Interrogating the Metabolomic Profile of Amyotrophic Lateral Sclerosis in the Post-Mortem Human Brain by Infrared Matrix-Assisted Laser Desorption Electrospray Ionization (IR-MALDESI) Mass Spectrometry Imaging (MSI)
Abstract
1. Introduction
2. Materials and Methods
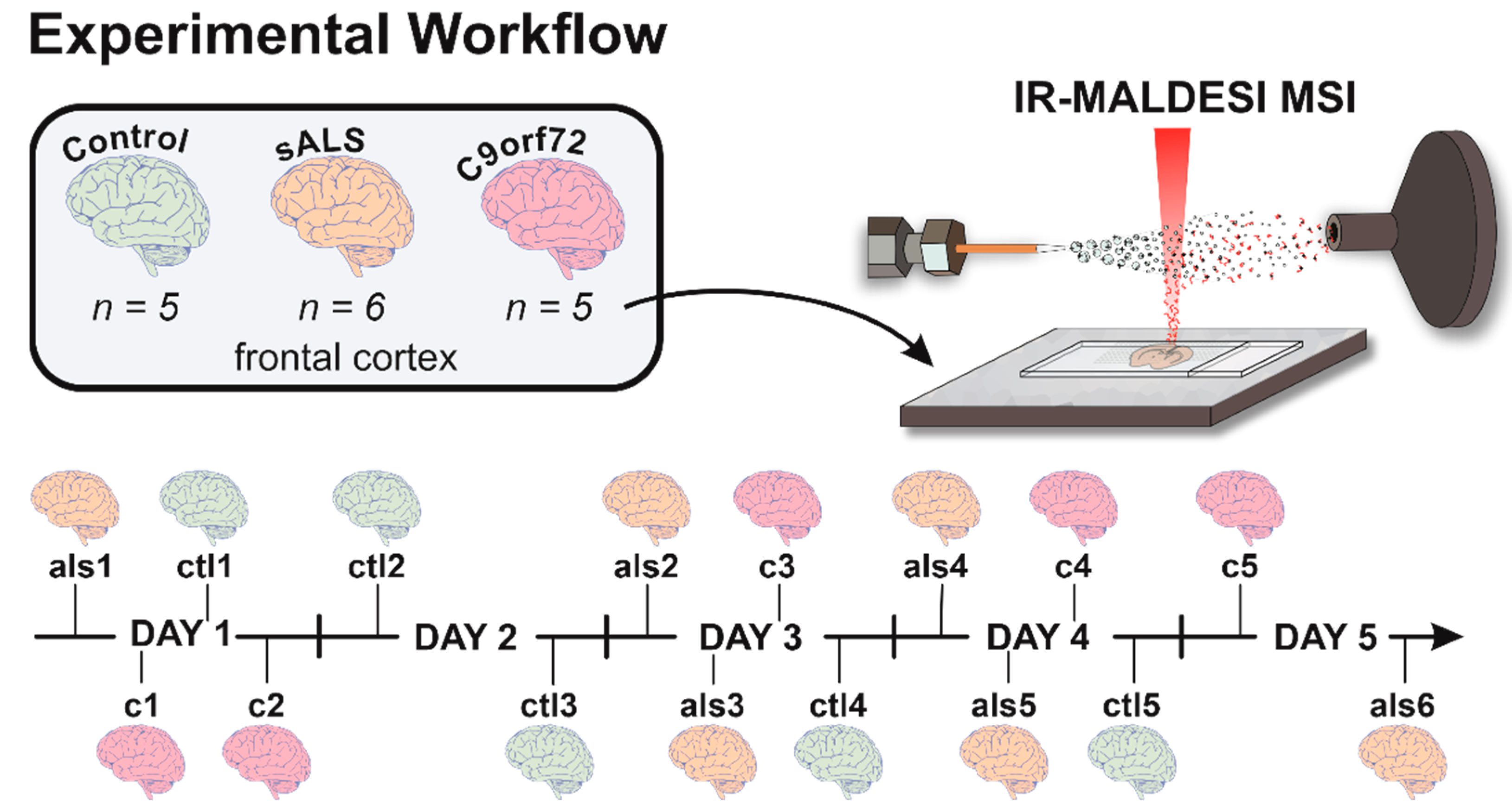
2.1. Case Details and Sample Preparation
2.2. IR-MALDESI MSI
2.3. IR-MALDESI MSI Data Analysis
2.4. Metabolomic Pathway Analysis and Proteomic Integration
2.5. Statistical Testing and Analysis
3. Results and Discussion
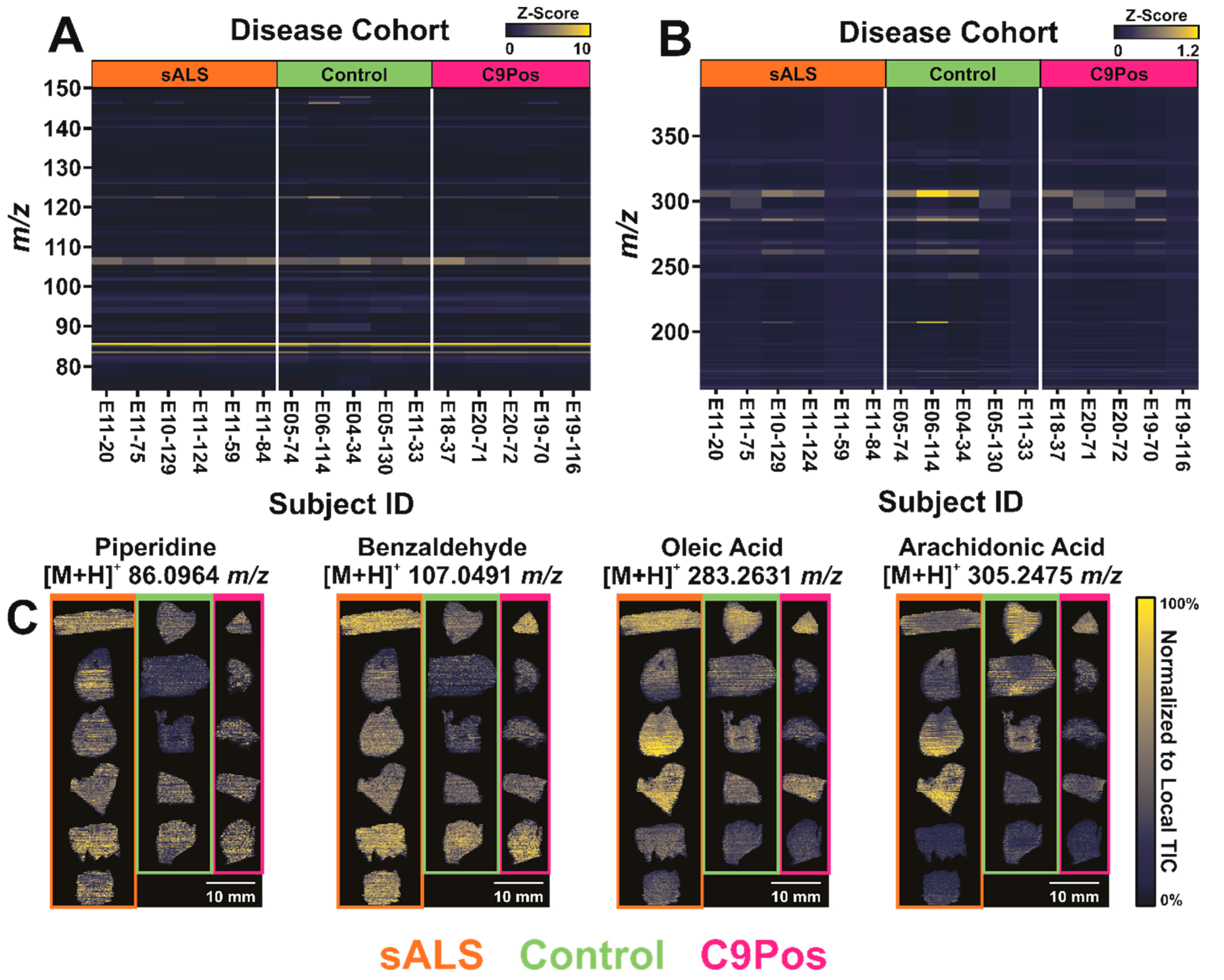
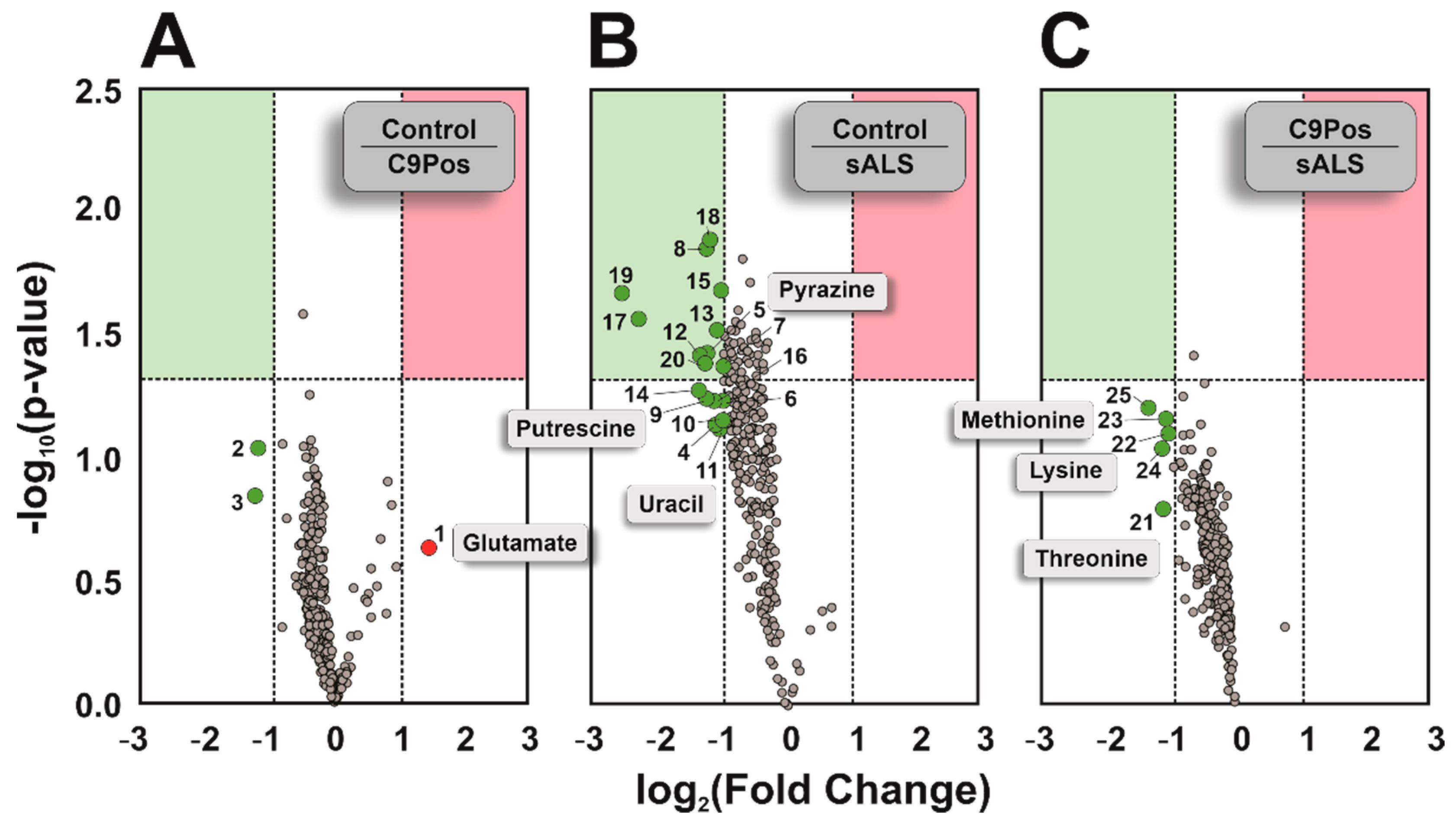
3.1. Comparing Metabolomic Detection between ALS Cohorts
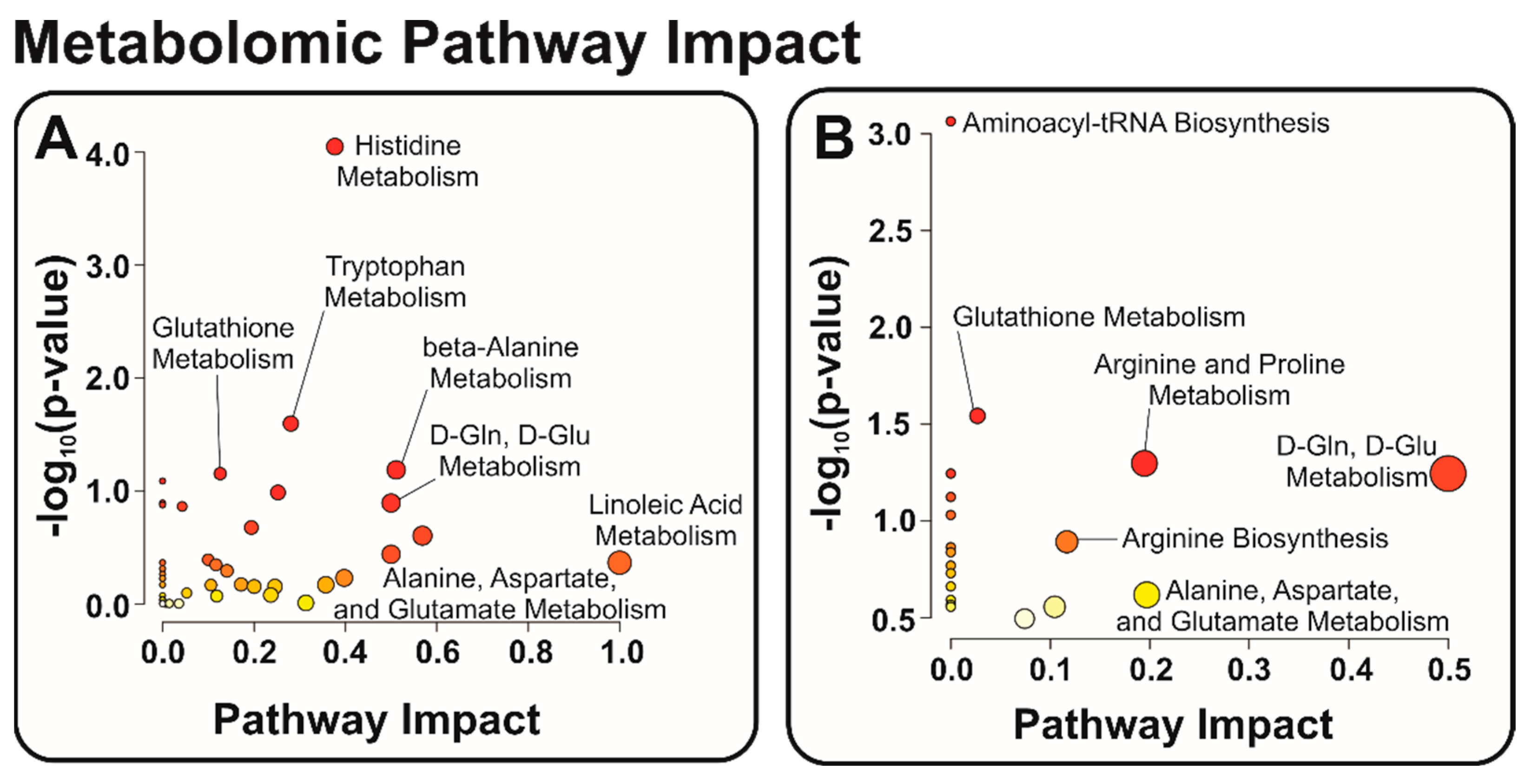
3.2. Identification of Relevant Metabolic Pathways
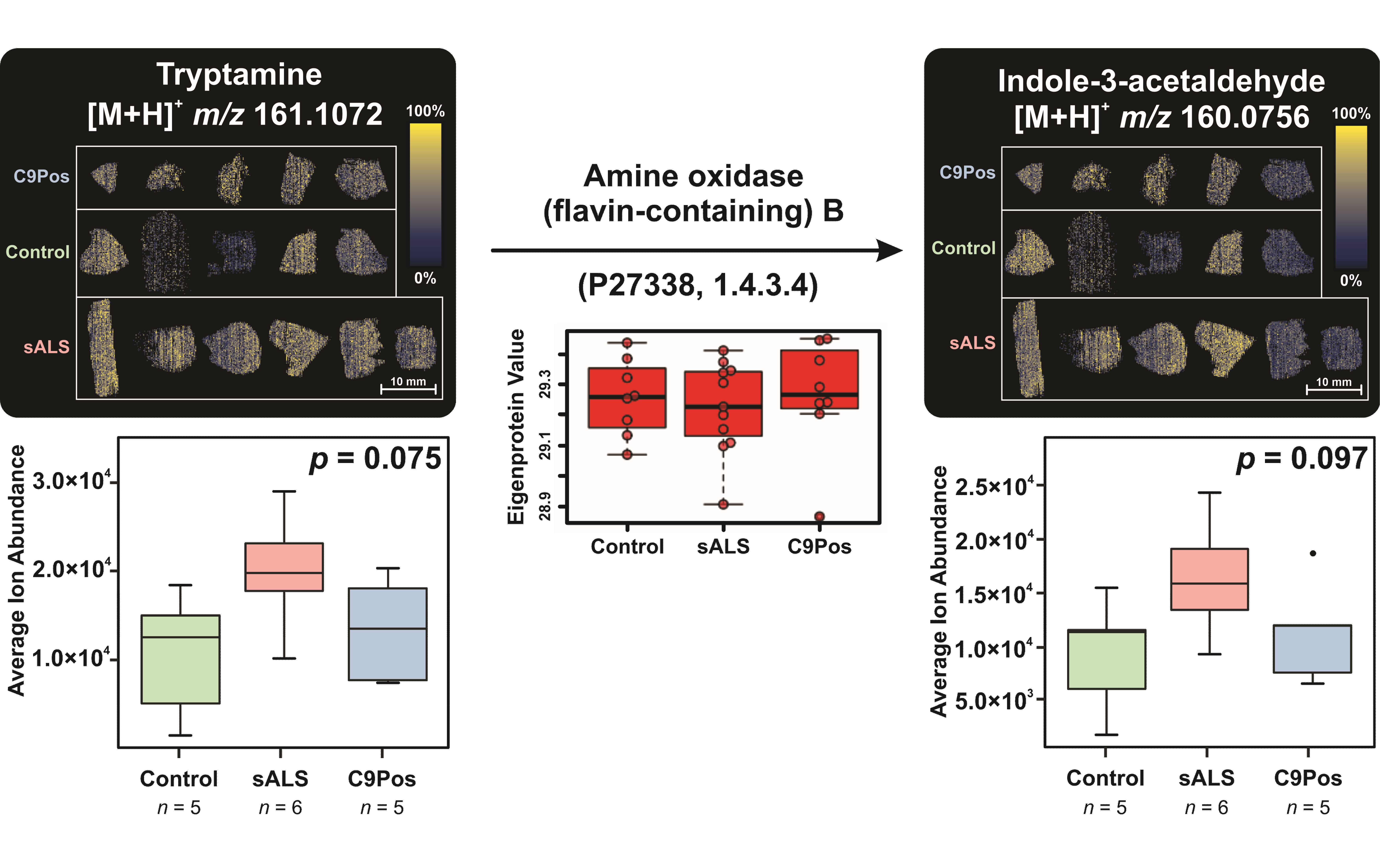
3.3. Multiomic Integration
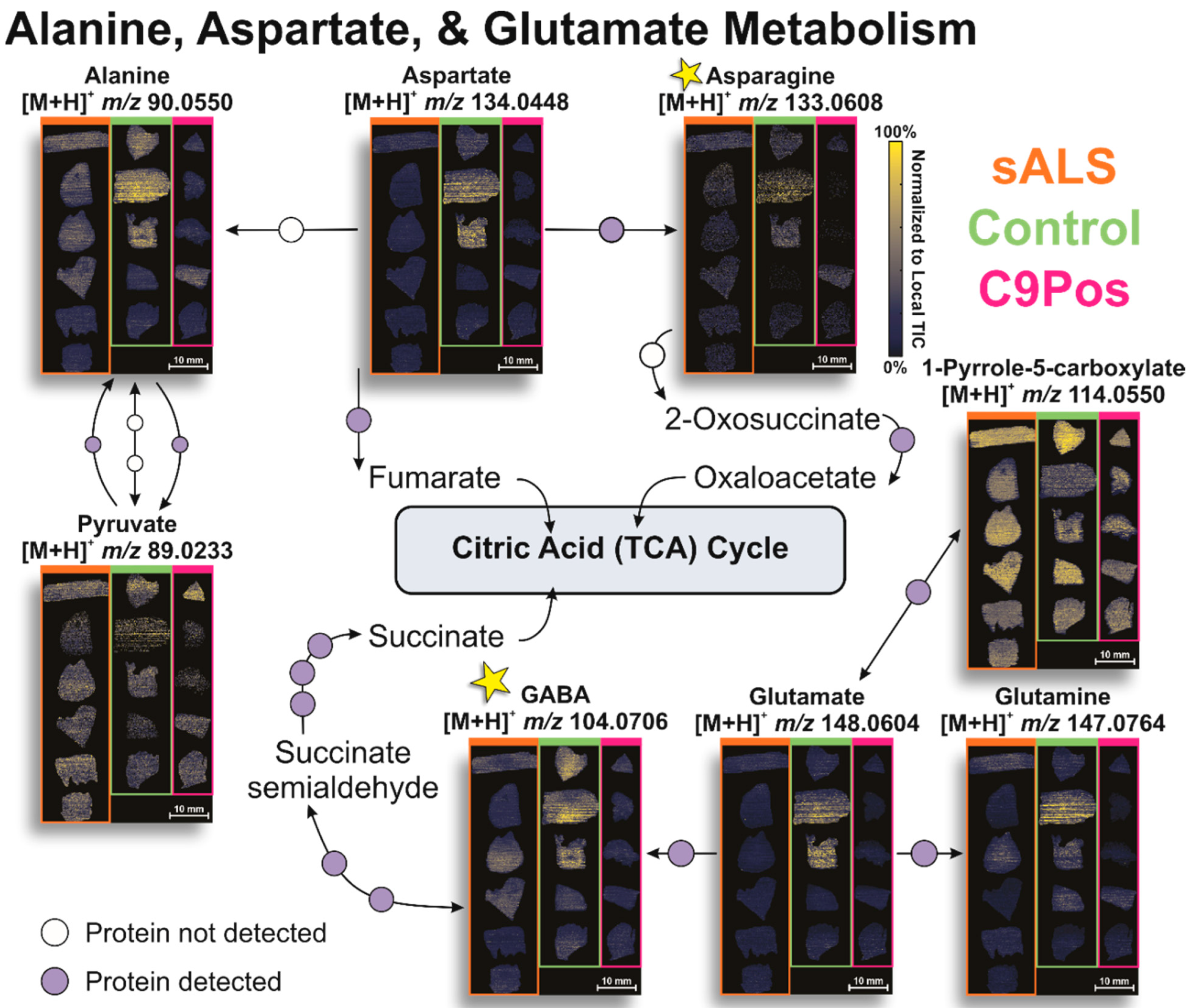
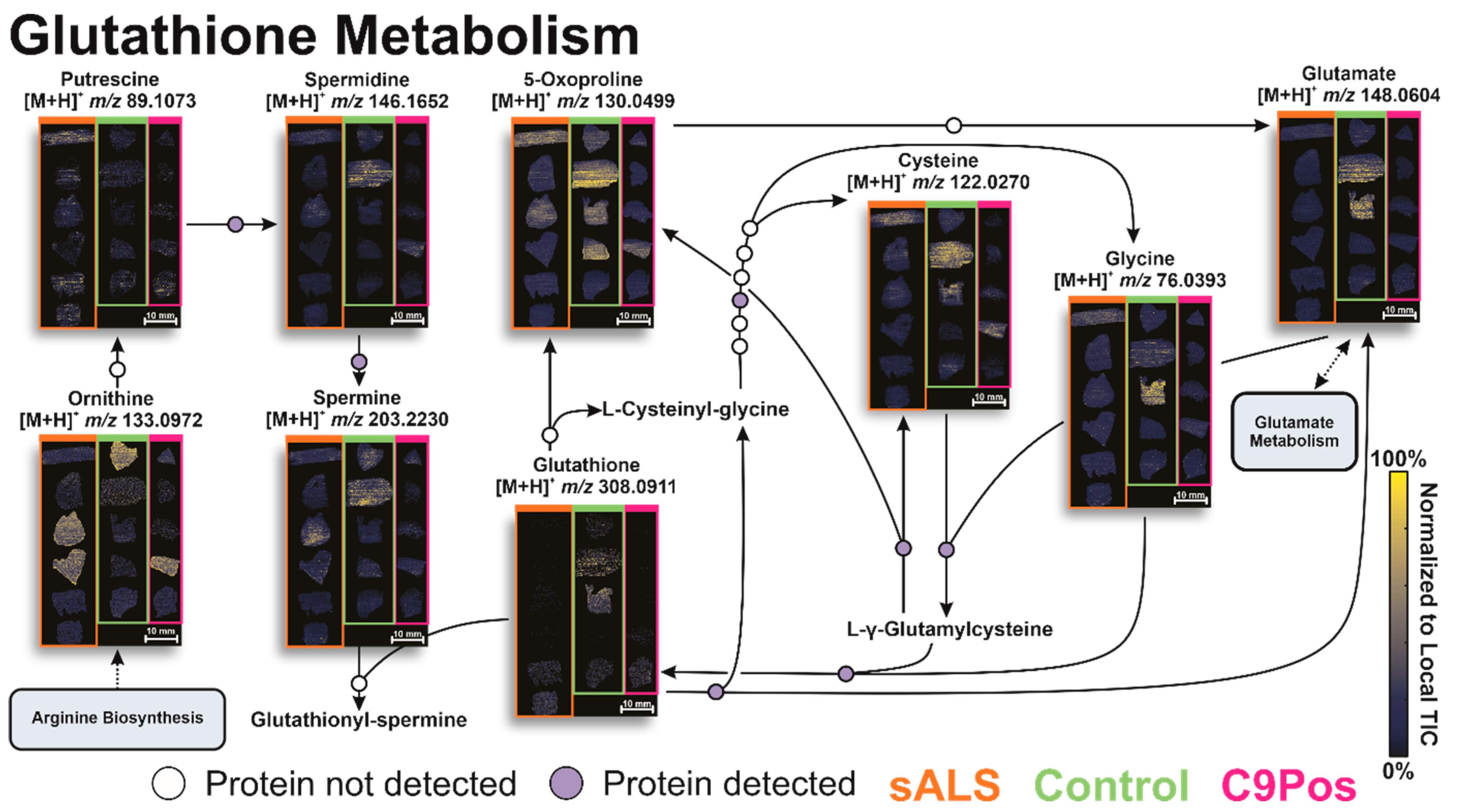
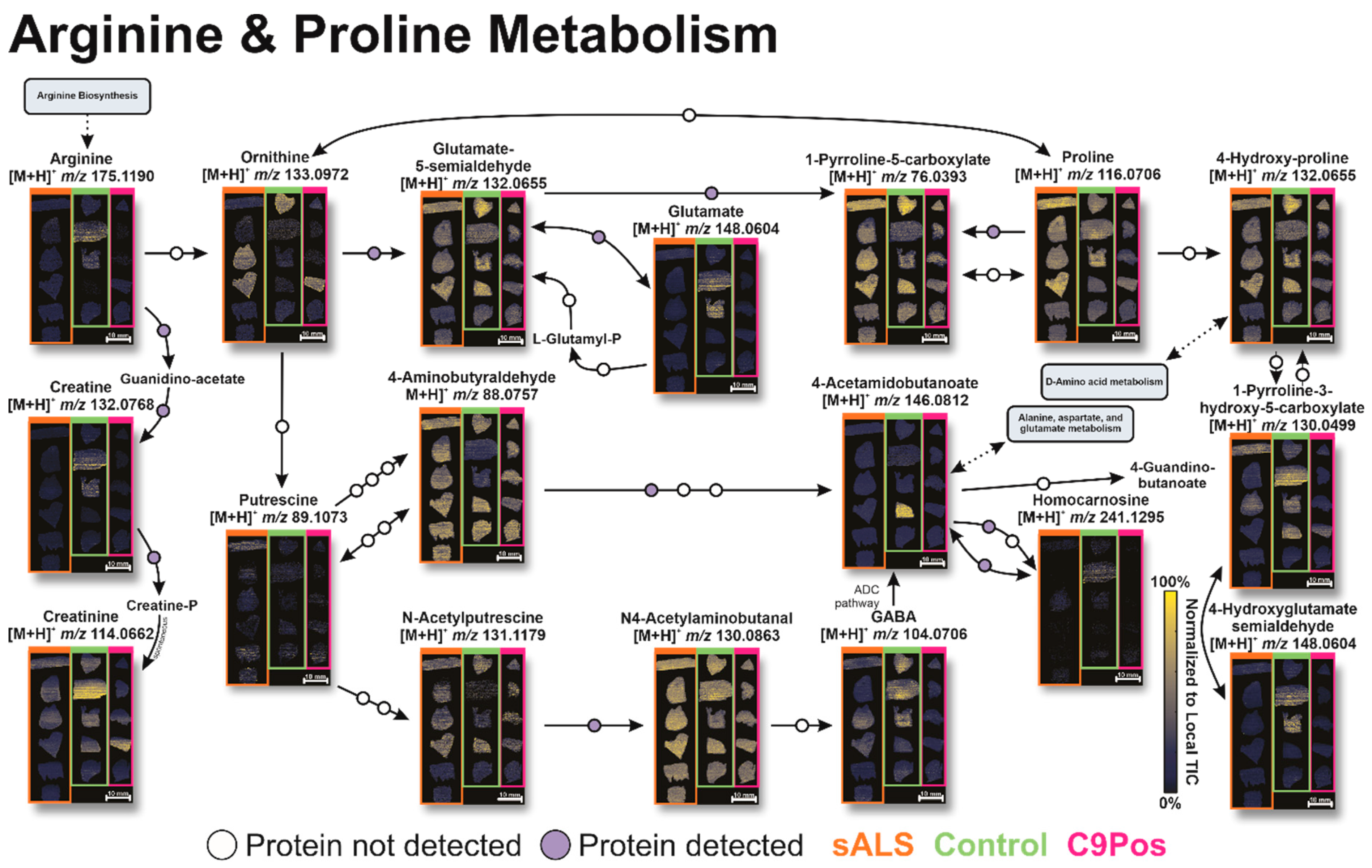
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mejzini, R.; Flynn, L.L.; Pitout, I.L.; Fletcher, S.; Wilton, S.D.; Akkari, P.A. ALS Genetics; Mechanisms, and Therapeutics: Where Are We Now? Front. Neurosci. 2019, 13, 1310. [Google Scholar] [CrossRef]
- Masrori, P.; Van Damme, P. Amyotrophic Lateral Sclerosis: A Clinical Review. Eur. J. Neurol. 2020, 27, 1918–1929. [Google Scholar] [CrossRef] [PubMed]
- Garbuzova-Davis, S.; Rodrigues, M.C.O.; Hernandez-Ontiveros, D.G.; Louis, M.K.; Willing, A.E.; Borlongan, C.V.; Sanberg, P.R. Amyotrophic Lateral Sclerosis: A Neurovascular Disease. Brain Res. 2011, 1398, 113–125. [Google Scholar] [CrossRef] [PubMed]
- Chiò, A.; Logroscino, G.; Traynor, B.J.; Collins, J.; Simeone, J.C.; Goldstein, L.A.; White, L.A. Global Systematic Review Global Epidemiology of Amyotrophic Lateral Sclerosis: A Systematic Review of the Published Literature. Neuroepidemiology 2013, 41, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Wijesekera, L.C.; Leigh, N. Amyotrophic Lateral Sclerosis. Orphanet J. Rare Dis. 2009, 4, 3. [Google Scholar] [CrossRef] [PubMed]
- Rothstein, J.D. Edaravone: A New Drug Approved for ALS. Cell 2017, 171, 725. [Google Scholar] [CrossRef]
- Mathis, S.; Goizet, C.; Soulages, A.; Vallat, J.M.; Le Masson, G. Genetics of Amyotrophic Lateral Sclerosis: A Review. J. Neurol. Sci. 2019, 399, 217–226. [Google Scholar] [CrossRef]
- Mathis, S.; Couratier, P.; Julian, A.; Vallat, J.M.; Corcia, P.; Le Masson, G. Management and Therapeutic Perspectives in Amyotrophic Lateral Sclerosis. Expert Rev. Neurother. 2017, 17, 263–276. [Google Scholar] [CrossRef]
- Yang, Q.; Jiao, B.; Shen, L. The Development of C9orf72-Related Amyotrophic Lateral Sclerosis and Frontotemporal Dementia Disorders. Front. Genet. 2020, 11, 1045. [Google Scholar] [CrossRef]
- Burberry, A.; Wells, M.F.; Limone, F.; Couto, A.; Smith, K.S.; Keaney, J.; Gillet, G.; van Gastel, N.; Wang, J.Y.; Pietilainen, O.; et al. C9orf72 Suppresses Systemic and Neural Inflammation Induced by Gut Bacteria. Nature 2020, 582, 89–94. [Google Scholar] [CrossRef]
- Haeusler, A.R.; Donnelly, C.J.; Periz, G.; Simko, E.A.J.; Shaw, P.G.; Kim, M.S.; Maragakis, N.J.; Troncoso, J.C.; Pandey, A.; Sattler, R.; et al. C9orf72 Nucleotide Repeat Structures Initiate Molecular Cascades of Disease. Nature 2014, 507, 195–200. [Google Scholar] [CrossRef] [PubMed]
- Mann, D.M.; Rollinson, S.; Robinson, A.; Callister, J.B.; Thompson, J.C.; Snowden, J.S.; Gendron, T.; Petrucelli, L.; Masuda-Suzukake, M.; Hasegawa, M.; et al. Dipeptide Repeat Proteins Are Present in the P62 Positive Inclusions in Patients with Frontotemporal Lobar Degeneration and Motor Neurone Disease Associated with Expansions in C9ORF72. Acta Neuropathol. Commun. 2013, 1, 68. [Google Scholar] [CrossRef] [PubMed]
- Mori, K.; Arzberger, T.; Grässer, F.A.; Gijselinck, I.; May, S.; Rentzsch, K.; Weng, S.M.; Schludi, M.H.; Van Der Zee, J.; Cruts, M.; et al. Bidirectional Transcripts of the Expanded C9orf72 Hexanucleotide Repeat Are Translated into Aggregating Dipeptide Repeat Proteins. Acta Neuropathol. 2013, 126, 881–893. [Google Scholar] [CrossRef] [PubMed]
- Freibaum, B.D.; Lu, Y.; Lopez-Gonzalez, R.; Kim, N.C.; Almeida, S.; Lee, K.H.; Badders, N.; Valentine, M.; Miller, B.L.; Wong, P.C.; et al. GGGGCC Repeat Expansion in C9orf72 Compromises Nucleocytoplasmic Transport. Nature 2015, 525, 129–133. [Google Scholar] [CrossRef]
- Boeynaems, S.; Bogaert, E.; Kovacs, D.; Konijnenberg, A.; Timmerman, E.; Volkov, A.; Guharoy, M.; De Decker, M.; Jaspers, T.; Ryan, V.H.; et al. Phase Separation of C9orf72 Dipeptide Repeats Perturbs Stress Granule Dynamics. Mol. Cell. 2017, 65, 1044–1055. [Google Scholar] [CrossRef]
- Chen, L.; Liu, B. Relationships between Stress Granules; Oxidative Stress, and Neurodegenerative Diseases. Oxidative Med. Cell. Longev. 2017, 2017, 1809592. [Google Scholar] [CrossRef]
- Borroni, B.; Premi, E.; Formenti, A.; Turrone, R.; Alberici, A.; Cottini, E.; Rizzetti, C.; Gasparotti, R.; Padovani, A. Structural and Functional Imaging Study in Dementia with Lewy Bodies and Parkinson’s Disease Dementia. Park. Relat. Disord. 2015, 21, 1049–1055. [Google Scholar] [CrossRef]
- Chollet, F.; Payoux, P. Functional Imaging for Neurodegenerative Diseases. Press. Med. 2022, 51, 104121. [Google Scholar] [CrossRef]
- Gaetani, L.; Paolini Paoletti, F.; Bellomo, G.; Mancini, A.; Simoni, S.; Di Filippo, M.; Parnetti, L. CSF and Blood Biomarkers in Neuroinflammatory and Neurodegenerative Diseases: Implications for Treatment. Trends Pharmacol. Sci. 2020, 41, 1023–1037. [Google Scholar] [CrossRef]
- van den Bos, M.A.J.; Geevasinga, N.; Higashihara, M.; Menon, P.; Vucic, S. Pathophysiology and Diagnosis of ALS: Insights from Advances in Neurophysiological Techniques. Int. J. Mol. Sci. 2019, 20, 2818. [Google Scholar] [CrossRef]
- Brooks, B.R.; Miller, R.G.; Swash, M.; Munsat, T.L. El Escorial revisited: Revised criteria for the diagnosis of amyotrophic lateral sclerosis. Amyotroph. Lateral Scler. Front. Degener. 2009, 1, 293–299. [Google Scholar] [CrossRef]
- Geevasinga, N.; Loy, C.T.; Menon, P.; de Carvalho, M.; Swash, M.; Schrooten, M.; Van Damme, P.; Gawel, M.; Sonoo, M.; Higashihara, M.; et al. Awaji criteria improves the diagnostic sensitivity in amyotrophic lateral sclerosis: A systematic review using individual patient data. Clin. Neurophysiol. 2016, 127, 2684–2691. [Google Scholar] [CrossRef] [PubMed]
- Cellura, E.; Spataro, R.; Taiello, A.C.; La Bella, V. Factors affecting the diagnostic delay in amyotrophic lateral sclerosis. Clin. Neurol. Neurosurg. 2012, 114, 550–554. [Google Scholar] [CrossRef]
- Nzwalo, H.; De Abreu, D.; Swash, M.; Pinto, S.; De Carvalho, M. Delayed diagnosis in ALS: The problem continues. J. Neurol. Sci. 2014, 343, 173–175. [Google Scholar] [CrossRef]
- Kori, M.; Aydln, B.; Unal, S.; Arga, K.Y.; Kazan, D. Metabolic Biomarkers and Neurodegeneration: A Pathway Enrichment Analysis of Alzheimer’s Disease; Parkinson’s Disease, and Amyotrophic Lateral Sclerosis. OMICS 2016, 20, 645–661. [Google Scholar] [CrossRef]
- Bakkar, N.; Boehringer, A.; Bowser, R. Use of biomarkers in ALS drug development and clinical trials. Brain Res. 2015, 1607, 94–107. [Google Scholar] [CrossRef] [PubMed]
- Schmacher-Schuh, A.; Bieger, A.; Borelli, W.V.; Portley, M.K.; Awad, P.S.; Bandres-Ciga, S. Advances in proteomic and metabolomic profiling of neurodegenerative diseases. Front. Neurol. 2022, 12, 792227. [Google Scholar] [CrossRef] [PubMed]
- Katzeff, J.S.; Bright, F.; Phan, K.; Kril, J.J.; Ittner, L.M.; Kassiou, M.; Hodges, J.R.; Piguet, O.; Kiernan, M.C.; Halliday, G.M.; et al. Biomarker discovery and development for frontotemporal dementia and amyotrophic lateral sclerosis. Brain 2022, 145, 1598–1609. [Google Scholar] [CrossRef]
- Zhang, A.; Sun, H.; Yan, G.; Wang, P.; Wang, X. Review Article Metabolomics for Biomarker Discovery: Moving to the Clinic. Biomed. Res. J. 2014, 2015, 354671. [Google Scholar] [CrossRef]
- Crutchfield, C.A.; Thomas, S.N.; Sokoll, L.J.; Chan, D.W. Advances in mass spectrometry-based clinical biomarker discovery. Clin. Proteom. 2014, 13, 1. [Google Scholar] [CrossRef]
- Dueñas, M.E.; Larson, E.A.; Lee, Y.J. Toward mass spectrometry imaging in the metabolomics scale: Increasing metabolic coverage through multiple on-tissue chemical modifications. Front. Plant Sci. 2019, 10, 860. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Li, J.; Jianjun Li, C. Recent advances in mass spectrometry imaging for multiomics application in neurology. J. Comp. Neurol. 2019, 527, 2158–2169. [Google Scholar] [CrossRef] [PubMed]
- Buchberger, A.R.; Delaney, K.; Johnson, J.; Li, L. Mass Spectrometry Imaging: A Review of Emerging Advancements and Future Insights. Anal. Chem. 2017, 90, 240–265. [Google Scholar] [CrossRef]
- Jang, H.J.; Le, M.U.T.; Park, J.H.; Chung, C.G.; Shon, J.G.; Lee, G.S.; Moon, J.H.; Lee, S.B.; Choi, J.S.; Lee, T.G.; et al. Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry Imaging of Phospholipid Changes in a Drosophila Model of Early Amyotrophic Lateral Sclerosis. J. Am. Soc. Mass Spectrom. 2021, 32, 2536–2545. [Google Scholar] [CrossRef]
- Hanrieder, J.; Ewing, A.G. Spatial Elucidation of Spinal Cord Lipid- and Metabolite-Regulations in Amyotrophic Lateral Sclerosis. Sci. Rep. 2014, 4, 5266. [Google Scholar] [CrossRef] [PubMed]
- Bagley, M.C.; Garrard, K.P.; Muddiman, D.C. The development and application of matrix assisted laser desorption electrospray ionization: The teenage years. Mass Spectrom. Rev. 2021, 1–32. [Google Scholar] [CrossRef]
- Bokhart, M.T.; Muddiman, D.C. Infrared matrix-assisted laser desorption electrospray ionization mass spectrometry imaging analysis of biospecimens. Analyst 2016, 141, 5236. [Google Scholar] [CrossRef]
- Dixon, R.B.; Muddiman, D.C. Study of the Ionization Mechanism in Hybrid Laser Based Desorption Techniques. Analyst 2010, 135, 880–882. [Google Scholar] [CrossRef]
- Robichaud, G.; Barry, J.A.; Muddiman, D.C. IR-MALDESI Mass Spectrometry Imaging of Biological Tissue Sections Using Ice as a Matrix. J. Am. Soc. Mass Spectrom. 2014, 25, 319–328. [Google Scholar] [CrossRef]
- Tu, A.; Muddiman, D.C. Internal Energy Deposition in Infrared Matrix-Assisted Laser Desorption Electrospray Ionization With and Without the Use of Ice as a Matrix. J. Am. Soc. Mass Spectrom. 2019, 30, 2380–2391. [Google Scholar] [CrossRef]
- Pace, C.L.; Horman, B.; Patisaul, H.; Muddiman, D.C. Analysis of neurotransmitters in rat placenta exposed to flame retardants using IR-MALDESI mass spectrometry imaging. Anal. Bioanal. Chem. 2020, 412, 3745–3752. [Google Scholar] [CrossRef] [PubMed]
- Bai, H.; Linder, K.E.; Muddiman, D.C. Three-dimensional (3D) imaging of lipids in skin tissues with infrared matrix-assisted laser desorption electrospray ionization (MALDESI) mass spectrometry. Anal. Bioanal. Chem. 2021, 413, 2793–2801. [Google Scholar] [CrossRef] [PubMed]
- Mellinger, A.L.; Garrard, K.P.; Khodjaniyazova, S.; Rabbani, Z.N.; Gamcsik, M.P.; Muddiman, D.C. Multiple Infusion Start Time Mass Spectrometry Imaging of Dynamic SIL-Glutathione Biosynthesis Using Infrared Matrix-Assisted Laser Desorption Electrospray Ionization. J. Prot. Res. 2022, 21, 747–757. [Google Scholar] [CrossRef]
- Bagley, M.C.; Pace, C.L.; Ekelöf, M.; Muddiman, D.C. Infrared matrix-assisted laser desorption electrospray ionization (IR-MALDESI) mass spectrometry imaging analysis of endogenous metabolites in cherry tomatoes. Analyst 2020, 145, 5516. [Google Scholar] [CrossRef] [PubMed]
- Knizner, K.T.; Bagley, M.C.; Garrard, K.P.; Hauschild, J.P.; Pu, F.; Elsen, N.L.; Williams, J.D.; Muddiman, D.C. Optimized C-Trap Timing of an Orbitrap 240 Mass Spectrometer for High-Throughput Screening and Native MS by IR-MALDESI. J. Am. Soc. Mass Spectrom. 2022, 33, 328–334. [Google Scholar] [CrossRef]
- Kibbe, R.R.; Mellinger, A.L.; Muddiman, D.C. Novel Matrix Strategies for Improved Ionization and Spatial Resolution Using IR-MALDESI Mass Spectrometry Imaging. J. Mass Spectrom. 2022, 57, e4875. [Google Scholar] [CrossRef]
- Umoh, M.E.; Dammer, E.B.; Dai, J.; Duong, D.M.; Lah, J.J.; Levey, A.I.; Gearing, M.; Glass, J.D.; Seyfried, N.T. A Proteomic Network Approach across the ALS-FTD Disease Spectrum Resolves Clinical Phenotypes and Genetic Vulnerability in Human Brain. EMBO Mol. Med. 2018, 10, 48–62. [Google Scholar] [CrossRef]
- Ekelöf, M.; Manni, J.; Nazari, M.; Bokhart, M.; Muddiman, D.C. Characterization of a Novel Miniaturized Burst-Mode Infrared Laser System for IR-MALDESI Mass Spectrometry Imaging. Anal. Bioanal. Chem. 2018, 410, 2395–2402. [Google Scholar] [CrossRef]
- Garrard, K.P.; Ekelöf, M.; Khodjaniyazova, S.; Bagley, M.C.; Muddiman, D.C. A Versatile Platform for Mass Spectrometry Imaging of Arbitrary Spatial Patterns. J. Am. Soc. Mas Spectrom. 2020, 31, 2547–2552. [Google Scholar] [CrossRef]
- Race, A.M.; Styles, I.B.; Bunch, J. Inclusive sharing of mass spectrometry imaging data requires a converter for all. J. Prot. 2012, 75, 5111–5112. [Google Scholar] [CrossRef]
- Chambers, M.C.; MacLean, B.; Burke, R.; Amodei, D.; Ruderman, D.L.; Neumann, S.; Gatto, L.; Fischer, B.; Pratt, B.; Egertson, J.; et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 2012, 30, 918–920. [Google Scholar] [CrossRef] [PubMed]
- Palmer, A.; Phapale, P.; Chernyavsky, I.; Lavigine, R.; Fay, D.; Tarasov, A.; Koalev, V.; Fuchser, J.; Nikolenko, S.; Pineau, C.; et al. FDR-controlled metabolite annotation for high-resolution imaging mass spectrometry. Nat. Methods 2017, 14, 57–60. [Google Scholar] [CrossRef] [PubMed]
- Robichaud, G.; Garrard, K.P.; Barry, J.A.; Muddiman, D.C. MSiReader: An open-source interface to view and analyze high resolving power MS imaging files on matlab platform. J. Am. Soc. Mass Spectrom. 2013, 24, 718–721. [Google Scholar] [CrossRef] [PubMed]
- Bokhart, M.T.; Nazari, M.; Garrard, K.P.; Muddiman, D.C. MSiReader v1.0: Evolving Open-Source Mass Spectrometry Imaging Software for Targeted and Untargeted Analyses. J. Am. Soc. Mass Spectrom. 2018, 29, 8–16. [Google Scholar] [CrossRef]
- Tu, A.; Muddiman, D.C. Systematic evaluation of repeatability of IR-MALDESI-MS and normalization strategies for correcting the analytical variation and improving image quality. Anal. Bioanal. Chem. 2019, 411, 5729–5743. [Google Scholar] [CrossRef]
- Plotly Technologies Inc. Collaborative Data Science; Plotly Technologies Inc.: Montreal, QC, Canada, 2015; Available online: https://plot.ly (accessed on 14 March 2022).
- Pang, Z.; Chong, J.; Zhou, G.; Anderson De Lima Morais, D.; Chang, L.; Barrette, M.; Gauthier, C.; Jacques, P.E.; Li, S.; Xia, J. MetaboAnalyst 5.0: Narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 2021, 49, W388–W396. [Google Scholar] [CrossRef]
- The UniProt Consortium. UniProt: The universal protein knowledgebase in 2021. Nucleic Acids Res. 2021, 49, D1. [Google Scholar]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Lanznaster, D.; Bruno, C.; Bourgeais, J.; Emond, P.; Zemmoura, I.; Lefèvre, A.; Reynier, P.; Eymieux, S.; Blanchard, E.; Vourc’h, P.; et al. Metabolic Profile and Pathological Alterations in the Muscle of Patients with Early-Stage Amyotrophic Lateral Sclerosis. Biomedicines. 2022, 10, 1307. [Google Scholar] [CrossRef]
- Schousboe, A.; Scafidi, S.; Bak, L.K.; Waagepetersen, H.S.; McKenna, M.C. Glutamate Metabolism in the Brain Focusing on Astrocytes. Adv. Neurobiol. 2014, 11, 13–30. [Google Scholar] [CrossRef]
- Schulz, J.B.; Lindenau, J.; Seyfried, J.; Dichgans, J. Glutathione; oxidative stress and neurodegeneration. Eur. J. Biochem. 2000, 267, 4904–4911. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.X.; Wang, Y.; Qin, Z.H. Molecular mechanisms of excitotoxicity and their relevance to pathogenesis of neuro-degenerative diseases. Acta Pharmacol. Sin. 2009, 30, 379–387. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, G.; Calcagno, E.; Tartari, S.; Rizzardini, M.; Invernizzi, R.W.; Cantoni, L. Glutamate and glutathione interplay in a motor neuronal model of amyotrophic lateral sclerosis reveals altered energy metabolism. Neurobiol. Dis. 2011, 43, 346–355. [Google Scholar] [CrossRef] [PubMed]
- Patin, F.; Corcia, P.; Vourc’h, P.; Nadal-Desbarats, L.; Baranek, T.; Goossens, J.F.; Marouillat, S.; Dessein, A.-F.; Descat, A.; Blandine; et al. Omics to Explore Amyotrophic Lateral Sclerosis Evolution: The Central Role of Arginine and Proline Metabolism. Mol. Neurobiol. 2017, 54, 5361–5374. [Google Scholar] [CrossRef] [PubMed]
- Bereman, M.S.; Kirkwood, K.I.; Sabaretnam, T.; Furlong, S.; Rowe, D.B.; Guillemin, G.J.; Mellinger, A.L.; Muddiman, D.C. Metabolite Profiling Reveals Predictive Biomarkers and the Absence of β-Methyl Amino-L-alanine in Plasma from Indi-viduals Diagnosed with Amyotrophic Lateral Sclerosis. J. Prot. Res. 2020, 19, 3285. [Google Scholar] [CrossRef] [PubMed]
- Schomburg, I.; Chang, A.; Hofmann, O.; Ebeling, C.; Ehrentreich, F.; Schomburg, D. BRENDA: A resource for enzyme data and metabolic information. Trends Biochem. Sci. 2002, 27, 54–56. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sohn, A.L.; Ping, L.; Glass, J.D.; Seyfried, N.T.; Hector, E.C.; Muddiman, D.C. Interrogating the Metabolomic Profile of Amyotrophic Lateral Sclerosis in the Post-Mortem Human Brain by Infrared Matrix-Assisted Laser Desorption Electrospray Ionization (IR-MALDESI) Mass Spectrometry Imaging (MSI). Metabolites 2022, 12, 1096. https://doi.org/10.3390/metabo12111096
Sohn AL, Ping L, Glass JD, Seyfried NT, Hector EC, Muddiman DC. Interrogating the Metabolomic Profile of Amyotrophic Lateral Sclerosis in the Post-Mortem Human Brain by Infrared Matrix-Assisted Laser Desorption Electrospray Ionization (IR-MALDESI) Mass Spectrometry Imaging (MSI). Metabolites. 2022; 12(11):1096. https://doi.org/10.3390/metabo12111096
Chicago/Turabian StyleSohn, Alexandria L., Lingyan Ping, Jonathan D. Glass, Nicholas T. Seyfried, Emily C. Hector, and David C. Muddiman. 2022. "Interrogating the Metabolomic Profile of Amyotrophic Lateral Sclerosis in the Post-Mortem Human Brain by Infrared Matrix-Assisted Laser Desorption Electrospray Ionization (IR-MALDESI) Mass Spectrometry Imaging (MSI)" Metabolites 12, no. 11: 1096. https://doi.org/10.3390/metabo12111096
APA StyleSohn, A. L., Ping, L., Glass, J. D., Seyfried, N. T., Hector, E. C., & Muddiman, D. C. (2022). Interrogating the Metabolomic Profile of Amyotrophic Lateral Sclerosis in the Post-Mortem Human Brain by Infrared Matrix-Assisted Laser Desorption Electrospray Ionization (IR-MALDESI) Mass Spectrometry Imaging (MSI). Metabolites, 12(11), 1096. https://doi.org/10.3390/metabo12111096






