Structural Thermokinetic Modelling
Abstract
1. Introduction
2. Results
2.1. Structural Thermokinetic Modelling
2.2. Metabolic Effects of Gene Expression Changes
2.3. Enzyme Synergisms and Epistasis
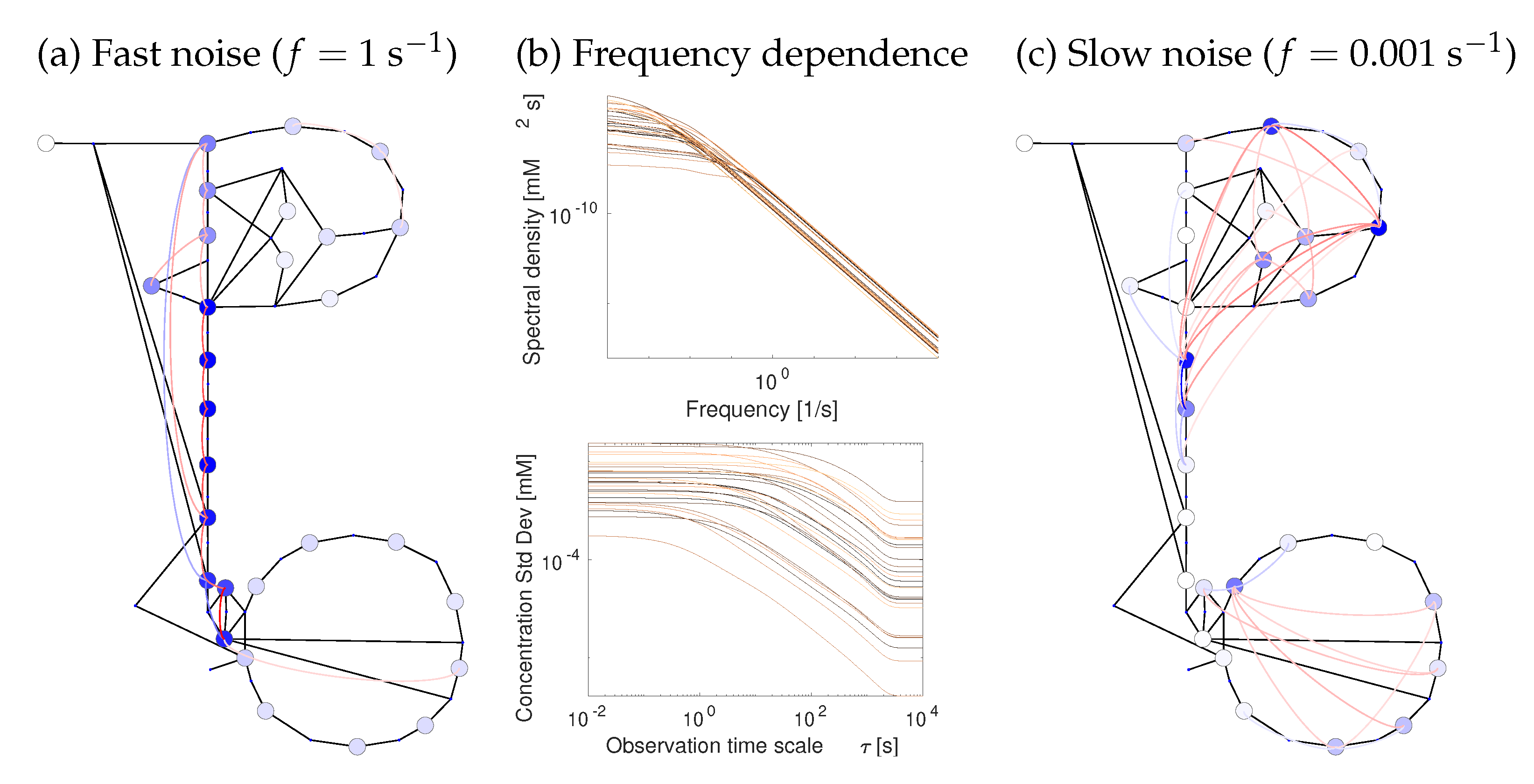
2.4. Uncertain States and Metabolic Fluctuations
2.5. Network Structure and Thermodynamic Forces Shape Metabolic Dynamics
3. Discussion
4. Materials and Methods
4.1. Constructing Kinetic Metabolic Models
4.2. Elasticities and Their Dependence on Thermodynamic Forces
4.3. Model Construction by STM
- Cell growth. To model metabolism in growing cells, the formulae must be adapted. For balanced growth at a cell growth rate , all cellular compounds must be reproduced continuously, so the fluxes follow a mass balance condition with an extra dilution term. Metabolite concentrations and fluxes are tightly coupled by this equation, and in model construction they must be chosen together. In the kinetic model, a dilution term must be added to the ODE of each metabolite, and the Jacobian matrix (which also appears in formulae for the control matrices) contains an extra term . Moreover, conserved moieties in such models must always vanish because otherwise they would be diluted, thus preventing a steady state.
- Kinetic constants as basic variables. Instead of choosing saturation values and concentrations and then computing the constants , we may treat the constants as basic variables and compute the saturation values from them. The distributions for ratios and for saturation values are obviously related. If is uniformly distributed in , the ratio shows a probability density function , i.e., follows a logistic distribution with location parameter 0 and scale parameter 1 (see Supplementary Section S5.1). In practice, we may conveniently sample saturation values from uniform or beta distributions, while kinetic constants may be sampled from sufficiently narrow gamma distributions or from similar log-normal distributions (for the choice of probability distributions, see Supplementary Text 3.1). By sampling saturation values not within but in a smaller range, one may avoid full saturation, and one may sample , , and values around known experimental values.
- Multiple steady states. To build a model with multiple steady states, in the metabolic state phase we choose one set of equilibrium constants, but several sets of concentrations and fluxes for the different steady states; in the kinetics phase the constants , and and velocity constants (geometric means of forward and backward catalytic constants) are chosen or sampled, for example, by parameter balancing [37,52,54]. Finally, enzyme concentrations for each state are computed by matching reaction rates from the rate laws to predefined fluxes.
4.4. Metabolic Control and Synergy Effects
4.5. Variability and Fluctuations in Cells
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| SKM | Structural kinetic modelling |
| MCT | Metabolic control theory |
| FBA | Flux balance analysis |
| MoMA | Minimisation of metabolic adjustment |
| ECM | Enzyme Cost Minimisation |
References
- Steuer, R.; Gross, T.; Selbig, J.; Blasius, B. Structural kinetic modeling of metabolic networks. Proc. Natl. Acad. Sci. USA 2006, 103, 11868–11873. [Google Scholar] [CrossRef] [PubMed]
- Grimbs, S.; Selbig, J.; Bulik, S.; Holzhütter, H.-G.; Steuer, R. The stability and robustness of metabolic states: Identifying stabilizing sites in metabolic networks. Mol. Syst. Biol. 2007, 3, 146. [Google Scholar] [CrossRef] [PubMed]
- Fendt, S.M.; Büscher, J.M.; Rudroff, F.; Picotti, P.; Zamboni, N.; Sauer, U. Tradeoff between enzyme and metabolite efficiency maintains metabolic homeostasis upon perturbations in enzyme capacity. Mol. Sys. Biol. 2010, 13, 356. [Google Scholar] [CrossRef] [PubMed]
- Feist, A.M.; Henry, C.S.; Reed, J.L.; Krummenacker, M.; Joyce, A.R.; Karp, P.D.; Broadbelt, L.J.; Hatzimanikatis, V.; Palsson, B.Ø. A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol. Syst. Biol. 2007, 3, 121. [Google Scholar] [CrossRef]
- Henry, C.S.; DeJongh, M.; Best, A.A.; Frybarger, P.M.; Linsay, B.; Stevens, R.L. High-throughput generation, optimization and analysis of genome-scale metabolic models. Nat. Biotechnol. 2010, 28, 977–982. [Google Scholar] [CrossRef]
- Fell, D.A.; Small, J.R. Fat synthesis in adipose tissue: An examination of stoichiometric constraints. Biochem. J. 1986, 238, 781–786. [Google Scholar] [CrossRef]
- Varma, A.; Palsson, B.Ø. Metabolic flux balancing: Basic concepts, scientific and practical use. Biotechnology 1994, 12, 994–998. [Google Scholar] [CrossRef]
- Segrè, D.; Vitkup, D.; Church, G.M. Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl. Acad. Sci. USA 2002, 99, 15112–15117. [Google Scholar] [CrossRef]
- Holzhütter, H.-G. The principle of flux minimization and its application to estimate stationary fluxes in metabolic networks. Eur. J. Biochem. 2004, 271, 2905–2922. [Google Scholar] [CrossRef]
- Shlomi, T.; Berkman, O.; Ruppin, E. Constraint-based modeling of perturbed organisms: A room for improvement. In Proceedings of the ISMB 2004, Glasgow, Scotland, 31 July–4 August 2004. [Google Scholar]
- Schuetz, R.; Kuepfer, L.; Sauer, U. Systematic evaluation of objective functions for predicting intracellular fluxes in Escherichia coli. Mol. Syst. Biol. 2007, 3, 119. [Google Scholar] [CrossRef]
- Price, N.D.; Schellenberger, J.; Palsson, B.Ø. Uniform sampling of steady-state flux spaces: Means to design experiments and to interpret enzymopathies. Biophys. J. 2004, 87, 2172–2186. [Google Scholar] [CrossRef] [PubMed]
- Beard, D.A.; Liang, S.; Qian, H. Energy balance for analysis of complex metabolic networks. Biophys. J. 2002, 83, 79–86. [Google Scholar] [CrossRef]
- Beard, D.A.; Babson, E.; Curtis, E.; Qian, H. Thermodynamic constraints for biochemical networks. J. Theor. Biol. 2004, 228, 327–333. [Google Scholar] [CrossRef]
- Kümmel, A.; Panke, S.; Heinemann, M. Putative regulatory sites unraveled by network-embedded thermodynamic analysis of metabolome data. Mol. Syst. Biol. 2006, 2, 2006.0034. [Google Scholar] [CrossRef] [PubMed]
- Henry, C.S.; Jankowski, M.D.; Broadbelt, L.J.; Hatzimanikatis, V. Genome-scale thermodynamic analysis of E. coli metabolism. Biophys. J. 2006, 90, 1453–1461. [Google Scholar] [CrossRef]
- Hoppe, A.; Hoffmann, S.; Holzhütter, H.-G. Including metabolite concentrations into flux-balance analysis: Thermodynamic realizability as a constraint on flux distributions in metabolic networks. BMC Syst. Biol. 2007, 1, 23. [Google Scholar] [CrossRef]
- Heinrich, R.; Schuster, S. The Regulation of Cellular Systems; Chapman & Hall: Chapel Hill, NC, USA, 1996. [Google Scholar]
- Reder, C. Metabolic control theory: A structural approach. J. Theor. Biol. 1988, 135, 175–201. [Google Scholar] [CrossRef]
- Hofmeyr, J.-H.S. Metabolic control analysis in a nutshell. In Proceedings of the ICSB 2001, Online, 4–7 November 2001; Available online: http://www.icsb2001.org/toc.html (accessed on 5 April 2021).
- Klipp, E.; Heinrich, R. Competition for enzymes in metabolic pathways: Implications for optimal distributions of enzyme concentrations and for the distribution of flux control. BioSystems 1999, 54, 1–14. [Google Scholar] [CrossRef]
- Liebermeister, W.; Klipp, E. Biochemical networks with uncertain parameters. IEE Proc. Sys. Biol. 2005, 152, 97–107. [Google Scholar] [CrossRef]
- Höfer, T.; Heinrich, R. A second-order approach to metabolic control analysis. J. Theor. Biol. 1993, 164, 85–102. [Google Scholar] [CrossRef]
- Liebermeister, W.; Klipp, E.; Schuster, S.; Heinrich, R. A theory of optimal differential gene expression. BioSystems 2004, 76, 261–278. [Google Scholar] [CrossRef] [PubMed]
- Ingalls, B.P. A frequency domain approach to sensitivity analysis of biochemical systems. J. Phys. Chem. B 2004, 108, 1143–1152. [Google Scholar] [CrossRef]
- Liebermeister, W. Response to temporal parameter fluctuations in biochemical networks. J. Theor. Biol. 2005, 234, 423–438. [Google Scholar] [CrossRef] [PubMed]
- Liebermeister, W.; Baur, U.; Klipp, E. Biochemical network models simplified by balanced truncation. FEBS J. 2005, 272, 4034–4043. [Google Scholar] [CrossRef]
- Wang, L.; Birol, I.; Hatzimanikatis, V. Metabolic control analysis under uncertainty: Framework development and case studies. Biophys. J. 2004, 87, 3750–3763. [Google Scholar] [CrossRef]
- Wang, L.; Hatzimanikatis, V. Metabolic engineering under uncertainty. I: Framework development. Metab. Eng. 2006, 8, 133–141. [Google Scholar] [CrossRef]
- Wang, L.; Hatzimanikatis, V. Metabolic engineering under uncertainty. II: Analysis of yeast metabolism. Metab. Eng. 2006, 8, 142–159. [Google Scholar] [CrossRef]
- Soh, K.C.; Miskovic, L.; Hatzimanikatis, V. From network models to network responses: Integration of thermodynamic and kinetic properties of yeast genome-scale metabolic networks. FEMS Yeast Res. 2012, 12, 129–143. [Google Scholar] [CrossRef]
- Weilandt, D.R.; Salvy, P.; Masid, M.; Fengos, G.; Denhardt-Erikson, R.; Hosseini, Z.; Hatzimanikatis, V. Symbolic Kinetic Models in Python (SKiMpy): Intuitive modeling of large-scale biological kinetic models. bioRxiv 2022. [Google Scholar] [CrossRef]
- Tran, L.M.; Rizk, M.L.; Liao, J.C. Ensemble modeling of metabolic networks. Biophys J. 2008, 95, 5606–5617. [Google Scholar] [CrossRef]
- Jamshidi, N.; Palsson, B.Ø. Formulating genome-scale kinetic models in the post-genome era. Mol. Syst. Biol. 2008, 4, 171. [Google Scholar] [CrossRef] [PubMed]
- Jamshidi, N.; Palsson, B.Ø. Mass action stoichiometric simulation models: Incorporating kinetics and regulation into stoichiometric models. Biophys. J. 2010, 8, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Rivera, J.G.L.; Contador, C.A.; Asenjo, J.A.; Liao, J.C. Reducing the allowable kinetic space by constructing ensemble of dynamic models with the same steady-state flux. Metab. Eng. 2011, 13, 60–75. [Google Scholar] [CrossRef] [PubMed]
- Stanford, N.J.; Lubitz, T.; Smallbone, K.; Klipp, E.; Mendes, P.; Liebermeister, W. Systematic construction of kinetic models from genome-scale metabolic networks. PLoS ONE 2013, 8, e79195. [Google Scholar] [CrossRef] [PubMed]
- Murabito, E.; Verma, M.; Bekker, M.; Bellomo, D.; Westerhoff, H.V.; Teusink, B.; Steuer, R. Monte-Carlo modeling of the central carbon metabolism of Lactococcus lactis: Insights into metabolic regulation. PLoS ONE 2014, 9, e106453. [Google Scholar] [CrossRef]
- Reznik, E.; Segrè, D. On the stability of metabolic cycles. J. Theor. Biol. 2010, 266, 536–549. [Google Scholar] [CrossRef]
- Murabito, E.; Smallbone, K.; Swinton, J.; Westerhoff, H.V.; Steuer, R. A probabilistic approach to identify putative drug targets in biochemical networks. J. R. Soc. Interface 2011, 8, 880–895. [Google Scholar] [CrossRef]
- Girbig, D.; Grimbs, S.; Selbig, J. Systematic analysis of stability patterns in plant primary metabolism. PLoS ONE 2012, 7, e34686. [Google Scholar] [CrossRef]
- Murabito, E. Targeting breast cancer metabolism: A metabolic control analysis approach. Curr. Synth. Sys. Biol. 2013, 1, 104. [Google Scholar]
- Miesković, L.; Hatzimanikatis, V. Modeling of uncertainties in biochemical reactions. Biotechnol. Bioeng. 2011, 108, 413–423. [Google Scholar] [CrossRef]
- Beard, D.A.; Qian, H. Relationship between thermodynamic driving force and one-way fluxes in reversible processes. PLoS ONE 2007, 2, e144. [Google Scholar] [CrossRef] [PubMed]
- Wegscheider, R. Über simultane Gleichgewichte und die Beziehungen zwischen Thermodynamik und Reactionskinetik homogener Systeme. Z. Phys. Chem. 1902, 39, 257–303. [Google Scholar] [CrossRef]
- Liebermeister, W.; Uhlendorf, J.; Klipp, E. Modular rate laws for enzymatic reactions: Thermodynamics, elasticities, and implementation. Bioinformatics 2010, 26, 1528–1534. [Google Scholar] [CrossRef] [PubMed]
- Noor, E.; Bar-Even, A.; Flamholz, A.; Reznik, E.; Liebermeister, W.; Milo, R. Pathway thermodynamics highlights kinetic obstacles in central metabolism. PLoS Comput. Biol. 2014, 10, e100348. [Google Scholar] [CrossRef]
- Alves, R.; Savageau, M.A. Irreversibility in unbranched pathways: Preferred positions based on regulatory considerations. Biophys. J. 2001, 80, 1174–1185. [Google Scholar] [CrossRef][Green Version]
- Euler, C.; Mahadevan, R. On the design principles of metabolic flux sensing. Biophys. J. 2022, 121, 237–247. [Google Scholar] [CrossRef]
- Flamholz, A.; Noor, E.; Bar-Even, A.; Milo, R. eQuilibrator—The biochemical thermodynamics calculator. Nucleic Acids Res. 2012, 40, D770–D775. [Google Scholar] [CrossRef]
- Noor, E.; Flamholz, A.; Bar-Even, A.; Davidi, D.; Milo, R.; Liebermeister, W. The protein cost of metabolic fluxes: Prediction from enzymatic rate laws and cost minimization. PLoS Comput. Biol. 2016, 12, e1005167. [Google Scholar] [CrossRef]
- Lubitz, T.; Schulz, M.; Klipp, E.; Liebermeister, W. Parameter balancing for kinetic models of cell metabolism. J. Phys. Chem. B 2010, 114, 16298–16303. [Google Scholar] [CrossRef]
- Gille, C.; Bölling, C.; Hoppe, A.; Bulik, S.; Hoffmann, S.; Hübner, K.; Karlstädt, A.; Ganeshan, R.; König, M.; Rother, K.; et al. HepatoNet1: A comprehensive metabolic reconstruction of the human hepatocyte for the analysis of liver physiology. Mol. Syst. Biol. 2010, 6, 411. [Google Scholar] [CrossRef]
- Lubitz, T.; Liebermeister, W. Parameter balancing: Consistent parameter sets for kinetic metabolic models. Bioinformatics 2019, 35, 3857–3858. [Google Scholar] [CrossRef] [PubMed]
- Chassagnole, C.; Raïs, B.; Quentin, E.; Fell, D.A.; Mazat, J. An integrated study of threonine-pathway enzyme kinetics in Escherichia coli. Biochem. J. 2001, 356, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Novère, N.; Bornstein, B.; Broicher, A.; Courtot, M.; Donizelli, M.; Dharuri, H.; Li, L.; Sauro, H.; Schilstra, M.; Shapiro, B.; et al. BioModels Database: A free, centralized database of curated, published, quantitative kinetic models of biochemical and cellular systems. Nucleic Acids Res. 2006, 34, D689–D691. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Liebermeister, W. Optimal metabolic states in cells. bioRxiv 2018. [Google Scholar] [CrossRef]
- Liebermeister, W. Metabolic economics in kinetic models. arXiv 2014, arXiv:1404.5252. [Google Scholar]
- Teusink, B.; Walsh, M.C.; van Dam, K.; Westerhoff, H.V. The danger of metabolic pathways with turbo design. Trends Biochem. Sci. 1998, 23, 162–169. [Google Scholar] [CrossRef]
- Costanzo, M.; Baryshnikova, A.; Bellay, J.; Kim, Y.; Spear, E.D.; Sevier, C.S.; Ding, H.; Koh, J.L.Y.; Prinz, J.; Boone, C.; et al. The genetic landscape of a cell. Science 2010, 327, 425–431. [Google Scholar] [CrossRef]
- Elowitz, M.B.; Levine, A.J.; Siggia, E.D.; Swain, P.S. Stochastic gene expression in a single cell. Science 2002, 297, 1183. [Google Scholar] [CrossRef]
- Swain, P.S.; Elowitz, M.B.; Siggia, E.D. Intrinsic and extrinsic contributions to stochasticity in gene expression. Proc. Natl. Acad. Sci. USA 2002, 99, 12795. [Google Scholar] [CrossRef]
- Gillespie, D.T. The chemical Langevin equation. J. Chem. Phys. 2000, 113, 297–306. [Google Scholar] [CrossRef]
- Pellegrini, M.; Marcotte, E.M.; Thompson, M.J.; Eisenberg, D.; Yeates, T.O. Assigning protein functions by comparative genome analysis: Protein phylogenetic profiles. Proc. Natl. Acad. Sci. USA 1996, 96, 4285–4288. [Google Scholar] [CrossRef] [PubMed]
- Tong, A.H.Y.; Lesage, G.; Bader, G.; Ding, H.; Xu, H.; Xin, X.; Young, J.; Burd, C.G.; Munro, S.; Sander, C.; et al. Global mapping of the yeast genetic interaction network. Science 2004, 303, 808–813. [Google Scholar] [CrossRef] [PubMed]
- Steuer, R.; Kurths, J.; Fiehn, O.; Weckwerth, W. Observing and interpreting correlations in metabolomics networks. Bioinformatics 2003, 19, 1019–1026. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, R.; Reder, C. Metabolic control analysis of relaxation processes. J. Theor. Biol. 1991, 151, 343–350. [Google Scholar] [CrossRef]
- Liebermeister, W. Predicting physiological concentrations of metabolites from their molecular structure. J. Comp. Biol. 2005, 12, 1307–1315. [Google Scholar] [CrossRef]
- Liebermeister, W.; Klipp, E. Bringing metabolic networks to life: Convenience rate law and thermodynamic constraints. Theor. Biol. Med. Mod. 2006, 3, 41. [Google Scholar] [CrossRef] [PubMed]
- Bar-Even, A.; Noor, E.; Savir, Y.; Liebermeister, W.; Davidi, D.; Tawfik, D.S.; Milo, R. The moderately efficient enzyme: Evolutionary and physicochemical trends shaping enzyme parameters. Biochemistry 2011, 21, 4402–4410. [Google Scholar] [CrossRef]
- Goldberg, R.N. Thermodynamics of enzyme-catalyzed reactions: Part 6—1999 update. J. Phys. Chem. Ref. Data 1999, 28, 931. [Google Scholar] [CrossRef][Green Version]
- Schomburg, I.; Chang, A.; Ebeling, C.; Gremse, M.; Heldt, C.; Huhn, G.; Schomburg, D. BRENDA, the enzyme database: Updates and major new developments. Nucleic Acids Res. 2004, 32, D431–D433. [Google Scholar] [CrossRef]
- Bulik, S.; Grimbs, S.; Huthmacher, C.; Selbig, J.; Holzhütter, H.-G. Kinetic hybrid models composed of mechanistic and simplified enzymatic rate laws: A promising method for speeding up the kinetic modelling of complex metabolic networks. FEBS J. 2009, 276, 410. [Google Scholar] [CrossRef]
- Liebermeister, W.; Noor, E. Model balancing: A search for in-vivo kinetic constants and consistent metabolic states. Metabolites 2021, 11, 749. [Google Scholar] [CrossRef] [PubMed]
- Klipp, E.; Heinrich, R. Evolutionary optimization of enzyme kinetic parameters; effect of constraints. J. Theor. Biol. 1994, 171, 309–323. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, R.; Rapoport, T.A. A linear steady-state treatment of enzymatic chains. general properties, control and effector strength. Eur. J. Biochem. 1974, 42, 89–95. [Google Scholar] [CrossRef]
- Savageau, M.A. Biochemical systems analysis. III. Dynamic solutions using a power-law approximation. J. Theor. Biol. 1970, 26, 215–226. [Google Scholar] [CrossRef]
- Visser, D.; Heijnen, J.J. Dynamic simulation and metabolic re-design of a branched pathway using linlog kinetics. Metab. Eng. 2003, 5, 164–176. [Google Scholar] [CrossRef]
- Ederer, M.; Gilles, E.D. Thermodynamically feasible kinetic models of reaction networks. Biophys. J. 2007, 92, 1846–1857. [Google Scholar] [CrossRef] [PubMed]
- Haldane, J.B.S. Enzymes; Longmans, Green and Co.: London, UK, 1930. [Google Scholar]
- Smallbone, K.; Simeonidis, E.; Swainston, N.; Mendes, P. Towards a genome-scale kinetic model of cellular metabolism. BMC Syst. Biol. 2010, 4, 6. [Google Scholar] [CrossRef]
- Li, P.; Dada, J.O.; Jameson, D.; Spasic, I.; Swainston, N.; Carroll, K.; Paton, N.W.; Gaskell, S.J.; Westerhoff, H.V.; Broomhead, D.S.; et al. Systematic integration of experimental data and models in systems biology. BMC Bioinform. 2010, 11, 582. [Google Scholar] [CrossRef]
- Flamholz, A.; Noor, E.; Bar-Even, A.; Liebermeister, W.; Milo, R. Glycolytic strategy as a tradeoff between energy yield and protein cost. Proc. Natl. Acad. Sci. USA 2013, 110, 10039–10044. [Google Scholar] [CrossRef]
- Noor, E.; Flamholz, A.; Liebermeister, W.; Bar-Even, A.; Milo, R. A note on the kinetics of enzyme action: A decomposition that highlights thermodynamic effects. FEBS Lett. 2013, 587, 2772–2777. [Google Scholar] [CrossRef] [PubMed]
- Price, N.D.; Thiele, I.; Palsson, B.Ø. Candidate states of Helicobacter pylori genome-scale metabolic network upon application of loop law thermodynamic constraints. Biophys. J. 2006, 90, 3919–3928. [Google Scholar] [CrossRef] [PubMed]
- Jaynes, E.T. Information theory and statistical mechanics. Phys. Rev. 1957, 106, 620–630. [Google Scholar] [CrossRef]
- Segrè, D.; DeLuna, A.; Church, G.M.; Kishony, R. Modular epistasis in yeast metabolism. Nat. Genet. 2005, 37, 77–83. [Google Scholar] [CrossRef]
- Schuster, S.; Schuster, R. A generalization of Wegscheider’s condition. Implications for properties of steady states and for quasi-steady-state approximation. J. Math. Chem. 1989, 3, 25–42. [Google Scholar] [CrossRef]
- Chait, R.; Craney, A.; Kishony, R. Antibiotic interactions that select against resistance. Nature 2007, 446, 668–671. [Google Scholar] [CrossRef]
- Bader, B.W.; Kolda, T.G. Tensor Toolbox for MATLAB, Version 3.2.1. 2021. Available online: www.tensortoolbox.org (accessed on 5 April 2021).
- Bader, B.W.; Kolda, T.G. Efficient MATLAB computations with sparse and factored tensors. SIAM J. Sci. Comput. 2007, 30, 205–231. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liebermeister, W. Structural Thermokinetic Modelling. Metabolites 2022, 12, 434. https://doi.org/10.3390/metabo12050434
Liebermeister W. Structural Thermokinetic Modelling. Metabolites. 2022; 12(5):434. https://doi.org/10.3390/metabo12050434
Chicago/Turabian StyleLiebermeister, Wolfram. 2022. "Structural Thermokinetic Modelling" Metabolites 12, no. 5: 434. https://doi.org/10.3390/metabo12050434
APA StyleLiebermeister, W. (2022). Structural Thermokinetic Modelling. Metabolites, 12(5), 434. https://doi.org/10.3390/metabo12050434





