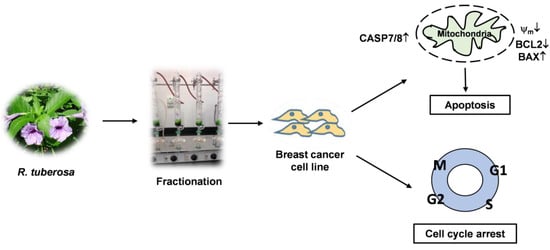
Ruellia tuberosa Ethyl Acetate Leaf Extract Induces Apoptosis and Cell Cycle Arrest in Human Breast Cancer Cell Line, MCF-7
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Extraction
2.2. Cell Culture and Treatment
2.3. Cell Proliferation Assay
2.4. Cell Staining
2.5. Detection of DNA Ladder
2.6. Apoptosis Assay
2.7. Measurement of Mitochondrial Membrane Potential
2.8. Cell Cycle Analysis
2.9. Isolation of RNA and RT-PCR
2.10. Protein Extraction and Western Blot Analysis
2.11. Statistical Analysis
3. Results
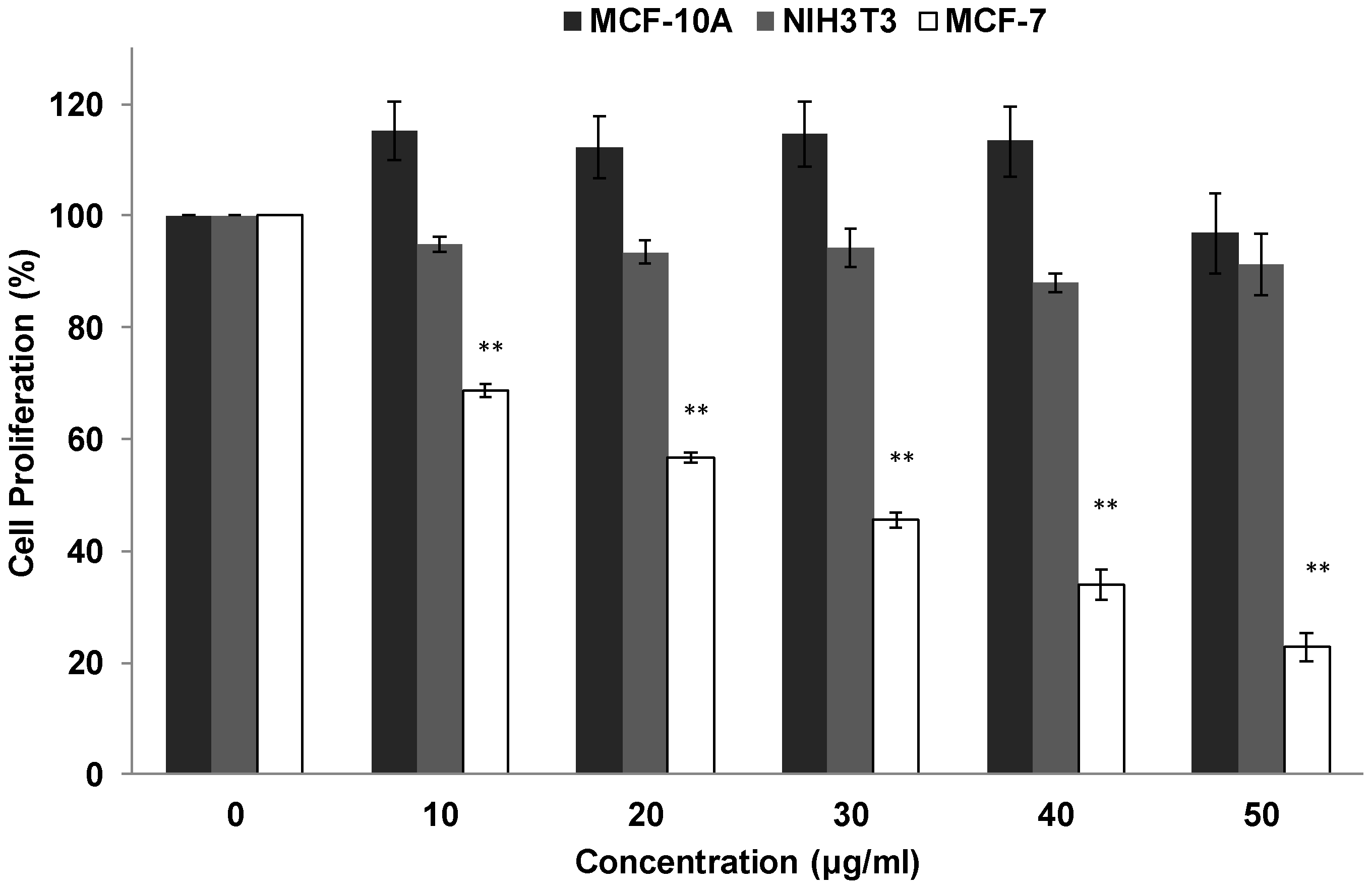
3.1. Antiproliferative Effects of RTEAL
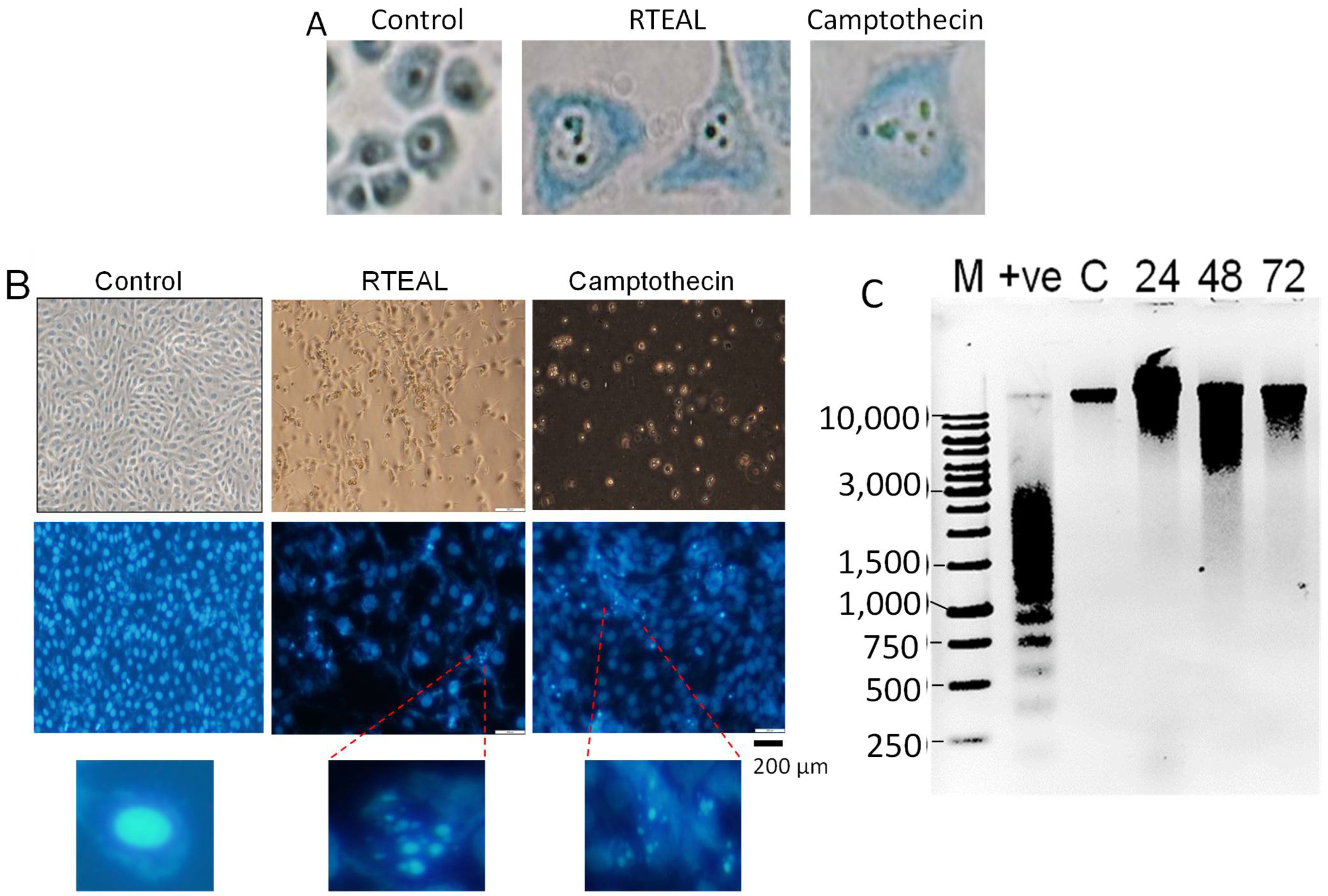
3.2. Effects of RTEAL on Cell Morphology and DNA Fragmentation
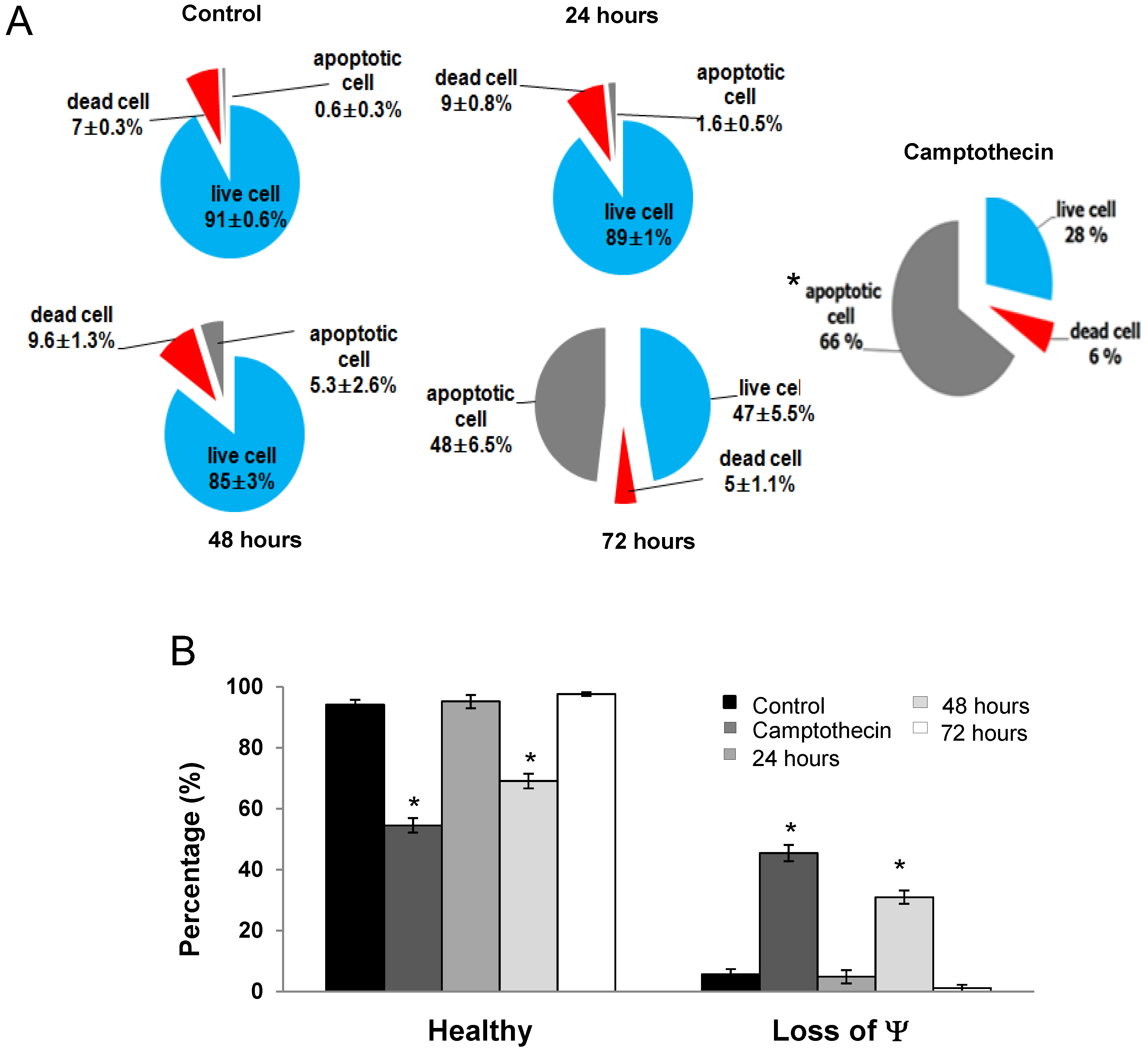
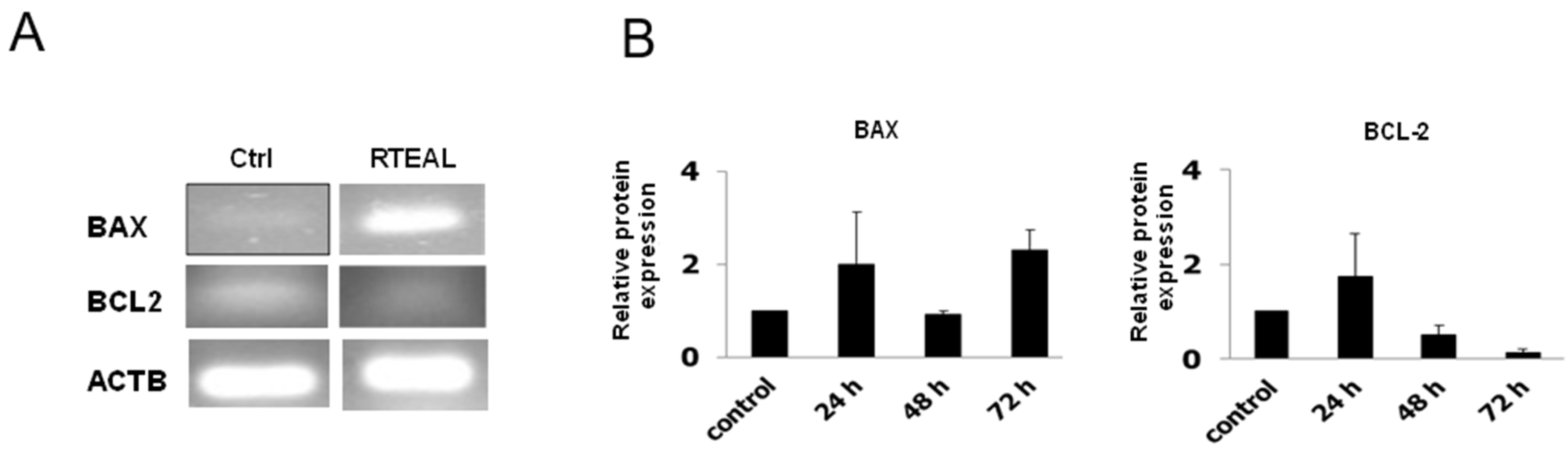
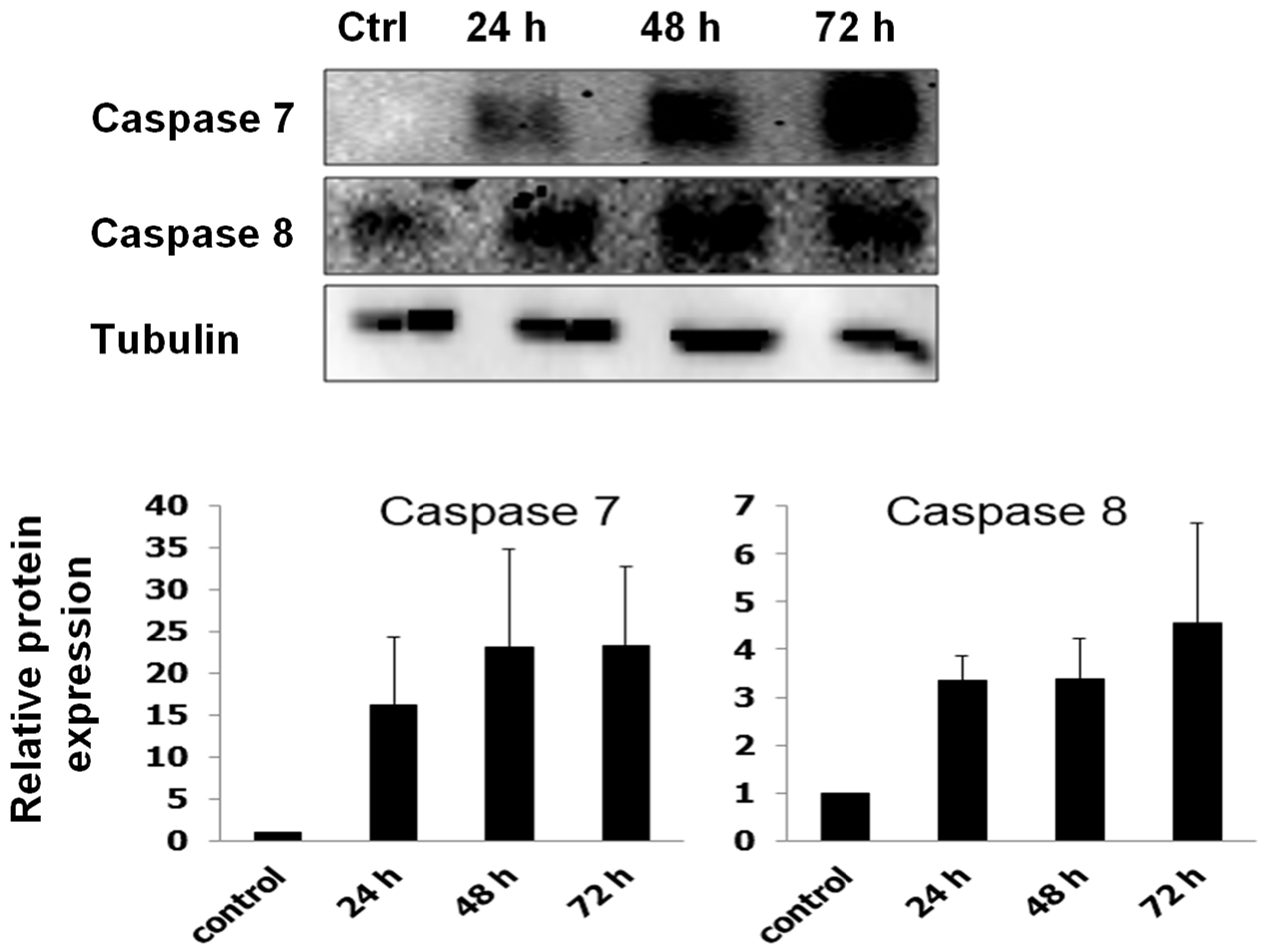
3.3. Induction of Apoptosis
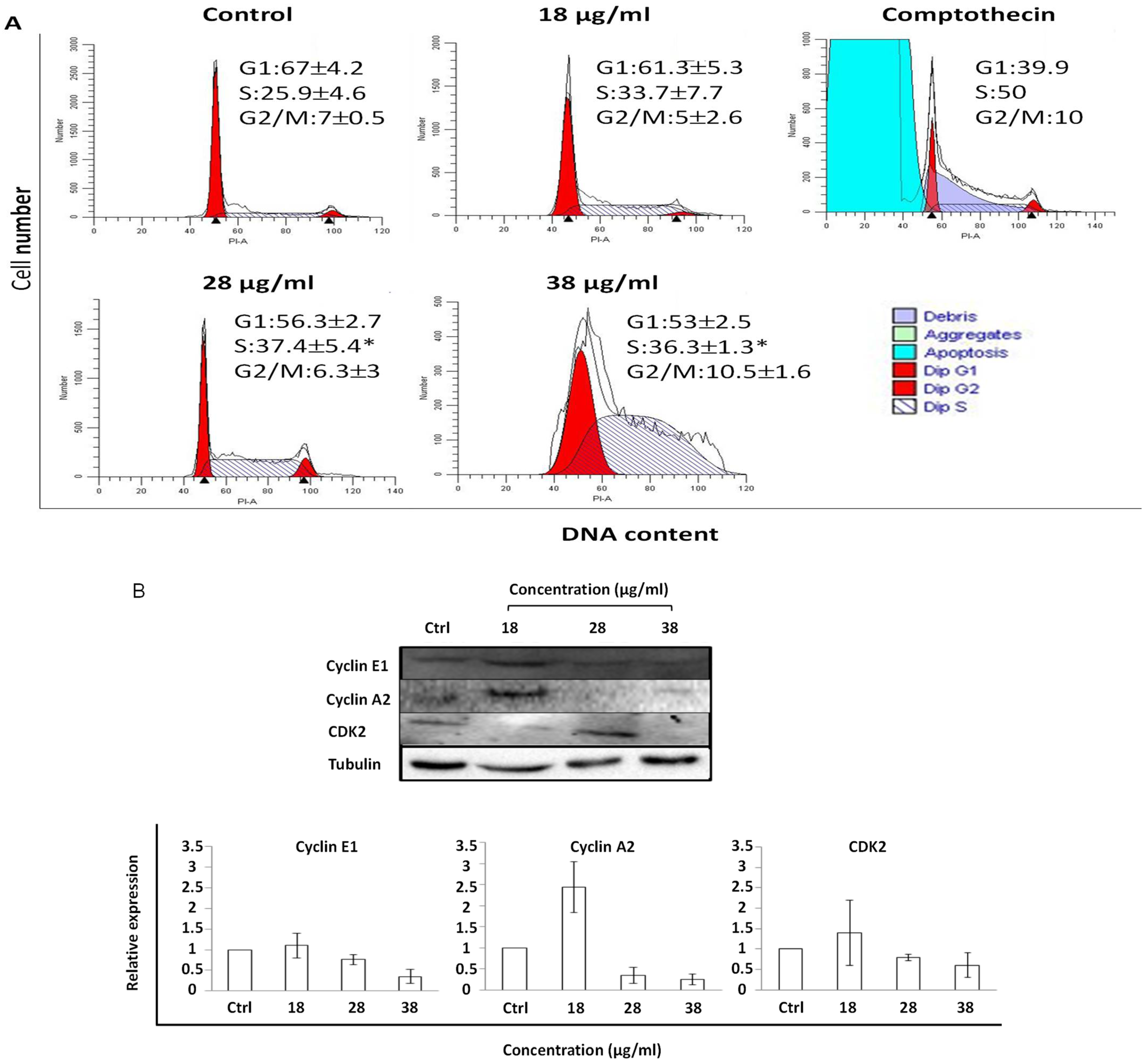
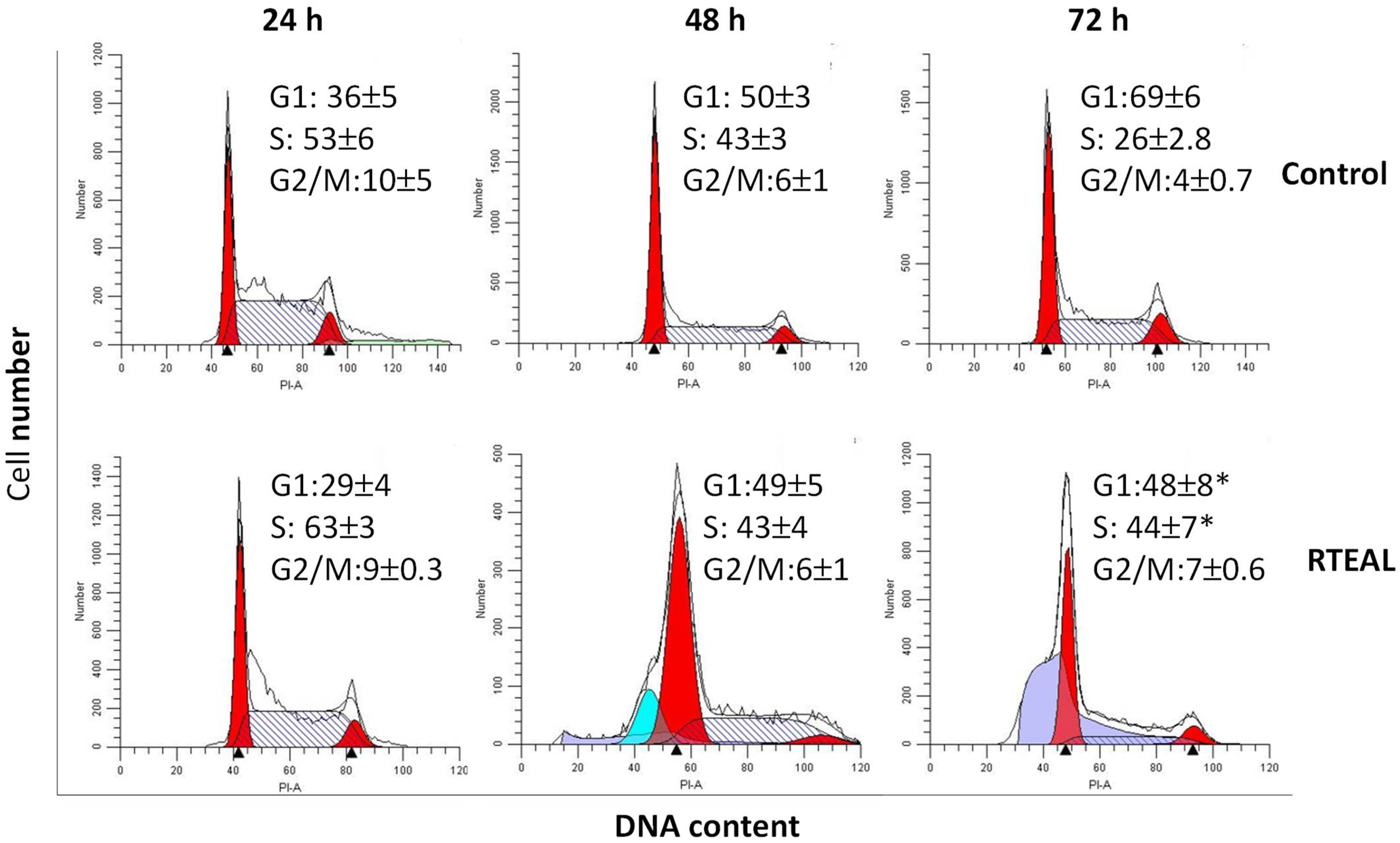
3.4. Perturbation of Cell Cycle
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pucci, C.; Martinelli, C.; Ciofani, G. Innovative approaches for cancer treatment: Current perspectives and new challenges. Ecancermedicalscience 2019, 13, 961. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Cancer. 2022. Available online: https://www.who.int/news-room/fact-sheets/detail/cancer (accessed on 29 April 2022).
- Turner, K.M.; Yeo, S.K.; Holm, T.M.; Shaughnessy, E.; Guan, J.L. Heterogeneity within molecular subtypes of breast cancer. Am. J. Physiol.-Cell Physiol. 2021, 321, C343–C354. [Google Scholar] [CrossRef] [PubMed]
- Testa, U.; Castelli, G.; Pelosi, E. Breast cancer: A molecularly heterogenous disease needing subtype-specific treatments. Med. Sci. 2020, 8, 18. [Google Scholar] [CrossRef]
- Nouri, Z.; Fakhri, S.; Nouri, K.; Wallace, C.E.; Farzaei, M.H.; Bishayee, A. Targeting multiple signaling pathways in cancer: The rutin therapeutic approach. Cancers 2020, 12, 2276. [Google Scholar] [CrossRef]
- De Melo Gagliato, D.; Gonzalez-Angulo, A.M. Targeting multiple pathways in breast cancer. Breast Cancer Manag. 2014, 3, 87–101. [Google Scholar] [CrossRef]
- Jan, R.; Chaudhry, G.E. Understanding apoptosis and apoptotic pathways targeted cancer therapeutics. Adv. Pharm. Bull. 2019, 9, 205–218. [Google Scholar] [CrossRef]
- Alimbetov, D.; Askarova, S.; Umbayev, B.; Davis, T.; Kipling, D. Pharmacological targeting of cell cycle, apoptotic and cell adhesion signaling pathways implicated in chemoresistance of cancer Cells. Int. J. Mol. Sci. 2018, 19, 1690. [Google Scholar] [CrossRef]
- Haque, A.; Brazeau, D.; Amin, A.R. Perspectives on natural compounds in chemoprevention and treatment of cancer: An update with new promising compounds. Eur. J. Cancer 2021, 149, 165–183. [Google Scholar] [CrossRef]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J. Nat. Prod. 2020, 83, 770–803. [Google Scholar] [CrossRef]
- Sun, L.R.; Zhou, W.; Zhang, H.M.; Guo, Q.S.; Yang, W.; Li, B.J.; Sun, Z.H.; Gao, S.H.; Cui, R.J. Modulation of multiple signaling pathways of the plant-derived natural products in cancer. Front. Oncol. 2019, 9, 1153. [Google Scholar] [CrossRef]
- Yuan, L.; Cai, Y.; Zhang, L.; Liu, S.; Li, P.; Li, X. Promoting apoptosis, a promising way to treat breast cancer with natural products: A comprehensive review. Front. Pharmacol. 2022, 12, 801662. [Google Scholar] [CrossRef] [PubMed]
- Arirudran, B.; Saraswathy, A.; Krishnamurthy, V. Pharmacognostic and preliminary phytochemical studies on Ruellia tuberosa L. (Whole plant). Pharmacogn. J. 2011, 3, 29–34. [Google Scholar] [CrossRef]
- Chen, F.A.; Wu, A.B.; Shieh, P.; Kuo, D.H.; Hsieh, C.Y. Evaluation of the antioxidant activity of Ruellia tuberosa. Food Chem. 2006, 94, 14–18. [Google Scholar] [CrossRef]
- Chothani, D.L.; Mishra, H. In vitro anti-oxidant activity of Ruellia tuberosa root extracts. Free. Radic. Antioxid. 2012, 2, 38–44. [Google Scholar] [CrossRef]
- Afzal, K.; Uzair, M.; Chaudhary, B.A.; Ahmad, A.; Afzal, S.; Saadullah, M. Genus Ruellia: Pharmacological and phytochemical importance in ethnopharmacology. Acta Pol. Pharm.-Drug Res. 2015, 72, 821–827. [Google Scholar]
- Alam, M.A.; Subhan, N.; Awal, M.A.; Alam, M.S.; Sarder, M.; Nahar, L.; Sarker, S.D. Atinociceptive and anti-inflammatory properties of Ruellia tuberosa. Pharma. Biol. 2009, 47, 209–214. [Google Scholar] [CrossRef]
- Jiorry, J.R.S.; Cheong, B.E. Metabolic fingerprinting of Sabah Ruellia tuberosa plant extracts for the identification of potential anticancer compounds. Short Commun. Biotech. 2017, 4, 75–87. Available online: https://www.ums.edu.my/ipbv2/files/Metabolic-Fingerprinting-of-Sabah-Ruellia-tuberosa-Plant-Extracts-for-the-Identification-of-Potential-Anticancer-Compounds.pdf (accessed on 3 July 2022).
- Lin, C.-F.; Huang, Y.-L.; Cheng, L.-Y.; Sheu, S.-J.; Chen, C.-C. Bioactive flavonoids from Ruellia tuberosa. J. Chin. Med. 2006, 17, 103–109. [Google Scholar]
- Mohan, V.; Rajendrakumar, N.; Vasantha, K. GC-MS Analysis of bioactive components of tubers of Ruellia tuberosa L. (Acanthaceae). Am. J. Phytomed. Clin. Ther. 2014, 2, 209–216. [Google Scholar]
- Sánchez-Quesada, C.; Gutiérrez-Santiago, F.; Rodríguez-García, C.; Gaforio, J.J. Synergistic effect of squalene and hydroxytyrosol on highly invasive MDA-MB-231 breast cancer cells. Nutrients 2022, 14, 255. [Google Scholar] [CrossRef]
- Pitchai, D.; Roy, A.; Ignatius, C. In vitro evaluation of anticancer potentials of lupeol isolated from Elephantopus scaber L. on MCF-7 cell line. J. Adv. Pharm. Technol. Res. 2014, 5, 179–184. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Gao, Z.; Chen, K.; Zhuo, Q.; Chen, M.; Wang, J.; Lai, X.; Wang, L. Lupeol inhibits the proliferation and migration of MDA-MB-231 breast cancer cells via a novel crosstalk mechanism between autophagy and the EMT. Food Funct. 2022, 13, 4967. [Google Scholar] [CrossRef] [PubMed]
- Cheong, B.E.; Waslim, M.Z.; Lem, F.F.; Teoh, P.L. Antioxidant and anti-proliferative activities of Sabah Ruellia tuberosa. J. Appl. Pharma. Sci. 2013, 3, 20–24. [Google Scholar] [CrossRef]
- Dey, S.; Roy, S.; Deb, N.; Kumar Sen, K.; Besra, S.E. Anticarcinogenic activity of Ruellia tuberosa L. (Acanthaceae) leaf extract on hepatoma cell line and increased superoxide dismutase activity on macrophage cell lysate. Int. J. Pharm. Pharm. Sci. 2015, 5, 854–861. [Google Scholar]
- Teoh, P.L.; Cheng, A.Y.F.; Liau, M.; Lem, F.F.; Kaling, G.P.; Chua, F.N.; Cheong, B.E. Chemical composition and cytotoxic properties of Clinacanthus nutans root extracts. Pharm. Biol. 2017, 55, 394–401. [Google Scholar] [CrossRef]
- Teoh, P.L.; Liau, M.; Cheong, B.E. Phyla nodiflora L. extracts induce apoptosis and cell cycle arrest in human breast cancer cell line, MCF-7. Nutr. Cancer 2019, 71, 668–675. [Google Scholar] [CrossRef]
- Walker, P.R.; Leblanc, J.; Carson, C.; Ribecco, M.; Sikorska, M. Neither caspase-3 nor DNA fragmentation factor is required for high molecular weight DNA degradation in apoptosis. Ann. N. Y. Acad. Sci. 1999, 887, 48–59. [Google Scholar] [CrossRef]
- Iglesias-Guimarais, V.; Gil-Guiñon, E.; Gabernet, G.; García-Belinchón, M.; Sánchez-Osuna, M.; Casanelles, E.; Comella, J.X.; Yuste, V.J. Apoptotic DNA degradation into oligonucleosomal fragments, but not apoptotic nuclear morphology, relies on a cytosolic pool of DFF40/CAD endonuclease. J. Biol. Chem. 2012, 287, 7766–7779. [Google Scholar] [CrossRef]
- Susin, S.A.; Lorenzo, H.K.; Zamzami, N.; Marzo, I.; Snow, B.E.; Brothers, G.M.; Mangion, J.; Jacotot, E.; Costantini, P.; Loeffler, M.; et al. Molecular characterization of mitochondrial apoptosis-inducing factor. Nature 1999, 397, 441–446. [Google Scholar] [CrossRef]
- Kitazumi, I.; Tsukahara, M. Regulation of DNA fragmentation: The role of caspases and phosphorylation. FEBS J. 2011, 278, 427–441. [Google Scholar] [CrossRef]
- Lalier, L.; Vallette, F.; Manon, S. Bcl-2 family members and the mitochondrial import machineries: The roads to death. Biomolecules 2022, 12, 162. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, S.; Eguchi, Y.; Kamiike, W.; Funahashi, Y.; Mignon, A.; Lacronique, V.; Matsuda, H.; Tsujimoto, Y. Bcl-2 prevents apoptotic mitochondrial dysfunction by regulating proton flux. Proc. Natl. Acad. Sci. USA 1998, 95, 1455–1459. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Hu, J.; Xu, X.; Gao, X.; Wang, Y.; Zhu, S. Sodium azide induces mitochondria-mediated apoptosis in PC12 cells through Pgc-1α-associated signaling pathway. Mol. Med. Rep. 2019, 19, 2211–2219. [Google Scholar] [CrossRef] [PubMed]
- Ly, J.D.; Grubb, D.R.; Lawen, A. The mitochondrial membrane potential (Δψm) in apoptosis; an update. Apoptosis 2003, 8, 115–128. [Google Scholar] [CrossRef]
- Kantari, C.; Walczak, H. Caspase-8 and bid: Caught in the act between death receptors and mitochondria. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2011, 1813, 558–563. [Google Scholar] [CrossRef]
- Wu, Y.; Zhao, D.; Zhuang, J.; Zhang, F.; Xu, C. Caspase-8 and caspase-9 functioned differently at different stages of the cyclic stretch-induced apoptosis in human periodontal ligament cells. PLoS ONE 2016, 11, e0168268. [Google Scholar] [CrossRef]
- McComb, S.; Chan, P.K.; Guinot, A.; Hartmannsdottir, H.; Jenni, S.; Dobay, M.P.; Bourquin, J.P.; Bornhauser, B.C. Efficient apoptosis requires feedback amplification of upstream apoptotic signals by effector caspase-3 or -7. Sci. Adv. 2019, 5, eaau9433. [Google Scholar] [CrossRef]
- Walsh, J.G.; Cullen, S.P.; Sheridan, C.; Lüthi, A.U.; Gerner, C.; Martin, S.J. Executioner caspase-3 and caspase-7 are functionally distinct proteases. Proc. Natl. Acad. Sci. USA 2008, 105, 12815–12819. [Google Scholar] [CrossRef]
- Lamkanfi, M.; Kanneganti, T.D. Caspase-7: A protease involved in apoptosis and inflammation. Int. J. Biochem. Cell Biol. 2010, 42, 21–24. [Google Scholar] [CrossRef]
- Foo, J.B.; Yazan, L.S.; Tor, Y.S.; Armania, N.; Ismail, N.; Imam, M.U.; Yeap, S.K.; Cheah, Y.K.; Abdullah, R.; Ismail, M. Induction of cell cycle arrest and apoptosis in caspase-3 deficient MCF-7 cells by Dillenia suffruticosa root extract via multiple signaling pathways. BMC Complement. Altern. Med. 2014, 14, 197. [Google Scholar] [CrossRef]
- Sakle, N.S.; More, S.A.; Mokale, S.N. Chemomodulatory effects of Alysicarpus vaginalis extract via mitochondria-dependent apoptosis and necroptosis in breast cancer. Nutr. Cancer 2020, 72, 1243–1253. [Google Scholar] [CrossRef] [PubMed]
- Lim, S.; Kaldis, P. Cdks, cyclins and CKIs: Roles beyond cell cycle regulation. Development 2013, 140, 3079–3093. [Google Scholar] [CrossRef] [PubMed]
- Ding, L.; Cao, J.; Lin, W.; Chen, H.; Xiong, X.; Ao, H.; Yu, M.; Lin, J.; Cui, Q. The roles of cyclin-dependent kinases in cell-cycle progression and therapeutic strategies in human breast cancer. Int. J. Mol. Sci. 2020, 21, 1960. [Google Scholar] [CrossRef]
- Bailon-Moscoso, N.; Cevallos-Solorzano, G.; Romero-Benavides, J.C.; Orellana, M.I.R. Natural compounds as modulators of cell cycle arrest: Application for anticancer chemotherapies. Curr. Genom. 2017, 18, 106–131. [Google Scholar] [CrossRef] [PubMed]
- Thu, K.L.; Soria-Bretones, I.; Mak, T.W.; Cescon, D.W. Targeting the cell cycle in breast cancer: Towards the next phase. Cell Cycle 2018, 17, 1871–1885. [Google Scholar] [CrossRef] [PubMed]
- Milioli, H.H.; Alexandrou, S.; Lim, E.; Caldon, C.E. Cyclin E1 and cyclin E2 in ER+ breast cancer: Prospects as biomarkers and therapeutic targets. Endocr.-Relat. Cancer 2020, 27, R93–R112. [Google Scholar] [CrossRef]
- Guerrero Llobet, S.; van der Vegt, B.; Jongeneel, E.; Bense, R.D.; Zwager, M.C.; Schröder, C.P.; Everts, M.; Fehrmann, R.S.; de Bock, G.H.; van Vugt, M.A. Cyclin E expression is associated with high levels of replication stress in triple-negative breast cancer. NPJ Breast Cancer 2020, 6, 40. [Google Scholar] [CrossRef]
- Moore, N.L.; Edwards, D.P.; Weigel, N.L. Cyclin A2 and its associated kinase activity are required for optimal induction of progesterone receptor target genes in breast cancer cells. J. Steroid Biochem. Mol. Biol. 2014, 144 Pt B, 471–482. [Google Scholar] [CrossRef][Green Version]
- Gordon, S.; Plüddemann, A. Macrophage clearance of apoptotic cells: A critical assessment. Front. Immunol. 2018, 9, 127. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lem, F.F.; Cheong, B.E.; Teoh, P.L. Ruellia tuberosa Ethyl Acetate Leaf Extract Induces Apoptosis and Cell Cycle Arrest in Human Breast Cancer Cell Line, MCF-7. Sci. Pharm. 2022, 90, 44. https://doi.org/10.3390/scipharm90030044
Lem FF, Cheong BE, Teoh PL. Ruellia tuberosa Ethyl Acetate Leaf Extract Induces Apoptosis and Cell Cycle Arrest in Human Breast Cancer Cell Line, MCF-7. Scientia Pharmaceutica. 2022; 90(3):44. https://doi.org/10.3390/scipharm90030044
Chicago/Turabian StyleLem, Fui Fui, Bo Eng Cheong, and Peik Lin Teoh. 2022. "Ruellia tuberosa Ethyl Acetate Leaf Extract Induces Apoptosis and Cell Cycle Arrest in Human Breast Cancer Cell Line, MCF-7" Scientia Pharmaceutica 90, no. 3: 44. https://doi.org/10.3390/scipharm90030044
APA StyleLem, F. F., Cheong, B. E., & Teoh, P. L. (2022). Ruellia tuberosa Ethyl Acetate Leaf Extract Induces Apoptosis and Cell Cycle Arrest in Human Breast Cancer Cell Line, MCF-7. Scientia Pharmaceutica, 90(3), 44. https://doi.org/10.3390/scipharm90030044