mGWAS-Explorer: Linking SNPs, Genes, Metabolites, and Diseases for Functional Insights
Abstract
:1. Introduction
- Comprehensive collection and deep annotation of SNP–metabolite associations based on data from the 65 mGWAS to date.
- Support for SNP-based, gene-based, and metabolite-based network generation to facilitate interpreting results.
- Powerful network visual analytics system facilitating interactive exploration and built-in topological and functional enrichment analysis.
2. Results
2.1. Overview of the Curated mGWAS Datasets
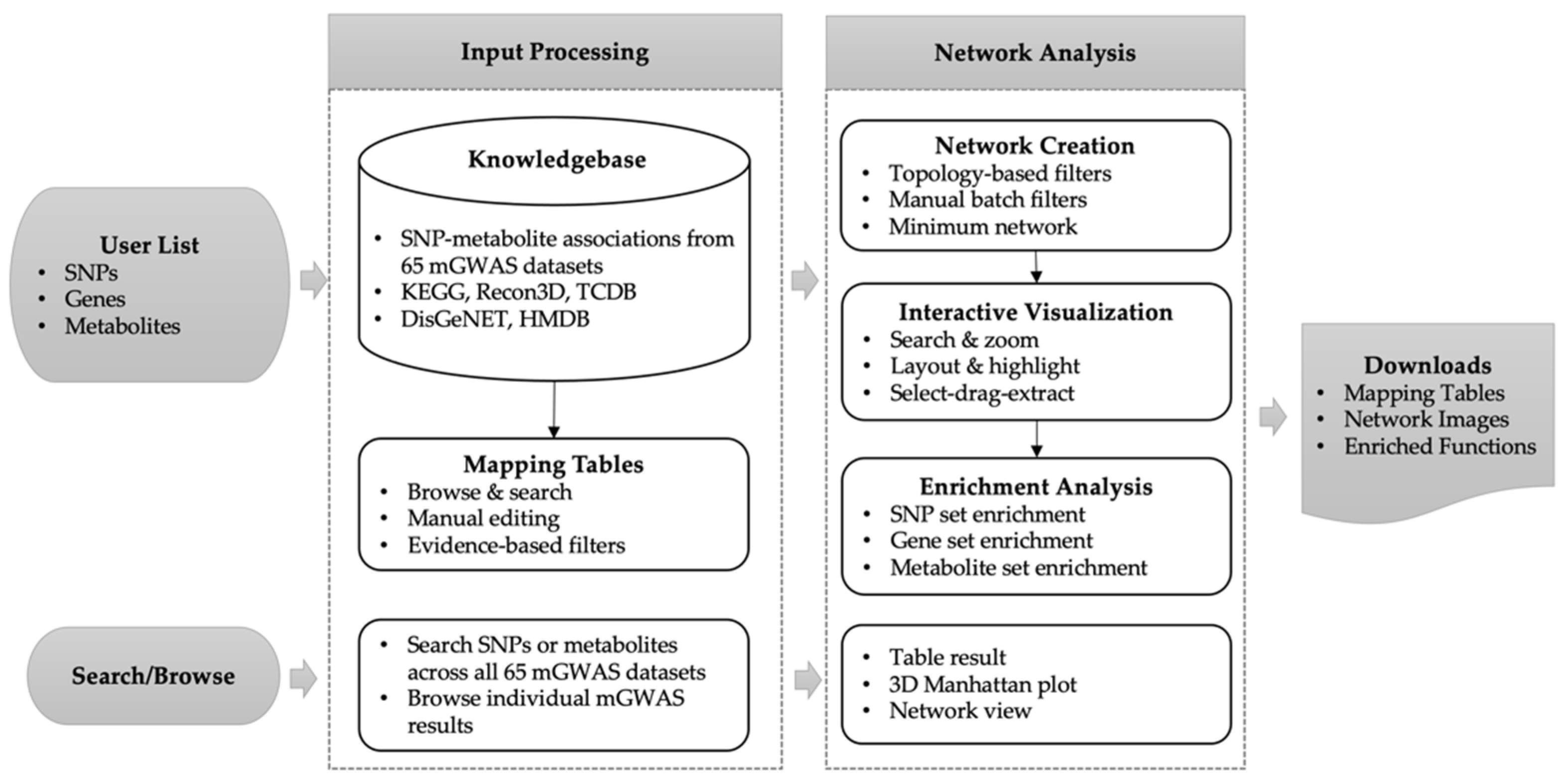
2.2. Overview of the mGWAS-Explorer
2.3. Analysis Workflow
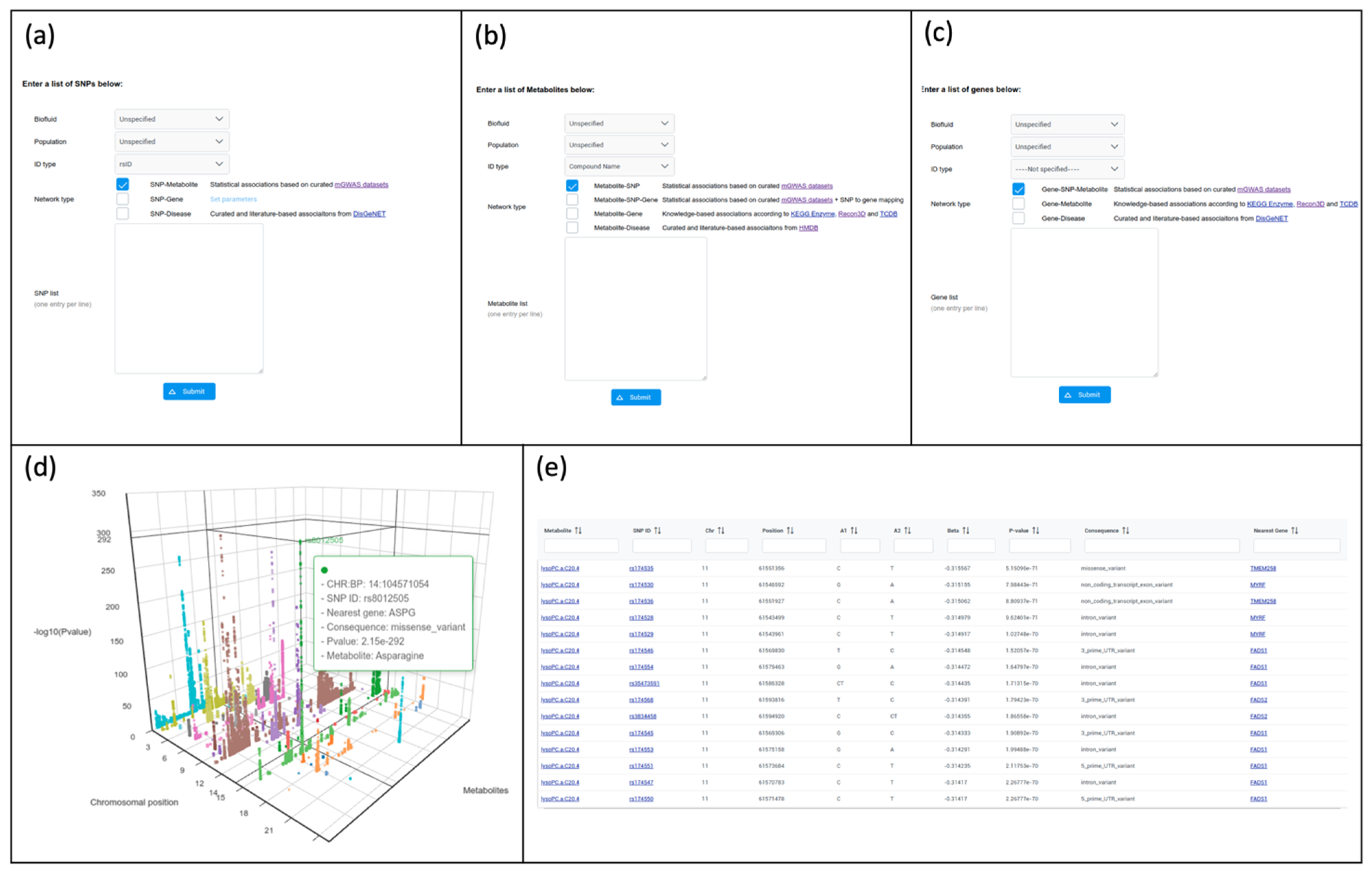
2.3.1. Search and Browse
2.3.2. From SNPs to Networks
2.3.3. From Metabolites to Networks
2.3.4. From Genes to Networks
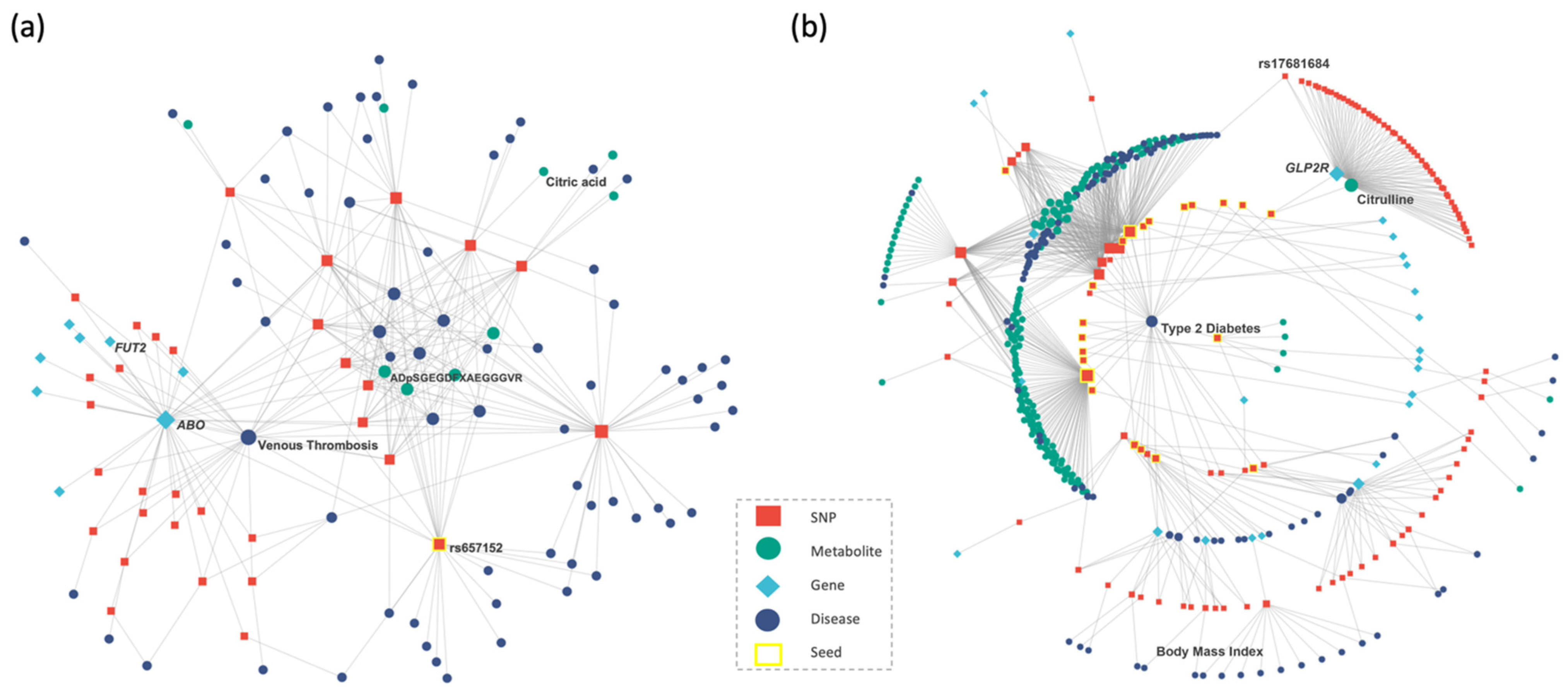
2.4. Case Studies
2.4.1. COVID-19 Case Study
2.4.2. Type 2 Diabetes Case Study
2.5. Comparison with Other Tools
- Metabolomics GWAS Server: http://metabolomics.helmholtz-muenchen.de/gwas/ (accessed on 1 May 2022).
- PheWeb: https://github.com/statgen/pheweb (accessed on 1 May 2022)
- NETMAGE: https://hdpm.biomedinfolab.com/netmage/ (accessed on 1 May 2022) (accept PheWAS summary statistics)
- GePhEx: https://gephex.ega-archive.org/ (accessed on 1 May 2022).
3. Discussion
4. Materials and Methods
4.1. Knowledgebase Curation
4.2. Input Processing and Connection Identification
4.3. Implementation
4.4. Data Collection for Case Studies
4.5. Network Visual Analytics
4.5.1. Network Creation and Customization
4.5.2. Functional Enrichment Analysis
4.5.3. Other Advanced Features
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Visscher, P.M.; Wray, N.R.; Zhang, Q.; Sklar, P.; McCarthy, M.I.; Brown, M.A.; Yang, J. 10 years of GWAS discovery: Biology, function, and translation. Am. J. Hum. Genet. 2017, 101, 5–22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cano-Gamez, E.; Trynka, G. From GWAS to Function: Using Functional Genomics to Identify the Mechanisms Underlying Complex Diseases. Front. Genet. 2020, 11, 424. [Google Scholar] [CrossRef] [PubMed]
- Kastenmüller, G.; Raffler, J.; Gieger, C.; Suhre, K. Genetics of human metabolism: An update. Hum. Mol. Genet. 2015, 24, R93–R101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hagenbeek, F.A.; Pool, R.; van Dongen, J.; Draisma, H.H.M.; Jan Hottenga, J.; Willemsen, G.; Abdellaoui, A.; Fedko, I.O.; den Braber, A.; Visser, P.J.; et al. Heritability estimates for 361 blood metabolites across 40 genome-wide association studies. Nat. Commun. 2020, 11, 39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gieger, C.; Geistlinger, L.; Altmaier, E.; De Angelis, M.H.; Kronenberg, F.; Meitinger, T.; Mewes, H.-W.; Wichmann, H.-E.; Weinberger, K.M.; Adamski, J. Genetics meets metabolomics: A genome-wide association study of metabolite profiles in human serum. PLoS Genet. 2008, 4, e1000282. [Google Scholar] [CrossRef] [Green Version]
- Gallois, A.; Mefford, J.; Ko, A.; Vaysse, A.; Julienne, H.; Ala-Korpela, M.; Laakso, M.; Zaitlen, N.; Pajukanta, P.; Aschard, H. A comprehensive study of metabolite genetics reveals strong pleiotropy and heterogeneity across time and context. Nat. Commun. 2019, 10, 4788. [Google Scholar] [CrossRef] [Green Version]
- Lotta, L.A.; Pietzner, M.; Stewart, I.D.; Wittemans, L.B.L.; Li, C.; Bonelli, R.; Raffler, J.; Biggs, E.K.; Oliver-Williams, C.; Auyeung, V.P.W.; et al. A cross-platform approach identifies genetic regulators of human metabolism and health. Nat. Genet. 2021, 53, 54–64. [Google Scholar] [CrossRef]
- Solovieff, N.; Cotsapas, C.; Lee, P.H.; Purcell, S.M.; Smoller, J.W. Pleiotropy in complex traits: Challenges and strategies. Nat. Rev. Genet. 2013, 14, 483. [Google Scholar] [CrossRef] [Green Version]
- Visscher, P.M.; Yang, J. A plethora of pleiotropy across complex traits. Nat. Genet. 2016, 48, 707. [Google Scholar] [CrossRef]
- Watanabe, K.; Stringer, S.; Frei, O.; Umićević Mirkov, M.; de Leeuw, C.; Polderman, T.J.C.; van der Sluis, S.; Andreassen, O.A.; Neale, B.M.; Posthuma, D. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 2019, 51, 1339–1348. [Google Scholar] [CrossRef]
- Weighill, D.; Jones, P.; Bleker, C.; Ranjan, P.; Shah, M.; Zhao, N.; Martin, M.; DiFazio, S.; Macaya-Sanz, D.; Schmutz, J.; et al. Multi-Phenotype Association Decomposition: Unraveling Complex Gene-Phenotype Relationships. Front. Genet. 2019, 10, 417. [Google Scholar] [CrossRef] [PubMed]
- Julienne, H.; Laville, V.; McCaw, Z.R.; He, Z.; Guillemot, V.; Lasry, C.; Ziyatdinov, A.; Nerin, C.; Vaysse, A.; Lechat, P.; et al. Multitrait GWAS to connect disease variants and biological mechanisms. PLoS Genet. 2021, 17, e1009713. [Google Scholar] [CrossRef] [PubMed]
- Bulik-Sullivan, B.K.; Loh, P.R.; Finucane, H.K.; Ripke, S.; Yang, J.; Patterson, N.; Daly, M.J.; Price, A.L.; Neale, B.M. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 2015, 47, 291–295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giambartolomei, C.; Vukcevic, D.; Schadt, E.E.; Franke, L.; Hingorani, A.D.; Wallace, C.; Plagnol, V. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet. 2014, 10, e1004383. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Majumdar, A.; Haldar, T.; Bhattacharya, S.; Witte, J.S. An efficient Bayesian meta-analysis approach for studying cross-phenotype genetic associations. PLoS Genet. 2018, 14, e1007139. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, B.; Pouget, J.G.; Slowikowski, K.; Stahl, E.; Lee, C.H.; Diogo, D.; Hu, X.; Park, Y.R.; Kim, E.; Gregersen, P.K.; et al. A method to decipher pleiotropy by detecting underlying heterogeneity driven by hidden subgroups applied to autoimmune and neuropsychiatric diseases. Nat. Genet. 2016, 48, 803–810. [Google Scholar] [CrossRef] [PubMed]
- Trochet, H.; Pirinen, M.; Band, G.; Jostins, L.; McVean, G.; Spencer, C.C.A. Bayesian meta-analysis across genome-wide association studies of diverse phenotypes. Genet. Epidemiol. 2019, 43, 532–547. [Google Scholar] [CrossRef] [Green Version]
- Li, X.; Zhu, X. Cross-Phenotype Association Analysis Using Summary Statistics from GWAS. Methods Mol. Biol. 2017, 1666, 455–467. [Google Scholar] [CrossRef]
- Arnold, M.; Raffler, J.; Pfeufer, A.; Suhre, K.; Kastenmüller, G. SNiPA: An interactive, genetic variant-centered annotation browser. Bioinformatics 2014, 31, 1334–1336. [Google Scholar] [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.; Thormann, A.; Flicek, P.; Cunningham, F. The ensembl variant effect predictor. Genome Biol. 2016, 17, 122. [Google Scholar] [CrossRef] [Green Version]
- Carlin, D.E.; Fong, S.H.; Qin, Y.; Jia, T.; Huang, J.K.; Bao, B.; Zhang, C.; Ideker, T. A Fast and Flexible Framework for Network-Assisted Genomic Association. iScience 2019, 16, 155–161. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bastarache, L.; Denny, J.C.; Roden, D.M. Phenome-Wide Association Studies. JAMA 2022, 327, 75–76. [Google Scholar] [CrossRef]
- Tanha, H.M.; Sathyanarayanan, A.; Nyholt, D.R. Genetic overlap and causality between blood metabolites and migraine. Am. J. Hum. Genet. 2021, 108, 2086–2098. [Google Scholar] [CrossRef] [PubMed]
- Kaur, Y.; Wang, D.X.; Liu, H.Y.; Meyre, D. Comprehensive identification of pleiotropic loci for body fat distribution using the NHGRI-EBI Catalog of published genome-wide association studies. Obes. Rev. 2019, 20, 385–406. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Rist, P.M.; Daghlas, I.; Giulianini, F.; Kurth, T.; Chasman, D.I. A genome-wide cross-phenotype meta-analysis of the association of blood pressure with migraine. Nat. Commun. 2020, 11, 3368. [Google Scholar] [CrossRef] [PubMed]
- George, G.; Huang, Y.; Gan, S.; Nar, A.S.; Ha, J.; Venkatesan, R.; Mohan, V.; Wang, H.; Brown, A.; Palmer, C.N.A.; et al. iPheGWAS: An intelligent computational framework to integrate and visualise genome-phenome wide association results. bioRxiv 2022. [Google Scholar] [CrossRef]
- Buniello, A.; MacArthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E.; et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef] [Green Version]
- Kamat, M.A.; Blackshaw, J.A.; Young, R.; Surendran, P.; Burgess, S.; Danesh, J.; Butterworth, A.S.; Staley, J.R. PhenoScanner V2: An expanded tool for searching human genotype-phenotype associations. Bioinformatics 2019, 35, 4851–4853. [Google Scholar] [CrossRef] [Green Version]
- Elsworth, B.; Lyon, M.; Alexander, T.; Liu, Y.; Matthews, P.; Hallett, J.; Bates, P.; Palmer, T.; Haberland, V.; Smith, G.D.; et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv 2020. [Google Scholar] [CrossRef]
- Ghoussaini, M.; Mountjoy, E.; Carmona, M.; Peat, G.; Schmidt, E.M.; Hercules, A.; Fumis, L.; Miranda, A.; Carvalho-Silva, D.; Buniello, A.; et al. Open Targets Genetics: Systematic identification of trait-associated genes using large-scale genetics and functional genomics. Nucleic Acids Res. 2021, 49, D1311–D1320. [Google Scholar] [CrossRef]
- Shashkova, T.I.; Pakhomov, E.D.; Gorev, D.D.; Karssen, L.C.; Joshi, P.K.; Aulchenko, Y.S. PheLiGe: An interactive database of billions of human genotype-phenotype associations. Nucleic Acids Res. 2021, 49, D1347–D1350. [Google Scholar] [CrossRef] [PubMed]
- Piñero, J.; Ramírez-Anguita, J.M.; Saüch-Pitarch, J.; Ronzano, F.; Centeno, E.; Sanz, F.; Furlong, L.I. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res 2020, 48, D845–D855. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, S.-Y.; Fauman, E.B.; Petersen, A.-K.; Krumsiek, J.; Santos, R.; Huang, J.; Arnold, M.; Erte, I.; Forgetta, V.; Yang, T.-P. An atlas of genetic influences on human blood metabolites. Nat. Genet. 2014, 46, 543. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Raffler, J.; Friedrich, N.; Arnold, M.; Kacprowski, T.; Rueedi, R.; Altmaier, E.; Bergmann, S.; Budde, K.; Gieger, C.; Homuth, G.; et al. Genome-Wide Association Study with Targeted and Non-targeted NMR Metabolomics Identifies 15 Novel Loci of Urinary Human Metabolic Individuality. PLoS Genet. 2015, 11, e1005487. [Google Scholar] [CrossRef] [Green Version]
- Gagliano Taliun, S.A.; VandeHaar, P.; Boughton, A.P.; Welch, R.P.; Taliun, D.; Schmidt, E.M.; Zhou, W.; Nielsen, J.B.; Willer, C.J.; Lee, S.; et al. Exploring and visualizing large-scale genetic associations by using PheWeb. Nat. Genet. 2020, 52, 550–552. [Google Scholar] [CrossRef]
- Wang, L.; Balmat, T.J.; Antonia, A.L.; Constantine, F.J.; Henao, R.; Burke, T.W.; Ingham, A.; McClain, M.T.; Tsalik, E.L.; Ko, E.R.; et al. An atlas connecting shared genetic architecture of human diseases and molecular phenotypes provides insight into COVID-19 susceptibility. Genome Med. 2021, 13, 83. [Google Scholar] [CrossRef]
- Sriram, V.; Shivakumar, M.; Jung, S.H.; Nam, Y.; Bang, L.; Verma, A.; Lee, S.; Choe, E.K.; Kim, D. NETMAGE: A human disease phenotype map generator for the network-based visualization of phenome-wide association study results. Gigascience 2022, 11, giac002. [Google Scholar] [CrossRef]
- Strayer, N.; Shirey-Rice, J.K.; Shyr, Y.; Denny, J.C.; Pulley, J.M.; Xu, Y. PheWAS-ME: A web-app for interactive exploration of multimorbidity patterns in PheWAS. Bioinformatics 2021, 37, 1778–1780. [Google Scholar] [CrossRef]
- George, G.; Gan, S.; Huang, Y.; Appleby, P.; Nar, A.S.; Venkatesan, R.; Mohan, V.; Palmer, C.N.A.; Doney, A.S.F. PheGWAS: A new dimension to visualize GWAS across multiple phenotypes. Bioinformatics 2020, 36, 2500–2505. [Google Scholar] [CrossRef]
- Zhu, Z.; Anttila, V.; Smoller, J.W.; Lee, P.H. Statistical power and utility of meta-analysis methods for cross-phenotype genome-wide association studies. PLoS ONE 2018, 13, e0193256. [Google Scholar] [CrossRef] [Green Version]
- Lee, B.; Zhang, S.; Poleksic, A.; Xie, L. Heterogeneous Multi-Layered Network Model for Omics Data Integration and Analysis. Front. Genet. 2019, 10, 1381. [Google Scholar] [CrossRef] [PubMed]
- Sadegh, S.; Skelton, J.; Anastasi, E.; Bernett, J.; Blumenthal, D.B.; Galindez, G.; Salgado-Albarrán, M.; Lazareva, O.; Flanagan, K.; Cockell, S.; et al. Network medicine for disease module identification and drug repurposing with the NeDRex platform. Nat. Commun. 2021, 12, 6848. [Google Scholar] [CrossRef] [PubMed]
- Petersen, A.-K.; Krumsiek, J.; Wägele, B.; Theis, F.J.; Wichmann, H.-E.; Gieger, C.; Suhre, K. On the hypothesis-free testing of metabolite ratios in genome-wide and metabolome-wide association studies. BMC Bioinform. 2012, 13, 120. [Google Scholar] [CrossRef] [Green Version]
- Stacey, D.; Fauman, E.B.; Ziemek, D.; Sun, B.B.; Harshfield, E.L.; Wood, A.M.; Butterworth, A.S.; Suhre, K.; Paul, D.S. ProGeM: A framework for the prioritization of candidate causal genes at molecular quantitative trait loci. Nucleic Acids Res. 2018, 47, e3. [Google Scholar] [CrossRef] [PubMed]
- Kousathanas, A.; Pairo-Castineira, E.; Rawlik, K.; Stuckey, A.; Odhams, C.A.; Walker, S.; Russell, C.D.; Malinauskas, T.; Wu, Y.; Millar, J.; et al. Whole genome sequencing reveals host factors underlying critical COVID-19. Nature 2022. [Google Scholar] [CrossRef]
- Pairo-Castineira, E.; Clohisey, S.; Klaric, L.; Bretherick, A.D.; Rawlik, K.; Pasko, D.; Walker, S.; Parkinson, N.; Fourman, M.H.; Russell, C.D.; et al. Genetic mechanisms of critical illness in COVID-19. Nature 2021, 591, 92–98. [Google Scholar] [CrossRef]
- Ellinghaus, D.; Degenhardt, F.; Bujanda, L.; Buti, M.; Albillos, A.; Invernizzi, P.; Fernández, J.; Prati, D.; Baselli, G.; Asselta, R.; et al. Genomewide Association Study of Severe COVID-19 with Respiratory Failure. N. Engl. J. Med. 2020, 383, 1522–1534. [Google Scholar] [CrossRef]
- Mapping the human genetic architecture of COVID-19. Nature 2021, 600, 472–477. [CrossRef]
- Zhang, Q.; Bastard, P.; Liu, Z.; Le Pen, J.; Moncada-Velez, M.; Chen, J.; Ogishi, M.; Sabli, I.K.D.; Hodeib, S.; Korol, C.; et al. Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science 2020, 370, eabd4570. [Google Scholar] [CrossRef]
- Sindelar, M.; Stancliffe, E.; Schwaiger-Haber, M.; Anbukumar, D.S.; Adkins-Travis, K.; Goss, C.W.; O’Halloran, J.A.; Mudd, P.A.; Liu, W.C.; Albrecht, R.A.; et al. Longitudinal metabolomics of human plasma reveals prognostic markers of COVID-19 disease severity. Cell Rep. Med. 2021, 2, 100369. [Google Scholar] [CrossRef]
- Shi, D.; Yan, R.; Lv, L.; Jiang, H.; Lu, Y.; Sheng, J.; Xie, J.; Wu, W.; Xia, J.; Xu, K.; et al. The serum metabolome of COVID-19 patients is distinctive and predictive. Metab. Clin. Exp. 2021, 118, 154739. [Google Scholar] [CrossRef] [PubMed]
- Timmann, C.; Thye, T.; Vens, M.; Evans, J.; May, J.; Ehmen, C.; Sievertsen, J.; Muntau, B.; Ruge, G.; Loag, W.; et al. Genome-wide association study indicates two novel resistance loci for severe malaria. Nature 2012, 489, 443–446. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Cai, Y.; Xu, A.D. Association study of polymorphisms in the ABO gene and their gene-gene interactions with ischemic stroke in Chinese population. J. Clin. Lab. Anal. 2018, 32, e22329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Germain, M.; Saut, N.; Oudot-Mellakh, T.; Letenneur, L.; Dupuy, A.M.; Bertrand, M.; Alessi, M.C.; Lambert, J.C.; Zelenika, D.; Emmerich, J.; et al. Caution in interpreting results from imputation analysis when linkage disequilibrium extends over a large distance: A case study on venous thrombosis. PLoS ONE 2012, 7, e38538. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Yang, Y.; Huang, H.; Li, D.; Gu, D.; Lu, X.; Zhang, Z.; Liu, L.; Liu, T.; Liu, Y.; et al. Relationship between the ABO Blood Group and the Coronavirus Disease 2019 (COVID-19) Susceptibility. Clin. Infect. Dis. 2021, 73, 328–331. [Google Scholar] [CrossRef]
- Goel, R.; Bloch, E.M.; Pirenne, F.; Al-Riyami, A.Z.; Crowe, E.; Dau, L.; Land, K.; Townsend, M.; Jecko, T.; Rahimi-Levene, N.; et al. ABO blood group and COVID-19: A review on behalf of the ISBT COVID-19 Working Group. Vox Sang. 2021, 116, 849–861. [Google Scholar] [CrossRef]
- Kattula, S.; Byrnes, J.R.; Wolberg, A.S. Fibrinogen and Fibrin in Hemostasis and Thrombosis. Arterioscler. Thromb. Vasc. Biol. 2017, 37, e13–e21. [Google Scholar] [CrossRef] [Green Version]
- Malas, M.B.; Naazie, I.N.; Elsayed, N.; Mathlouthi, A.; Marmor, R.; Clary, B. Thromboembolism risk of COVID-19 is high and associated with a higher risk of mortality: A systematic review and meta-analysis. EClinicalMedicine 2020, 29, 100639. [Google Scholar] [CrossRef]
- Lange, C.; Wolf, J.; Auw-Haedrich, C.; Schlecht, A.; Boneva, S.; Lapp, T.; Horres, R.; Agostini, H.; Martin, G.; Reinhard, T.; et al. Expression of the COVID-19 receptor ACE2 in the human conjunctiva. J. Med. Virol. 2020, 92, 2081–2086. [Google Scholar] [CrossRef]
- Mankelow, T.J.; Singleton, B.K.; Moura, P.L.; Stevens-Hernandez, C.J.; Cogan, N.M.; Gyorffy, G.; Kupzig, S.; Nichols, L.; Asby, C.; Pooley, J.; et al. Blood group type A secretors are associated with a higher risk of COVID-19 cardiovascular disease complications. EJHaem 2021, 2, 175–187. [Google Scholar] [CrossRef]
- Langenberg, C.; Lotta, L.A. Genomic insights into the causes of type 2 diabetes. Lancet 2018, 391, 2463–2474. [Google Scholar] [CrossRef]
- Fuchsberger, C.; Flannick, J.; Teslovich, T.M.; Mahajan, A.; Agarwala, V.; Gaulton, K.J.; Ma, C.; Fontanillas, P.; Moutsianas, L.; McCarthy, D.J.; et al. The genetic architecture of type 2 diabetes. Nature 2016, 536, 41–47. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Scott, R.A.; Scott, L.J.; Mägi, R.; Marullo, L.; Gaulton, K.J.; Kaakinen, M.; Pervjakova, N.; Pers, T.H.; Johnson, A.D.; Eicher, J.D.; et al. An Expanded Genome-Wide Association Study of Type 2 Diabetes in Europeans. Diabetes 2017, 66, 2888–2902. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.; Chen, C.; Ke, X.; Xiong, L.; Shi, Y.; Li, J.; Tan, X.; Ye, S. Analysis of circulating cholesterol levels as a mediator of an association between ABO blood group and coronary heart disease. Circulation. Cardiovasc. Genet. 2014, 7, 43–48. [Google Scholar] [CrossRef]
- Li, S.; Schooling, C.M. A phenome-wide association study of ABO blood groups. BMC Med. 2020, 18, 334. [Google Scholar] [CrossRef]
- Meo, S.A.; Rouq, F.A.; Suraya, F.; Zaidi, S.Z. Association of ABO and Rh blood groups with type 2 diabetes mellitus. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 237–242. [Google Scholar]
- Fagherazzi, G.; Gusto, G.; Clavel-Chapelon, F.; Balkau, B.; Bonnet, F. ABO and Rhesus blood groups and risk of type 2 diabetes: Evidence from the large E3N cohort study. Diabetologia 2015, 58, 519–522. [Google Scholar] [CrossRef] [Green Version]
- Yahaya, T.O.; Oladele, E.O.; Mshelia, M.B.; Sifau, M.O.; Fashola, O.D.; Bunza, M.; Nathaniel, J. Influence of ABO blood groups and demographic characteristics on the prevalence of type 2 diabetes in Lagos, southwest Nigeria. Bull. Natl. Res. Cent. 2021, 45, 144. [Google Scholar] [CrossRef]
- Zhu, L.; She, Z.G.; Cheng, X.; Qin, J.J.; Zhang, X.J.; Cai, J.; Lei, F.; Wang, H.; Xie, J.; Wang, W.; et al. Association of Blood Glucose Control and Outcomes in Patients with COVID-19 and Pre-existing Type 2 Diabetes. Cell Metab. 2020, 31, 1068–1077.e1063. [Google Scholar] [CrossRef]
- Metwally, A.A.; Mehta, P.; Johnson, B.S.; Nagarjuna, A.; Snyder, M.P. COVID-19-Induced New-Onset Diabetes: Trends and Technologies. Diabetes 2021, 70, 2733–2744. [Google Scholar] [CrossRef]
- Rajpal, A.; Rahimi, L.; Ismail-Beigi, F. Factors leading to high morbidity and mortality of COVID-19 in patients with type 2 diabetes. J. Diabetes 2020, 12, 895–908. [Google Scholar] [CrossRef] [PubMed]
- Farré, X.; Spataro, N.; Haziza, F.; Rambla, J.; Navarro, A. Genome-phenome explorer (GePhEx): A tool for the visualization and interpretation of phenotypic relationships supported by genetic evidence. Bioinformatics 2019, 36, 890–896. [Google Scholar] [CrossRef] [PubMed]
- Hemani, G.; Zheng, J.; Elsworth, B.; Wade, K.H.; Haberland, V.; Baird, D.; Laurin, C.; Burgess, S.; Bowden, J.; Langdon, R.; et al. The MR-Base platform supports systematic causal inference across the human phenome. eLife 2018, 7, e34408. [Google Scholar] [CrossRef] [PubMed]
- Williams, N.C.; O’Neill, L.A.J. A Role for the Krebs Cycle Intermediate Citrate in Metabolic Reprogramming in Innate Immunity and Inflammation. Front. Immunol. 2018, 9, 141. [Google Scholar] [CrossRef] [Green Version]
- Martínez-Reyes, I.; Chandel, N.S. Mitochondrial TCA cycle metabolites control physiology and disease. Nat. Commun. 2020, 11, 102. [Google Scholar] [CrossRef] [Green Version]
- Delanghe, J.R.; De Buyzere, M.L.; Speeckaert, M.M. Genetic Polymorphisms in the Host and COVID-19 Infection. Adv. Exp. Med. Biol. 2021, 1318, 109–118. [Google Scholar] [CrossRef]
- Matzhold, E.M.; Berghold, A.; Bemelmans, M.K.B.; Banfi, C.; Stelzl, E.; Kessler, H.H.; Steinmetz, I.; Krause, R.; Wurzer, H.; Schlenke, P.; et al. Lewis and ABO histo-blood types and the secretor status of patients hospitalized with COVID-19 implicate a role for ABO antibodies in susceptibility to infection with SARS-CoV-2. Transfusion 2021, 61, 2736–2745. [Google Scholar] [CrossRef]
- Lindesmith, L.; Moe, C.; Marionneau, S.; Ruvoen, N.; Jiang, X.; Lindblad, L.; Stewart, P.; LePendu, J.; Baric, R. Human susceptibility and resistance to Norwalk virus infection. Nat. Med. 2003, 9, 548–553. [Google Scholar] [CrossRef]
- Payne, D.C.; Currier, R.L.; Staat, M.A.; Sahni, L.C.; Selvarangan, R.; Halasa, N.B.; Englund, J.A.; Weinberg, G.A.; Boom, J.A.; Szilagyi, P.G.; et al. Epidemiologic Association between FUT2 Secretor Status and Severe Rotavirus Gastroenteritis in Children in the United States. JAMA Pediatrics 2015, 169, 1040–1045. [Google Scholar] [CrossRef] [Green Version]
- Cunningham, F.; Allen, J.E.; Allen, J.; Alvarez-Jarreta, J.; Amode, M.R.; Armean, I.M.; Austine-Orimoloye, O.; Azov, A.G.; Barnes, I.; Bennett, R.; et al. Ensembl 2022. Nucleic Acids Res. 2021, 50, D988–D995. [Google Scholar] [CrossRef]
- Ferrer-Admetlla, A.; Sikora, M.; Laayouni, H.; Esteve, A.; Roubinet, F.; Blancher, A.; Calafell, F.; Bertranpetit, J.; Casals, F. A natural history of FUT2 polymorphism in humans. Mol. Biol. Evol. 2009, 26, 1993–2003. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ward, L.D.; Kellis, M. HaploReg v4: Systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 2015, 44, D877–D881. [Google Scholar] [CrossRef]
- Wishart, D.S.; Guo, A.; Oler, E.; Wang, F.; Anjum, A.; Peters, H.; Dizon, R.; Sayeeda, Z.; Tian, S.; Lee, B.L.; et al. HMDB 5.0: The Human Metabolome Database for 2022. Nucleic Acids Res. 2022, 50, D622–D631. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M. Enzyme annotation and metabolic reconstruction using KEGG. Protein Funct. Predict. Methods Protoc. 2017, 1611, 135–145. [Google Scholar]
- Saier, M.H.; Reddy, V.S.; Moreno-Hagelsieb, G.; Hendargo, K.J.; Zhang, Y.; Iddamsetty, V.; Lam, K.J.K.; Tian, N.; Russum, S.; Wang, J.; et al. The Transporter Classification Database (TCDB): 2021 update. Nucleic Acids Res. 2021, 49, D461–D467. [Google Scholar] [CrossRef]
- Brunk, E.; Sahoo, S.; Zielinski, D.C.; Altunkaya, A.; Dräger, A.; Mih, N.; Gatto, F.; Nilsson, A.; Preciat Gonzalez, G.A.; Aurich, M.K.; et al. Recon3D enables a three-dimensional view of gene variation in human metabolism. Nat. Biotechnol. 2018, 36, 272–281. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [Green Version]
- Breuer, K.; Foroushani, A.K.; Laird, M.R.; Chen, C.; Sribnaia, A.; Lo, R.; Winsor, G.L.; Hancock, R.E.; Brinkman, F.S.; Lynn, D.J. InnateDB: Systems biology of innate immunity and beyond—Recent updates and continuing curation. Nucleic Acids Res. 2013, 41, D1228–D1233. [Google Scholar] [CrossRef]
- Rolland, T.; Taşan, M.; Charloteaux, B.; Pevzner, S.J.; Zhong, Q.; Sahni, N.; Yi, S.; Lemmens, I.; Fontanillo, C.; Mosca, R.J.C. A proteome-scale map of the human interactome network. Cell 2014, 159, 1212–1226. [Google Scholar] [CrossRef] [Green Version]
- Lee, B.T.; Barber, G.P.; Benet-Pagès, A.; Casper, J.; Clawson, H.; Diekhans, M.; Fischer, C.; Gonzalez, J.N.; Hinrichs, A.S.; Lee, C.M.; et al. The UCSC Genome Browser database: 2022 update. Nucleic Acids Res. 2022, 50, D1115–D1122. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Akhmedov, M.; Kedaigle, A.; Chong, R.E.; Montemanni, R.; Bertoni, F.; Fraenkel, E.; Kwee, I. PCSF: An R-package for network-based interpretation of high-throughput data. PLoS Comput. Biol. 2017, 13, e1005694. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Csardi, G.; Nepusz, T.J.I. The igraph software package for complex network research. InterJournal Complex Syst. 1695, 5, 1–9. [Google Scholar]
- UniProt Consortium, T. UniProt: The universal protein knowledgebase. Nucleic Acids Res. 2018, 46, 2699. [Google Scholar] [CrossRef] [Green Version]
- Landrum, M.J.; Kattman, B.L. ClinVar at five years: Delivering on the promise. Hum. Mutat. 2018, 39, 1623–1630. [Google Scholar] [CrossRef]
- MacArthur, J.; Bowler, E.; Cerezo, M.; Gil, L.; Hall, P.; Hastings, E.; Junkins, H.; McMahon, A.; Milano, A.; Morales, J.; et al. The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 2017, 45, D896–D901. [Google Scholar] [CrossRef]
- Li, M.J.; Liu, Z.; Wang, P.; Wong, M.P.; Nelson, M.R.; Kocher, J.P.; Yeager, M.; Sham, P.C.; Chanock, S.J.; Xia, Z.; et al. GWASdb v2: An update database for human genetic variants identified by genome-wide association studies. Nucleic Acids Res. 2016, 44, D869–D876. [Google Scholar] [CrossRef] [Green Version]
- Luck, K.; Kim, D.K.; Lambourne, L.; Spirohn, K.; Begg, B.E.; Bian, W.; Brignall, R.; Cafarelli, T.; Campos-Laborie, F.J.; Charloteaux, B.; et al. A reference map of the human binary protein interactome. Nature 2020, 580, 402–408. [Google Scholar] [CrossRef]
- Boyle, E.I.; Weng, S.; Gollub, J.; Jin, H.; Botstein, D.; Cherry, J.M.; Sherlock, G. GO::TermFinder—Open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 2004, 20, 3710–3715. [Google Scholar] [CrossRef] [Green Version]
| Sample Type | Study # | * Metabolite # | ** Metabolite Ratio # | SNP # | SNP–Metabolite Associations # |
|---|---|---|---|---|---|
| Blood | 57 | 3992 | 1265 | 67,570 | 30,3090 |
| Urine | 5 | 271 | 1123 | 6877 | 9647 |
| Saliva | 1 | 14 | 0 | 1364 | 1454 |
| Cerebrospinal fluid (CSF) | 1 | 15 | 0 | 1178 | 1182 |
| Mitochondria | 1 | 0 | 390 | 194 | 404 |
| Sum (unique) | 65 | 4147 | 2388 | 73,737 | 313,720 |
| Tool Name | mGWAS-Explorer | Metabolomics GWAS Server | PheWeb | NETMAGE | GePhEx |
|---|---|---|---|---|---|
| Data input and processing | |||||
| SNP | √ | √ | √ | √ | √ |
| LD proxy search | √ | √ | − | − | √ |
| Gene | √ | √ | √ | − | √ |
| Metabolite | √ | √ | √ | − | √ |
| Enrichment analysis | |||||
| SNP-set | √ | − | − | − | − |
| Gene-set | √ | − | − | − | √ |
| Metabolite-set | √ | − | − | − | − |
| Cross-phenotype exploration | √ | √ | √ | √ | √ |
| Visual analytics | |||||
| Network visualization | +++ | − | − | + | − |
| Network customization | +++ | − | − | + | − |
| Integration with PPI network | √ | − | − | − | − |
| Subnetwork extraction | √ | − | − | √ | − |
| Topology-based filtering | √ | − | − | − | − |
| 3D Manhattan plot | √ | − | − | − | − |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chang, L.; Zhou, G.; Ou, H.; Xia, J. mGWAS-Explorer: Linking SNPs, Genes, Metabolites, and Diseases for Functional Insights. Metabolites 2022, 12, 526. https://doi.org/10.3390/metabo12060526
Chang L, Zhou G, Ou H, Xia J. mGWAS-Explorer: Linking SNPs, Genes, Metabolites, and Diseases for Functional Insights. Metabolites. 2022; 12(6):526. https://doi.org/10.3390/metabo12060526
Chicago/Turabian StyleChang, Le, Guangyan Zhou, Huiting Ou, and Jianguo Xia. 2022. "mGWAS-Explorer: Linking SNPs, Genes, Metabolites, and Diseases for Functional Insights" Metabolites 12, no. 6: 526. https://doi.org/10.3390/metabo12060526
APA StyleChang, L., Zhou, G., Ou, H., & Xia, J. (2022). mGWAS-Explorer: Linking SNPs, Genes, Metabolites, and Diseases for Functional Insights. Metabolites, 12(6), 526. https://doi.org/10.3390/metabo12060526








