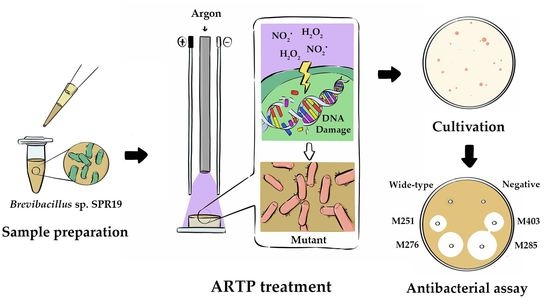
Enhanced Antibacterial Activity of Brevibacillus sp. SPR19 by Atmospheric and Room Temperature Plasma Mutagenesis (ARTP)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Microorganisms and Culture Conditions
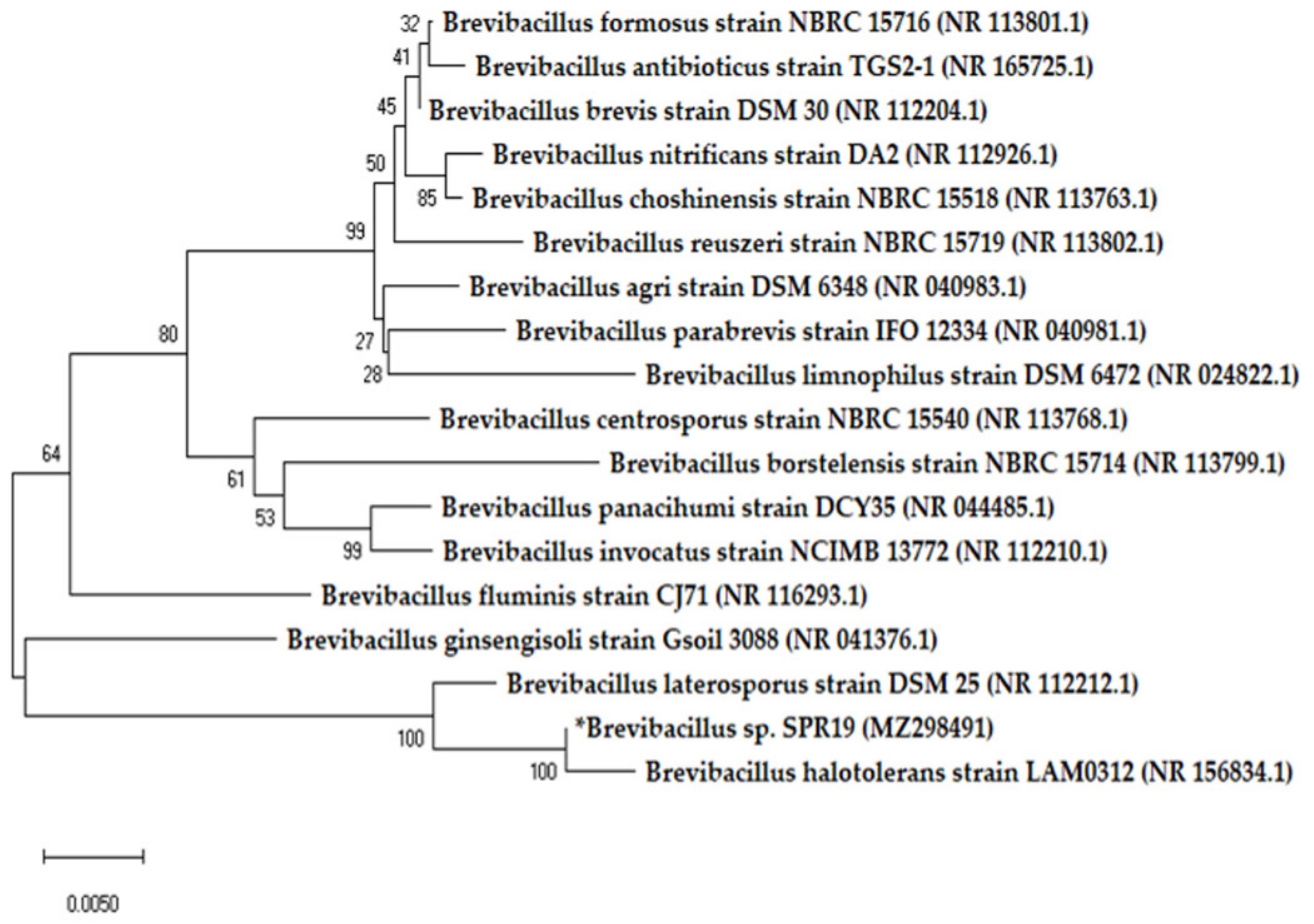
2.2. Strain Identification
2.3. Growth Curve and Production Kinetics of Antibacterial Substances
2.4. Atmospheric and Room Temperature Plasma (ARTP) Mutagenesis
2.5. Determination of Reactive Oxygen Species (ROS) and Reactive Nitrogen Species (RNS) Concentration
2.6. Antibacterial Assay by Agar Overlay and Agar Well Diffusion Methods
2.7. Strain Stability
2.8. Drug Susceptibility Study
2.9. Scanning Electron Microscope (SEM)
2.10. Comparison of Growth Curve and Production Kinetics of Antibacterial Substances
2.11. Determination of Minimum Inhibitory Concentration (MIC) and Minimum Bactericidal Concentration (MBC)
2.12. Stability of the Antibacterial Substances
2.12.1. Effect of Temperature
2.12.2. Effect of Proteolytic Enzyme
2.12.3. Effect of Surfactant
2.12.4. Effect of pH
2.13. Statistical Analysis
3. Results and Discussion
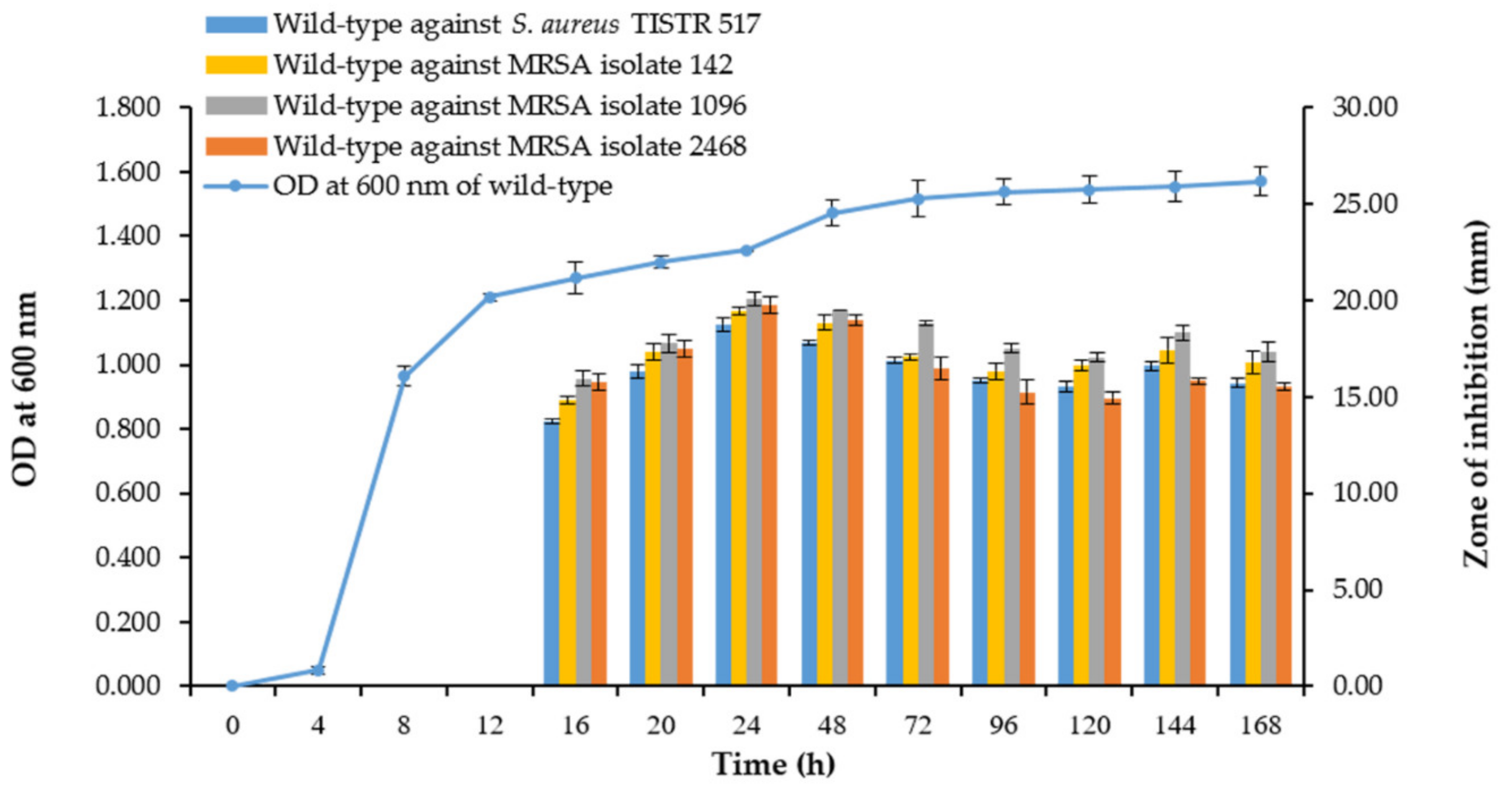
3.1. Characteristic, Growth Curve, and Production Kinetics of SPR19
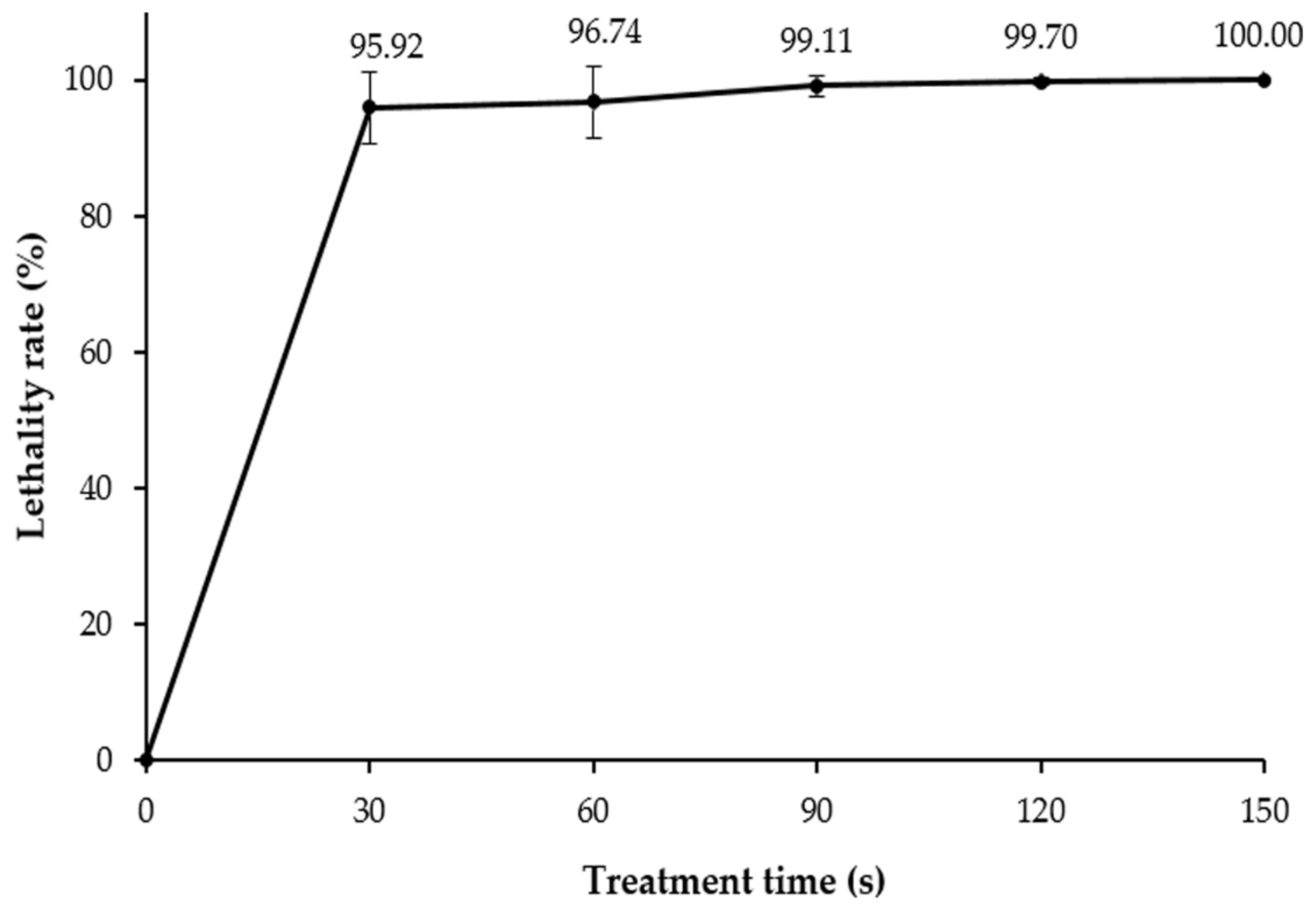
3.2. ARTP Mutagenesis and Lethality Rate
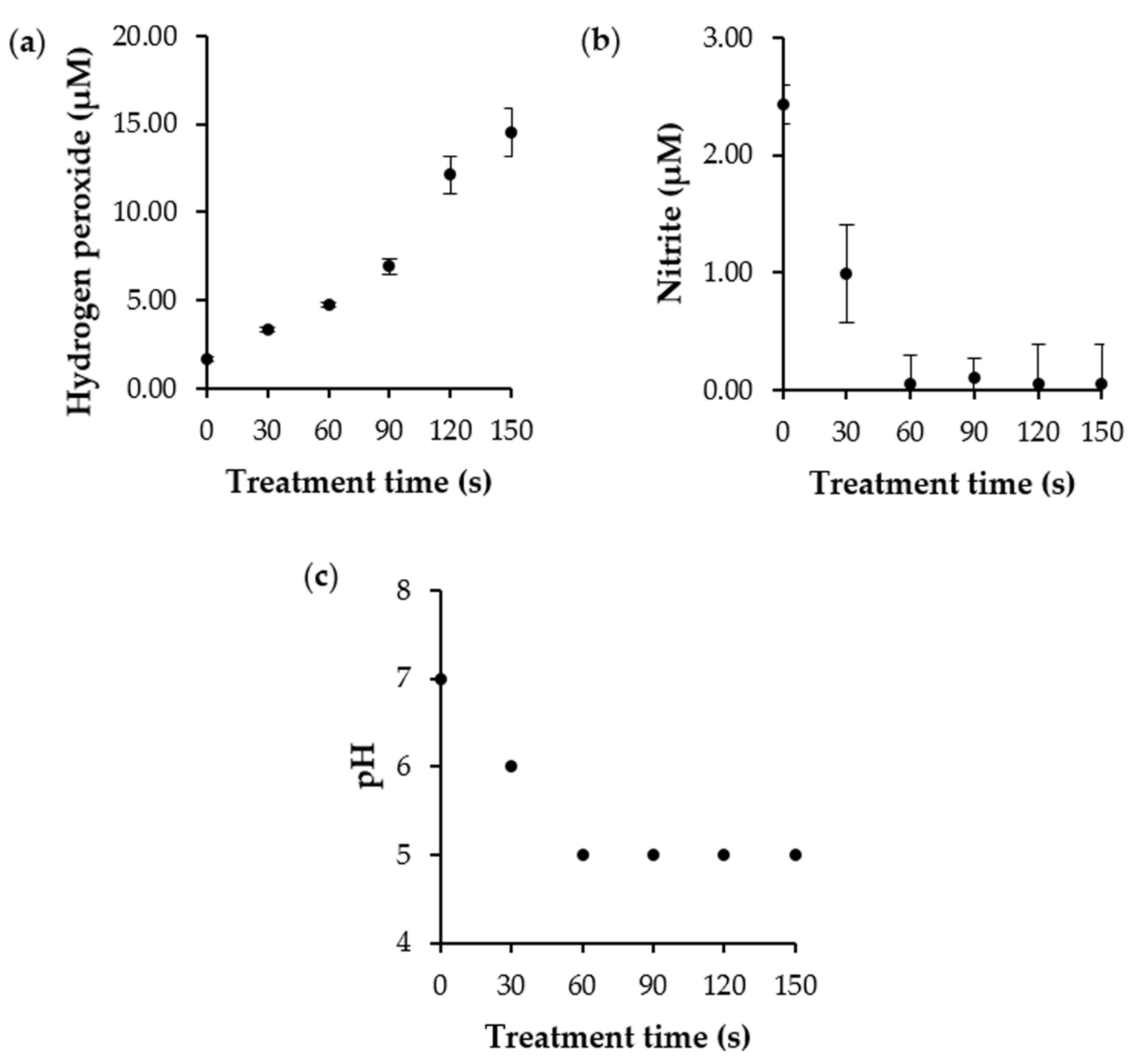
3.3. Determination of ROS and RNS Concentration
3.4. Determination of the Antibacterial Activity of the Mutants
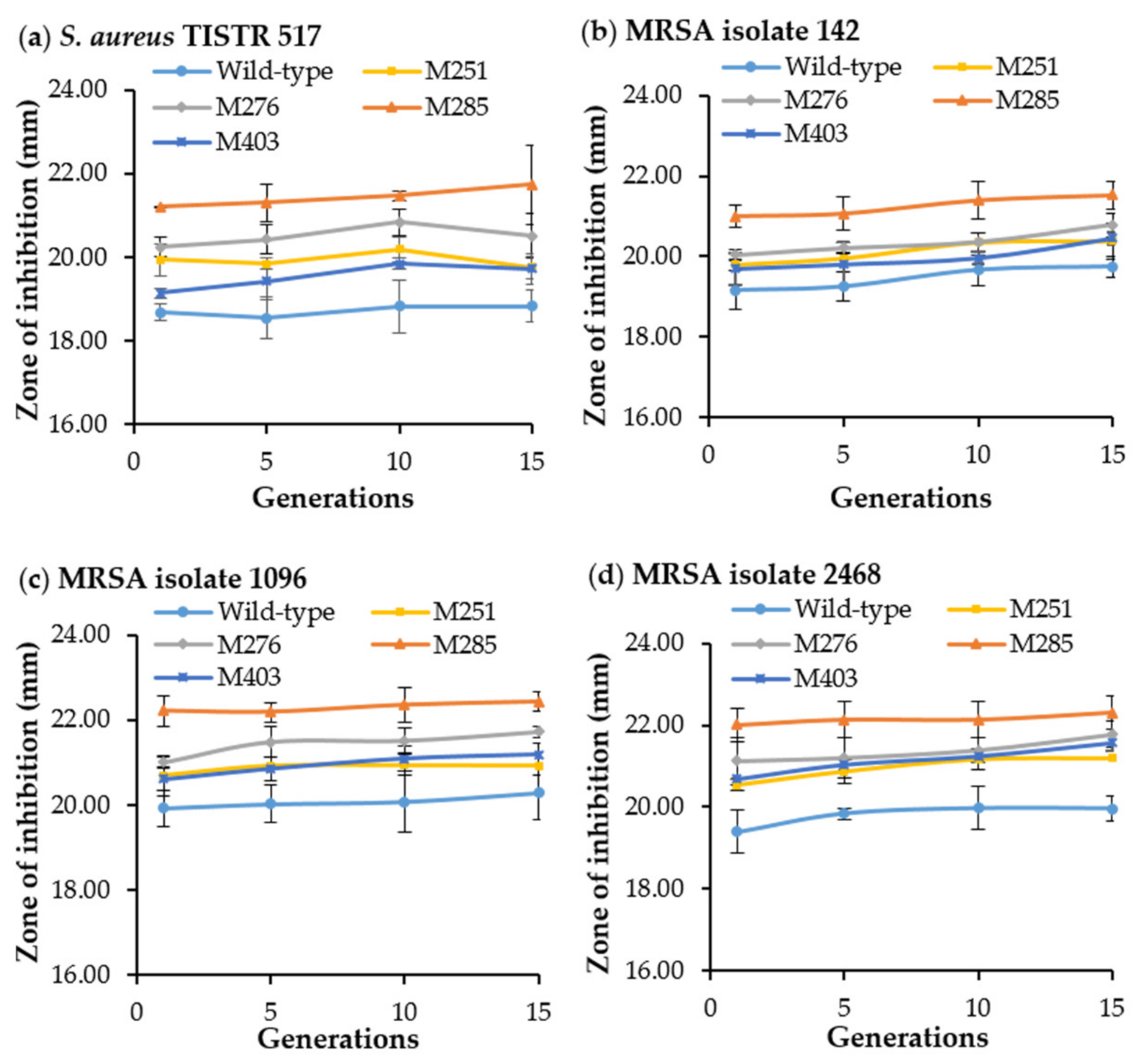
3.5. Strain Stability
3.6. Drug Susceptibility Study
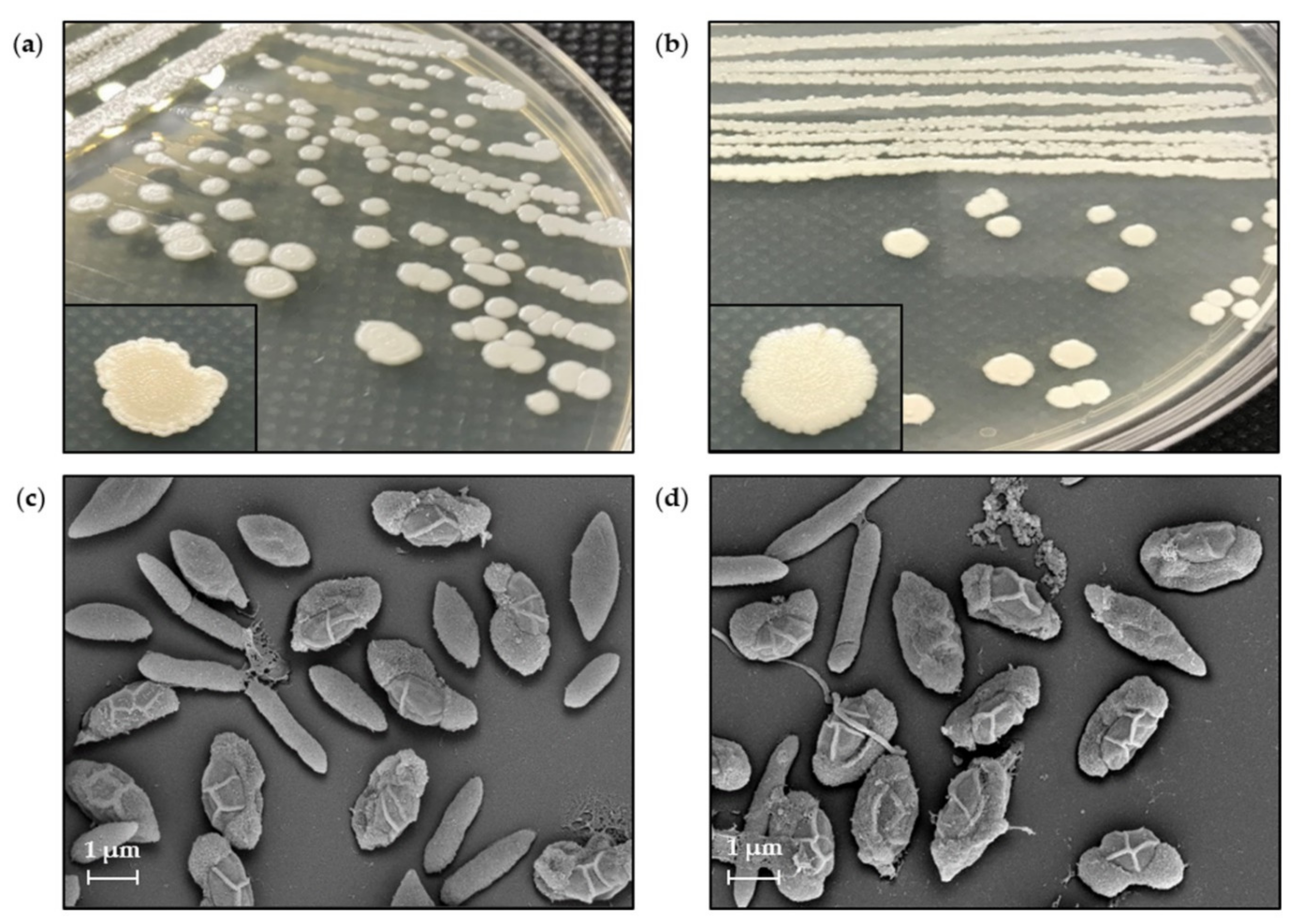
3.7. Morphological Characteristics by SEM
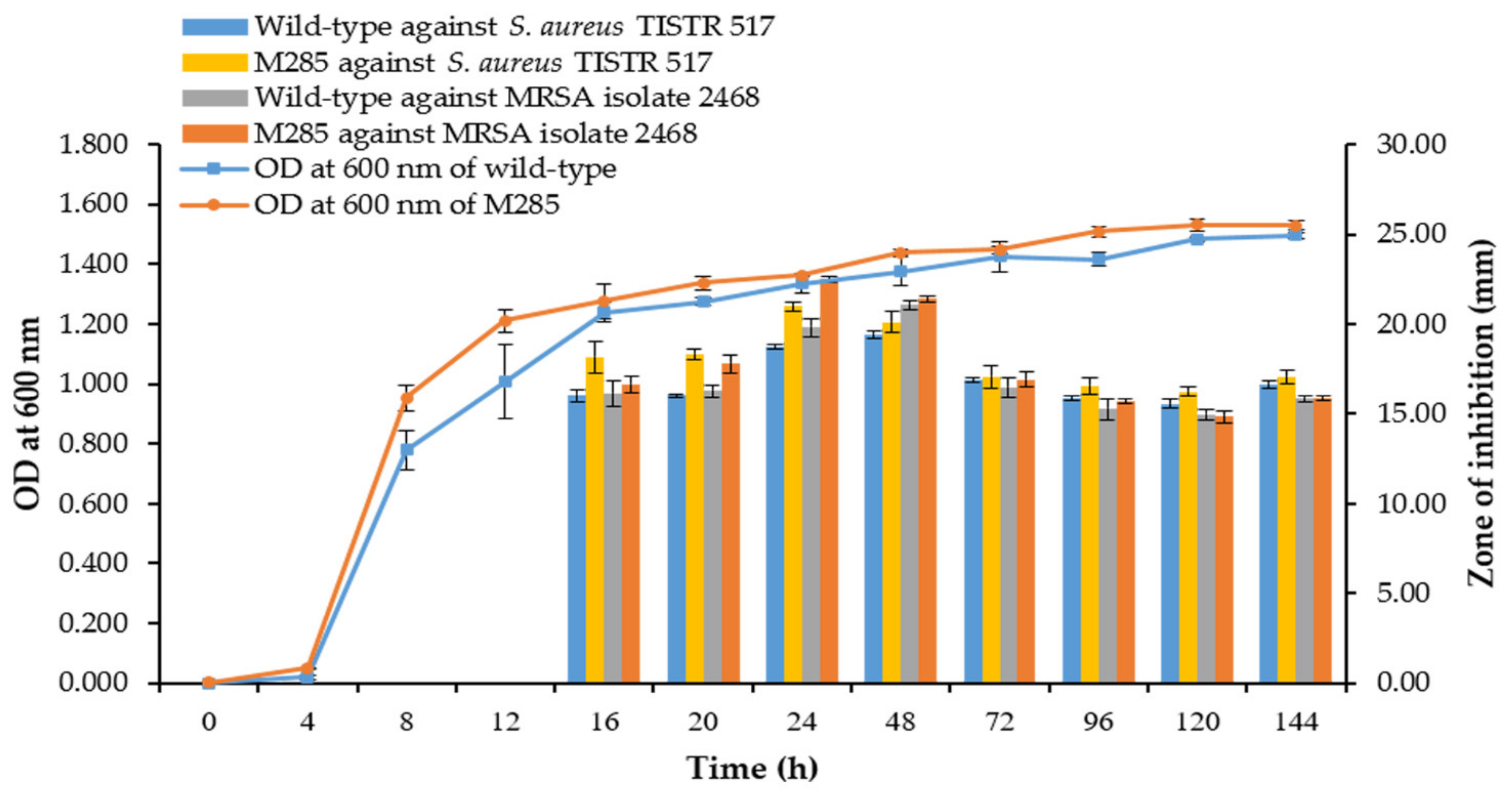
3.8. Comparison of Growth Curve and Kinetics of Antibacterial Production
3.9. Determination of MIC and MBC
3.10. Stability of the Antibacterial Substances
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kotra, L.P. xPharm: The Comprehensive Pharmacology Reference Infectious Diseases; Elsevier: Amsterdam, The Netherlands, 2007; pp. 1–2. [Google Scholar] [CrossRef]
- Hutchings, M.I.; Truman, A.W.; Wilkinson, B. Antibiotics: Past, present and future. Curr. Opin. Microbiol. 2019, 51, 72–80. [Google Scholar] [CrossRef] [PubMed]
- McGuinness, W.A.; Malachowa, N.; DeLeo, F.R. Vancomycin resistance in Staphylococcus aureus. Yale J. Biol. Med. 2017, 90, 269–281. [Google Scholar] [PubMed]
- Centers for Disease Control and Prevention. Antibiotic Resistance Threats in the United States. 2019. Available online: https://stacks.cdc.gov/view/cdc/82532/ (accessed on 19 February 2022).
- Ventola, C.L. The Antibiotic Resistance Crisis: Part 1: Causes and threats. Pharm. Ther. 2015, 40, 277–283. [Google Scholar]
- Da Cunha, B.R.; Fonseca, L.P.; Calado, C.R.C. Antibiotic Discovery: Where Have We Come from, Where Do We Go? Antibiotics 2019, 8, 45. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Procópio, R.E.D.L.; da Silva, I.R.; Martins, M.K.; de Azevedo, J.L.; de Araújo, J.M. Antibiotics produced by Streptomyces. Braz. J. Infect. Dis. 2012, 16, 466–471. [Google Scholar] [CrossRef] [Green Version]
- Sumi, C.D.; Yang, B.W.; Yeo, I.-C.; Hahm, Y.T. Antimicrobial peptides of the genus Bacillus: A new era for antibiotics. Can. J. Microbiol. 2015, 61, 93–103. [Google Scholar] [CrossRef]
- Okafor, N. Screening for productive strains and strain improvement in biotechnological organisms. In Modern Industrial Microbiology and Biotechnology Strain Improvement; Science Publishers: Enfield, NH, USA, 2007; Volume 3, pp. 125–170. [Google Scholar]
- Ottenheim, C.; Nawrath, M.; Wu, J.C. Microbial mutagenesis by atmospheric and room-temperature plasma (ARTP): The latest development. Bioresour. Bioprocess. 2018, 5, 12. [Google Scholar] [CrossRef] [Green Version]
- Xu, J.-Z.; Zhang, W.-G. Menaquinone-7 production from maize meal hydrolysate by Bacillus isolates with diphenylamine and analogue resistance. J. Zhejiang Univ. Sci. B 2017, 18, 462–473. [Google Scholar] [CrossRef] [Green Version]
- Zhu, L.; Xu, Q.; Jiang, L.; Huang, H.; Li, S. Polydiacetylene-Based High-Throughput Screen for Surfactin Producing Strains of Bacillus subtilis. PLoS ONE 2014, 9, e88207. [Google Scholar] [CrossRef] [Green Version]
- Li, G.; Li, H.-P.; Wang, L.-Y.; Wang, S.; Zhao, H.-X.; Sun, W.-T.; Xing, X.-H.; Bao, C.-Y. Genetic effects of radio-frequency, atmospheric-pressure glow discharges with helium. Appl. Phys. Lett. 2008, 92, 221504. [Google Scholar] [CrossRef] [Green Version]
- Arjunan, K.P.; Sharma, V.K.; Ptasinska, S. Effects of Atmospheric Pressure Plasmas on Isolated and Cellular DNA—A Review. Int. J. Mol. Sci. 2015, 16, 2971–3016. [Google Scholar] [CrossRef] [Green Version]
- Jiang, H.; Zou, J.; Cheng, H.; Fang, J.; Huang, G. Purification, Characterization, and Mode of Action of Pentocin JL-1, a Novel Bacteriocin Isolated from Lactobacillus pentosus, against Drug-Resistant Staphylococcus aureus. BioMed Res. Int. 2017, 2017, 7657190. [Google Scholar] [CrossRef] [Green Version]
- Tripathi, N.; Sapra, A. Gram Staining, in StatPearls Gram Staining; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Ashby, G.K. Simplified Schaeffer spore stain. Science 1938, 87, 443. [Google Scholar] [CrossRef]
- Zhang, J.; Yang, Y.; Yang, H.; Bu, Y.; Yi, H.; Zhang, L.; Han, X.; Ai, L. Purification and Partial Characterization of Bacteriocin Lac-B23, a Novel Bacteriocin Production by Lactobacillus plantarum J23, Isolated From Chinese Traditional Fermented Milk. Front. Microbiol. 2018, 9, 2165. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Goh, H.F.; Philip, K. Purification and Characterization of Bacteriocin Produced by Weissella confusa A3 of Dairy Origin. PLoS ONE 2015, 10, e0140434. [Google Scholar] [CrossRef]
- Liu, K.; Fang, H.; Cui, F.; Nyabako, B.A.; Tao, T.; Zan, X.; Chen, H.; Sun, W. ARTP mutation and adaptive laboratory evolution improve probiotic performance of Bacillus coagulans. Appl. Microbiol. Biotechnol. 2020, 104, 6363–6373. [Google Scholar] [CrossRef]
- Halder, D.; Mandal, M.; Chatterjee, S.S.; Pal, N.K.; Mandal, S. Indigenous Probiotic Lactobacillus Isolates Presenting Antibiotic like Activity against Human Pathogenic Bacteria. Biomedicines 2017, 5, 31. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Huang, Z.; Li, G.; Zhao, H.; Xing, X.; Sun, W.; Li, H.; Gou, Z.; Bao, C. Novel mutation breeding method for Streptomyces avermitilis using an atmospheric pressure glow discharge plasma. J. Appl. Microbiol. 2010, 108, 851–858. [Google Scholar] [CrossRef]
- Bauer, A.W.; Kirby, W.M.; Sherris, J.C.; Turck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 1966, 45, 493–496. [Google Scholar] [CrossRef]
- Daba, H.; Pandian, S.; Gosselin, J.F.; Simard, R.E.; Huang, J.; Lacroix, C. Detection and activity of a bacteriocin produced by Leuconostoc mesenteroides. Appl. Environ. Microbiol. 1991, 57, 3450–3455. [Google Scholar] [CrossRef] [Green Version]
- Chalasani, A.G.; Dhanarajan, G.; Nema, S.; Sen, R.; Roy, U. An Antimicrobial Metabolite from Bacillus sp.: Significant Activity Against Pathogenic Bacteria Including Multidrug-Resistant Clinical Strains. Front. Microbiol. 2015, 6, 1335. [Google Scholar] [CrossRef] [Green Version]
- Song, J.; Wang, Y.; Song, Y.; Zhao, B.; Wang, H.; Zhou, S.; Kong, D.; Guo, X.; Li, Y.; He, M.; et al. Brevibacillus halotolerans sp. nov., isolated from saline soil of a paddy field. Int. J. Syst. Evol. Microbiol. 2017, 67, 772–777. [Google Scholar] [CrossRef]
- Adrio, J.L.; Demain, A.L. Genetic improvement of processes yielding microbial products. FEMS Microbiol. Rev. 2006, 30, 187–214. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, X.-F.; Li, H.-P.; Wang, L.-Y.; Zhang, C.; Xing, X.-H.; Bao, C.-Y. Atmospheric and room temperature plasma (ARTP) as a new powerful mutagenesis tool. Appl. Microbiol. Biotechnol. 2014, 98, 5387–5396. [Google Scholar] [CrossRef]
- Zhang, X.; Zhang, C.; Zhou, Q.-Q.; Zhang, X.-F.; Wang, L.-Y.; Chang, H.-B.; Li, H.-P.; Oda, Y.; Xing, X.-H. Quantitative evaluation of DNA damage and mutation rate by atmospheric and room-temperature plasma (ARTP) and conventional mutagenesis. Appl. Microbiol. Biotechnol. 2015, 99, 5639–5646. [Google Scholar] [CrossRef]
- Wende, K.; Bekeschus, S.; Schmidt, A.; Jatsch, L.; Hasse, S.; Weltmann, K.; Masur, K.; von Woedtke, T. Risk assessment of a cold argon plasma jet in respect to its mutagenicity. Mutat. Res. Toxicol. Environ. Mutagen. 2016, 798–799, 48–54. [Google Scholar] [CrossRef]
- Adhikari, B.C.; Lamichhane, P.; Lim, J.S.; Nguyen, L.N.; Choi, E.H. Generation of reactive species by naturally sucked air in the Ar plasma jet. Results Phys. 2021, 30, 104863. [Google Scholar] [CrossRef]
- Oehmigen, K.; Hähnel, M.; Brandenburg, R.; Wilke, C.; Weltmann, K.-D.; Von Woedtke, T. The Role of Acidification for Antimicrobial Activity of Atmospheric Pressure Plasma in Liquids. Plasma Process. Polym. 2010, 7, 250–257. [Google Scholar] [CrossRef]
- Bolouki, N.; Kuan, W.-H.; Huang, Y.-Y.; Hsieh, J.-H. Characterizations of a Plasma-Water System Generated by Repetitive Microsecond Pulsed Discharge with Air, Nitrogen, Oxygen, and Argon Gases Species. Appl. Sci. 2021, 11, 6158. [Google Scholar] [CrossRef]
- Ghimire, B.; Szili, E.J.; Patenall, B.L.; Lamichhane, P.; Gaur, N.; Robson, A.J.; Trivedi, D.; Thet, N.T.; Jenkins, A.T.A.; Choi, E.H.; et al. Enhancement of hydrogen peroxide production from an atmospheric pressure argon plasma jet and implications to the antibacterial activity of plasma activated water. Plasma Sources Sci. Technol. 2021, 30, 035009. [Google Scholar] [CrossRef]
- Wang, L.; Zhao, H.; He, D.; Wu, Y.; Jin, L.; Li, G.; Su, N.; Li, H.; Xing, X.-H. Insights into the molecular-level effects of atmospheric and room-temperature plasma on mononucleotides and single-stranded homo- and hetero-oligonucleotides. Sci. Rep. 2020, 10, 14298. [Google Scholar] [CrossRef] [PubMed]
- Dizdaroglu, M.; Jaruga, P. Mechanisms of free radical-induced damage to DNA. Free Radic. Res. 2012, 46, 382–419. [Google Scholar] [CrossRef]
- Chatterjee, N.; Walker, G.C. Mechanisms of DNA damage, repair, and mutagenesis. Environ. Mol. Mutagen. 2017, 58, 235–263. [Google Scholar] [CrossRef] [Green Version]
- Cao, X.; Luo, Z.; Zeng, W.; Xu, S.; Zhao, L.; Zhou, J. Enhanced avermectin production by Streptomyces avermitilis ATCC 31267 using high-throughput screening aided by fluorescence-activated cell sorting. Appl. Microbiol. Biotechnol. 2018, 102, 703–712. [Google Scholar] [CrossRef]
- Zhang, C.; Qin, J.; Dai, Y.; Mu, W.; Zhang, T. Atmospheric and room temperature plasma (ARTP) mutagenesis enables xylitol over-production with yeast Candida tropicalis. J. Biotechnol. 2019, 296, 7–13. [Google Scholar] [CrossRef]
- Mitra, S.; Nguyen, L.N.; Akter, M.; Park, G.; Choi, E.H.; Kaushik, N.K. Impact of ROS Generated by Chemical, Physical, and Plasma Techniques on Cancer Attenuation. Cancers 2019, 11, 1030. [Google Scholar] [CrossRef] [Green Version]
- Liu, T.; Huang, Z.; Gui, X.; Xiang, W.; Jin, Y.; Chen, J.; Zhao, J. Multi-omics Comparative Analysis of Streptomyces Mutants Obtained by Iterative Atmosphere and Room-Temperature Plasma Mutagenesis. Front. Microbiol. 2021, 11, 630309. [Google Scholar] [CrossRef]
- Yang, X.; Yousef, A.E. Antimicrobial peptides produced by Brevibacillus spp.: Structure, classification and bioactivity: A mini review. World J. Microbiol. Biotechnol. 2018, 34, 57. [Google Scholar] [CrossRef]
- Choyam, S.; Jain, P.M.; Kammara, R. Characterization of a Potent New-Generation Antimicrobial Peptide of Bacillus. Front. Microbiol. 2021, 12, 710741. [Google Scholar] [CrossRef]
| Isolates | S. aureus TISTR 517 (mm) | MRSA Isolate 142 (mm) | MRSA Isolate 1096 (mm) | MRSA Isolate 2468 (mm) |
|---|---|---|---|---|
| Wild-type | 18.72 ± 1.54 | 19.45 ± 1.39 | 20.07 ± 1.67 | 19.79 ± 1.46 |
| M30 | 18.34 ± 0.47 | 18.91 ± 0.48 | 19.84 ± 0.61 | 19.87 ± 0.27 |
| M64 | 17.65 ± 0.34 | 18.12 ± 0.34 | 19.44 ± 0.26 | 18.99 ± 0.13 |
| M71 | 18.80 ± 0.35 | 18.90 ± 0.35 | 19.99 ± 0.59 | 19.87 ± 0.14 |
| M95 | 17.51 ± 0.65 | 18.46 ± 0.54 | 18.45 ± 0.84 | 18.51 ± 0.27 |
| M97 | 18.27 ± 0.23 | 18.92 ± 0.47 | 18.66 ± 0.09 | 19.04 ± 0.50 |
| M151 | 18.24 ± 0.00 | 19.45 ± 0.00 | 19.57 ± 0.25 | 19.71 ± 0.14 |
| M152 | 18.56 ± 0.28 | 19.69 ± 0.25 | 20.07 ± 0.00 | 20.04 ± 0.24 |
| M156 | 19.00 ± 0.48 | 19.71 ± 0.26 | 20.99 ± 0.90 | 20.62 ± 0.95 |
| M169 | 19.55 ± 0.73 | 20.33 ± 0.31 * | 21.08 ± 0.59 | 20.71 ± 0.71 |
| M187 | 19.64 ± 0.33 * | 19.98 ± 0.46 | 21.07 ± 0.33 * | 19.98 ± 0.84 |
| M196 | 20.09 ± 0.73 | 20.86 ± 0.44 * | 22.08 ± 0.45 * | 21.35 ± 0.71 * |
| M251 | 19.86 ± 0.56 * | 21.37 ± 0.75 * | 22.10 ± 0.29 * | 21.40 ± 0.73 * |
| M252 | 19.22 ± 0.90 | 20.55 ± 0.28 * | 20.90 ± 0.50 | 19.89 ± 0.76 |
| M270 | 19.87 ± 0.91 | 21.19 ± 0.66 * | 21.28 ± 0.92 | 20.50 ± 0.15 |
| M276 | 20.89 ± 0.71 * | 21.10 ± 0.57 * | 21.46 ± 0.30 * | 20.86 ± 0.48 * |
| M285 | 20.97 ± 0.39 * | 21.46 ± 0.58* | 22.76 ± 0.89* | 22.48 ± 1.01 * |
| M332 | 19.62 ± 1.01 | 19.90 ± 0.68 | 20.33 ± 0.27 | 20.25 ± 0.83 |
| M334 | 19.35 ± 0.31 | 19.81 ± 0.42 | 19.10 ± 0.65 | 19.88 ± 0.55 |
| M347 | 18.73 ± 0.27 | 19.37 ± 0.56 | 19.09 ± 0.15 | 19.44 ± 0.66 |
| M368 | 19.53 ± 0.53 | 20.36 ± 0.63 | 20.34 ± 0.27 | 20.23 ± 0.56 |
| M371 | 20.07 ± 0.28 * | 20.55 ± 0.48 * | 21.42 ± 0.50 * | 20.59 ± 0.46 |
| M374 | 19.62 ± 0.42 * | 20.27 ± 0.49 | 20.78 ± 0.17 | 20.41 ± 0.31 |
| M396 | 19.62 ± 0.42 * | 20.46 ± 0.43 * | 20.52 ± 0.67 | 20.50 ± 0.42 |
| M402 | 18.55 ± 0.62 | 19.72 ± 0.73 | 19.72 ± 0.67 | 19.88 ± 0.31 |
| M403 | 20.16 ± 0.15 * | 20.91 ± 0.32 * | 21.24 ± 0.43 * | 20.77 ± 0.16 * |
| Vancomycin (30 µg) | 21.77 ± 0.13 | 23.06 ± 0.16 | 24.18 ± 0.29 | 25.46 ± 0.68 |
| Cefoxitin (30 µg) | 30.60 ± 0.40 | 0.00 ± 0.00 | 0.00 ± 0.00 | 0.00 ± 0.00 |
| Oxacillin (1 µg) | 27.70 ± 0.35 | 0.00 ± 0.00 | 0.00 ± 0.00 | 0.00 ± 0.00 |
| Antibiotics | Wild-Type (mm) | M251 (mm) | M276 (mm) | M285 (mm) |
|---|---|---|---|---|
| Cefoxitin (30 µg) | 43.01 ± 1.92 | 45.38 ± 0.59 | 43.35 ± 0.29 | 46.74 ± 0.00 |
| Ceftriaxone (30 µg) | 44.03 ± 0.78 | 45.72 ± 0.00 | 46.23 ± 1.02 * | 46.57 ± 0.78 * |
| Ciprofloxacin (5 µg) | 46.57 ± 1.06 | 46.91 ± 0.29 | 48.26 ± 1.76 | 48.94 ± 0.29 |
| Doxycycline (30 µg) | 54.86 ± 0.88 | 56.90 ± 3.52 | 56.73 ± 1.55 | 58.59 ± 1.06 |
| Erythromycin (15 µg) | 46.57 ± 0.78 | 46.91 ± 0.29 | 46.74 ± 1.34 | 48.60 ± 0.29 * |
| Gentamycin (10 µg) | 26.08 ± 0.29 | 25.40 ± 1.34 | 24.89 ± 1.34 | 27.60 ± 0.78 |
| Imipenem (10 µg) | 44.20 ± 0.51 | 46.91 ± 2.05 | 45.89 ± 1.28 | 47.92 ± 0.78 * |
| Vancomycin (30 µg) | 26.25 ± 0.29 | 26.92 ± 1.02 | 27.60 ± 0.59 * | 28.62 ± 0.59 * |
| Isolates | Antibacterial Activity (AU/mL) | |||
|---|---|---|---|---|
| MIC | MBC | |||
| Wild-Type | M285 | Wild-Type | M285 | |
| S. aureus TISTR 517 | 80.00 | 160.00 | 80.00 | 160.00 |
| MRSA isolate 142 | 80.00 | 160.00 | 40.00 | 40.00 |
| MRSA isolate 1096 | 80.00 | 160.00 | 40.00 | 80.00 |
| MRSA isolate 2468 | 80.00 | 160.00 | 40.00 | 80.00 |
| Conditions | % Remaining Activity | |||
|---|---|---|---|---|
| S. aureus TISTR 517 | MRSA Isolate 2468 | |||
| Wild-Type | M285 | Wild-Type | M285 | |
| Untreated sample | 100.00 ± 3.28 | 100.00 ± 2.45 | 100.00 ± 2.23 | 100.00 ± 1.37 |
| Thermal stability | ||||
| Sample at 60 °C, 1 h | 97.07 ± 1.42 * | 94.35 ± 1.49 * | 96.22 ± 2.16 | 94.47 ± 3.47 |
| Sample at 80 °C, 1 h | 94.10 ± 1.41 * | 92.96 ± 0.10 * | 89.62 ± 1.67 * | 93.54 ± 4.03 |
| Sample at 100 °C, 1 h | 91.13 ± 0.20 * | 90.61 ± 1.63 * | 88.21 ± 0.77 * | 91.18 ± 3.23 * |
| Sample at 121 °C, 15 psi, 15 min (autoclave) | 95.59 ± 1.38 * | 92.00 ± 2.26 * | 91.52 ± 1.36 * | 92.54 ± 0.56 * |
| Proteolytic enzyme stability | ||||
| Proteinase K (1 mg/mL) | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * |
| Sample + Proteinase K (1 mg/mL) | 99.03 ± 1.67 | 92.13 ± 1.60 * | 99.06 ± 1.63 | 95.34 ± 2.10 |
| Trypsin (1 mg/mL) | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * |
| Sample + Trypsin (1 mg/mL) | 100.54 ± 3.83 | 95.83 ± 1.39 * | 99.05 ± 0.83 | 99.07 ± 1.60 |
| α-chymotrypsin (1 mg/mL) | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * |
| Sample + α-chymotrypsin (1 mg/mL) | 97.54 ± 2.27 | 93.98 ± 2.12 * | 97.12 ± 1.47 * | 97.67 ± 2.13 |
| Surfactant stability | ||||
| 1% SDS | 113.93 ± 1.74 * | 111.98 ± 1.09 * | 114.02 ± 1.01 * | 110.33 ± 2.93 * |
| Sample + 1% SDS | 114.44 ± 1.93 * | 111.51 ± 2.66 * | 111.63 ± 4.01 * | 110.33 ± 3.54 * |
| 1% Triton X-100 | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * | 0.00 ± 0.00 * |
| Sample + 1% Triton X | 109.48 ± 3.60 * | 107.66 ± 0.81 * | 109.68 ± 1.81 * | 107.04 ± 1.41 * |
| pH stability | ||||
| pH 1 | 103.51 ± 3.52 | 101.46 ± 2.93 | 100.51 ± 0.87 | 102.82 ± 2.44 |
| pH 2 | 101.52 ± 2.62 | 99.05 ± 2.22 | 100.58 ± 3.71 | 100.48 ± 2.91 |
| pH 3 | 102.01 ± 2.32 | 100.03 ± 2.90 | 102.09 ± 6.22 | 100.48 ± 1.62 |
| pH 4 | 100.54 ± 3.08 | 100.51 ± 3.01 | 100.18 ± 6.45 | 100.95 ± 2.14 |
| pH 5 | 102.03 ± 3.78 | 101.47 ± 2.55 | 101.05 ± 3.11 | 99.54 ± 0.80 |
| pH 6 | 102.01 ± 2.32 | 100.50 ± 1.67 | 101.52 ± 2.62 | 100.47 ± 0.81 |
| pH 7 | 100.51 ± 0.87 | 100.49 ± 0.85 | 100.51 ± 0.87 | 100.47 ± 0.81 |
| pH 8 | 99.54 ± 0.79 | 100.49 ± 0.85 | 100.48 ± 2.14 | 100.01 ± 1.40 |
| pH 9 | 95.36 ± 0.72 * | 100.99 ± 0.86 | 100.48 ± 1.62 | 100.93 ± 0.81 |
| pH 10 | 95.83 ± 1.33 * | 100.49 ± 0.85 | 100.93 ± 0.81 | 100.93 ± 0.81 |
| pH 11 | 94.89 ± 0.77 * | 100.01 ± 1.47 | 100.46 ± 0.80 | 100.93 ± 0.81 |
| pH 12 | 93.49 ± 0.76 * | 100.50 ± 0.86 | 100.00 ± 0.00 | 100.93 ± 1.60 |
| pH 13 | 93.03 ± 1.29 * | 100.49 ± 0.85 | 100.47 ± 0.81 | 100.46 ± 0.80 |
| pH 14 | 92.10 ± 0.76 * | 100.00 ± 0.00 | 100.46 ± 0.80 | 100.00 ± 1.39 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Songnaka, N.; Nisoa, M.; Atipairin, A.; Wanganuttara, T.; Chinnawong, T. Enhanced Antibacterial Activity of Brevibacillus sp. SPR19 by Atmospheric and Room Temperature Plasma Mutagenesis (ARTP). Sci. Pharm. 2022, 90, 23. https://doi.org/10.3390/scipharm90020023
Songnaka N, Nisoa M, Atipairin A, Wanganuttara T, Chinnawong T. Enhanced Antibacterial Activity of Brevibacillus sp. SPR19 by Atmospheric and Room Temperature Plasma Mutagenesis (ARTP). Scientia Pharmaceutica. 2022; 90(2):23. https://doi.org/10.3390/scipharm90020023
Chicago/Turabian StyleSongnaka, Nuttapon, Mudtorlep Nisoa, Apichart Atipairin, Thamonwan Wanganuttara, and Thapanee Chinnawong. 2022. "Enhanced Antibacterial Activity of Brevibacillus sp. SPR19 by Atmospheric and Room Temperature Plasma Mutagenesis (ARTP)" Scientia Pharmaceutica 90, no. 2: 23. https://doi.org/10.3390/scipharm90020023
APA StyleSongnaka, N., Nisoa, M., Atipairin, A., Wanganuttara, T., & Chinnawong, T. (2022). Enhanced Antibacterial Activity of Brevibacillus sp. SPR19 by Atmospheric and Room Temperature Plasma Mutagenesis (ARTP). Scientia Pharmaceutica, 90(2), 23. https://doi.org/10.3390/scipharm90020023