |
| ACE-inhibitor/ARB n (%) | 102 (91.9%) | 44 (88.0%) | 0.814 |
| NT-proBNP (median; Q) (pg/mL) | 278 (241; 301) | 359 (182; 881) | 0.029 |
| Troponin T (ng/L) (median; Q) (ng/mL) | 0.01 (0.01; 0.01) | 0.26 (0.04; 2.33) | <0.001 |
| Creatine kinase (U/L) (median; Q) | 85 (55; 137) | 248 (131; 569) | <0.001 |
| Baseline WMSI (mean ± SD) | 1.29 ± 0.43 | 1.34 ± 0.54 | 0.611 |
| Follow-up WMSI (mean ± SD) | 1.29 ± 0.46 | 1.34 ± 0.64 | 0.716 |
| FUP events n (%) | 30 (27.0%) | 18 (36.0%) | 0.268 |
| Hospitalization n (%) | 16 (14.4%) | 7 (14.0%) | |
| Acute MI n (%) | 6 (5.4%) | 2 (4.0%) | |
| Revascularization n (%) | 6 (5.4%) | 7 (14%) | |
| Death n (%) | 2 (1.8%) | 2 (4.0%) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Giurgea, G.-A.; Zlabinger, K.; Gugerell, A.; Lukovic, D.; Syeda, B.; Mandic, L.; Pavo, N.; Mester-Tonczar, J.; Traxler-Weidenauer, D.; Spannbauer, A.; et al. Multimarker Approach to Identify Patients with Coronary Artery Disease at High Risk for Subsequent Cardiac Adverse Events: The Multi-Biomarker Study. Biomolecules 2020, 10, 909. https://doi.org/10.3390/biom10060909
Giurgea G-A, Zlabinger K, Gugerell A, Lukovic D, Syeda B, Mandic L, Pavo N, Mester-Tonczar J, Traxler-Weidenauer D, Spannbauer A, et al. Multimarker Approach to Identify Patients with Coronary Artery Disease at High Risk for Subsequent Cardiac Adverse Events: The Multi-Biomarker Study. Biomolecules. 2020; 10(6):909. https://doi.org/10.3390/biom10060909
Chicago/Turabian StyleGiurgea, Georgiana-Aura, Katrin Zlabinger, Alfred Gugerell, Dominika Lukovic, Bonni Syeda, Ljubica Mandic, Noemi Pavo, Julia Mester-Tonczar, Denise Traxler-Weidenauer, Andreas Spannbauer, and et al. 2020. "Multimarker Approach to Identify Patients with Coronary Artery Disease at High Risk for Subsequent Cardiac Adverse Events: The Multi-Biomarker Study" Biomolecules 10, no. 6: 909. https://doi.org/10.3390/biom10060909
APA StyleGiurgea, G.-A., Zlabinger, K., Gugerell, A., Lukovic, D., Syeda, B., Mandic, L., Pavo, N., Mester-Tonczar, J., Traxler-Weidenauer, D., Spannbauer, A., Kastner, N., Müller, C., Anvari, A., Bergler-Klein, J., & Gyöngyösi, M. (2020). Multimarker Approach to Identify Patients with Coronary Artery Disease at High Risk for Subsequent Cardiac Adverse Events: The Multi-Biomarker Study. Biomolecules, 10(6), 909. https://doi.org/10.3390/biom10060909