Production of a New Cyclic Depsipeptide by the Culture Broth of Staphylococcus sp. Isolated from Corallina officinalis L.
Abstract
1. Introduction
2. Results and Discussion
2.1. Isolation Method of Compounds 1–7
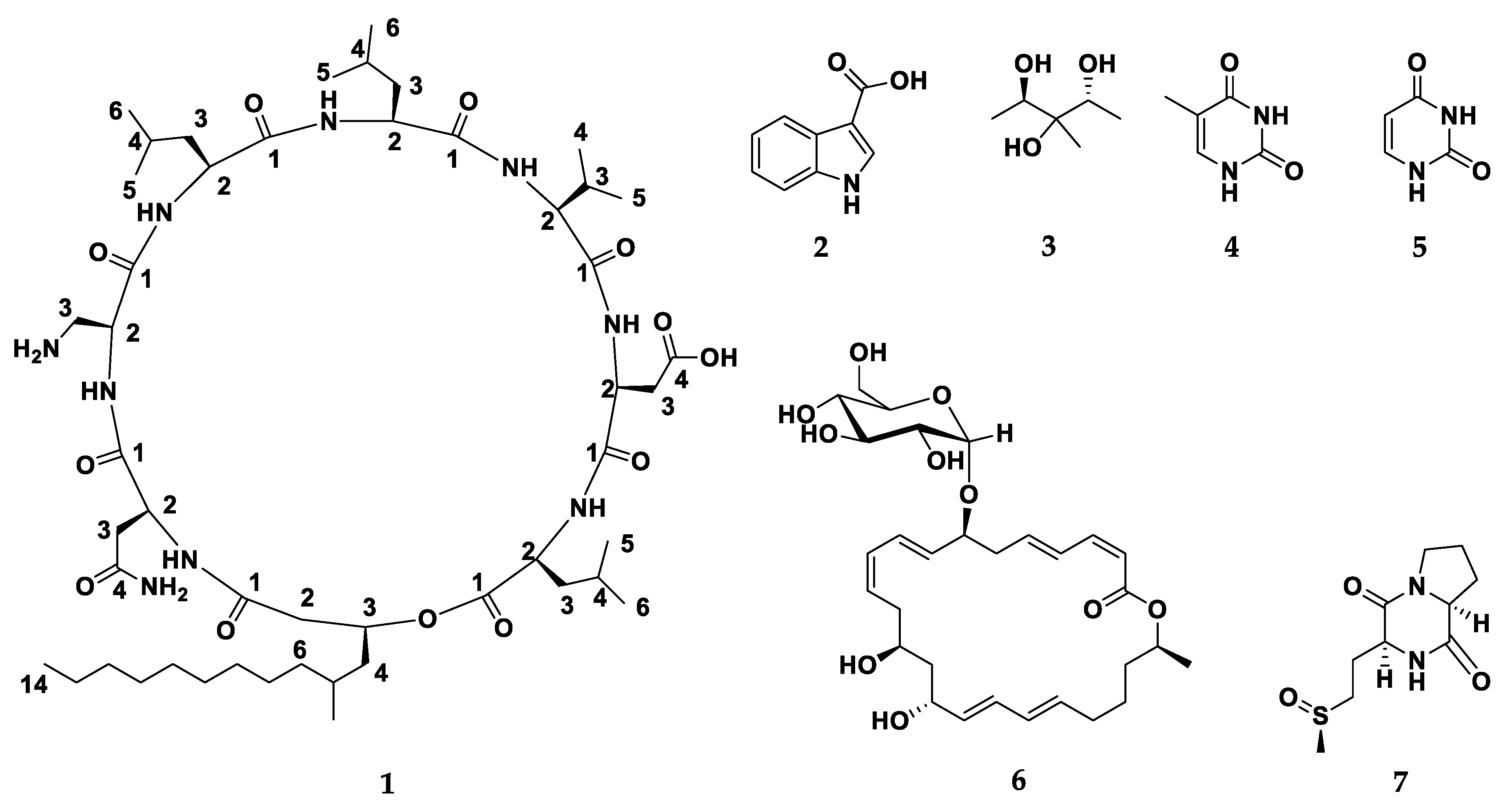
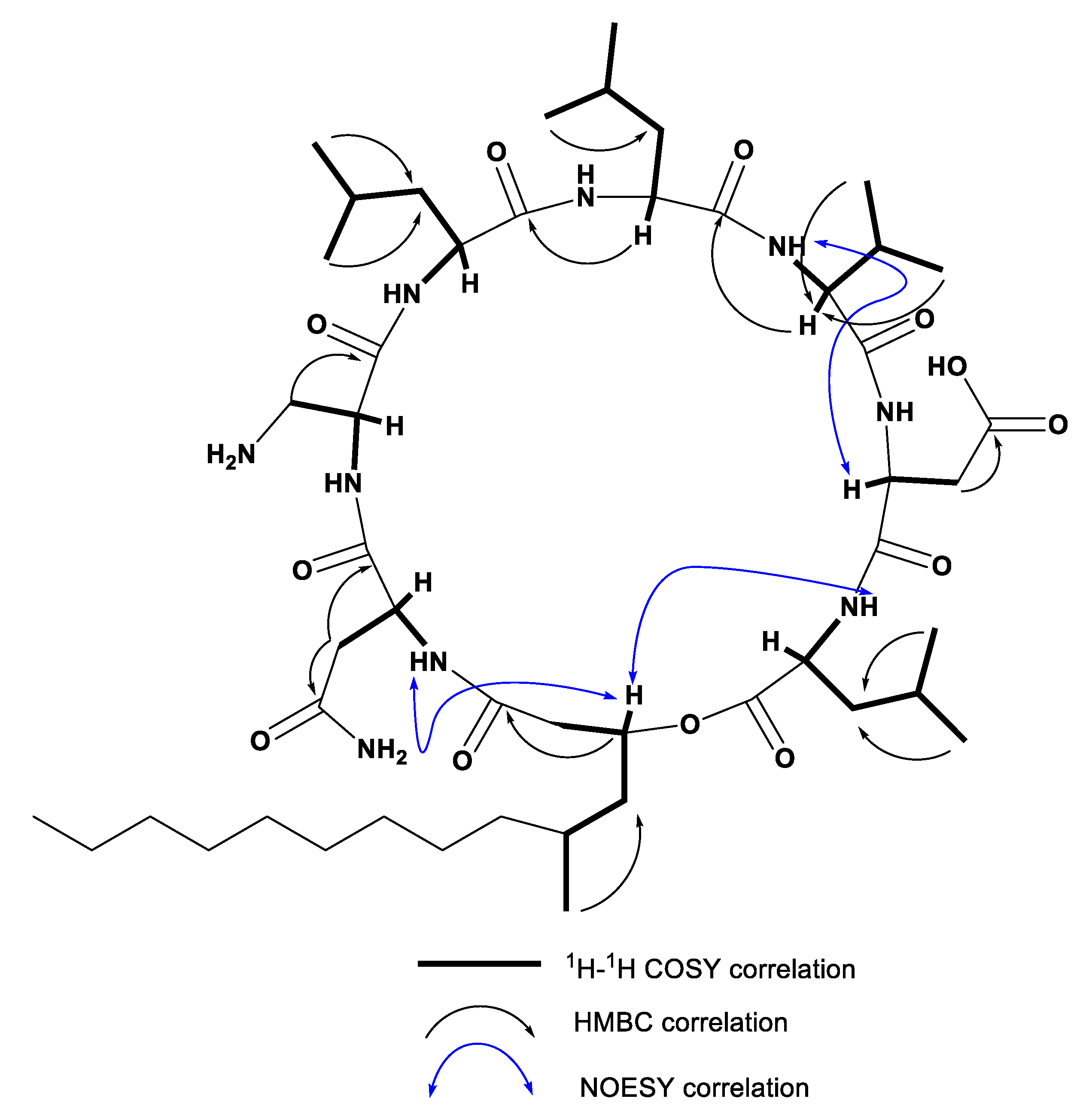
2.2. Characterization of Compounds 1–7 Structures
2.3. Biological Activities of the Pure Compounds 1–7
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Biological Materials
3.3. Purification of Compounds 1–7
3.4. Configuration of Amino Acids
3.5. Antimicrobial Activity of Compounds 1–7
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Newman, D.J.; Cragg, G.M.; Snader, K.M. Natural products as sources of new drugs over the period 1981–2002. J. Nat. Prod. 2003, 66, 1022–1037. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335. [Google Scholar] [CrossRef] [PubMed]
- Harvey, A.L. Natural products in drug discovery. Drug Discov. Today 2008, 13, 894–901. [Google Scholar] [CrossRef] [PubMed]
- Fenical, W. Chemical studies of marine bacteria: Developing a new resource. Chem. Rev. 1993, 93, 1673–1683. [Google Scholar] [CrossRef]
- Blunt, J.W.; Carroll, A.R.; Copp, B.R.; Davis, R.A.; Keyzers, R.A.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2018, 35, 8–53. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2010, 27, 165–237. [Google Scholar] [CrossRef] [PubMed]
- Faulkner, D.J. Marine natural products. Nat. Prod. Rep. 2002, 19, 1–48. [Google Scholar] [PubMed]
- Fenical, W.; Jensen, P.R. Marine microorganisms: A new biomedical resource. In Pharmaceutical and Bioactive Natural Products; Springer: Berlin/Heidelberg, Germany, 1993; pp. 419–457. [Google Scholar]
- Lu, X.; Cao, X.; Liu, X.; Jiao, B. Marine microbes-derived anti-bacterial agents. Mini Rev. Med. Chem. 2010, 10, 1077–1090. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Keyzers, R.A.; Munro, M.H.G.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2017, 34, 235–294. [Google Scholar] [CrossRef] [PubMed]
- Andryukov, B.G.; Mikhaylov, V.V.; Besednova, N.N.; Zaporozhets, T.S.; Bynina, M.P.; Matosova, E.V. The Bacteriocinogenic Potential of Marine Microorganisms. Russ. J. Mar. Biol. 2018, 44, 433–441. [Google Scholar] [CrossRef]
- Chen, E.; Chen, Q.; Chen, S.; Xu, B.; Ju, J.; Wang, H. Mathermycin, a Lantibiotic from the Marine Actinomycete Marinactinospora thermotolerans SCSIO 00652. Appl. Environ. Microbiol. 2017, 83, e00926-17. [Google Scholar] [CrossRef] [PubMed]
- Andryukov, B.; Mikhailov, V.; Besednova, N. The Biotechnological Potential of Secondary Metabolites from Marine Bacteria. J. Mar. Sci. Eng. 2019, 7, 176. [Google Scholar] [CrossRef]
- Bohringer, N.; Fisch, K.M.; Schillo, D.; Bara, R.; Hertzer, C.; Grein, F.; Eisenbarth, J.H.; Kaligis, F.; Schneider, T.; Wagele, H.; et al. Antimicrobial Potential of Bacteria Associated with Marine Sea Slugs from North Sulawesi, Indonesia. Front. Microbiol. 2017, 8, 1092. [Google Scholar] [CrossRef] [PubMed]
- Felnagle, E.A.; Jackson, E.E.; Chan, Y.A.; Podevels, A.M.; Berti, A.D.; McMahon, M.D.; Thomas, M.G. Nonribosomal peptide synthetases involved in the production of medically relevant natural products. Mol. Pharm. 2008, 5, 191–211. [Google Scholar] [CrossRef] [PubMed]
- Miller, B.R.; Gulick, A.M. Structural Biology of Nonribosomal Peptide Synthetases. Methods Mol. Biol. 2016, 1401, 3–29. [Google Scholar] [PubMed]
- Elhady, S.S.; Al-Abd, A.M.; El-Halawany, A.M.; Alahdal, A.M.; Hassanean, H.A.; Ahmed, S.A. Antiproliferative Scalarane-Based Metabolites from the Red Sea Sponge Hyrtios erectus. Mar. Drugs 2016, 14, 130. [Google Scholar] [CrossRef] [PubMed]
- Alahdal, A.; Asfour, H.; Ahmed, S.; Noor, A.; Al-Abd, A.; Elfaky, M.; Elhady, S. Anti-Helicobacter, Antitubercular and Cytotoxic Activities of Scalaranes from the Red Sea Sponge Hyrtios erectus. Molecules 2018, 23, 978. [Google Scholar] [CrossRef] [PubMed]
- Asfour, H.Z.; Awan, Z.A.; Bagalagel, A.A.; Elfaky, M.A.; Abdelhameed, R.F.A.; Elhady, S.S. Large-Scale Production of Bioactive Terrein by Aspergillus terreus Strain S020 Isolated from the Saudi Coast of the Red Sea. Biomolecules 2019, 9, 480. [Google Scholar] [CrossRef] [PubMed]
- Khedr, A.I.M.; Mohamed, G.A.; Orabi, M.A.A.; Ibrahim, S.R.M.; Yamada, K. Staphylopeptide A, a new cyclic tetrapeptide from culture broth of Staphylococcus sp. Phytochem. Lett. 2015, 13, 11–14. [Google Scholar] [CrossRef]
- Li, G.-Q.; Deng, Z.-W.; Li, J.; Fu, H.-Z.; Lin, W.-H. Chemical constituents from starfish Asterias rollestoni. J. Chin. Pharm. Sci. 2004, 13, 81–86. [Google Scholar]
- Tang, J.; Gao, H.; Hong, K.; Jiang, M.; Zhou, G.; Wang, N.; Yao, X. Secondary metabolites from a mangrove bacterium Bacillus sp. Chin. J. Med. Chem. 2008, 18, 206–209. [Google Scholar]
- Kitajima, J.; Ishikawa, T.; TANAKA, T.; IDA, Y. Water-soluble constitutents of fennel. IX. glucides and nucleosides. Chem. Pharm. Bull. 1999, 47, 988–992. [Google Scholar] [CrossRef]
- Yang, X.-Q.; Yang, Y.-B.; Zhou, H.; He, G.-W.; Zhao, L.-X.; Xu, L.-H.; Ding, Z.-T. New megastigmane glycoside and alkaloids from Streptomyces sp. YIM 63342. Nat. Prod. Res. 2013, 27, 1191–1196. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.-J.; Lee, S.; Lee, C.-H.; Kim, W.-G. Macrolactins O–R, glycosylated 24-membered lactones from Bacillus sp. AH159-1. J. Nat. Prod. 2007, 70, 1632–1635. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, S.R.; Min, C.C.; Teuscher, F.; Ebel, R.; Kakoschke, C.; Lin, W.; Wray, V.; Edrada-Ebel, R.; Proksch, P. Callyaerins A–F and H, new cytotoxic cyclic peptides from the Indonesian marine sponge Callyspongia aerizusa. Biorg. Med. Chem. 2010, 18, 4947–4956. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, S.R.M.; Edrada-Ebel, R.; Mohamed, G.A.; Youssef, D.T.A.; Wray, V.; Proksch, P. Callyaerin G, a new cytotoxic cyclic peptide from the marine sponge Callyspongia aerizusa. Arkivoc 2008, 164–171. [Google Scholar] [CrossRef]
- Kiehlbauch, J.A.; Hannett, G.E.; Salfinger, M.; Archinal, W.; Monserrat, C.; Carlyn, C. Use of the National Committee for Clinical Laboratory Standards guidelines for disk diffusion susceptibility testing in New York state laboratories. J. Clin. Microbiol. 2000, 38, 3341–3348. [Google Scholar] [PubMed]
- Bonev, B.; Hooper, J.; Parisot, J. Principles of assessing bacterial susceptibility to antibiotics using the agar diffusion method. J. Antimicrob. Chemother. 2008, 61, 1295–1301. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, J.H.; Turnidge, J.D. Susceptibility test methods: Dilution and disk diffusion methods. In Manual of Clinical Microbiology, 11th ed.; American Society of Microbiology: Washington, DC, USA, 2015; pp. 1253–1273. [Google Scholar]
| Position | δH (m, J in Hz) | δC | Position | δH (m, J in Hz) | δC |
|---|---|---|---|---|---|
| L-Leu1 | 2 | 4.62 m | 52.5 (CH) | ||
| 1 | - | 173.4 (C) | 3 | 2.10 m | 40.0 (CH2) |
| 2 | 5.05 m | 52.7 (CH) | 4 | 1.97 m | 25.1 (CH) |
| 3 | 1.97, 2.10 m | 39.5 (CH2) | 5 | 1.02 d (6.6) | 23.4 (CH3) |
| 4 | 1.97 m | 24.9 (CH) | 6 | 0.95 d (6.6) | 21.5 (CH3) |
| 5 | 0.91 d (6.6) | 21.3 (CH3) | NH | 9.58 br s | - |
| 6 | 0.85 d (6.6) | 22.8 (CH3) | A2Pr | ||
| NH | 8.41 br s | 1 | - | 173.8 (C) | |
| L-Asp | 2 | 5.64 m | 51.5 (CH) | ||
| 1 | - | 172 (C) | 3 | 3.62 dd (15.8, 8.8) 3.40 dd (15.8, 4.8) | 37.3 (CH2) |
| 2 | 4.83 m | 53.5 (CH) | NH | 9.20 br s | - |
| 3 | 1.97, 2.67 m | 33.7 (CH2) | NH2 | Not observed | - |
| 4 | 175.6 (C) | L-Asn | |||
| NH | 8.83 (1H, brs) | - | 1 | - | 173.6 (C) |
| L-Val | 2 | 4.90 m | 55.0 (CH) | ||
| 1 | - | 172.5 (C) | 3 | 2.67,2.90 m | 35.0 (CH2) |
| 2 | 4.70 t (6.6) | 61.0 (CH) | 4 | - | 175.5 (C) |
| 3 | 2.67 m | 28.5 (CH) | NH | 8.91 br s | - |
| 4 | 1.17 d (6.6) | 19.5 (CH3) | NH2 | Not observed | - |
| 5 | 1.13 d (6.6) | 18.7 (CH3) | HMTDA | ||
| NH | 9.48 br s | - | 1 | - | 171.7 (C) |
| L-Leu2 | 2 | 2.94, 2.90 m | 43.0 (CH2) | ||
| 1 | - | 172.4 (C) | 3 | 5.64 m | 72.5 (CH) |
| 2 | 4.98 q (7.9) | 52.5 (CH) | 4 | 1.76 m | 42.6 (CH2) |
| 3 | 2.10 m | 39.6 (CH2) | 5 | 1.40 m | 25.4 (CH) |
| 4 | 1.97 m | 25.3 (CH) | 5Me | 0.93 d (6.6) | 23.6 (CH3) |
| 5 | 0.96 d (6.6) | 23.2 (CH3) | 6 | 1.29 (overlapped) | 32.0 (CH2) |
| 6 | 0.92 d (6.6) | 21.8 (CH3) | 7–12 | 1.22 m | 28.7–29.0 (CH2) |
| NH | 8.75 br s | - | 13 | 1.33 (overlapped) | 22.9 (CH2) |
| L-Leu3 | 14 | 0.81 t (6.9) | 14.4 (CH3) | ||
| 1 | - | 174.8 (C) |

| Fragments | m/z |
|---|---|
| [Leu3–A2Pr–Asn–HMTDA–Leu1–Asp–Val–Leu2]+ | 994.7 |
| [Leu3–A2Pr–Asn–HMTDA–Leu1–Asp–Val]+ | 881.6 |
| [A2Pr–Asn–HMTDA–Leu1–Asp]+ | 669.6 |
| [A2Pr–Asn–HMTDA–Leu1]+ | 554.6 |
| [A2Pr–Asn–HMTDA]+ | 441.4 |
| [A2Pr–Asn]+ | 201.2 |
| Compound | Inhibition Zone (mm, 100 µg/disc) | |||||
|---|---|---|---|---|---|---|
| S. aureus | E. coli | P. aeruginosa | S. commune | P. crustosum | A. niger | |
| 1 | 13 | 11 | NA | 11 | NA | NA |
| 2 | 11 | 9 | NA | 13 | NA | NA |
| 3 | 10 | 13 | NA | 12 | NA | NA |
| 4 | 10 | 12 | NA | 11 | NA | NA |
| 5 | 12 | 11 | NA | 10 | NA | NA |
| 6 | 20 | 21 | 25 | 23 | 18 | 16 |
| 7 | 13 | 10 | NA | 13 | NA | NA |
| Ciprofloxacin a | 23 | 24 | 29 | |||
| Nystatin b | 23 | 20 | 19 | |||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdelhameed, R.F.A.; Elhady, S.S.; Noor, A.O.; Almasri, D.M.; Bagalagel, A.A.; Maatooq, G.T.; Khedr, A.I.M.; Yamada, K. Production of a New Cyclic Depsipeptide by the Culture Broth of Staphylococcus sp. Isolated from Corallina officinalis L. Metabolites 2019, 9, 273. https://doi.org/10.3390/metabo9110273
Abdelhameed RFA, Elhady SS, Noor AO, Almasri DM, Bagalagel AA, Maatooq GT, Khedr AIM, Yamada K. Production of a New Cyclic Depsipeptide by the Culture Broth of Staphylococcus sp. Isolated from Corallina officinalis L. Metabolites. 2019; 9(11):273. https://doi.org/10.3390/metabo9110273
Chicago/Turabian StyleAbdelhameed, Reda F. A., Sameh S. Elhady, Ahmad O. Noor, Diena M. Almasri, Alaa A. Bagalagel, Galal T. Maatooq, Amgad I. M. Khedr, and Koji Yamada. 2019. "Production of a New Cyclic Depsipeptide by the Culture Broth of Staphylococcus sp. Isolated from Corallina officinalis L." Metabolites 9, no. 11: 273. https://doi.org/10.3390/metabo9110273
APA StyleAbdelhameed, R. F. A., Elhady, S. S., Noor, A. O., Almasri, D. M., Bagalagel, A. A., Maatooq, G. T., Khedr, A. I. M., & Yamada, K. (2019). Production of a New Cyclic Depsipeptide by the Culture Broth of Staphylococcus sp. Isolated from Corallina officinalis L. Metabolites, 9(11), 273. https://doi.org/10.3390/metabo9110273






