Effects of SLCO1B1 Genetic Variant on Metabolite Profile in Participants on Simvastatin Treatment
Abstract
:1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Clinical and Laboratory Measurements
2.3. Metabolomics Analysis
2.4. Genotyping
2.5. Statistical Analysis
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chou, R.; Cantor, A.; Tracy, D.T.; Wagner, J.; Ahmed, A.Y.; Rongwei, F.; Maros, F. Statin use for the primary prevention of cardiovascular disease in adults updated evidence report and systematic review for the US preventive services task force. JAMA 2022, 328, 754–771. [Google Scholar] [CrossRef]
- Swerdlow, D.I.; Preiss, D.; Kuchenbaecker, K.B.; Holmes, M.V.; Engmann, J.E.L.; Shah, T.; Sofat, R.; Stender, S.; Johnson, P.C.D.; Scott, R.A.; et al. HMG-coenzyme A reductase inhibition, type 2 diabetes, and bodyweight: Evidence from genetic analysis and randomised trials. Lancet 2015, 385, 351–361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, G.; Shi, M.; Mosley, J.D.; Weng, C.; Zhang, Y.; Lee, M.T.M.; Jarvik, G.P.; Hakonarson, H.; Namjou-Khales, B.; Sleiman, P.; et al. A Mendelian Randomization approach using 3-HMG-coenzyme-A reductase gene variation to evaluate the association of statin-induced low-density lipoprotein cholesterol lowering with non-cardiovascular disease phenotypes. JAMA 2021, 4, e2112820. [Google Scholar] [CrossRef]
- Cederberg, H.; Stančáková, A.; Yaluri, N.; Modi, S.; Kuusisto, J.; Laakso, M. Increased risk of diabetes with statin treatment is associated with impaired insulin sensitivity and insulin secretion: A 6-year follow-up study of the METSIM cohort. Diabetologia 2015, 58, 1109–1117. [Google Scholar] [CrossRef] [Green Version]
- Laakso, M.; Kuusisto, J. Diabetes secondary to treatment with statins. Curr. Diab. Rep. 2017, 17, 10. [Google Scholar] [CrossRef]
- Joy, T.R.; Hegele, R. Narrative review: Statin-related myopathy. Ann. Intern. Med. 2009, 150, 858–868. [Google Scholar] [CrossRef] [Green Version]
- Niemi, M.; Pasanen, M.K.; Neuvonen, P.J. Organic anion transporting polypeptide 1B1: A genetically polymorphic transporter of major importance for hepatic drug uptake. Pharmacol. Rev. 2011, 63, 157–181. [Google Scholar] [CrossRef] [Green Version]
- Ramsey, L.B.; Gong, L.; Lee, S.; Wagner, J.B.; Zhou, X.; Sangkuhl, K.; Adams, S.M.; Straka, R.J.; Empey, P.E.; Boone, E.C.; et al. PharmVar GeneFocus: SLCO1B1. Clin. Pharmacol. Ther. 2022. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, L.B.; Johnson, S.G.; Caudle, K.E.; Haidar, C.E.; Voora, D.; Wilke, R.A.; Maxwell, W.D.; McLeod, H.L.; Krauss, R.M.; Roden, D.M.; et al. The clinical pharmacogenetics implementation consortium guideline for SLCO1B1 and simvastatin-induced myopathy: 2014 update. Clin. Pharmacol. Ther. 2014, 96, 423–428. [Google Scholar] [CrossRef]
- Choi, H.Y.; Bae, K.S.; Cho, S.H.; Ghim, J.L.; Choe, S.; Jung, J.A.; Jin, S.J.; Kim, H.S.; Lim, H.S. Impact of CYP2D6, CYP3A5, CYP2C19, CYP2A6, SLCO1B1, ABCB1, and ABCG2 gene polymorphisms on the pharmacokinetics of simvastatin and simvastatin acid. Pharm. Genom. 2015, 25, 595–608. [Google Scholar] [CrossRef]
- Licito, A.; Marotta, G.; Battaglia, M.; Benincasa, G.; Mentone, L.; Grillo, M.R.; Lucia, V.D.; Leonardi, G.; Bignucolo, A.; Mentone, L.; et al. Assessment of pharmacogenomic SLCO1B1 assay for prediction of neuromuscular pain in type 2 diabetes mellitus and cardiovascular patients: Preliminary results. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 469–477. [Google Scholar] [PubMed]
- Wojtyniak, J.-G.; Selzer, D.; Schwab, M.; Lehr, T. Physiologically Based Precision Dosing Approach for Drug-Drug-Gene Interactions: A Simvastatin Network Analysis. Clin. Pharmacol. Ther. 2021, 109, 201–211. [Google Scholar] [CrossRef]
- Mykkänen, A.J.; Taskinen, S.; Neuvonen, M.; Paile-Hyvärinen, M.; Tarkiainen, E.K.; Lilius, T.; Tapaninen, T.; Backman, J.T.; Tornio, A.; Niemi, M. Genomewide association study of simvastatin pharmacokinetics. Clin. Pharmacol. Ther. 2022, 112, 676–686. [Google Scholar] [CrossRef] [PubMed]
- Peyser, B.; Perry, E.P.; Singh, K.; Gill, R.D.; Mehan, M.R.; Haga, S.B.; Musty, M.D..; Milazzo, N.A.; Savard, D.; Li, Y.J.; et al. Effects of delivering SLCO1B1 pharmacogenetic information in randomized trial and observational settings. Circ. Genom. Precis. Med. 2018, 11, e002228. [Google Scholar] [CrossRef] [Green Version]
- Meyer, U.A.; Ulrich, M.; Zanger, U.M.; Schwab, M. Omics and drug response. Annu. Rev. Pharmacol. Toxicol. 2013, 53, 475–503. [Google Scholar] [CrossRef]
- Laakso, M.; Kuusisto, J.; Stančáková, A.; Kuulasmaa, T.; Pajukanta, P.; Lusis, A.J.; Collins, F.S.; Mohlke, K.L.; Boehnke, M. The Metabolic Syndrome in Men study: A resource for studies of metabolic and cardiovascular diseases. J. Lipid Res. 2017, 58, 481–493. [Google Scholar] [CrossRef] [Green Version]
- Stancáková, A.; Javorský, M.; Kuulasmaa, T.; Haffner, S.M.; Kuusisto, J.; Laakso, M. Changes in insulin sensitivity and insulin release in relation to glycemia and glucose tolerance in 6,414 Finnish men. Diabetes 2009, 38, 1212–1221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care 2013, 36, S67–S74. [Google Scholar] [CrossRef] [Green Version]
- Stančáková, A.; Civelek, A.; Saleem, N.K.; Soininen,, P.; Kangas, A.J.; Cederberg, H.; Paananen, J.; Pihlajamäki, J.; Bonnycastle,, L.L.; Morken, M.A.; et al. Hyperglycemia and a common variant of GCKR are associated with the levels of eight amino acids in 9369 Finnish men. Diabetes 2012, 61, 1895–1902. [Google Scholar]
- Yin, X.; Chan, L.S.; Bose, D.; Jackson, A.U.; VandeHaar, P.; Locke, A.E.; Fuchsberger, C.; Stringham, H.M.; Welch, R.; Yu, K.; et al. Genome-wide association studies of metabolites in Finnish men identify disease-relevant loci. Nat. Commun. 2022, 13, 1644. [Google Scholar] [CrossRef] [PubMed]
- Fernandes Silva, L.; Ravi, R.; Vangipurapu, J.; Laakso, M. Metabolite signature of simvastatin treatment involves multiple metabolic pathways. Metabolites 2022, 12, 753. [Google Scholar] [CrossRef]
- Stančáková, A.; Kuulasmaa, T.; Kuusisto, J.; Mohlke, K.L.; Collins, F.S.; Boehnke, M.; Laakso, M. Genetic risk scores in the prediction of plasma glucose, impaired insulin secretion, insulin resistance and incident type 2 diabetes in the METSIM study. Diabetologia 2017, 60, 1722–1730. [Google Scholar] [CrossRef] [Green Version]
- Suhre, K.; Shin, S.Y.; Petersen, A.K.; Mohney, R.P.; Meredith, D.; Wägele, B.; Altmaier, E.; Deloukas, P.; Erdmann, J.; Grundberg, E.; et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 2011, 477, 54–60. [Google Scholar] [CrossRef] [Green Version]
- Johnson, A.D.; Kavousi, M.; Smith, A.V.; Chen, M.H.; Dehghan, A.; Aspelund, T.; Lin, J.P.; Van Duijn, C.M.; Harris, T.B.; Cupples, L.A.; et al. Genome-wide association meta-analysis for total serum bilirubin levels. Hum. Mol. Genet. 2009, 18, 2700–2710. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yee, S.; Giacomini, M.; Hsueh, C.-H.; Weitz, D.; Liang, X.; Goswami, S.; Kinchen, J.; Coelho, A.; Zur, A.; Mertsch, K.; et al. Metabolomic and genome-wide association studies reveal potential endogenous biomarkers for OATP1B1. Clin. Pharmacol. Ther. 2016, 100, 524–536. [Google Scholar] [CrossRef]
- Yousri, N.A.; Fakhro, K.A.; Robay, A.; Rodriguez-Flores, J.L.; Mohney, R.P.; Zeriri, H.; Odeh, T.; Kader, S.A.; Aldous, E.K.; Thareja, G.; et al. Whole-exome sequencing identifies common and rare variant metabolic QTLs in a Middle Eastern population. Nat. Commun. 2018, 9, 333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wanders, R.J.; Komen, J.; Kemp, S. Fatty acid omega-oxidation as a rescue pathway for fatty acid oxidation disorders in humans. FEBS J. 2011, 278, 182–194. [Google Scholar] [CrossRef] [PubMed]
- Kelson, T.L.; Secor McVoy, J.R.; Rizzo, W.B. Human liver fatty aldehyde dehydrogenase: Microsomal localization, purification, and biochemical characterization. Biochim. Biophys. Acta 1997, 1335, 99–110. [Google Scholar] [CrossRef] [PubMed]
- Reddy, J.K. Nonalcoholic steatosis and steatohepatitis III. Peroxisomal β-oxidation, PPARα, and steatohepatitis. Am. J. Physiol. Gastrointest. 2001, 281, G1333–G1339. [Google Scholar] [CrossRef]
- Mortensen, P.B. Mechanisms of dicarboxylic aciduria and dicarboxylic acid metabolism. In Fatty Acid Oxidation: Clinical, Biochemical and Molecular Aspects; Alan R. Liss, Inc.: New York, NY, USA, 1990; pp. 249–257. [Google Scholar]
- Park, H.-S.; Jang, J.E.; Ko, M.S.; Woo, S.H.; Kim, B.J.; Kim, H.S.; Park, H.S.; Park, I.-S.; Koh, E.H.; Lee, K.-U. Statins Increase Mitochondrial and Peroxisomal Fatty Acid Oxidation in the Liver and Prevent Non-Alcoholic Steatohepatitis in Mice. Diabetes Metab. J. 2016, 40, 376–385. [Google Scholar] [CrossRef] [Green Version]
- Pasanen, M.K.; Neuvonen, M.; Neuvonen, P.J.; Niemi, M. SLCO1B1 polymorphism markedly affects the pharmacokinetics of simvastatin acid. Pharm. Genom. 2006, 16, 873–879. [Google Scholar] [CrossRef] [PubMed]
- Ramsey, L.B.; Bruun, G.H.; Yang, W.; Treviño, L.R.; Vattathil, S.; Scheet, P.; Cheng, C.; Rosner, G.L.; Giacomini, K.M.; Fan, Y.; et al. Rare versus common variants in pharmacogenetics: SLCO1B1 variation and methotrexate disposition. Genome Res. 2012, 22, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Goldstein, J.L.; Brown, M.S. The LDL receptor. Arterioscler. Thromb. Vasc. Biol. 2009, 29, 431–438. [Google Scholar] [CrossRef] [Green Version]
- Xu, X.; Zhang, A.; Halquist, M.S.; Yuan, X.; Henderson, S.C.; Dewey, W.L.; Li, P.L.; Li, N.; Zhang, F. Simvastatin promotes NPC1-mediated free cholesterol efflux from lysosomes through CYP7A1/LXRα signalling pathway in oxLDL-loaded macrophages. J. Cell. Mol. Med. 2017, 21, 364–374. [Google Scholar] [CrossRef]
- Fu, Z.D.; Cui, J.Y.; Klaassen, C.D. Atorvastatin induces bile acid-synthetic enzyme Cyp7a1 by suppressing FXR signaling in both liver and intestine in mice. J. Lipid Res. 2014, 55, 2576–2586. [Google Scholar] [CrossRef] [Green Version]
- Rizzolo, D.; Buckley, K.; Kong, B.; Zhan, L.; Shen, J.; Stofan, M.; Brinker, A.; Goedken, M.; Buckley, B.; Guo, G.L. Bile acid homeostasis in a cholesterol 7α-hydroxylase and sterol 27-hydroxylase double knockout mouse model. Hepatology 2019, 70, 389–402. [Google Scholar] [CrossRef]
- Chambers, K.F.; Day, P.E.; Aboufarrag, H.T.; Kroon, P.A. Polyphenol effects on cholesterol metabolism via bile acid biosynthesis, CYP7A1: A review. Nutrients 2019, 11, 2588. [Google Scholar] [CrossRef] [Green Version]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Hillard, C.J. Circulating endocannabinoids: From whence do they come and where are they coing? Neuropsychopharmacol. Rev. 2018, 43, 155–172. [Google Scholar] [CrossRef] [Green Version]
- Chiang, J.Y. Bile acid metabolism and signaling. Compr. Physiol. 2013, 3, 1191–1212. [Google Scholar]
- Hagenbuch, B.; Gui, C. OATP1B1 transports steroid sulfates and phase II sulfate conjugates Xenobiotic transporters of the human organic anion transporting polypeptides (OATP) family. Xenobiotica 2008, 38, 778–801. [Google Scholar] [CrossRef]
- Tóth, B.; Jani, M.; Beéry, E.; Heslop, T.; Bayliss, M.; Kitteringham, N.R.; Park, B.K.; Weaver, R.J.; Krajcsi, P. Human OATP1B1 (SLCO1B1) transports sulfated bile acids and bile salts with particular efficiency. Toxicol. In Vitro 2018, 52, 189–194. [Google Scholar] [CrossRef]
- Kalliokoski, A.; Niemi, M. Impact of OATP transporters on pharmacokinetics. Br. J. Pharmacol. 2009, 158, 693–705. [Google Scholar] [CrossRef] [PubMed]

| Clinical and Laboratory Characteristics | Participants without Statin Treatment | Participants on Simvastatin Treatment | p * | ||||
|---|---|---|---|---|---|---|---|
| rs4149056 TT (n = 814) | rs4149056 CC + CT (n = 554) | rs4149056 TT (n = 841) | rs4149056 CC + CT (n = 532) | ||||
| Mean ± SD | Mean ± SD | p | Mean ± SD | Mean ± SD | p | ||
| Age (years) | 59.65 ± 6.74 | 58.81 ± 7.04 | 0.028 | 59.53 ± 7.1 | 59.67 ± 7.14 | 0.707 | 0.046 |
| Body mass index (kg/m2) | 27.32 ± 3.73 | 27.13 ± 3.85 | 0.300 | 27.29 ± 3.74 | 27.51 ± 4.11 | 0.387 | 0.112 |
| Waist (cm) | 98.51 ± 10.43 | 97.57 ± 10.37 | 0.096 | 98.81 ± 10.55 | 99.31 ± 11.03 | 0.430 | 0.007 |
| Systolic blood pressure (mmHg) | 140.99 ± 16.77 | 140.9 ± 16.82 | 0.907 | 137.5 ± 16.16 | 138.66 ± 15.34 | 0.143 | 0.022 |
| Fasting plasma glucose (mmol/L) | 5.66 ± 0.51 | 5.63 ± 0.49 | 0.309 | 5.78 ± 0.47 | 5.80 ± 0.49 | 0.336 | 3.5 × 10−9 |
| 2 h plasma glucose (mmol/L) | 6.14 ± 1.7 | 6.06 ± 1.76 | 0.329 | 6.46 ± 1.74 | 6.35 ± 1.75 | 0.265 | 0.005 |
| LDL cholesterol (mmol/L) | 3.62 ± 0.8 | 3.59 ± 0.79 | 0.522 | 2.68 ± 0.72 | 2.74 ± 0.71 | 0.133 | 9.1 × 10−66 |
| Total triglycerides (mmol/L) | 1.41 ± 0.78 | 1.52 ± 1.79 | 0.365 | 1.41 ± 0.73 | 1.43 ± 0.73 | 0.360 | 0.320 |
| ALT (U/L) | 29.21 ± 17.63 | 29.32 ± 22.24 | 0.616 | 32.29 ± 18.06 | 32.89 ± 16.69 | 0.385 | 0.003 |
| Fasting plasma insulin (mU/L) | 8.55 ± 6.31 | 8.0 ± 5.53 | 0.067 | 9.53 ± 6.98 | 9.42 ± 6.57 | 0.870 | 0.0001 |
| Fasting plasma proinsulin (pmol/L) | 13.88 ± 6.92 | 13.43 ± 6.1 | 0.241 | 15.04 ± 7.49 | 14.85 ± 7.52 | 0.454 | 0.0006 |
| hS-CRP (mg/L) | 2.42 ± 3.22 | 2.57 ± 3.66 | 0.145 | 1.99 ± 3.87 | 1.72 ± 2.18 | 0.825 | 5.0 × 10−06 |
| Matsuda ISI comp (mg/dL. mU/L) | 6.9 ± 4.28 | 7.34 ± 4.41 | 0.061 | 5.74 ± 3.58 | 5.73 ± 3.37 | 0.776 | 3.0 × 10−11 |
| Disposition index | 162.05 ± 76.56 | 167.35 ± 88.51 | 0.228 | 154.15 ± 66.96 | 153.99 ± 65.18 | 0.818 | 0.005 |
| No Statin | Simvastatin | Novel (Ref.) | |||||
| Metabolite | Beta | p | Beta | p | Subclass | p comp | |
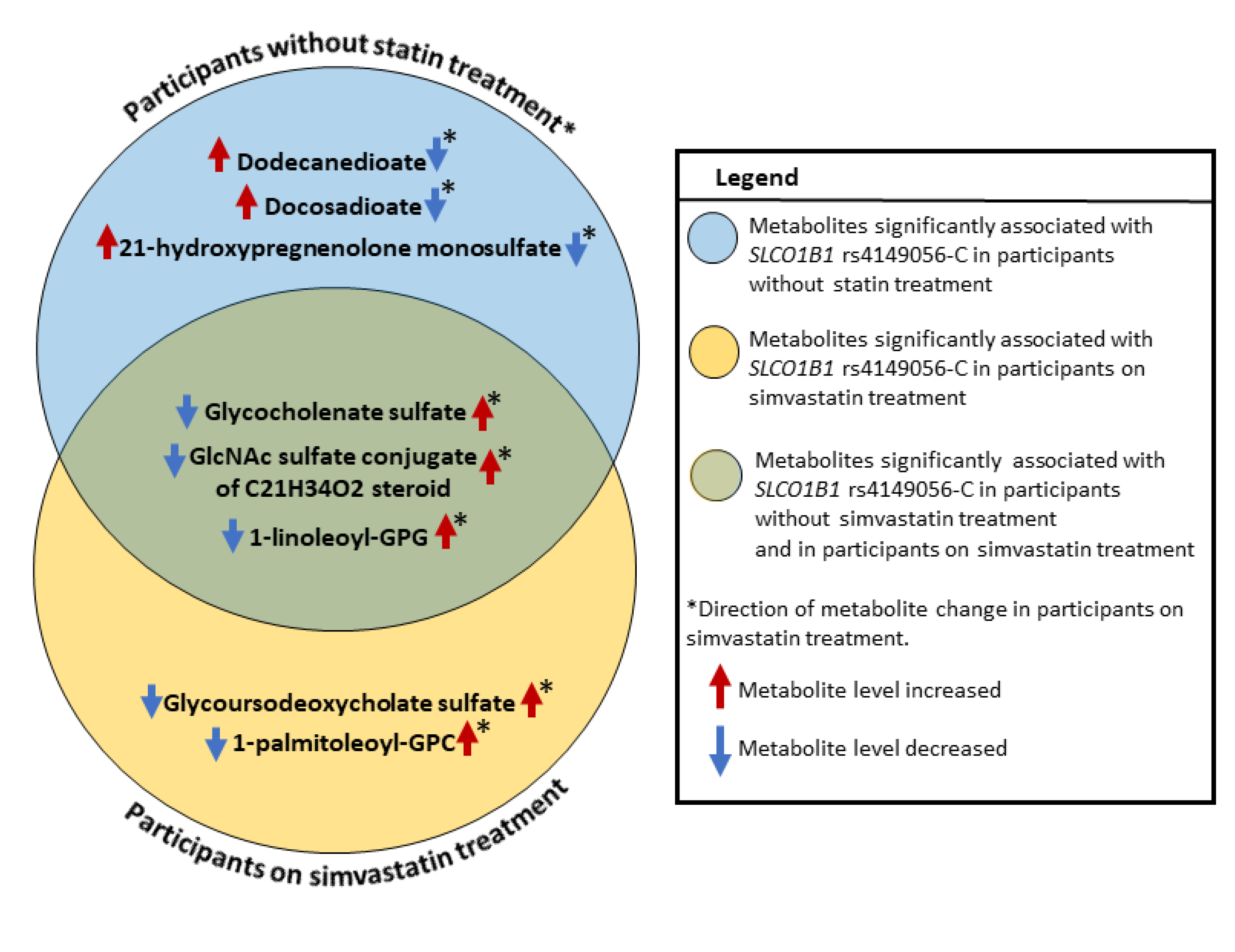
| 21-hydroxypregnenolone monosulfate (1) | 0.211 | 1.9 × 10−6 | 0.112 | 0.003 | Pregnenolone Steroids | 0.008 | Yes |
| Docosadioate (C22-DC) | 0.110 | 4.7 × 10−5 | 0.025 | 0.351 | Fatty Acid, Dicarboxylate | 0.026 | Yes |
| Dodecanedioate (C12-DC) | 0.115 | 1.9 × 10−5 | 0.012 | 0.659 | Fatty Acid, Dicarboxylate | 0.006 | Yes |
| Comparison between statistically significant associations of SLCO1B1 rs4149056-C with metabolites in participants with simvastatin treatment (n = 1368) and not statistically significant in participants with simvastatin treatment (n = 1373) | |||||||
| No Statin | Simvastatin | ||||||
| Metabolite | Beta | p | Beta | p | Subclass | p comp | |
| Glycoursodeoxycholic acid sulfate (1) | 0.036 | 0.307 | 0.174 | 1.4 × 10−7 | Secondary Bile Acid Metabolism | 0.0003 | Yes |
| 1-palmitoleoyl-GPC (16:1) | 0.029 | 0.280 | 0.111 | 3.5 × 10−5 | Lysophospholipid | 0.031 | No (16) |
| Comparison between statistically significant associations of SLCO1B1 rs4149056-C with metabolites in participants without statin treatment (n = 1368) and in participants with simvastatin treatment (n = 1373) | |||||||
| No Statin | Simvastatin | ||||||
| Metabolite | Beta | p | Beta | p | Subclass | p comp | |
| Glycocholenate sulfate | 0.352 | 4.1 × 10−41 | 0.416 | 1.4 × 10−58 | Secondary Bile Acid Metabolism | 0.049 | No (16.35) |
| GlcNAc sulfate conjugate of C21H34O2 steroid | 0.293 | 1.2 × 10−18 | 0.389 | 8.2 × 10−36 | Partially Characterized Molecules | 0.005 | Yes |
| 1-linoleoyl-GPG (18:2) | 0.189 | 1.8 × 10−8 | 0.284 | 2.5 × 10−22 | Lysophospholipid | 0.009 | Yes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernandes Silva, L.; Ravi, R.; Vangipurapu, J.; Oravilahti, A.; Laakso, M. Effects of SLCO1B1 Genetic Variant on Metabolite Profile in Participants on Simvastatin Treatment. Metabolites 2022, 12, 1159. https://doi.org/10.3390/metabo12121159
Fernandes Silva L, Ravi R, Vangipurapu J, Oravilahti A, Laakso M. Effects of SLCO1B1 Genetic Variant on Metabolite Profile in Participants on Simvastatin Treatment. Metabolites. 2022; 12(12):1159. https://doi.org/10.3390/metabo12121159
Chicago/Turabian StyleFernandes Silva, Lilian, Rowmika Ravi, Jagadish Vangipurapu, Anniina Oravilahti, and Markku Laakso. 2022. "Effects of SLCO1B1 Genetic Variant on Metabolite Profile in Participants on Simvastatin Treatment" Metabolites 12, no. 12: 1159. https://doi.org/10.3390/metabo12121159
APA StyleFernandes Silva, L., Ravi, R., Vangipurapu, J., Oravilahti, A., & Laakso, M. (2022). Effects of SLCO1B1 Genetic Variant on Metabolite Profile in Participants on Simvastatin Treatment. Metabolites, 12(12), 1159. https://doi.org/10.3390/metabo12121159






