Human Gut Microbiota in Coronary Artery Disease: A Systematic Review and Meta-Analysis †
Abstract
1. Introduction
2. Materials and Methods
2.1. Systematic Review
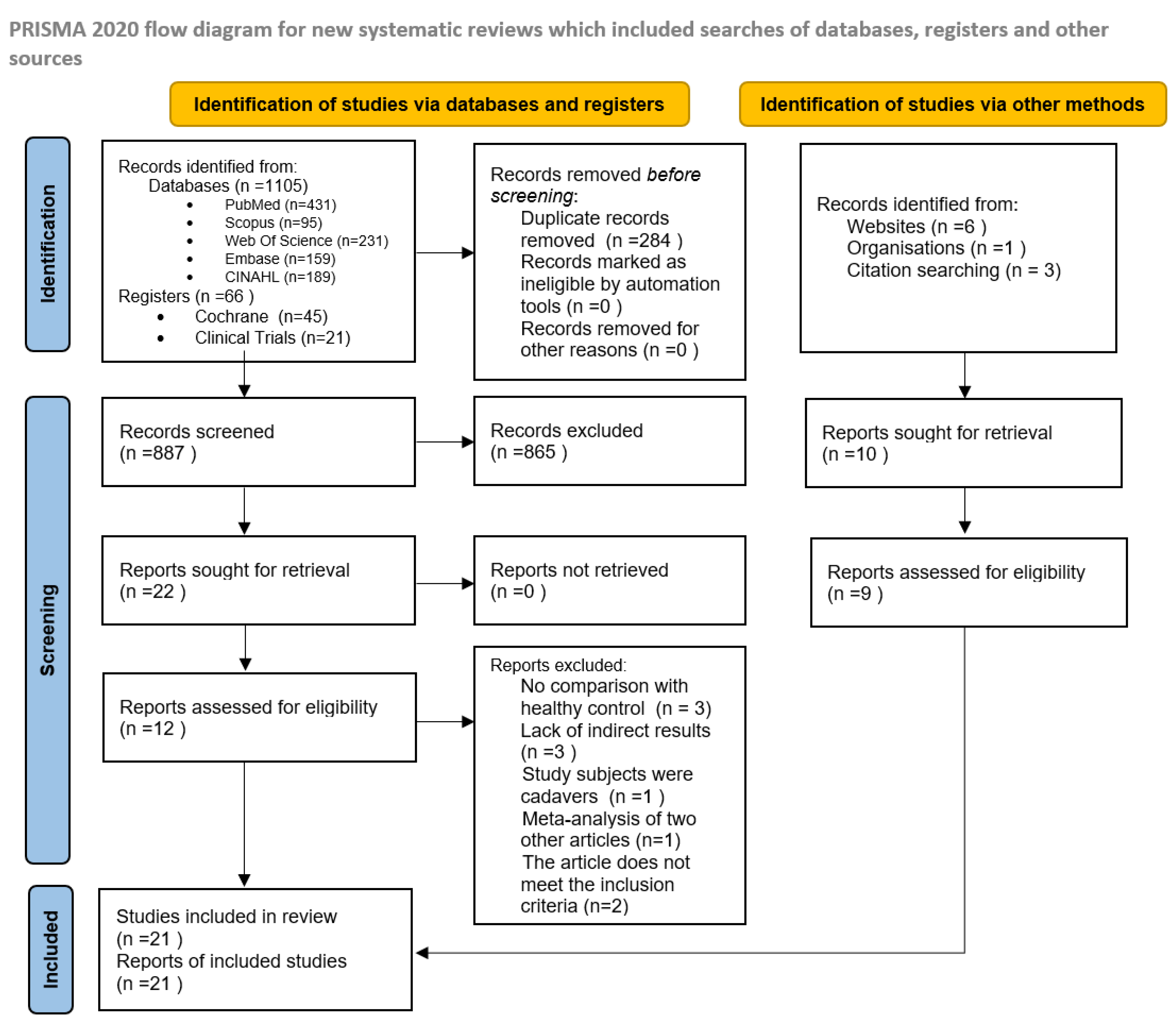
2.1.1. Search Strategy
2.1.2. Inclusion and Exclusion Criteria
2.1.3. Assessment of Eligibility and Data Extraction
2.2. Meta-Analysis
2.2.1. Readings Preparation and zOTU Picking
2.2.2. Differential Abundance Testing
2.2.3. Statistical Analysis
3. Results
3.1. Characteristics of Included Studies
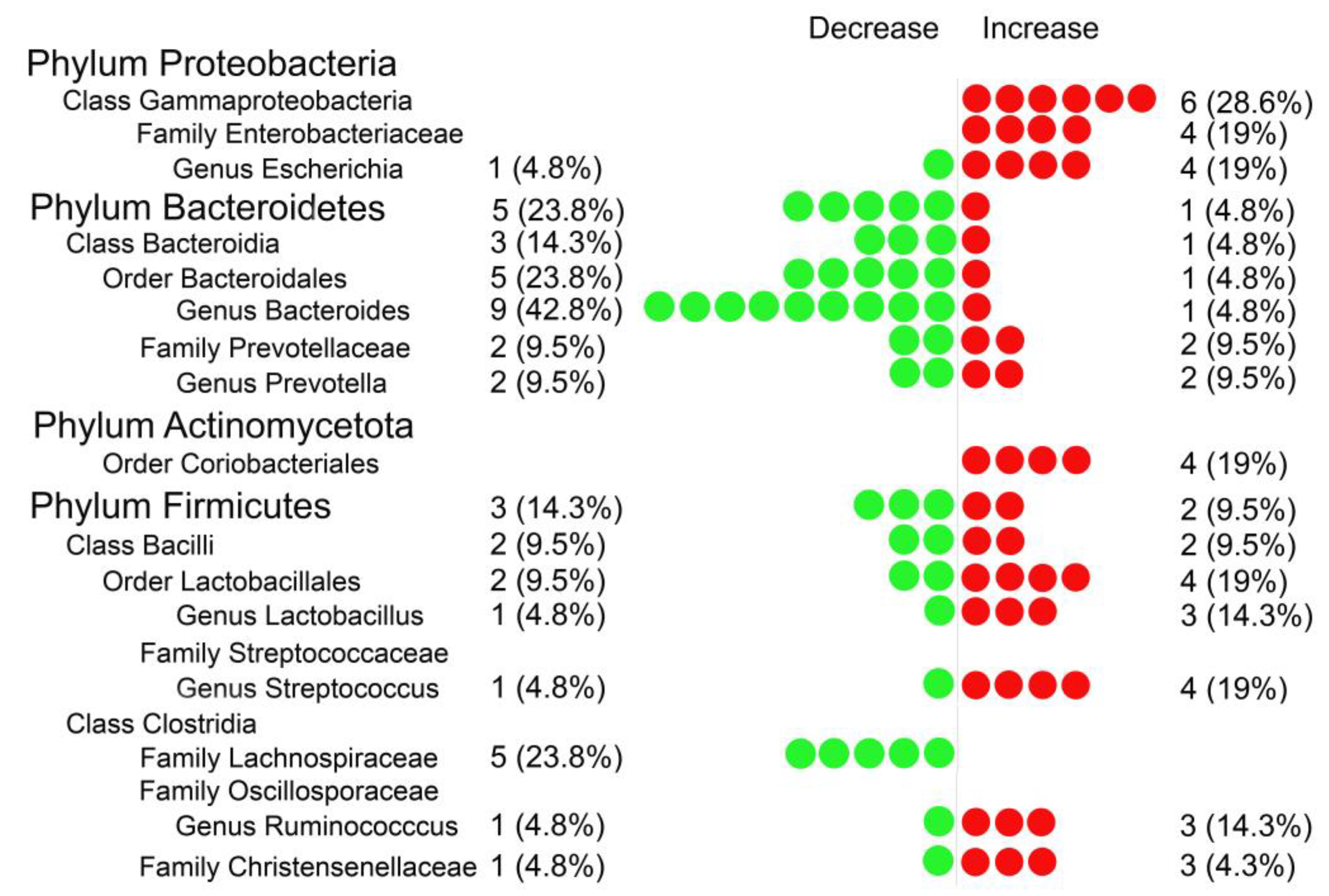
3.2. Systematic Review Results
| RoB NOS | 7 | 10 | 7 | 10 | 8 |
| Diversity α β | − | No data | + | + | No data |
| − | No data | − | + | No data | |
| Major outcome- CAD relative abundance | Bacteroidales▼ Gammaproteobacteria▲ Enterobacteriaceae▲ Prevotella ▲ | Lactobacillales▲ Bacteroides▼ | Gammaproteobacteria▲ Firmicutes▼ Lactobacillales▼ Enterobacteriaceae ▲ Lachnospiraceae▼ Prevotella▼ Bacilli▼ Christensellaceae▲ Prevotellaceae ▼ | Bacteroidetes▲ Firmicutes ▼ Bacteroidales▲ Coriobacteriales▲ Christensellaceae▲ Prevotellaceae▲ | Enterobacteriaceae ▲ Escherichia/Shigella ▲ |
| Major exclusion criteria | patients less than 19 years, pregnancy abnormal kidney function | Patients with systemic diseases: hepatic, renal, collagen, malignancy | other identifiable etiologies of coronary thrombi, active infection during admission. | heart failure, structural heart disease, history of antibiotic use within 1 month, serious dysfunction of liver or kidney | antibiotics, probiotics, decompensated chronic diseases, oncological diseases |
| Major inclusion criteria CAD Control | No prior heart attacks or strokes, and no antibiotic use within three months before enrollment | -coronary risk factors: hypertension, diabetes, and/or dyslipidemia, | no previous history of cardiovascular disease, no evidence of active infection | coronary arteries without stenosis | over 50 years, without cardiovascular disease and acute or acute exacerbations of chronic diseases |
| acute coronary syndrome (STEMI or NSTEMI) | -stable angina pectoris, old myocardial infarction, PCI or CABG for m in 6 months interval | ECG criteria of STEMI, angiographically proven coronary thrombi | ≥50% stenosis in at least one main coronary artery | CAD confirmed by anamnestic data, results of daily ECG Holter monitoring, coronary angiography. | |
| Sample size | CAD 19 Ctrl 19 | CAD 39 Ctrl 30 | CAD 22 Ctrl 20 | CAD 186 Ctrl 123 | CAD 29 Ctrl 30 |
| Technique | 16S rRNA Illumina MiSeq | T-RFLP | 16S rRNA V3-V4 Illumina Miseq | 16S rRNA V3-V4 Illumina MiSeq | 16S rRNA V3-V4 Illumina MiSeq |
| Study design | Case-control | Cross-sectional | Case-control | Cross-sectional | Cross-sectional |
| First author/year | Alhmoud T., et al. [38] 2019 | Emoto T., et al. [39] 2016 | Kwun J., et al. [40] 2020 | Zheng Y, et al. [41] 2020 | Ivashkin, V., et al. [42] 2019 |
| RoB NOS | 7 | 9 | 8 | 9 | 7 |
| Diversity α β | No data | No data | + | No data | No data |
| No data | − | − | − | No data | |
| Major outcome- CAD relative abundance | Bacteroidetes ▼ Lactobacillales ▲ | Escherichia coli ▲ R. gnavus ▲ Bacteroides ▼ Streptococcus ▲ | Enterococcus ▲ Lactobacillus ▲ Bacteroides ▼ Fusobacterium ▼ Dorea ▼ Streptococcus ▼ | Bacteroides vulgatus ▼ Bacteroides dorei ▼ Faecalibacterium ▲ prausnitzii Prevotella copri ▲ | Bacteroides ▼ |
| Major exclusion criteria | acute coronary syndrome systemic disease, including hepatic, renal, collagen disease and malignancy, antibiotic treatment | ongoing infectious diseases, cancer, renal, or hepatic failure, stroke, use of antibiotics within 1 month of sample collection. | kidney dialysis acute infection, gastrointestinal diseases cancer, treatments with antibiotics or probiotics within one month | Patients with: acute coronary syndrome, with systemic disease: including hepatic, renal, collagen disease and malignancy, antibiotics | heart failure, renal and hepatic disease, malignancies, inflammatory disease |
| Major inclusion criteria CAD Control | no history of coronary or another vascular disease, no symptoms indicating angina, no ischemic abnormality in ECG | asymptomatic, no history of CAD, renal failure, systemic disease, and stroke | No data | Patients with coronary risk factors: hypertension, diabetes, dyslipidemia Without a present or past history of coronary or other vascular diseases | Patients with coronary risk factors |
| stable angina, old myocardial infarction, PCI or CABG at least 6 months interval, 75% stenosis of coronary artery | confirmed by coronary angiography, and ≥50% stenosis in single or multiple vessels | coronary angiography or coronary computed tomography angioplasty | stable angina, old myocardial infarction, PCI or CABG ≥ 6 months before the present study. >75% stenosis. | CAD confirmed by a coronary angiography | |
| Sample size | CAD 39 Ctrl 50 | CAD 218 Ctrl 187 | CAD 67 Ctrl 17 | CAD 30 Ctrl 30 | CAD 11 Ctrl 10 |
| Technique | T-RFLP | Shotgun sequen- cing | 16S rRNA V4-V5 Illumina Miseq | 16S rRNA V3-V4 Illumina MiSeq | 16S rRNA V3-V4 Illumina MiSeq |
| Study design | Case-control | Cross-sectional | Cross-sectional | Cross-sectional | Cross-sectional |
| First author/year | Emoto T. et al. [43] 2016 | Jie Z.,et al. [44] 2017 | Liu Z., et al. [45] 2019 | Yoshida N. et al. [46] 2018 | Yoshida N., et al. [47] 2019 |
| RoB NOS | 10 | 10 | 7 | 9 | |
| Diversity α β | + | + | No data | + | |
| + | + | + | + | ||
| Major outcome- CAD relative abundance | Bacteroides ▼ Escherichia ▼ Desulfovibrio ▲ Parabacteroides ▲ Streptococcus ▲ Lacobacillus ▲ | Gamaaproteobacteria ▲ Enterobacteriaceae ▲ Prevotellaceae ▼ Escherichia-Shigella ▲ Fusobacterium ▲ Streptococcus ▲ Bacilli ▼ Lactobacillus ▼ Lactobacillales ▼ Coriobacteriales ▲ | Ruminococcus Gnavus ▲ Lachnospiraceae ▼ Ruminococcus Gauvreauii group ▼ | Bacteroidetes ▼ Bacteroidia ▼ Bacilli ▲ Gammaproteobacteria ▲ Bacteroidales ▼ Lactobacillales ▲ | |
| Major exclusion criteria | No history of unstable angina, myocardial infarction, stroke, cancers, coronary revascularization. | history of GIT surgery or organic disease, history of stroke, hypertension, diabetes, kidney disease infection within one the month of the study or the use of a probiotic, antacid, antibiotic | prior gastrointestinal surgery, the current administration of antibiotics or probiotics, history of IBD or auto-immune diseases, | history of acute or chronic intestinal disease, active cancer aged below 30 and over 80, history of acute coronary syndrome or typical angina (HC) | |
| Major inclusion criteria CAD Control | Without significant stenosis in coronary arteries | healthy volunteers from the hospital health examination center | abnormal peripheral endothelial dysfunction without CAD based on clinical history, non-invasive stress testing, and coronary imaging studies | randomly selected. sex and age-matched to CAD group, without CAD | |
| SA and AMI criteria. The coronary angiography was performed on all patients. | coronary angiography, ECG changes | history of PCI or CABG, coronary arteries diagnosed by coronary angiography or computed tomography coronary angiography | aged 30–79, hospitalization 12–18 months before the evaluation for elective PCI ACS: STEMI, NSTEMI, UA | ||
| Sample size | CAD 141 Ctrl 49 | CAD 60 Ctrl 30 | CAD 88 Ctrl 114 | CAD 169 Ctrl 166 | |
| Technique | 16S rRNA Illumina HiSeq | Phusion High-Fidelity PCR Master Mix | 16S rDNA IM-TORNADO | 16S rRNA | |
| Study design | Cross-sectional | Cross-sectional | Cross-sectional | Cross-sectional | |
| First author/year | Dong Ch,, et al., 2021 preprint [48] | Gao J., et al. [49] 2020 | Toya T, et al. [50] 2021 | Sawicka- Śmiarowska et al. [27] 2021 | |
| RoB NOS | 10 | 10 | 10 | 10 | |
| Diversity α β | + | No data | + | + | |
| − | + | + | + | ||
| Major outcome- CAD relative abundance | Prevotellaceae ▲ Fusobacterium ▼ Bacteroides ▼ Parabacteroides ▲ | Lachnospiraceae ▼ R. gnavus ▲ | Firmicutes ▲ Bacteroides ▼ Bacteroidetes ▼ Bacteroidia ▲ | Bacteroides ▼ Bacilli ▲ Firmicutes ▼ Gammaprotebacteria ▲ Lachnospiracea ▼ Escherichia-Shigella ▲ Lactobacillus ▲ Bacteroidia ▼ Lactobacillales ▲ Bacteroidales ▼ Coriobacteriales ▲ Christensellaceae ▼ Streptococcus ▲ | |
| Major exclusion criteria | IBD, hepatitis B or cirrhosis, cancer, organ failure, exposure to probiotics or prebiotics within one month; receiving treatment with antibiotics, | gastrointestinal surgery, current administration of antibiotics and a probiotic, history of IBD and auto-immune diseases | probiotics, antibiotics within a month before sample gastrointestinal surgery;history of alcohol abuse, diabetes, gastrointestinal disease | cancer, infectious diseases: IBD, antibiotic or probiotic consumption within 1 month before sample collection | |
| Major inclusion criteria CAD Control | participants with no evidence of stenosis in the coronary artery | healthy volunteers, normal or <50% stenosis in coronary arteries | healthy volunteers | no history of CAD and other diseases from the exclusion criteria | |
| coronary angiography, >50% stenosis | ≥50% stenosis in at least one main coronary artery, patients with ACS | CAD confirmed by coronary angiography and CABG or PCI, residents of southern China, 50–85 years. | CAD confirmed by a coronary angiography | ||
| Sample size | CAD 45 Ctrl 19 | CAD 53 Ctrl 53 | CAD 29 Ctrl 34 | CAD 70 Ctrl 98 | |
| Technique | 16S rRNA V4 Ion Torrent | 16S rDNA V3-V5 IM-Tornado | 16S rRNA V3-V5 Illumina Miseq | 16S rRNA V4 Illumina MiSeq | |
| Study design | Cross-sectional | Cross-sectional | Cross-sectional | Cross-sectional | |
| First author/year | Hu Jl, et al. [51] 2021 | Toya T, et al. [52] 2020 | Cui L., et al. [53] 2017 | Zhu Q.et al. [54] 2018 | |
| RoB NOS | 9 | 7 | 7 | ||
| Diversity α β | + | No data | No data | ||
| + | + | No data | |||
| Major outcome- CAD relative abundance | Firmicutes ▲ Bacteroides ▲ Bacteroidetes ▼ Gammaproteobacteria ▲ Bacteroidia ▼ Desulfovibrio ▲ Prevotella ▼ Bacteroidales ▼ Christensellaceae ▲ | Lachnospiraceae▼ | Bacteroidetes ▼ Bacteroidales ▼ Coriobacteriales ▲ | ||
| Major exclusion criteria | antibiotics, probiotics, or prebiotics for at least 3 months before sampling, acute and chronic inflammatory diseases, tumors | Prior gastrointestinal surgery, the current administration of antibiotics, IBD, malignancy, auto-immune disease | renal disease, malignancy, ongoing infectious disease, hepatic disease, use of antibiotics within four weeks before sample collection. | ||
| Major inclusion criteria CAD Control | Tibetan native residents family members of the patients | Without CAD based on clinical history, non-invasive stress testing, and coronary imaging studies | healthy volunteers | ||
| Tibetan native residents coronary artery stenosis >50% ages 40–70 years | typical symptoms, the ECG pattern, cardiac enzyme raise, coronary angiography | CAD confirmed by coronary angiography, and patients with ≥50% stenosis in single or multiple vessels | |||
| Sample size | CAD 18 Ctrl 23 | CAD 19 Ctrl 25 | CAD 15 Ctrl 15 | ||
| Technique | 16S rRNA regions Illumina Hiseq | 16S rRNA V3-V4 Illumina Miseq | 16S rRNA V3-V4 Illumina Miseq | ||
| Study design | Cross-sectional | Case-control study | Cross-sectional | ||
| First author/year | Liu F. et al., 2020 [55] | Chiu et al., 2022 [56] | Choroszy et. al. [57] 2022 | ||
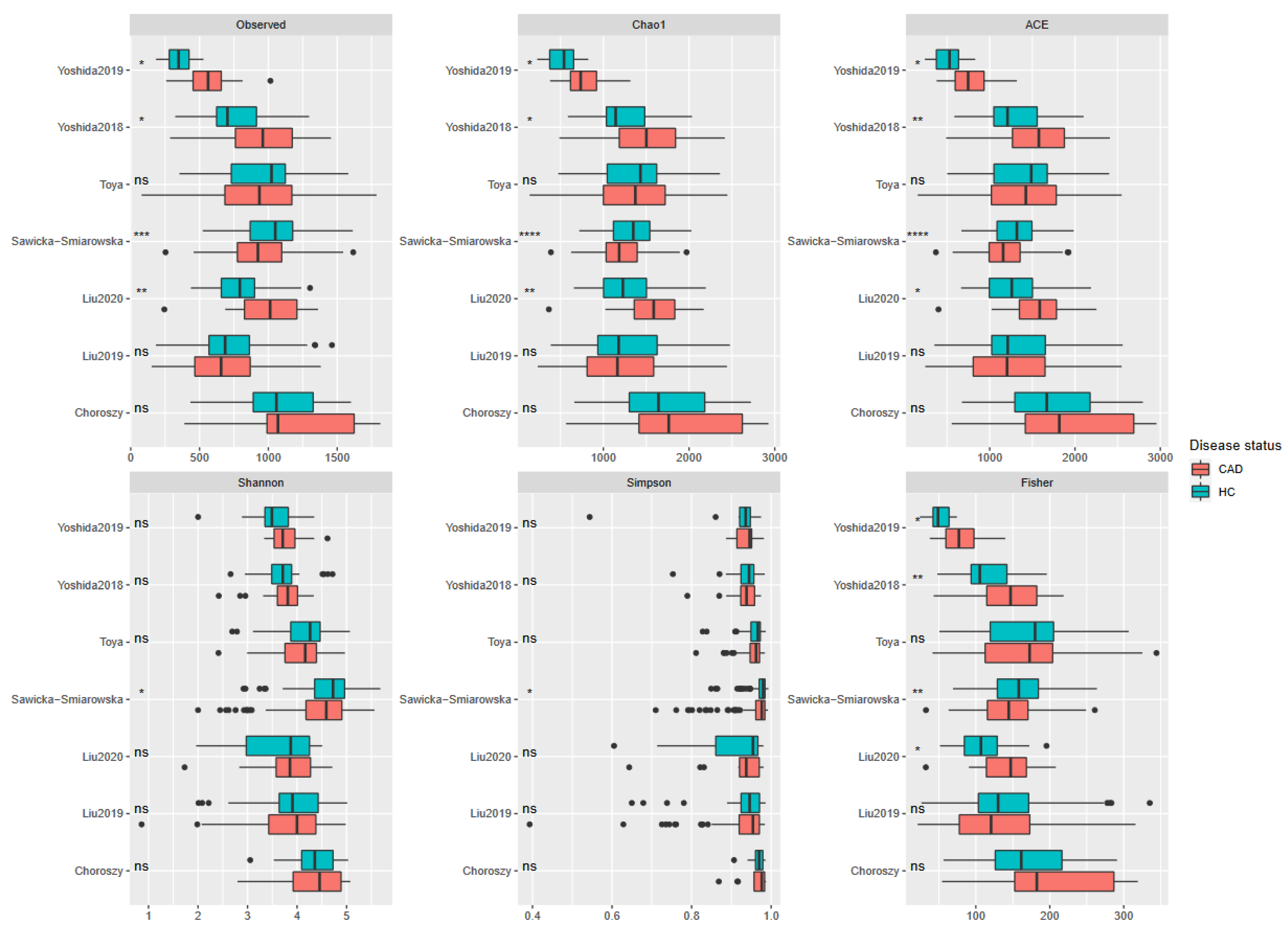
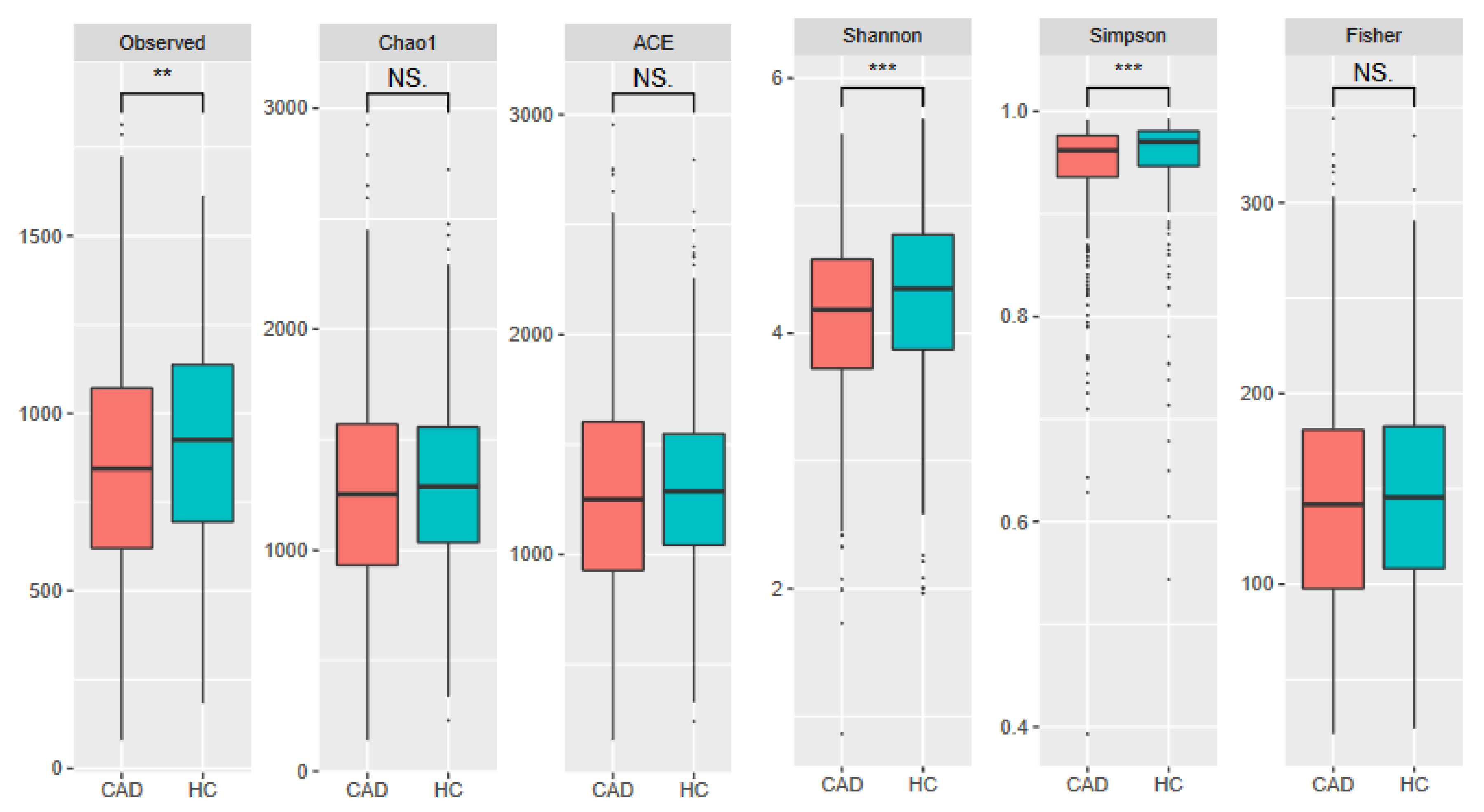
3.3. Meta-Analysis Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Medina-Leyte, D.J.; Zepeda-García, O.; Domínguez-Pérez, M.; González-Garrido, A.; Villarreal-Molina, T.; Jacobo-Albavera, L. Endothelial Dysfunction, Inflammation and Coronary Artery Disease: Potential Biomarkers and Promising Therapeutical Approaches. Int. J. Mol. Sci. 2021, 22, 3850. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.A.; Hashim, M.J.; Mustafa, H.; Baniyas, M.Y.; Al Suwaidi, S.K.B.M.; AlKatheeri, R.; Alblooshi, F.M.K.; Almatrooshi, M.E.A.H.; Alzaabi, M.E.H.; Al Darmaki, R.S.; et al. Global Epidemiology of Ischemic Heart Disease: Results from the Global Burden of Disease Study. Cureus 2020, 12, e9349. [Google Scholar] [CrossRef] [PubMed]
- Maiuolo, J.; Carresi, C.; Gliozzi, M.; Mollace, R.; Scarano, F.; Scicchitano, M.; Macrì, R.; Nucera, S.; Bosco, F.; Oppedisano, F.; et al. The Contribution of Gut Microbiota and Endothelial Dysfunction in the Development of Arterial Hypertension in Animal Models and in Humans. Int. J. Mol. Sci. 2022, 23, 3698. [Google Scholar] [CrossRef] [PubMed]
- Thursby, E.; Juge, N. Introduction to the Human Gut Microbiota. Biochem. J. 2017, 474, 1823–1836. [Google Scholar] [CrossRef] [PubMed]
- Lynch, S.V.; Pedersen, O. The Human Intestinal Microbiome in Health and Disease. N. Engl. J. Med. 2016, 375, 2369–2379. [Google Scholar] [CrossRef] [PubMed]
- Afzaal, M.; Saeed, F.; Shah, Y.A.; Hussain, M.; Rabail, R.; Socol, C.T.; Hassoun, A.; Pateiro, M.; Lorenzo, J.M.; Rusu, A.V.; et al. Human Gut Microbiota in Health and Disease: Unveiling the Relationship. Front. Microbiol. 2022, 13, 999001. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Pedersen, O. Gut Microbiota in Human Metabolic Health and Disease. Nat. Rev. Microbiol. 2021, 19, 55–71. [Google Scholar] [CrossRef]
- Ghosh, S.; Whitley, C.S.; Haribabu, B.; Jala, V.R. Regulation of Intestinal Barrier Function by Microbial Metabolites. Cmgh 2021, 11, 1463–1482. [Google Scholar] [CrossRef]
- Yoo, J.Y.; Sniffen, S.; McGill Percy, K.C.; Pallaval, V.B.; Chidipi, B. Gut Dysbiosis and Immune System in Atherosclerotic Cardiovascular Disease (ACVD). Microorganisms 2022, 10, 108. [Google Scholar] [CrossRef]
- Baizabal-Carvallo, J.F.; Alonso-Juarez, M. The Link between Gut Dysbiosis and Neuroinflammation in Parkinson’s Disease. Neuroscience 2020, 432, 160–173. [Google Scholar] [CrossRef]
- Janeiro, M.H.; Ramírez, M.J.; Solas, M. Dysbiosis and Alzheimer’s Disease: Cause or Treatment Opportunity? Cell Mol. Neurobiol. 2022, 42, 377–387. [Google Scholar] [CrossRef] [PubMed]
- Halverson, T.; Alagiakrishnan, K. Gut Microbes in Neurocognitive and Mental Health Disorders. Ann. Med. 2020, 52, 423–443. [Google Scholar] [CrossRef] [PubMed]
- Zhou, A.; Yuan, Y.; Yang, M.; Huang, Y.; Li, X.; Li, S.; Yang, S.; Tang, B. Crosstalk Between the Gut Microbiota and Epithelial Cells Under Physiological and Infectious Conditions. Front. Cell Infect. Microbiol. 2022, 12, 832672. [Google Scholar] [CrossRef]
- Fachi, J.L.; Felipe, J.d.S.; Pral, L.P.; da Silva, B.K.; Corrêa, R.O.; de Andrade, M.C.P.; da Fonseca, D.M.; Basso, P.J.; Câmara, N.O.S.; de Sales e Souza, É.L.; et al. Butyrate Protects Mice from Clostridium Difficile-Induced Colitis through an HIF-1-Dependent Mechanism. Cell Rep. 2019, 27, 750–761.e7. [Google Scholar] [CrossRef]
- Venkatesh, M.; Mukherjee, S.; Wang, H.; Li, H.; Sun, K.; Benechet, A.P.; Qiu, Z.; Maher, L.; Redinbo, M.R.; Phillips, R.S.; et al. Symbiotic Bacterial Metabolites Regulate Gastrointestinal Barrier Function via the Xenobiotic Sensor PXR and Toll-like Receptor 4. Immunity 2014, 41, 296–310. [Google Scholar] [CrossRef]
- Louis, P.; Flint, H.J. Formation of Propionate and Butyrate by the Human Colonic Microbiota. Environ. Microbiol. 2017, 19, 29–41. [Google Scholar] [CrossRef]
- Ghosh, T.S.; Das, M.; Jeffery, I.B.; O’Toole, P.W. Adjusting for Age Improves Identification of Gut Microbiome Alterations in Multiple Diseases. Elife 2020, 9, e50240. [Google Scholar] [CrossRef]
- Chelakkot, C.; Ghim, J.; Ryu, S.H. Mechanisms Regulating Intestinal Barrier Integrity and Its Pathological Implications. Exp. Mol. Med. 2018, 50, 1–9. [Google Scholar] [CrossRef]
- Violi, F.; Cammisotto, V.; Bartimoccia, S.; Pignatelli, P.; Carnevale, R.; Nocella, C. Gut-Derived Low-Grade Endotoxaemia, Atherothrombosis and Cardiovascular Disease. Nat. Rev. Cardiol. 2022, 0123456789. [Google Scholar] [CrossRef]
- Akash, M.S.H.; Fiayyaz, F.; Rehman, K.; Sabir, S.; Rasool, M.H. Gut Microbiota and Metabolic Disorders: Advances in Therapeutic Interventions. Crit. Rev. Immunol. 2019, 39, 223–237. [Google Scholar] [CrossRef]
- Zhang, B.; Wang, X.; Xia, R.; Li, C. Gut Microbiota in Coronary Artery Disease: A Friend or Foe? Biosci. Rep 2020, 40, BSR20200454. [Google Scholar] [CrossRef] [PubMed]
- Hasan, N.; Yang, H. Factors Affecting the Composition of the Gut Microbiota, and Its Modulation. PeerJ 2019, 7, e7502. [Google Scholar] [CrossRef] [PubMed]
- Boers, S.A.; Jansen, R.; Hays, J.P. Understanding and Overcoming the Pitfalls and Biases of Next-Generation Sequencing (NGS) Methods for Use in the Routine Clinical Microbiological Diagnostic Laboratory. Eur. J. Clin. Microbiol. Infect. Dis. Off. Publ. Eur. Soc. Clin. Microbiol. 2019, 38, 1059–1070. [Google Scholar] [CrossRef] [PubMed]
- Nearing, J.T.; Douglas, G.M.; Hayes, M.G.; MacDonald, J.; Desai, D.K.; Allward, N.; Jones, C.M.A.; Wright, R.J.; Dhanani, A.S.; Comeau, A.M.; et al. Microbiome Differential Abundance Methods Produce Different Results across 38 Datasets. Nat. Commun. 2022, 13, 342. [Google Scholar] [CrossRef]
- Köster, J.; Rahmann, S. Snakemake-a Scalable Bioinformatics Workflow Engine. Bioinformatics 2012, 28, 2520–2522. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Sawicka-Śmiarowska, E.; Bondarczuk, K.; Bauer, W.; Niemira, M.; Szalkowska, A.; Raczkowska, J.; Kwasniewski, M.; Tarasiuk, E.; Dubatowka, M.; Lapinska, M.; et al. Gut Microbiome in Chronic Coronary Syndrome Patients. J. Clin. Med. 2021, 10, 5074. [Google Scholar] [CrossRef]
- RC, E. Search and Clustering Orders of Magnitude Faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef]
- Edgar, R. UNOISE2: Improved Error-Correction for Illumina 16S and ITS Amplicon Sequencing. bioRxiv 2016, 081257. [Google Scholar] [CrossRef]
- Murali, A.; Bhargava, A.; Wright, E.S. IDTAXA: A Novel Approach for Accurate Taxonomic Classification of Microbiome Sequences. Microbiome 2018, 6, 140. [Google Scholar] [CrossRef]
- Chaumeil, P.A.; Mussig, A.J.; Hugenholtz, P.; Parks, D.H. GTDB-Tk: A Toolkit to Classify Genomes with the Genome Taxonomy Database. Bioinformatics 2020, 36, 1925–1927. [Google Scholar] [CrossRef] [PubMed]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.; Peddada, S. Das Analysis of Compositions of Microbiomes with Bias Correction. Nat. Commun. 2020, 11, 3514. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, A.J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; Mcglinn, D.; Minchin, P.R.; Hara, R.B.O.; Simpson, G.L.; Solymos, P.; et al. Vegan. Encycl. Food Agric. Ethics 2019, 2, 2395–2396. [Google Scholar] [CrossRef]
- Agresti, A. Generalized Odds Ratios for Ordinal Data. Biometrics 1980, 36, 59. [Google Scholar] [CrossRef]
- Package, T.; Generalized, T.; Ratios, O.; Rcpp, L. Package ‘Genodds’. 2021. [software].
- Balduzzi, S.; Rücker, G.; Schwarzer, G. How to Perform a Meta-Analysis with R: A Practical Tutorial. Evid. Based. Ment. Health 2019, 22, 153–160. [Google Scholar] [CrossRef]
- Alhmoud, T.; Kumar, A.; Lo, C.-C.; Al-Sadi, R.; Clegg, S.; Alomari, I.; Zmeili, T.; Gleasne, C.D.; Mcmurry, K.; Dichosa, A.E.K.; et al. Investigating Intestinal Permeability and Gut Microbiota Roles in Acute Coronary Syndrome Patients. Hum. Microbiome J. 2019, 13, 100059. [Google Scholar] [CrossRef]
- Emoto, T.; Yamashita, T.; Kobayashi, T.; Sasaki, N.; Hirota, Y.; Hayashi, T.; So, A.; Kasahara, K.; Yodoi, K.; Matsumoto, T.; et al. Characterization of Gut Microbiota Profiles in Coronary Artery Disease Patients Using Data Mining Analysis of Terminal Restriction Fragment Length Polymorphism: Gut Microbiota Could Be a Diagnostic Marker of Coronary Artery Disease. Hear. Vessel. 2016, 32, 39–46. [Google Scholar] [CrossRef] [PubMed]
- Kwun, J.S.; Kang, S.H.; Lee, H.J.; Park, H.K.; Lee, W.J.; Yoon, C.H.; Suh, J.W.; Cho, Y.S.; Youn, T.J.; Chae, I.H. Comparison of Thrombus, Gut, and Oral Microbiomes in Korean Patients with ST-Elevation Myocardial Infarction: A Case–Control Study. Exp. Mol. Med. 2020, 52, 2069–2079. [Google Scholar] [CrossRef]
- Zheng, Y.Y.; Wu, T.T.; Liu, Z.Q.; Li, A.; Guo, Q.Q.; Ma, Y.Y.; Zhang, Z.L.; Xun, Y.L.; Zhang, J.C.; Wang, W.R.; et al. Gut Microbiome-Based Diagnostic Model to Predict Coronary Artery Disease. J. Agric. Food Chem. 2020, 68, 3548–3557. [Google Scholar] [CrossRef] [PubMed]
- Ivashkin, V.T.; Kashukh, Y.A. Impact of L-Carnitine and Phosphatidylcholine Containing Products on the Proatherogenic Metabolite TMAO Production and Gut Microbiome Changes in Patients with Coronary Artery Disease. Vopr. Pitan. 2019, 88, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Emoto, T.; Yamashita, T.; Sasaki, N.; Hirota, Y.; Hayashi, T.; So, A.; Kasahara, K.; Yodoi, K.; Matsumoto, T.; Mizoguchi, T.; et al. Analysis of Gut Microbiota in Coronary Artery Disease Patients: A Possible Link between Gut Microbiota and Coronary Artery Disease. J. Atheroscler. Thromb. 2016, 23, 908–921. [Google Scholar] [CrossRef] [PubMed]
- Jie, Z.; Xia, H.; Zhong, S.-L.; Feng, Q.; Li, S.; Liang, S.; Zhong, H.; Liu, Z.; Gao, Y.; Zhao, H.; et al. The Gut Microbiome in Atherosclerotic Cardiovascular Disease. Nat. Commun. 2017, 8, 845. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Li, J.; Liu, H.; Tang, Y.; Zhan, Q.; Lai, W.; Ao, L.; Meng, X.; Ren, H.; Xu, D.; et al. The Intestinal Microbiota Associated with Cardiac Valve Calcification Differs from That of Coronary Artery Disease. Atherosclerosis 2019, 284, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, N.; Emoto, T.; Yamashita, T.; Watanabe, H.; Hayashi, T.; Tabata, T.; Hoshi, N.; Hatano, N.; Ozawa, G.; Sasaki, N.; et al. Bacteroides Vulgatus and Bacteroides Dorei Reduce Gut Microbial Lipopolysaccharide Production and Inhibit Atherosclerosis. Circulation 2018, 138, 2486–2498. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, N.; Sasaki, K.; Sasaki, D.; Yamashita, T.; Fukuda, H.; Hayashi, T.; Tabata, T.; Osawa, R.; Hirata, K.I.; Kondo, A. Effect of Resistant Starch on the Gut Microbiota and Its Metabolites in Patients with Coronary Artery Disease. J. Atheroscler. Thromb. 2019, 26, 705–719. [Google Scholar] [CrossRef]
- Dong, C.; He, Z.; Zhu, Q.; Liu, J.; Gao, F.; Li, K.; Sun, S.; Liu, Q.; Wang, Y.; Tang, Y.; et al. Correlation Network Analyses Based on Metagenomics and Multi-Type Metabolomic Data Identified Biomarkers of Coronary Artery Disease. Res. Sq. 2020, preprint. [Google Scholar] [CrossRef]
- Gao, J.; Yan, K.T.; Wang, J.X.; Dou, J.; Wang, J.; Ren, M.; Ma, J.; Zhang, X.; Liu, Y. Gut Microbial Taxa as Potential Predictive Biomarkers for Acute Coronary Syndrome and Post-STEMI Cardiovascular Events. Sci. Rep. 2020, 10, 2639. [Google Scholar] [CrossRef]
- Toya, T.; Ozcan, I.; Corban, M.T.; Sara, J.D.; Marietta, E.V.; Ahmad, A.; Horwath, I.E.; Loeffler, D.L.; Murray, J.A.; Lerman, L.O.; et al. Compositional Change of Gut Microbiome and Osteocalcin Expressing Endothelial Progenitor Cells in Patients with Coronary Artery Disease. PLoS ONE 2021, 16, e0249187. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.L.; Yao, Z.F.; Tang, M.N.; Tang, C.; Zhao, X.F.; Su, X.; Lu, D.B.; Li, Q.R.; Wang, Z.S.; Yan, Y.; et al. Gut Microbiota Community Shift with Severity of Coronary Artery Disease. Engineering 2021, 7, 1715–1724. [Google Scholar] [CrossRef]
- Toya, T.; Corban, M.T.; Marrietta, E.; Horwath, I.E.; Lerman, L.O.; Murray, J.A.; Lerman, A. Coronary Artery Disease Is Associated with an Altered Gut Microbiome Composition. PLoS ONE 2020, 15, e0227147. [Google Scholar] [CrossRef]
- Cui, L.; Zhao, T.; Hu, H.; Zhang, W.; Hua, X. Association Study of Gut Flora in Coronary Heart Disease through High-Throughput Sequencing. Biomed Res. Int. 2017, 2017, 3796359. [Google Scholar] [CrossRef]
- Zhu, Q.; Gao, R.; Zhang, Y.; Pan, D.; Zhu, Y.; Zhang, X.; Yang, R.; Jiang, R.; Xu, Y.; Qin, H. Dysbiosis Signatures of Gut Microbiota in Coronary Artery Disease. Physiol. Genomics. 2018, 50, 893–903. [Google Scholar] [CrossRef]
- Liu, F.; Fan, C.; Zhang, L.; Li, Y.; Hou, H.; Ma, Y.; Fan, J.; Tan, Y.; Wu, T.; Jia, S.; et al. Alterations of Gut Microbiome in Tibetan Patients With Coronary Heart Disease. Front. Cell Infect. Microbiol. 2020, 10, 373. [Google Scholar] [CrossRef]
- Chiu, F.-C.; Tsai, C.-F.; Huang, P.-S.; Shih, C.-Y.; Tsai, M.-H.; Hwang, J.-J.; Wang, Y.-C.; Chuang, E.Y.; Tsai, C.-T.; Chang, S.-N. The Gut Microbiome, Seleno-Compounds, and Acute Myocardial Infarction. J. Clin. Med. 2022, 11, 1462. [Google Scholar] [CrossRef]
- Choroszy, M.; Litwinowicz, K.; Łaczmanski, Ł.; Roleder, T.; Sobieszczanska, B. Co-Toxicity of Bacterial Metabolites, to Vascular Endothelial Cells in Coronary Arterial Disease Accompanied by Gut Dysbiosis. Nutrients 2022, 14, 424. [Google Scholar] [CrossRef]
- Xu, J.; Yang, Y.J. Implications of Gut Microbiome on Coronary Artery Disease. Cardiovasc. Diagn. Ther. 2020, 10, 869–880. [Google Scholar] [CrossRef]
- Olvera-Rosales, L.-B.; Cruz-Guerrero, A.-E.; Ramírez-Moreno, E.; Quintero-Lira, A.; Contreras-López, E.; Jaimez-Ordaz, J.; Castañeda-Ovando, A.; Añorve-Morga, J.; Calderón-Ramos, Z.-G.; Arias-Rico, J.; et al. Impact of the Gut Microbiota Balance on the Health-Disease Relationship: The Importance of Consuming Probiotics and Prebiotics. Foods 2021, 10, 1261. [Google Scholar] [CrossRef]
- Liu, J.; Tan, Y.; Cheng, H.; Zhang, D.; Feng, W.; Peng, C. Functions of Gut Microbiota Metabolites, Current Status and Future Perspectives. Aging Dis. 2022, 13, 1106–1126. [Google Scholar] [CrossRef]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. What Is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms 2019, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- Kazemian, N.; Mahmoudi, M.; Halperin, F.; Wu, J.C.; Pakpour, S. Gut Microbiota and Cardiovascular Disease: Opportunities and Challenges. Microbiome 2020, 8, 36. [Google Scholar] [CrossRef]
- Thomas, F.; Hehemann, J.H.; Rebuffet, E.; Czjzek, M.; Michel, G. Environmental and Gut Bacteroidetes: The Food Connection. Front. Microbiol. 2011, 2, 93. [Google Scholar] [CrossRef]
- Kriaa, A.; Bourgin, M.; Potiron, A.; Mkaouar, H.; Jablaoui, A.; Gerard, P.; Maguin, E.; Rhimi, M. Microbial Impact on Cholesterol and Bile Acid Metabolism: Current Status and Future Prospects. J. Lipid Res. 2019, 60, 323–332. [Google Scholar] [CrossRef]
- Troy, E.B.; Kasper, D.L. Beneficial Effects of Bacteroides Fragilis Polysaccharides on the Immune System. Front. Biosci. 2010, 15, 25–34. [Google Scholar] [CrossRef]
- Wexler, A.G.; Goodman, A.L. An Insider’s Perspective: Bacteroides as a Window into the Microbiome. Nat. Microbiol. 2017 25 2017, 2, 17026. [Google Scholar] [CrossRef]
- Zheng, D.; Liwinski, T.; Elinav, E. Interaction between Microbiota and Immunity in Health and Disease. Cell Res. 2020, 30, 492–506. [Google Scholar] [CrossRef]
- Yoshida, N.; Yamashita, T.; Kishino, S.; Watanabe, H.; Sasaki, K.; Sasaki, D.; Tabata, T.; Sugiyama, Y.; Kitamura, N.; Saito, Y.; et al. A Possible Beneficial Effect of Bacteroides on Faecal Lipopolysaccharide Activity and Cardiovascular Diseases. Sci. Rep. 2020, 10, 13009. [Google Scholar] [CrossRef]
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.-M.; et al. Enterotypes of the Human Gut Microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef]
- Vacca, M.; Celano, G.; Calabrese, F.M.; Portincasa, P.; Gobbetti, M.; Angelis, M. De The Controversial Role of Human Gut Lachnospiraceae. Microorganisms 2020, 8, 573. [Google Scholar] [CrossRef]
- Heeney, D.D.; Gareau, M.G.; Marco, M.L. Intestinal Lactobacillus in Health and Disease, a Driver or Just along for the Ride? Curr. Opin. Biotechnol. 2018, 49, 140–147. [Google Scholar] [CrossRef]
- Kechagia, M.; Basoulis, D.; Konstantopoulou, S.; Dimitriadi, D.; Gyftopoulou, K.; Skarmoutsou, N.; Fakiri, E.M. Health Benefits of Probiotics: A Review. ISRN Nutr. 2013, 2013, 481651. [Google Scholar] [CrossRef] [PubMed]
- Ferrarese, R.; Ceresola, E.R.; Preti, A.; Canducci, F. Probiotics, Prebiotics and Synbiotics for Weight Loss and Metabolic Syndrome in the Microbiome Era. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 7588–7605. [Google Scholar] [CrossRef] [PubMed]
- Queipo-Ortuño, M.I.; Seoane, L.M.; Murri, M.; Pardo, M.; Gomez-Zumaquero, J.M.; Cardona, F.; Casanueva, F.; Tinahones, F.J. Gut Microbiota Composition in Male Rat Models under Different Nutritional Status and Physical Activity and Its Association with Serum Leptin and Ghrelin Levels. PLoS ONE 2013, 8, e65465. [Google Scholar] [CrossRef]
- Grigor’eva, I.N. Gallstone Disease, Obesity and the Firmicutes/Bacteroidetes Ratio as a Possible Biomarker of Gut Dysbiosis. J. Pers. Med. 2020, 11, 13. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Li, L.; Zhou, X. Comparative Analysis of the Gut Microbiota in Distinct Statin Response Patients in East China. J. Microbiol. 2018, 56, 886–892. [Google Scholar] [CrossRef] [PubMed]
- Puurunen, V.-P.; Kiviniemi, A.; Lepojärvi, S.; Piira, O.-P.; Hedberg, P.; Junttila, J.; Ukkola, O.; Huikuri, H. Leptin Predicts Short-Term Major Adverse Cardiac Events in Patients with Coronary Artery Disease. Ann. Med. 2017, 49, 448–454. [Google Scholar] [CrossRef]
- Yuan, M.-J.; Li, W.; Zhong, P. Research Progress of Ghrelin on Cardiovascular Disease. Biosci. Rep. 2021, 41, BSR20203387. [Google Scholar] [CrossRef]
- Niknam, M.; Liaghat, T.; Zarghami, M.; Akrami, M.; Shahnematollahi, S.M.; Ahmadipour, A.; Moazzen, F.; Soltanabadi, S. Ghrelin and Ghrelin/Total Cholesterol Ratio as Independent Predictors for Coronary Artery Disease: A Systematic Review and Meta-Analysis. J. Investig. Med. 2022, 70, 759–765. [Google Scholar] [CrossRef]
- Torres-Fuentes, C.; Golubeva, A.V.; Zhdanov, A.V.; Wallace, S.; Arboleya, S.; Papkovsky, D.B.; Aidy, S.E.; Ross, P.; Roy, B.L.; Stanton, C.; et al. Short-Chain Fatty Acids and Microbiota Metabolites Attenuate Ghrelin Receptor Signaling. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2019, 33, 13546–13559. [Google Scholar] [CrossRef]
- Hoyles, L.; Jiménez-Pranteda, M.L.; Chilloux, J.; Brial, F.; Myridakis, A.; Aranias, T.; Magnan, C.; Gibson, G.R.; Sanderson, J.D.; Nicholson, J.K.; et al. Metabolic Retroconversion of Trimethylamine N-Oxide and the Gut Microbiota. Microbiome 2018, 6, 73. [Google Scholar] [CrossRef]
- Dalla Via, A.; Gargari, G.; Taverniti, V.; Rondini, G.; Velardi, I.; Gambaro, V.; Visconti, G.L.; De Vitis, V.; Gardana, C.; Ragg, E.; et al. Urinary TMAO Levels Are Associated with the Taxonomic Composition of the Gut Microbiota and with the Choline TMA-Lyase Gene (CutC) Harbored by Enterobacteriaceae. Nutrients 2019, 12, 62. [Google Scholar] [CrossRef] [PubMed]
- Cosola, C.; Rocchetti, M.T.; Cupisti, A.; Gesualdo, L. Microbiota Metabolites: Pivotal Players of Cardiovascular Damage in Chronic Kidney Disease. Pharmacol. Res. 2018, 130, 132–142. [Google Scholar] [CrossRef] [PubMed]
- Hashizume-Takizawa, T.; Yamaguchi, Y.; Kobayashi, R.; Shinozaki-Kuwahara, N.; Saito, M.; Kurita-Ochiai, T. Oral Challenge with Streptococcus Sanguinis Induces Aortic Inflammation and Accelerates Atherosclerosis in Spontaneously Hyperlipidemic Mice. Biochem. Biophys. Res. Commun. 2019, 520, 507–513. [Google Scholar] [CrossRef] [PubMed]
- Sayols-Baixeras, S.; Dekkers, K.F.; Hammar, U.; Baldanzi, G.; Lin, Y.-T.; Ahmad, S.; Nguyen, D.; Varotsis, G.; Pita, S.; Nielsen, N.; et al. Streptococcus Species Abundance in the Gut Is Linked to Subclinical Coronary Atherosclerosis in 8,973 Participants from the SCAPIS Cohort. medRxiv 2022. [Google Scholar] [CrossRef]
- Karlsson, F.H.; Fåk, F.; Nookaew, I.; Tremaroli, V.; Fagerberg, B.; Petranovic, D.; Bäckhed, F.; Nielsen, J. Symptomatic Atherosclerosis Is Associated with an Altered Gut Metagenome. Nat. Commun. 2012, 3, 1245. [Google Scholar] [CrossRef] [PubMed]
- Claus, S.P.; Ellero, S.L.; Berger, B.; Krause, L.; Bruttin, A.; Molina, J.; Paris, A.; Want, E.J.; de Waziers, I.; Cloarec, O.; et al. Colonization-Induced Host-Gut Microbial Metabolic Interaction. MBio 2011, 2, e00271-10. [Google Scholar] [CrossRef] [PubMed]
- Lahti, L.; Salonen, A.; Kekkonen, R.A.; Salojärvi, J.; Jalanka-Tuovinen, J.; Palva, A.; Orešič, M.; de Vos, W.M. Associations between the Human Intestinal Microbiota, Lactobacillus Rhamnosus GG and Serum Lipids Indicated by Integrated Analysis of High-Throughput Profiling Data. PeerJ 2013, 1, e32. [Google Scholar] [CrossRef] [PubMed]
- Martínez, I.; Wallace, G.; Zhang, C.; Legge, R.; Benson, A.K.; Carr, T.P.; Moriyama, E.N.; Walter, J. Diet-Induced Metabolic Improvements in a Hamster Model of Hypercholesterolemia Are Strongly Linked to Alterations of the Gut Microbiota. Appl. Environ. Microbiol. 2009, 75, 4175–4184. [Google Scholar] [CrossRef]
- Bosco, N.; Noti, M. The Aging Gut Microbiome and Its Impact on Host Immunity. Genes Immun. 2021, 22, 289–303. [Google Scholar] [CrossRef]
- Kim, Y.S.; Unno, T.; Kim, B.Y.; Park, M.S. Sex Differences in Gut Microbiota. World J. Mens. Health 2020, 38, 48–60. [Google Scholar] [CrossRef]
- Gao, X.; Zhang, M.; Xue, J.; Huang, J.; Zhuang, R.; Zhou, X.; Zhang, H.; Fu, Q.; Hao, Y. Body Mass Index Differences in the Gut Microbiota Are Gender Specific. Front. Microbiol. 2018, 9, 1250. [Google Scholar] [CrossRef] [PubMed]
- Han, J.-L.; Lin, H.-L. Intestinal Microbiota and Type 2 Diabetes: From Mechanism Insights to Therapeutic Perspective. World J. Gastroenterol. 2014, 20, 17737–17745. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choroszy, M.; Litwinowicz, K.; Bednarz, R.; Roleder, T.; Lerman, A.; Toya, T.; Kamiński, K.; Sawicka-Śmiarowska, E.; Niemira, M.; Sobieszczańska, B. Human Gut Microbiota in Coronary Artery Disease: A Systematic Review and Meta-Analysis. Metabolites 2022, 12, 1165. https://doi.org/10.3390/metabo12121165
Choroszy M, Litwinowicz K, Bednarz R, Roleder T, Lerman A, Toya T, Kamiński K, Sawicka-Śmiarowska E, Niemira M, Sobieszczańska B. Human Gut Microbiota in Coronary Artery Disease: A Systematic Review and Meta-Analysis. Metabolites. 2022; 12(12):1165. https://doi.org/10.3390/metabo12121165
Chicago/Turabian StyleChoroszy, Marcin, Kamil Litwinowicz, Robert Bednarz, Tomasz Roleder, Amir Lerman, Takumi Toya, Karol Kamiński, Emilia Sawicka-Śmiarowska, Magdalena Niemira, and Beata Sobieszczańska. 2022. "Human Gut Microbiota in Coronary Artery Disease: A Systematic Review and Meta-Analysis" Metabolites 12, no. 12: 1165. https://doi.org/10.3390/metabo12121165
APA StyleChoroszy, M., Litwinowicz, K., Bednarz, R., Roleder, T., Lerman, A., Toya, T., Kamiński, K., Sawicka-Śmiarowska, E., Niemira, M., & Sobieszczańska, B. (2022). Human Gut Microbiota in Coronary Artery Disease: A Systematic Review and Meta-Analysis. Metabolites, 12(12), 1165. https://doi.org/10.3390/metabo12121165









