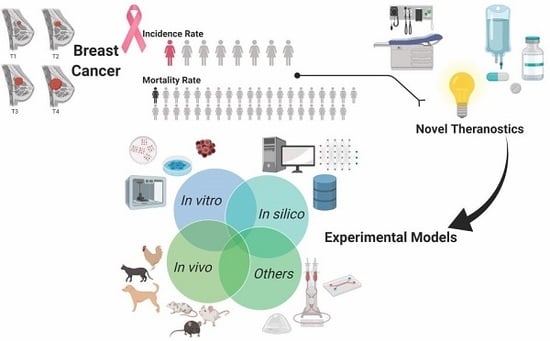
Experimental Models as Refined Translational Tools for Breast Cancer Research
Abstract
1. Introduction
2. Animal Models in Breast Cancer
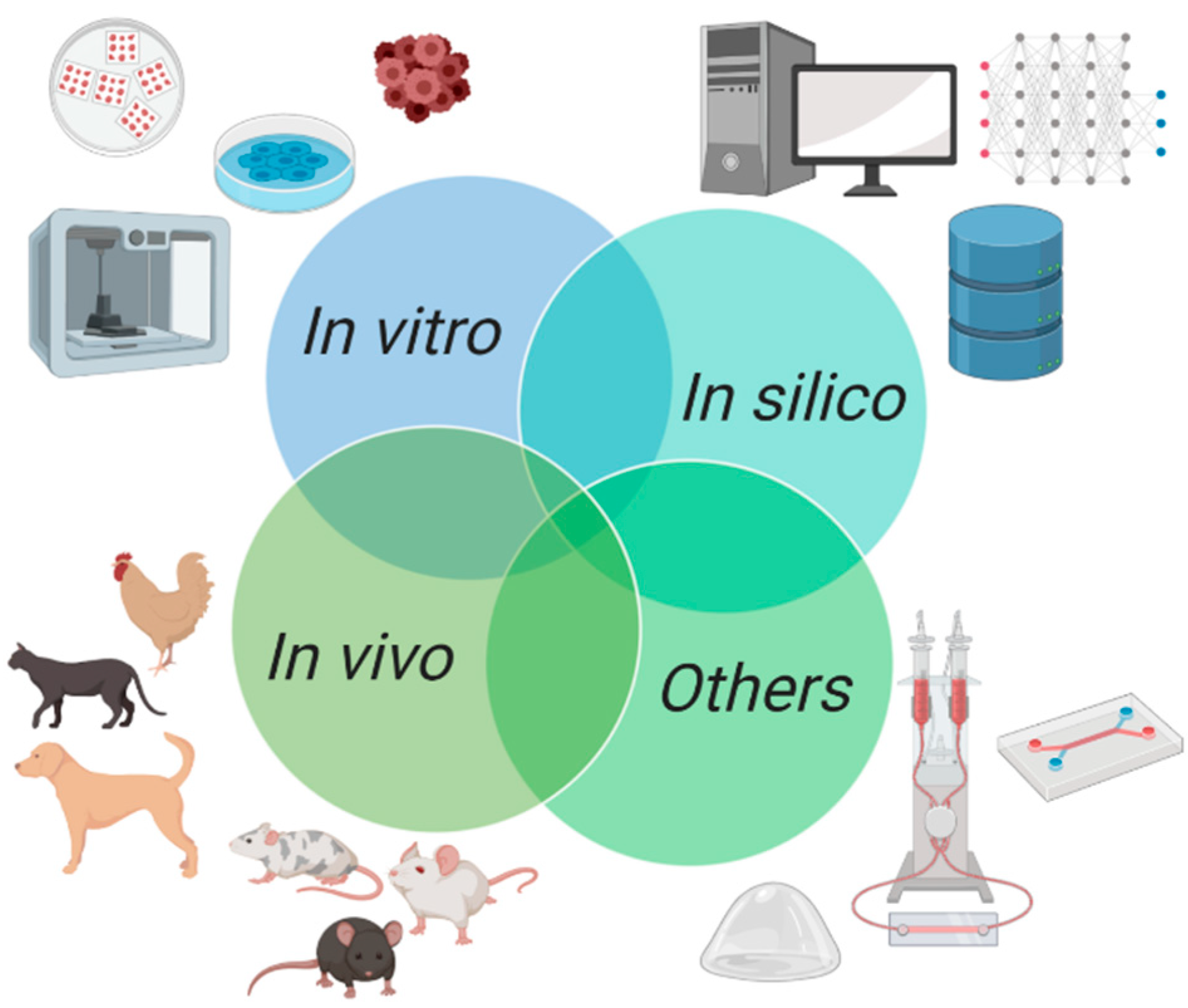
2.1. In Vitro Models in Breast Cancer
2.1.1. 2D Models
2.1.2. 3D Models
Tissue Slice Models
Organoids
Spheroids
Scaffold-Based Models
2.2. In Vivo Models in Breast Cancer
2.2.1. Canine and Feline Models
2.2.2. Murine Models
Chemically-Induced Models
Transplanted Tumors Models
Genetically Engineered Models
Radiation Models
2.3. In Silico Models in Breast Cancer
2.4. Other Models Helpful in Breast Cancer
3. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Carioli, G.; Malvezzi, M.; Rodriguez, T.; Bertuccio, P.; Negri, E.; La Vecchia, C. Trends and predictions to 2020 in breast cancer mortality in Europe. Breast 2017, 36, 89–95. [Google Scholar] [CrossRef] [PubMed]
- Malhotra, G.K.; Zhao, X.; Band, H.; Band, V. Histological, molecular and functional subtypes of breast cancers. Cancer Biol. Ther. 2010, 10, 955–960. [Google Scholar] [CrossRef] [PubMed]
- Bansal, C.; Singh, U.S.; Misra, S.; Sharma, K.L.; Tiwari, V.; Srivastava, A.N. Comparative evaluation of the modified Scarff-Bloom-Richardson grading system on breast carcinoma aspirates and histopathology. Cytojournal 2012, 9, 4. [Google Scholar] [CrossRef] [PubMed]
- Singletary, S.E.; Allred, C.; Ashley, P.; Bassett, L.W.; Berry, D.; Bland, K.I.; Borgen, P.I.; Clark, G.; Edge, S.B.; Hayes, D.F.; et al. Revision of the American Joint Committee on Cancer staging system for breast cancer. J. Clin. Oncol. 2002, 20, 3628–3636. [Google Scholar] [CrossRef]
- American Joint Committee on Cancer. AJCC Cancer Staging Manual, 7th ed.; Edge, S.B., Byrd, D.R., Compton, C.C., Fritz, A.G., Greene, F.L., Trotti, A., Eds.; Springer: Chicago, IL, USA, 2010; ISBN 978-0-387-88440-0. [Google Scholar]
- Sotiriou, C.; Pusztai, L. Gene-expression signatures in breast cancer. N. Engl. J. Med. 2009, 360, 790–800. [Google Scholar] [CrossRef]
- Li, J.; Chen, Z.; Su, K.; Zeng, J. Clinicopathological classification and traditional prognostic indicators of breast cancer. Int. J. Clin. Exp. Pathol. 2015, 8, 8500–8505. [Google Scholar]
- Rakha, E.A.; Reis-Filho, J.S.; Ellis, I.O. Combinatorial biomarker expression in breast cancer. Breast Cancer Res. Treat. 2010, 120, 293–308. [Google Scholar] [CrossRef]
- Prat, A.; Karginova, O.; Parker, J.S.; Fan, C.; He, X.; Bixby, L.; Harrell, J.C.; Roman, E.; Adamo, B.; Troester, M.; et al. Characterization of cell lines derived from breast cancers and normal mammary tissues for the study of the intrinsic molecular subtypes. Breast Cancer Res. Treat. 2013, 142, 237–255. [Google Scholar] [CrossRef]
- Eroles, P.; Bosch, A.; Perez-Fidalgo, J.A.; Lluch, A. Molecular biology in breast cancer: Intrinsic subtypes and signaling pathways. Cancer Treat. Rev. 2012, 38, 698–707. [Google Scholar] [CrossRef]
- Giovannelli, P.; Di Donato, M.; Galasso, G.; Di Zazzo, E.; Bilancio, A.; Migliaccio, A. The androgen receptor in breast cancer. Front. Endocrinol. (Lausanne) 2018, 9, 1–8. [Google Scholar] [CrossRef]
- Sunar, V.; Dogan, H.T.; Sarici, F.; Ates, O.; Akin, S.; Baspinar, B.; Aksoy, S.; Altundag, K. Association between androgen receptor status and prognosis in triple negative breast cancer. JBUON 2018, 23, 1325–1330. [Google Scholar]
- Gerratana, L.; Basile, D.; Buono, G.; De Placido, S.; Giuliano, M.; Minichillo, S.; Coinu, A.; Martorana, F.; De Santo, I.; Del Mastro, L.; et al. Androgen receptor in triple negative breast cancer: A potential target for the targetless subtype. Cancer Treat. Rev. 2018, 68, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Perou, C.M.; Borresen-Dale, A.-L. Systems biology and genomics of breast cancer. Cold Spring Harb. Perspect. Biol. 2011, 3. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Cheng, H.; Bai, Z.; Li, J. Breast cancer cell line classification and Its relevance with breast tumor subtyping. J. Cancer 2017, 8, 3131–3141. [Google Scholar] [CrossRef] [PubMed]
- Subik, K.; Lee, J.-F.; Baxter, L.; Strzepek, T.; Costello, D.; Crowley, P.; Xing, L.; Hung, M.-C.; Bonfiglio, T.; Hicks, D.G.; et al. The Expression Patterns of ER, PR, HER2, CK5/6, EGFR, Ki-67 and AR by Immunohistochemical Analysis in Breast Cancer Cell Lines. Breast Cancer (Auckl.) 2010, 4, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Goldhirsch, A.; Winer, E.P.; Coates, A.S.; Gelber, R.D.; Piccart-Gebhart, M.; Thurlimann, B.; Senn, H.-J. Personalizing the treatment of women with early breast cancer: Highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2013, 24, 2206–2223. [Google Scholar] [CrossRef]
- Vuong, D.; Simpson, P.T.; Green, B.; Cummings, M.C.; Lakhani, S.R. Molecular classification of breast cancer. Virchows Arch. 2014, 465, 1–14. [Google Scholar] [CrossRef]
- Holliday, D.L.; Speirs, V. Choosing the right cell line for breast cancer research. Breast Cancer Res. 2011, 13, 215. [Google Scholar] [CrossRef]
- American Cancer Society Breast Cancer: Treatment and Side Effects. Available online: https://www.cancer.org/cancer/breast-cancer.html (accessed on 7 December 2019).
- American Cancer Society. Breast Cancer: Treatment Guideline for Patients, Version VIII. J. Natl. Compr. Cancer Netw. 2006. [Google Scholar]
- Waks, A.G.; Winer, E.P. Breast Cancer Treatment: A Review. JAMA 2019, 321, 288–300. [Google Scholar] [CrossRef]
- Gradishar, W.J.; Anderson, B.O.; Balassanian, R.; Blair, S.L.; Burstein, H.J.; Cyr, A.; Elias, A.D.; Farrar, W.B.; Forero, A.; Giordano, S.H.; et al. Clinical practice guidelines in oncology. J. Natl. Compr. Cancer Netw. 2018, 16, 310–320. [Google Scholar] [CrossRef] [PubMed]
- Castaneda, S.A.; Strasser, J. Updates in the Treatment of Breast Cancer with Radiotherapy. Surg. Oncol. Clin. N. Am. 2017, 26, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Deng, X.; Wu, H.; Gao, F.; Su, Y.; Li, Q.; Liu, S.; Cai, J. Brachytherapy in the treatment of breast cancer. Int. J. Clin. Oncol. 2017, 22, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Giugliano, F.M.; Falivene, S.; Esposito, E.; Di Franco, R.; Muto, M.; D’Aiuto, M.; Muto, P. External radiotherapy for breast cancer in the elderly. Aging Clin. Exp. Res. 2017, 29, 149–157. [Google Scholar] [CrossRef]
- Meisel, J.L.; Venur, V.A.; Gnant, M.; Carey, L. Evolution of Targeted Therapy in Breast Cancer: Where Precision Medicine Began. In American Society of Clinical Oncology Educational Book; American Society of Clinical Oncology: Alexandria, VA, USA, 2018; Volume 38, pp. 78–86. [Google Scholar] [CrossRef]
- Munagala, R.; Aqil, F.; Gupta, R. Promising molecular targeted therapies in breast cancer. Indian J. Pharmacol. 2011, 43, 236–245. [Google Scholar] [CrossRef]
- Liyanage, P.Y.; Hettiarachchi, S.D.; Zhou, Y.; Ouhtit, A.; Seven, E.S.; Oztan, C.Y.; Celik, E.; Leblanc, R.M. Nanoparticle-mediated targeted drug delivery for breast cancer treatment. Biochim. Biophys. Acta Rev. Cancer 2019, 1871, 419–433. [Google Scholar] [CrossRef]
- Tray, N.; Adams, S.; Esteva, F.J. Antibody-drug conjugates in triple negative breast cancer. Future Oncol. 2018, 14, 2651–2661. [Google Scholar] [CrossRef]
- Perez-Garcia, J.; Munoz-Couselo, E.; Cortes, J.; Scaltriti, M. Therapeutic antibodies in breast cancer. Semin. Oncol. 2014, 41, 576–588. [Google Scholar] [CrossRef]
- Ernst, B.; Anderson, K.S. Immunotherapy for the treatment of breast cancer. Curr. Oncol. Rep. 2015, 17, 5. [Google Scholar] [CrossRef]
- Heimes, A.-S.; Schmidt, M. Atezolizumab for the treatment of triple-negative breast cancer. Expert Opin. Investig. Drugs 2019, 28, 1–5. [Google Scholar] [CrossRef]
- Schmid, P.; Adams, S.; Rugo, H.S.; Schneeweiss, A.; Barrios, C.H.; Iwata, H.; Dieras, V.; Hegg, R.; Im, S.-A.; Shaw Wright, G.; et al. Atezolizumab and Nab-Paclitaxel in Advanced Triple-Negative Breast Cancer. N. Engl. J. Med. 2018, 379, 2108–2121. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Xu, J.; Tang, L.; Guan, X. Breast cancer stem cell: The roles and therapeutic implications. Cell. Mol. Life Sci. 2017, 74, 951–966. [Google Scholar] [CrossRef] [PubMed]
- Giovannelli, P.; Di Donato, M.; Galasso, G.; Di Zazzo, E.; Medici, N.; Bilancio, A.; Migliaccio, A.; Castoria, G. Breast cancer stem cells: The role of sex steroid receptors. World J. Stem Cells 2019, 11, 594–603. [Google Scholar] [CrossRef] [PubMed]
- O’Conor, C.J.; Chen, T.; González, I.; Cao, D.; Peng, Y. Cancer stem cells in triple-negative breast cancer: A potential target and prognostic marker. Biomark. Med. 2018, 12, 813–820. [Google Scholar] [CrossRef] [PubMed]
- El-Abd, E.; Shalaby, E.; Matalkah, F. Animal Models of Breast Cancer. In Omics Approaches in Breast Cancer: Towards Next-Generation Diagnosis, Prognosis and Therapy; Springer: Berlin, Germany, 2014; pp. 297–314. [Google Scholar]
- Barré-Sinoussi, F.; Montagutelli, X. Animal models are essential to biological research: Issues and perspectives. Future Sci. OA 2015, 1, 4–6. [Google Scholar] [CrossRef] [PubMed]
- Burdall, S.E.; Hanby, A.M.; Lansdown, M.R.J.; Speirs, V. Breast cancer cell lines: Friend or foe? Breast Cancer Res. 2003, 5, 89–95. [Google Scholar] [CrossRef]
- Liu, K.; Newbury, P.A.; Glicksberg, B.S.; Zeng, W.Z.D.; Paithankar, S.; Andrechek, E.R.; Chen, B. Evaluating cell lines as models for metastatic breast cancer through integrative analysis of genomic data. Nat. Commun. 2019, 10, 2138. [Google Scholar] [CrossRef]
- Holen, I.; Speirs, V.; Morrissey, B.; Blyth, K. In vivo models in breast cancer research: Progress, challenges and future directions. Dis. Model. Mech. 2017, 10, 359–371. [Google Scholar] [CrossRef]
- Akbari Bazm, M.; Naseri, L.; Khazaei, M. Methods of inducing breast cancer in animal models: A systemic review. World Cancer Res. J. 2018, 5, 1182. [Google Scholar]
- Jean-Quartier, C.; Jeanquartier, F.; Jurisica, I.; Holzinger, A. In silico cancer research towards 3R. BMC Cancer 2018, 18, 1–12. [Google Scholar] [CrossRef]
- Colquitt, R.B.; Colquhoun, D.A.; Thiele, R.H. In silico modelling of physiologic systems. Best Pract. Res. Clin. Anaesthesiol. 2011, 25, 499–510. [Google Scholar] [CrossRef] [PubMed]
- Sneddon, L.U.; Halsey, L.G.; Bury, N.R. Considering aspects of the 3Rs principles within experimental animal biology. J. Exp. Biol. 2017, 220, 3007–3016. [Google Scholar] [CrossRef] [PubMed]
- Parker, R.M.A.; Browne, W.J. The place of experimental design and statistics in the 3Rs. Inst. Lab. Anim. Res. J. 2014, 55, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Franco, N.H.; Olsson, I.A.S. Scientists and the 3Rs: Attitudes to animal use in biomedical research and the effect of mandatory training in laboratory animal science. Lab. Anim. 2014, 48, 50–60. [Google Scholar] [CrossRef] [PubMed]
- Workman, P.; Aboagye, E.O.; Balkwill, F.; Balmain, A.; Bruder, G.; Chaplin, D.J.; Double, J.A.; Everitt, J.; Farningham, D.A.H.; Glennie, M.J.; et al. Guidelines for the welfare and use of animals in cancer research. Br. J. Cancer 2010, 102, 1555–1577. [Google Scholar] [CrossRef] [PubMed]
- Guillen, J. FELASA guidelines and recommendations. J. Am. Assoc. Lab. Anim. Sci. 2012, 51, 311–321. [Google Scholar]
- Russel, W.M.S.; Burch, R.L. The Principles of Humane Experimental Technique; Methuen & Co. Ltd.: London, UK, 1959. [Google Scholar]
- Tannenbaum, J.; Bennett, B.T. Russell and Burch’s 3Rs then and now: The need for clarity in definition and purpose. J. Am. Assoc. Lab. Anim. Sci. 2015, 54, 120–132. [Google Scholar]
- Council of Europe. European Convention for the Protection of Vertebrate Animals used for Experimental and Other Scientific Purposes; European Treaty Series No. 123; Council of Europe: Strasbourg, France, 1986. [Google Scholar]
- The European Parliament and the Council of the European Union. Directive 2010/63/EU of the European Parliament and of the Council of 22 September 2010 on the protection of animals used for scientific purposes Text with EEA relevance. Off. J. Eur. Union 2010, 53, 33–79. [Google Scholar] [CrossRef]
- European Animal Research Association. European Animal Research Association: Goals and Milestones. Available online: http://eara.eu/en/about-us/info-brochure/ (accessed on 8 December 2019).
- Cavanaugh, P.; Haier, J. Basic Tissue and Cell Culture in Cancer Research. In The Cancer Handbook; American Cancer Society: Atlanta, GA, USA, 2007; ISBN 9780470025079. [Google Scholar]
- Forozan, F.; Mahlamaki, E.H.; Monni, O.; Chen, Y.; Veldman, R.; Jiang, Y.; Gooden, G.C.; Ethier, S.P.; Kallioniemi, A.; Kallioniemi, O.P. Comparative genomic hybridization analysis of 38 breast cancer cell lines: A basis for interpreting complementary DNA microarray data. Cancer Res. 2000, 60, 4519–4525. [Google Scholar]
- Lasfargues, E.Y.; Ozzello, L. Cultivation of Human Breast Carcinomas. JNCI J. Natl. Cancer Inst. 1958, 21, 1131–1147. [Google Scholar] [CrossRef]
- Soule, H.D.; Vazquez, J.; Long, A.; Albert, S.; Brennan, M. A Human Cell Line From a Pleural Effusion Derived From a Breast Carcinoma2. JNCI J. Natl. Cancer Inst. 1973, 51, 1409–1416. [Google Scholar] [CrossRef] [PubMed]
- International Cell Line Authentication Committee Naming a Cell Line. Available online: https://iclac.org/resources/cell-line-names/ (accessed on 9 December 2019).
- Cailleau, R.; Olive, M.; Cruciger, Q.V. Long-term human breast carcinoma cell lines of metastatic origin: Preliminary characterization. In Vitro 1978, 14, 911–915. [Google Scholar] [CrossRef] [PubMed]
- Band, V.; Zajchowski, D.; Swisshelm, K.; Trask, D.; Kulesa, V.; Cohen, C.; Connolly, J.; Sager, R. Tumor progression in four mammary epithelial cell lines derived from the same patient. Cancer Res. 1990, 50, 7351–7357. [Google Scholar] [PubMed]
- Ethier, S.P.; Mahacek, M.L.; Gullick, W.J.; Frank, T.S.; Weber, B.L. Differential isolation of normal luminal mammary epithelial cells and breast cancer cells from primary and metastatic sites using selective media. Cancer Res. 1993, 53, 627–635. [Google Scholar]
- Selenius, L.A.; Wallenberg Lundgren, M.; Jawad, R.; Danielsson, O.; Bjornstedt, M. The Cell Culture Medium Affects Growth, Phenotype Expression and the Response to Selenium Cytotoxicity in A549 and HepG2 Cells. Antioxidants 2019, 8, 130. [Google Scholar] [CrossRef]
- Arora, M. Cell Culture Media: A Review. Mater. Methods 2013, 3. [Google Scholar] [CrossRef]
- Garcia-Carpizo, V.; Ruiz-Llorente, S.; Sarmentero, J.; Gonzalez-Corpas, A.; Barrero, M.J. CREBBP/EP300 bromodomain inhibition affects the proliferation of AR-positive breast cancer cell lines. Mol. Cancer Res. 2019, 17, 720–730. [Google Scholar] [CrossRef]
- Barton, V.N.; D’Amato, N.C.; Gordon, M.A.; Lind, H.T.; Spoelstra, N.S.; Babbs, B.L.; Heinz, R.E.; Elias, A.; Jedlicka, P.; Jacobsen, B.M.; et al. Multiple Molecular Subtypes of Triple-Negative Breast Cancer Critically Rely on Androgen Receptor and Respond to Enzalutamide In Vivo. Mol. Cancer Ther. 2015, 14, 769–778. [Google Scholar] [CrossRef]
- Riaz, M.; van Jaarsveld, M.T.M.; Hollestelle, A.; Prager-van der Smissen, W.J.C.; Heine, A.A.J.; Boersma, A.W.M.; Liu, J.; Helmijr, J.; Ozturk, B.; Smid, M.; et al. miRNA expression profiling of 51 human breast cancer cell lines reveals subtype and driver mutation-specific miRNAs. Breast Cancer Res. 2013, 15, R33. [Google Scholar] [CrossRef]
- Neve, R.M.; Chin, K.; Fridlyand, J.; Yeh, J.; Baehner, F.L.; Fevr, T.; Clark, L.; Bayani, N.; Coppe, J.-P.; Tong, F.; et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 2006, 10, 515–527. [Google Scholar] [CrossRef]
- Hollestelle, A.; Nagel, J.H.A.; Smid, M.; Lam, S.; Elstrodt, F.; Wasielewski, M.; Ng, S.S.; French, P.J.; Peeters, J.K.; Rozendaal, M.J.; et al. Distinct gene mutation profiles among luminal-type and basal-type breast cancer cell lines. Breast Cancer Res. Treat. 2010, 121, 53–64. [Google Scholar] [CrossRef] [PubMed]
- Kao, J.; Salari, K.; Bocanegra, M.; Choi, Y.-L.; Girard, L.; Gandhi, J.; Kwei, K.A.; Hernandez-Boussard, T.; Wang, P.; Gazdar, A.F.; et al. Molecular profiling of breast cancer cell lines defines relevant tumor models and provides a resource for cancer gene discovery. PLoS ONE 2009, 4, e6146. [Google Scholar] [CrossRef] [PubMed]
- Lacroix, M.; Leclercq, G. Relevance of breast cancer cell lines as models for breast tumours: An update. Breast Cancer Res. Treat. 2004, 83, 249–289. [Google Scholar] [CrossRef] [PubMed]
- Speers, C.; Zhao, S.G.; Chandler, B.; Liu, M.; Wilder-Romans, K.; Olsen, E.; Nyati, S.; Ritter, C.; Alluri, P.G.; Kothari, V.; et al. Androgen receptor as a mediator and biomarker of radioresistance in triple-negative breast cancer. NPJ Breast Cancer 2017, 3, 1–10. [Google Scholar] [CrossRef]
- Kurebayashi, J.; Kurosumi, M.; Sonoo, H. A new human breast cancer cell line, KPL-1 secretes tumour-associated antigens and grows rapidly in female athymic nude mice. Br. J. Cancer 1995, 71, 845–853. [Google Scholar] [CrossRef]
- Briand, P.; Lykkesfeldt, A.E. An in vitro model of human breast carcinogenesis: Epigenetic aspects. Breast Cancer Res. Treat. 2001, 65, 179–187. [Google Scholar] [CrossRef]
- Micci, F.; Teixeira, M.R.; Heim, S. Complete cytogenetic characterization of the human breast cancer cell line MA11 combining G-banding, comparative genomic hybridization, multicolor fluorescence in situ hybridization, RxFISH, and chromosome-specific painting. Cancer Genet. Cytogenet. 2001, 131, 25–30. [Google Scholar] [CrossRef]
- Rye, P.D.; Norum, L.; Olsen, D.R.; Garman-Vik, S.; Kaul, S.; Fodstad, Ø. Brain metastasis model in athymic nude mice using a novel MUC1-secreting human breast-cancer cell line, MA11. Int. J. Cancer 1996, 68, 682–687. [Google Scholar] [CrossRef]
- Stadler, M.; Walter, S.; Walzl, A.; Kramer, N.; Unger, C.; Scherzer, M.; Unterleuthner, D.; Hengstschläger, M.; Krupitza, G.; Dolznig, H. Increased complexity in carcinomas: Analyzing and modeling the interaction of human cancer cells with their microenvironment. Semin. Cancer Biol. 2015, 35, 107–124. [Google Scholar] [CrossRef]
- Roberts, S.; Peyman, S.; Speirs, V. Current and Emerging 3D Models to Study Breast Cancer. In Breast Cancer Metastasis and Drug Resistance: Challenges and Progress; Ahmad, A., Ed.; Springer: Cham, Switzerland, 2019; pp. 413–427. ISBN 978-3-030-20301-6. [Google Scholar]
- Grosso, S.H.G.; Katayama, M.L.H.; Roela, R.A.; Nonogaki, S.; Soares, F.A.; Brentani, H.; Lima, L.; Folgueira, M.A.A.K.; Waitzberg, A.F.L.; Pasini, F.S.; et al. Breast cancer tissue slices as a model for evaluation of response to rapamycin. Cell Tissue Res. 2013, 352, 671–684. [Google Scholar] [CrossRef]
- Pennington, K.; Chu, Q.D.; Curiel, D.T.; Li, B.D.L.; Mathis, J.M. The utility of a tissue slice model system to determine breast cancer infectivity by oncolytic adenoviruses. J. Surg. Res. 2010, 163, 270–275. [Google Scholar] [CrossRef] [PubMed]
- Stoff-Khalili, M.A.; Stoff, A.; Rivera, A.A.; Banerjee, N.S.; Everts, M.; Young, S.; Siegal, G.P.; Richter, D.F.; Wang, M.; Dall, P.; et al. Preclinical evaluation of transcriptional targeting strategies for carcinoma of the breast in a tissue slice model system. Breast Cancer Res. 2005, 7, R1141–R1152. [Google Scholar] [CrossRef] [PubMed]
- Stoff-Khalili, M.A.; Stoff, A.; Rivera, A.A.; Mathis, J.M.; Everts, M.; Wang, M.; Kawakami, Y.; Waehler, R.; Mathews, Q.L.; Yamamoto, M.; et al. Gene transfer to carcinoma of the breast with fiber-modified adenoviral vectors in a tissue slice model system. Cancer Biol. Ther. 2005, 4, 1203–1220. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.A. Chapter 1—Organoids and mini-organs: Introduction, history, and potential. In Organs and Organoids; Davies, J.A., Lawrence, M.L., Eds.; Academic Press: Amsterdam, the Netherlands, 2018; pp. 3–23. ISBN 978-0-12-812636-3. [Google Scholar]
- Koledova, Z. 3D Coculture of Mammary Organoids with Fibrospheres: A Model for Studying Epithelial--Stromal Interactions During Mammary Branching Morphogenesis. In 3D Cell Culture: Methods and Protocols; Koledova, Z., Ed.; Springer: New York, NY, USA, 2017; pp. 107–124. ISBN 978-1-4939-7021-6. [Google Scholar]
- Djomehri, S.I.; Burman, B.; Gonzalez, M.E.; Takayama, S.; Kleer, C.G. A reproducible scaffold-free 3D organoid model to study neoplastic progression in breast cancer. J. Cell Commun. Signal. 2019, 13, 129–143. [Google Scholar] [CrossRef]
- Walsh, A.J.; Cook, R.S.; Sanders, M.E.; Aurisicchio, L.; Ciliberto, G.; Arteaga, C.L.; Skala, M.C. Quantitative Optical Imaging of Primary Tumor Organoid Metabolism Predicts Drug Response in Breast Cancer. Cancer Res. 2014. [Google Scholar] [CrossRef]
- Costa, E.C.; Moreira, A.F.; de Melo-Diogo, D.; Gaspar, V.M.; Carvalho, M.P.; Correia, I.J. 3D tumor spheroids: An overview on the tools and techniques used for their analysis. Biotechnol. Adv. 2016, 34, 1427–1441. [Google Scholar] [CrossRef]
- Rodallec, A.; Sicard, G.; Giacometti, S.; Carré, M.; Pourroy, B.; Bouquet, F.; Savina, A.; Lacarelle, B.; Ciccolini, J.; Fanciullino, R. From 3D spheroids to tumor bearing mice: Efficacy and distribution studies of trastuzumab-docetaxel immunoliposome in breast cancer. Int. J. Nanomed. 2018, 13, 6677–6688. [Google Scholar] [CrossRef]
- Ji, Y.; Li, J.; Zhao, J.; Shan, S.; Chu, C.C. A light-facilitated drug delivery system from a pseudo-protein/hyaluronic acid nanocomplex with improved anti-tumor effects. Nanoscale 2019, 11, 9987–10003. [Google Scholar] [CrossRef]
- Brown, M.J.; Bahsoun, S.; Morris, M.A.; Akam, E.C. Determining conditions for successful culture of multi-cellular 3D tumour spheroids to investigate the effect of mesenchymal stem cells on breast cancer cell invasiveness. Bioengineering 2019, 6, 101. [Google Scholar] [CrossRef]
- Jaganathan, H.; Gage, J.; Leonard, F.; Srinivasan, S.; Souza, G.R.; Dave, B.; Godin, B. Three-dimensional in vitro co-culture model of breast tumor using magnetic levitation. Sci. Rep. 2014, 4, 6468. [Google Scholar] [CrossRef]
- Leonard, F.; Godin, B. 3D In Vitro Model for Breast Cancer Research Using Magnetic Levitation and Bioprinting Method. Methods Mol. Biol. 2016, 1406, 239–251. [Google Scholar] [CrossRef]
- Almarshad, H.A.; Madhavan, M.; Hoshino, K. Focused Ion Beam-Based Milling, Imaging and Analysis of 3D Tumor Spheroids. In Proceedings of the 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 17–21 July 2018; pp. 4480–4483. [Google Scholar] [CrossRef]
- Ranamukhaarachchi, S.K.; Modi, R.N.; Han, A.; Velez, D.O.; Kumar, A.; Engler, A.J.; Fraley, S.I. Macromolecular crowding tunes 3D collagen architecture and cell morphogenesis. Biomater. Sci. 2019, 7, 618–633. [Google Scholar] [CrossRef] [PubMed]
- Reid, J.A.; Mollica, P.A.; Bruno, R.D.; Sachs, P.C. Consistent and reproducible cultures of large-scale 3D mammary epithelial structures using an accessible bioprinting platform. Breast Cancer Res. 2018, 20, 122. [Google Scholar] [CrossRef] [PubMed]
- Liverani, C.; De Vita, A.; Minardi, S.; Kang, Y.; Mercatali, L.; Amadori, D.; Bongiovanni, A.; La Manna, F.; Ibrahim, T.; Tasciotti, E. A biomimetic 3D model of hypoxia-driven cancer progression. Sci. Rep. 2019, 9, 12263. [Google Scholar] [CrossRef] [PubMed]
- Mollica, P.A.; Booth-Creech, E.N.; Reid, J.A.; Zamponi, M.; Sullivan, S.M.; Palmer, X.L.; Sachs, P.C.; Bruno, R.D. 3D bioprinted mammary organoids and tumoroids in human mammary derived ECM hydrogels. Acta Biomater. 2019, 95, 201–213. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, S.; Hamid, Q.; Sun, W.; Clyne, A.M. Bioprinting of 3D breast epithelial spheroids for human cancer models. Biofabrication 2019, 11, 25003. [Google Scholar] [CrossRef]
- Conn, P.M. Animal Models for the Study of Human Disease; Elsevier: Amsterdam, the Netherlands, 2013. [Google Scholar]
- Alvarado, A.; Faustino-Rocha, A.I.; Colaço, B.; Oliveira, P.A. Experimental mammary carcinogenesis—Rat models. Life Sci. 2017, 173, 116–134. [Google Scholar] [CrossRef]
- de las Mulas, J.M.; Reymundo, C. Animal models of human breast carcinoma: Canine and feline neoplasms. Rev. Oncol. 2000, 2, 274–281. [Google Scholar] [CrossRef]
- De Maria, R.; Olivero, M.; Iussich, S.; Nakaichi, M.; Murata, T.; Biolatti, B.; Di Renzo, M.F. Spontaneous Feline Mammary Carcinoma Is a Model of HER2 Overexpressing Poor Prognosis Human Breast Cancer. Cancer Res. 2005, 65, 907–912. [Google Scholar]
- Antuofermo, E.; Miller, M.A.; Pirino, S.; Xie, J.; Badve, S.; Mohammed, S.I. Spontaneous Mammary Intraepithelial Lesions in Dogs—A Model of Breast Cancer. Cancer Epidemiol. Biomark. Prev. 2007, 16, 2247–2256. [Google Scholar] [CrossRef]
- De Maria, R.; Maggiora, P.; Biolatti, B.; Prat, M.; Comoglio, P.M.; Castagnaro, M.; Di Renzo, M.F. Feline STK gene expression in mammary carcinomas. Oncogene 2002, 21, 1785–1790. [Google Scholar] [CrossRef] [PubMed]
- Uva, P.; Aurisicchio, L.; Watters, J.; Loboda, A.; Kulkarni, A.; Castle, J.; Palombo, F.; Viti, V.; Mesiti, G.; Zappulli, V.; et al. Comparative expression pathway analysis of human and canine mammary tumors. BMC Genom. 2009, 10, 135. [Google Scholar] [CrossRef]
- Lee, C.H.; Kim, W.H.; Lim, J.H.; Kang, M.S.; Kim, D.Y.K.O. Mutation and overexpression of p53 as a prognostic factor in canine mammary tumors. J. Vet. Sci. 2004, 5, 63–69. [Google Scholar] [CrossRef] [PubMed]
- MacEwen, E.G.; Patnaik, A.K.; Harvey, H.J.; Panko, W.B. Estrogen Receptors in Canine Mammary Tumors. Cancer Res. 1982, 42, 2255–2259. [Google Scholar] [PubMed]
- Pinho, S.S.; Carvalho, S.; Cabral, J.; Reis, C.A.; Gärtner, F. Canine tumors: A spontaneous animal model of human carcinogenesis. Transl. Res. 2012, 165–172. [Google Scholar] [CrossRef]
- Nieto, A.; Pérez-Alenza, M.D.; Del Castillo, N.; Tabanera, E.; Castaño, M.; Peña, L. BRCA1 Expression in Canine Mammary Dysplasias and Tumours: Relationship with Prognostic Variables. J. Comp. Pathol. 2003, 128, 260–268. [Google Scholar] [CrossRef]
- Klopfleisch, R.; Gruber, A.D. Increased Expression of BRCA2 and RAD51 in Lymph Node Metastases of Canine Mammary Adenocarcinomas. Vet. Pathol. 2009, 46, 416–422. [Google Scholar] [CrossRef]
- Klopfleisch, R.; Gruber, A.D. Differential expression of cell cycle regulators p21, p27 and p53 in metastasizing canine mammary adenocarcinomas versus normal mammary glands. Res. Vet. Sci. 2009, 87, 91–96. [Google Scholar] [CrossRef]
- European Commission. Seventh Report on the Statistics on the Number of Animals Used for Experimental and Other Scientific Purposes in the Member States of the European Union; European Commission: Brussels, Belgium.
- Understanding Animal Research What Is Animal Research? 10 Facts-Mouse. Available online: http://www.understandinganimalresearch.org.uk/animals/10-facts/mouse/ (accessed on 14 January 2020).
- Iannaccone, P.M.; Jacob, H.J. Rats! Dis. Model. Mech. 2009, 2, 206–210. [Google Scholar] [CrossRef]
- Clarke, R. Animal models of breast cancer: Their diversity and role in biomedical research. Breast Cancer Res. Treat. 1996, 39, 1–6. [Google Scholar] [CrossRef]
- Faustino-Rocha, A.I.; Ferreira, R.; Oliveira, P.A.; Gama, A.; Ginja, M. N-Methyl-N-nitrosourea as a mammary carcinogenic agent. Tumor Biol. 2015, 36, 9095–9117. [Google Scholar] [CrossRef] [PubMed]
- Fagundes, D.J.; Taha, M.O. Animal disease model: Choice’s criteria and current animals specimens. Acta Cir. Bras. 2004, 19, 59–65. [Google Scholar] [CrossRef]
- Kjell, J.; Olson, L. Rat models of spinal cord injury: From pathology to potential therapies. Dis. Model. Mech. 2016, 9, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Ellenbroek, B.; Youn, J. Rodent models in neuroscience research: Is it a rat race? Dis. Model. Mech. 2016, 9, 1079–1087. [Google Scholar] [CrossRef]
- Thompson, H.J.; Singh, M. Rat Models of Premalignant Breast Disease. J. Mamm. Gland Biol. Neop. 2000, 5, 409–420. [Google Scholar] [CrossRef]
- Nandi, S.; Guzman, R.C.; Yang, J. Hormones and mammary carcinogenesis in mice, rats, and humans: A unifying hypothesis. Proc. Natl. Acad. Sci. USA 1995, 92, 3650–3657. [Google Scholar] [CrossRef]
- Gould, M.N. Rodent models for the study of etiology, prevention and treatment of breast cancer. Semin. Cancer Biol. 1995, 6, 147–152. [Google Scholar] [CrossRef]
- Brekke, T.D.; Steele, K.A.; Mulley, J.F. Inbred or outbred? Genetic diversity in laboratory rodent colonies. G3 Genes Genomes Genet. 2018, 8, 679–686. [Google Scholar] [CrossRef]
- Eppig, J.T. Chapter 5—Mouse Strain and Genetic Nomenclature: An Abbreviated Guide. In The Mouse in Biomedical Research, 2nd ed.; Fox, J.G., Davisson, M.T., Quimby, F.W., Barthold, S.W., Newcomer, C.E., Smith, A.L., Eds.; Academic Press: Burlington, VT, USA, 2007; pp. 79–98. ISBN 978-0-12-369454-6. [Google Scholar]
- Jensen, V.S.; Porsgaard, T.; Lykkesfeldt, J.; Hvid, H. Rodent model choice has major impact on variability of standard preclinical readouts associated with diabetes and obesity research. Am. J. Transl. Res. 2016, 8, 3574–3584. [Google Scholar]
- Festing, M.F.W. Inbred strains should replace outbred stocks in toxicology, safety testing, and drug development. Toxicol. Pathol. 2010, 38, 681–690. [Google Scholar] [CrossRef]
- Gill, T.J. The use of randomly bred and genetically defined animals in biomedical research. Am. J. Pathol. 1980, 101, S21–S32. [Google Scholar] [PubMed]
- Herschkowitz, J.I.; Simin, K.; Weigman, V.J.; Mikaelian, I.; Usary, J.; Hu, Z.; Rasmussen, K.E.; Jones, L.P.; Assefnia, S.; Chandrasekharan, S.; et al. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol. 2007, 8, R76. [Google Scholar] [CrossRef] [PubMed]
- Matulka, L.A.; Wagner, K.U. Models of breast cancer. Drug Discov. Today Dis. Model. 2005, 2, 1–6. [Google Scholar] [CrossRef]
- Liu, Y.; Yin, T.; Feng, Y.; Cona, M.M.; Huang, G.; Liu, J.; Song, S.; Jiang, Y.; Xia, Q.; Swinnen, J.V.; et al. Mammalian models of chemically induced primary malignancies exploitable for imaging-based preclinical theragnostic research. Quant. Imaging Med. Surg. 2015, 5, 708–729. [Google Scholar] [CrossRef]
- Medina, D. Chemical carcinogenesis of rat and mouse mammary glands. Breast Dis. 2007, 28, 63–68. [Google Scholar] [CrossRef]
- Vargo-Gogola, T.; Rosen, J.M. Modelling breast cancer: One size does not fit all. Nat. Rev. Cancer 2007, 7, 659–672. [Google Scholar] [CrossRef]
- Trosko, J.E. Commentary: Is the concept of “tumor promotion” a useful paradigm? Mol. Carcinog. 2001, 30, 131–137. [Google Scholar] [CrossRef]
- Luch, A. Nature and nurture—lessons from chemical carcinogenesis. Nat. Rev. Cancer 2005, 5, 113–125. [Google Scholar] [CrossRef]
- Zarbl, H. Toxicogenomic analyses of genetic susceptibility to mammary gland carcinogenesis in rodents: Implications for human breast cancer. Breast Dis. 2007, 28, 87–105. [Google Scholar] [CrossRef]
- Santarelli, R.L.; Pierre, F.; Corpet, D.E. Processed meat and colorectal cancer: A review of epidemiologic and experimental evidence. Nutr. Cancer 2008, 60, 131–144. [Google Scholar] [CrossRef]
- Ward, M.H.; Cross, A.J.; Divan, H.; Kulldorff, M.; Nowell-Kadlubar, S.; Kadlubar, F.F.; Sinha, R. Processed meat intake, CYP2A6 activity and risk of colorectal adenoma. Carcinogenesis 2007, 28, 1210–1216. [Google Scholar] [CrossRef]
- Pohanish, R.P. Sittig’s Handbook of Toxic and Hazardous Chemicals and Carcinogens; Elsevier Science: Amsterdam, the Netherlands, 2011; ISBN 9781437778694. [Google Scholar]
- Agrawal, A.; Verma, P.; Goyal, P.K. Chemomodulatory Effects of Aegle Marmelos Against DMBA-Induced Skin Tumorigenesis in Swiss Albino Mice. Asian Pac. J. Cancer Prev. J Cancer Prev. 2010, 11, 1311–1314. [Google Scholar]
- Dimitrova-Shumkovska, J.; Veenman, L.; Ristoski, T.; Leschiner, S.; Gavish, M. Decreases in Binding Capacity of the Mitochondrial 18 kDa Translocator Protein Accompany Oxidative Stress and Pathological Signs in Rat Liver After DMBA Exposure. Toxicol. Pathol. 2010, 38, 957–968. [Google Scholar] [CrossRef]
- Loomis, T.A.; Hayes, A.W. Loomis’s Essentials of Toxicology; Academic Press: Amsterdam, the Netherlands, 1996; ISBN 9780080535630. [Google Scholar]
- Russo, J.; Russo, I.H. Atlas and Histologic Classification of Tumors of the Rat Mammary Gland. J. Mamm. Gland Biol. Neop. 2000, 5, 187–200. [Google Scholar] [CrossRef] [PubMed]
- Al-Dhaheri, W.S.; Hassouna, I.; Al-Salam, S.; Karam, S.M. Characterization of Breast Cancer Progression in the Rat. Ann. N. Y. Acad. Sci. 2008, 1138, 121–131. [Google Scholar] [CrossRef] [PubMed]
- Currier, N.; Solomon, S.E.; Demicco, E.G.; Chang, D.L.F.; Farago, M.; Ying, H.; Dominguez, I.; Sonenshein, G.E.; Cardiff, R.D.; Jim Xiao, Z.X.; et al. Oncogenic Signaling Pathways Activated in DMBA-Induced Mouse Mammary Tumors. Toxicol. Pathol. 2005, 33, 726–737. [Google Scholar] [CrossRef] [PubMed]
- Russo, J.; Russo, I.H. Differentiation and breast cancer development. In Breast Cancer; Bittar, E.E., Heppner, G., Peters, W.P., Visscher, D.W., Eds.; Elsevier: Amsterdam, the Netherlands, 1999; Volume 2, pp. 1–10. ISBN 1569-254X. [Google Scholar]
- Martinez, A.; Merchan, J.; Sala, M.L.; Renedo, G.; Fernandez-Pascual, J.; Bullón, A.; Blanes, A.; Renedo, G.; Bullón, A., Jr.; Bullón, F.; et al. Carcinogenesis y Nitrosoamidas. Patologia (Mex.) 1974, VII, 225–230. [Google Scholar]
- Murray, T.J.; Ucci, A.A.; Maffini, M.V.; Sonnenschein, C.; Soto, A.M. Histological analysis of low dose NMU effects in the rat mammary gland. BMC Cancer 2009, 9, 267. [Google Scholar] [CrossRef]
- Lyng, H.; Olsen, D.R.; Southon, T.E.; Rofstad, E.K. 31P-nuclear magnetic resonance spectroscopy in vivo of four human melanoma xenograft lines: Spin-lattice relaxation times. Br. J. Cancer 1993, 68, 1061–1070. [Google Scholar] [CrossRef][Green Version]
- Wu, I.; Wang, H.; Huso, D.; Wahl, R.L. Optimal definition of biological tumor volume using positron emission tomography in an animal model. EJNMMI Res. 2015, 5, 58. [Google Scholar] [CrossRef]
- Cotroneo, M.S.; Haag, J.D.; Zan, Y.; Lopez, C.C.; Thuwajit, P.; Petukhova, G.V.; Camerini-Otero, R.D.; Gendron-Fitzpatrick, A.; Griep, A.E.; Murphy, C.J.; et al. Characterizing a rat Brca2 knockout model. Oncogene 2007, 26, 1626–1635. [Google Scholar] [CrossRef] [PubMed]
- Zan, Y.; Haag, J.D.; Chen, K.-S.; Shepel, L.A.; Wigington, D.; Wang, Y.-R.; Hu, R.; Lopez-Guajardo, C.C.; Brose, H.L.; Porter, K.I.; et al. Production of knockout rats using ENU mutagenesis and a yeast-based screening assay. Nat. Biotechnol. 2003, 21, 645–651. [Google Scholar] [CrossRef] [PubMed]
- Bhardwaj, V.; Ankola, D.D.; Gupta, S.C.; Schneider, M.; Lehr, C.M.; Kumar, M.N.V.R. PLGA nanoparticles stabilized with cationic surfactant: Safety studies and application in oral delivery of paclitaxel to treat chemical-induced breast cancer in rat. Pharm. Res. 2009, 26, 2495–2503. [Google Scholar] [CrossRef] [PubMed]
- Gal, A.; Baba, A.; Miclaus, V.; Bouari, C.; Bolfă, P.; Borza, G.; Catoi, C. Comparative aspects regarding MNU-induced mammary carcinogenesis in immature Sprague-Dowley and Whistar rats. Bull. Univ. Agric. Sci. Vet. Med. Cluj-Napoca Vet. Med. 2011, 68, 159–163. [Google Scholar] [CrossRef]
- Alvarado, A.; Faustino-Rocha, A.I.; Ferreira, R.; Mendes, R.; Duarte, J.A.; Pires, M.J.; Colaço, B.; Oliveira, P.A. Prognostic factors in an exercised model of chemically-induced mammary cancer. Anticancer Res. 2016, 36, 2181–2188. [Google Scholar]
- Faustino-Rocha, A.I.; Gama, A.; Oliveira, P.A.; Alvarado, A.; Neuparth, M.J.; Ferreira, R.; Ginja, M. Effects of lifelong exercise training on mammary tumorigenesis induced by MNU in female Sprague–Dawley rats. Clin. Exp. Med. 2017, 17, 151–160. [Google Scholar] [CrossRef]
- Jiang, C.; Mitrenga, T.; Cutter, G.; Thompson, H. Pathogenic characterization of 1-methyl-1-nitrosourea-induced mammary carcinomas in the rat. Carcinogenesis 1998, 19, 223–227. [Google Scholar]
- Perše, M.; Cerar, A.; Injac, R.; Štrukelj, B. N-methylnitrosourea Induced Breast Cancer in Rat, the Histopathology of the Resulting Tumours and its Drawbacks as a Model. Pathol. Oncol. Res. 2009, 15, 115–121. [Google Scholar] [CrossRef]
- Takayama, S.; Thorgeirsson, U.P.; Adamson, R.H. Chemical carcinogenesis studies in nonhuman primates. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2008, 84, 176–188. [Google Scholar] [CrossRef]
- Yang, J.; Shikata, N.; Mizuoka, H.; Tsubura, A. Colon carcinogenesis in shrews by intrarectal infusion of N-methyl-N-nitrosourea. Cancer Lett. 1996, 110, 105–112. [Google Scholar] [CrossRef]
- Leung, W.K.; Wu, K.; Wong, C.Y.P.; Cheng, A.S.L.; Ching, A.K.K.; Chan, A.W.H.; Chong, W.W.S.; Go, M.Y.Y.; Yu, J.; To, K.-F.; et al. Transgenic cyclooxygenase-2 expression and high salt enhanced susceptibility to chemical-induced gastric cancer development in mice. Carcinogenesis 2008, 29, 1648–1654. [Google Scholar] [CrossRef] [PubMed]
- McCormick, D.L.; Adamowski, C.B.; Fiks, A.; Moon, R.C. Lifetime dose-response relationships for mammary tumor induction by a single administration of N-methyl-N-nitrosourea. Cancer Res. 1981, 41, 1690–1694. [Google Scholar]
- Russo, I.H.; Russo, J. Mammary gland neoplasia in long-term rodent studies. Environ. Health Perspect. 1996, 104, 938–967. [Google Scholar] [CrossRef] [PubMed]
- Russo, J.; Russo, I.H. Experimentally induced mammary tumors in rats. Breast Cancer Res. Treat. 1996, 39, 7–20. [Google Scholar] [CrossRef] [PubMed]
- Sternlicht, M.D. Key stages in mammary gland development: The cues that regulate ductal branching morphogenesis. Breast Cancer Res. 2006, 8, 201. [Google Scholar] [CrossRef]
- Russo, J. Significance of Rat mammary tumors for human risk assessment. Toxicol. Pathol. 2015, 43, 145–170. [Google Scholar] [CrossRef]
- Park, M.K.; Lee, C.H.; Lee, H. Mouse models of breast cancer in preclinical research. Lab. Anim. Res. 2018, 34, 160. [Google Scholar] [CrossRef]
- Ni, Y.; Wang, H.; Chen, F.; Li, J.; DeKeyzer, F.; Feng, Y.; Yu, J.; Bosmans, H.; Marchal, G. Tumor models and specific contrast agents for small animal imaging in oncology. Methods 2009, 48, 125–138. [Google Scholar] [CrossRef]
- Sano, D.; Myers, J.N. Xenograft models of head and neck cancers. Head Neck Oncol. 2009, 1, 32. [Google Scholar] [CrossRef]
- Forabosco, F.; Löhmus, M.; Rydhmer, L.; Sundström, L.F. Genetically modified farm animals and fish in agriculture: A review. Livest. Sci. 2013, 153, 1–9. [Google Scholar] [CrossRef]
- Doetschman, T. GI GEMs: Genetically engineered mouse models of gastrointestinal disease. Gastroenterology 2011, 140, 380–385.e2. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Workshop, E.C. Of mice and men—are mice relevant models for human disease? In Proceedings of the Outcomes of the European Commission Workshop ‘Are Mice Relevant Models for Human Disease?’, London, UK, 21 May 2010; Health Directorate, DG Research, European Commission: Brussels, Belgium, 2010; p. 10. [Google Scholar]
- Hoenerhoff, M.J.; Shibata, M.A.; Bode, A.; Green, J.E. Pathologic progression of mammary carcinomas in a C3(1)/SV40 T/t-antigen transgenic rat model of human triple-negative and Her2-positive breast cancer. Transgenic Res. 2011, 20, 247–259. [Google Scholar] [CrossRef]
- Mullins, L.J.; Brooker, G.; Mullins, J.J. Transgenesis in the Rat BT—Transgenesis Techniques: Principles and Protocols; Clarke, A.R., Ed.; Springer: Totowa, NJ, USA, 2002; pp. 255–270. ISBN 978-1-59259-178-7. [Google Scholar]
- Ito, N.; Hasegawa, R.; Sano, M.; Tamano, S.; Esumi, H.; Takayama, S.; Sugimura, T. A new colon and mammary carcinogen in cooked food, 2-amino-1-methyl-6-phenylimidazo[4,5-b]pyridine (PhIP). Carcinogenesis 1991, 12, 1503–1506. [Google Scholar] [CrossRef] [PubMed]
- Ronckers, C.M.; Erdmann, C.A.; Land, C.E. Radiation and breast cancer: A review of current evidence. Breast Cancer Res. 2005, 7, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Imaoka, T.; Nishimura, M.; Iizuka, D.; Daino, K.; Takabatake, T.; Okamoto, M.; Kakinuma, S.; Shimada, Y. Radiation-Induced Mammary Carcinogenesis in Rodent Models: What’s Different from Chemical Carcinogenesis? J. Radiat. Res. 2009, 50, 281–293. [Google Scholar] [CrossRef]
- Finerty, J.C.; Binhammer, R.T.; Schneider, M.; Cunningham, A.W.B. Neoplasms in Rats Exposed to Single-Dose Total-Body X Radiation2. J. Natl. Cancer Inst. 1953, 14, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Holtzman, S.; Stone, J.P.; Shellabarger, C.J. Synergism of diethylstilbestrol and radiation in mammary carcinogenesis in female F344 rats. J. Natl. Cancer Inst. 1979, 63, 1071–1074. [Google Scholar]
- Vogel, H.H.; Turner, J.E. Genetic Component in Rat Mammary Carcinogenesis. Radiat. Res. 1982, 89, 264–273. [Google Scholar] [CrossRef]
- Haag, J.D.; Hsu, L.-C.; Newton, M.A.; Gould, M.N. Allelic imbalance in mammary carcinomas induced by either 7,12-dimethylbenz[a]anthracene or ionizing radiation in rats carrying genes conferring differential susceptibilities to mammary carcinogenesis. Mol. Carcinog. 1996, 17, 134–143. [Google Scholar] [CrossRef]
- Bartstra, R.W.; Bentvelzen, P.A.J.; Zoetelief, J.; Mulder, A.H.; Broerse, J.J.; van Bekkum, D.W. Induction of Mammary Tumors in Rats by Single-Dose Gamma Irradiation at Different Ages. Radiat. Res. 1998, 150, 442–450. [Google Scholar] [CrossRef]
- Inano, H.; Suzuki, K.; Ishii-Ohba, H.; Ikeda, K.; Wakabayashi, K. Pregnancy-dependent initiation in tumorigenesis of Wistar rat mammary glands by 60Co-irradiation. Carcinogenesis 1991, 12, 1085–1090. [Google Scholar] [CrossRef] [PubMed]
- Shellabarger, C.J. Mammary Neoplastic Response of Lewis and Sprague-Dawley Female Rats to 7,12-Dimethylbenz(a)anthracene or X-ray. Cancer Res. 1972, 32, 883–885. [Google Scholar] [PubMed]
- Shellabarger, C.J.; Stone, J.P.; Holtzman, S. Rat Differences in Mammary Tumor Induction With Estrogen and Neutron Radiation. J. Natl. Cancer Inst. 1978, 61, 1505–1508. [Google Scholar] [CrossRef] [PubMed]
- Imaoka, T.; Nishimura, M.; Kakinuma, S.; Hatano, Y.; Ohmachi, Y.; Yoshinaga, S.; Kawano, A.; Maekawa, A.; Shimada, Y. High Relative Biologic Effectiveness of Carbon Ion Radiation on Induction of Rat Mammary Carcinoma and its Lack of H-ras and Tp53 Mutations. Int. J. Radiat. Oncol. 2007, 69, 194–203. [Google Scholar] [CrossRef]
- Ullrich, R.L.; Preston, R.J. Radiation Induced Mammary Cancer. J. Radiat. Res. 1991, 32, 104–109. [Google Scholar] [CrossRef]
- Mori, N.; Yamate, J.; Umesako, S.-I.; Hong, D.-P.; Okumoto, M.; Nakao, R. Preferential Induction of Mammary Tumors in p53 Hemizygous BALB/c Mice by Fractionated Irradiation of a Sub-lethal Dose of X-rays. J. Radiat. Res. 2003, 44, 249–254. [Google Scholar] [CrossRef][Green Version]
- Backlund, M.G.; Trasti, S.L.; Backlund, D.C.; Cressman, V.L.; Godfrey, V.; Koller, B.H. Impact of Ionizing Radiation and Genetic Background on Mammary Tumorigenesis in p53-deficient Mice. Cancer Res. 2001, 61, 6577–6582. [Google Scholar]
- Cressman, V.L.; Backlund, D.C.; Hicks, E.M.; Gowen, L.C.; Godfrey, V.; Koller, B.H. Mammary Tumor Formation in p53- and BRCA1-deficient Mice. Cell Growth Differ. 1999, 10, 1–10. [Google Scholar]
- Bassaganya-Riera, J.; Hontecillas, R.; Abedi, V.; Carbo, A.; Philipson, C.; Hoops, S. Computational Modeling; Elsevier Inc.: Amsterdam, the Netherlands, 2016; ISBN 9780128037157. [Google Scholar]
- Dimitrakopoulos, C.M.; Beerenwinkel, N. Computational approaches for the identification of cancer genes and pathways. Wiley Interdiscip. Rev. Syst. Biol. Med. 2017, 9, 1–18. [Google Scholar] [CrossRef]
- Fang, G.; Wang, W.; Paunic, V.; Heydari, H.; Costanzo, M.; Liu, X.; Liu, X.; VanderSluis, B.; Oately, B.; Steinbach, M.; et al. Discovering genetic interactions bridging pathways in genome-wide association studies. Nat. Commun. 2019, 10. [Google Scholar] [CrossRef]
- Dunning, A.M.; Michailidou, K.; Kuchenbaecker, K.B.; Thompson, D.; French, J.D.; Beesley, J.; Healey, C.S.; Kar, S.; Pooley, K.A.; Lopez-Knowles, E.; et al. Breast cancer risk variants at 6q25 display different phenotype associations and regulate ESR1, RMND1 and CCDC170. Nat. Genet. 2016, 48, 374–386. [Google Scholar] [CrossRef] [PubMed]
- Gitter, A.; Braunstein, A.; Pagnani, A.; Baldassi, C.; Borgs, C.; Chayes, J.; Zecchina, R.; Fraenkel, E. Sharing information to reconstruct patient-specific pathways in heterogeneous diseases. In Proceedings of the Pacific Symposium on Biocomputing 2014, Fairmont Orchid, HI, USA, 3–7 January 2014; pp. 39–50. [Google Scholar]
- Bao, S.; Zhao, H.; Yuan, J.; Fan, D.; Zhang, Z.; Su, J.; Zhou, M. Computational identification of mutator-derived lncRNA signatures of genome instability for improving the clinical outcome of cancers: A case study in breast cancer. Brief. Bioinform. 2019, 00, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, A.; Nielsen, J. Genome scale metabolic modeling of cancer. Metab. Eng. 2017, 43, 103–112. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, S.; Ozkan, A.; Rylander, M.N.; Woodward, W.A.; Vlachos, P. Mixture theory modeling for characterizing solute transport in breast tumor tissues. J. Biol. Eng. 2019, 13, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Benzekry, S.; Lamont, C.; Beheshti, A.; Tracz, A.; Ebos, J.M.L.; Hlatky, L.; Hahnfeldt, P. Classical Mathematical Models for Description and Prediction of Experimental Tumor Growth. PLoS Comput. Biol. 2014, 10. [Google Scholar] [CrossRef]
- Norton, L. A Gompertzian Model of Human Breast Cancer Growth. Cancer Res. 1988, 48, 7067–7071. [Google Scholar]
- Gardezi, S.J.S.; Elazab, A.; Lei, B.; Wang, T. Breast cancer detection and diagnosis using mammographic data: Systematic review. J. Med. Internet Res. 2019, 21, 1–22. [Google Scholar] [CrossRef]
- Bornefalk, H.; Hermansson, A.B. On the comparison of FROC curve in mammography CAD system. Med. Phys. 2005, 32, 412–417. [Google Scholar] [CrossRef]
- Tiwari, S.; Bhargava, R. Extracting knowledge from chemical imaging data using computational algorithms for digital cancer diagnosis. Yale J. Biol. Med. 2015, 88, 131–143. [Google Scholar]
- Fabian, H.; Thi, N.A.N.; Eiden, M.; Lasch, P.; Schmitt, J.; Naumann, D. Diagnosing benign and malignant lesions in breast tissue sections by using IR-microspectroscopy. Biochim. Biophys. Acta Biomembr. 2006, 1758, 874–882. [Google Scholar] [CrossRef][Green Version]
- Kirouac, D.C.; Du, J.Y.; Lahdenranta, J.; Overland, R.; Yarar, D.; Paragas, V.; Pace, E.; McDonagh, C.F.; Nielsen, U.B.; Onsum, M.D. Computational modeling of ERBB2-amplified breast cancer identifies combined ErbB2/3 blockade as superior to the combination of MEK and AKT inhibitors (Science Signaling 6: 288 (ra68)). Sci. Signal. 2014, 7, er5. [Google Scholar] [CrossRef]
- Madhukar, N.S.; Khade, P.K.; Huang, L.; Gayvert, K.; Galletti, G.; Stogniew, M.; Allen, J.E.; Giannakakou, P.; Elemento, O. A Bayesian machine learning approach for drug target identification using diverse data types. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, Z.; Mahmood, T.; Hassan, M.; Afzal, S.; Rafique, H.; Afzal, K.; Latip, J. Dexibuprofen amide derivatives as potential anticancer agents: Synthesis, in silico docking, bioevaluation, and molecular dynamic simulation. Drug Des. Dev. Ther. 2019, 13, 1643–1657. [Google Scholar] [CrossRef] [PubMed]
- Spinello, A.; Ritacco, I.; Magistrato, A. Recent advances in computational design of potent aromatase inhibitors: Open-eye on endocrine-resistant breast cancers. Expert Opin. Drug Discov. 2019, 14, 1065–1076. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Abramson, R.G.; Arlinghaus, L.R.; Chakravarthy, A.B.; Abramson, V.; Mayer, I.; Farley, J.; Delbeke, D.; Yankeelov, T.E. An algorithm for longitudinal registration of PET/CT images acquired during neoadjuvant chemotherapy in breast cancer: Preliminary results. EJNMMI Res. 2012, 2, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lu, W.; Wang, J.; Zhang, H.H. Computerized PET/CT image analysis in the evaluation of tumour response to therapy. Br. J. Radiol. 2015, 88, 10–12. [Google Scholar] [CrossRef]
- Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Shaw, K.R.M.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio Cancer Genomics Portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Uhlén, M.; Björling, E.; Agaton, C.; Szigyarto, C.A.K.; Amini, B.; Andersen, E.; Andersson, A.C.; Angelidou, P.; Asplund, A.; Asplund, C.; et al. A human protein atlas for normal and cancer tissues based on antibody proteomics. Mol. Cell. Proteom. 2005, 4, 1920–1932. [Google Scholar] [CrossRef]
- Subramanian, A.; Narayan, R.; Corsello, S.M.; Peck, D.D.; Natoli, T.E.; Lu, X.; Gould, J.; Davis, J.F.; Tubelli, A.A.; Asiedu, J.K.; et al. A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles. Cell 2017, 171, 1437–1452.e17. [Google Scholar] [CrossRef]
- Castellino, R.A. Computer aided detection (CAD): An overview. Cancer Imaging 2005, 5, 17–19. [Google Scholar] [CrossRef] [PubMed]
- Badgujar, N.V.; Tarapara, B.V.; Shah, F.D. Computational analysis of high-risk SNPs in human CHK2 gene responsible for hereditary breast cancer: A functional and structural impact. PLoS ONE 2019, 14, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Seo, J.-H.; Stranger, B.; McKenna, A.; Pe’er, I.; Laframboise, T.; Brown, M.; Tyekucheva, S.; Freedman, M.L. Integrative eQTL-based analyses reveal the biology of breast cancer risk loci. Cell 2013, 152, 633–641. [Google Scholar] [CrossRef] [PubMed]
- Chuang, H.Y.; Lee, E.; Liu, Y.T.; Lee, D.; Ideker, T. Network-based classification of breast cancer metastasis. Mol. Syst. Biol. 2007, 3, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Li, J.; Liu, E.; Kinnebrew, G.; Zhang, X.; Stover, D.; Huo, Y.; Zeng, Z.; Jiang, W.; Cheng, L.; et al. Identification of alternatively-activated pathways between primary breast cancer and liver metastatic cancer using microarray data. Genes (Basel) 2019, 10, 753. [Google Scholar] [CrossRef]
- Lai, X.; Geier, O.M.; Fleischer, T.; Garred, Y.; Borgen, E.; Funke, S.W.; Kumar, S.; Rognes, M.E.; Seierstad, T.; Børresen-Dale, A.L.; et al. Toward personalized computer simulation of breast cancer treatment: A multiscale pharmacokinetic and pharmacodynamic model informed by multitype patient data. Cancer Res. 2019, 79, 4293–4304. [Google Scholar] [CrossRef]
- Li, C.M.; Segars, W.P.; Tourassi, G.D.; Boone, J.M.; Dobbins, J.T. Methodology for generating a 3D computerized breast phantom from empirical data. Med. Phys. 2009, 36, 3122–3131. [Google Scholar] [CrossRef]
- Hsu, C.M.L.; Palmeri, M.L.; Segars, W.P.; Veress, A.I.; Dobbins, J.T. Generation of a suite of 3D computer-generated breast phantoms from a limited set of human subject data. Med. Phys. 2013, 40, 1–11. [Google Scholar] [CrossRef]
- Booth, M.E.; Nash, C.E.; Roberts, N.P.; Magee, D.R.; Treanor, D.; Hanby, A.M.; Speirs, V. 3-D tissue modelling and virtual pathology as new approaches to study ductal carcinoma in Situ. ATLA Altern. Lab. Anim. 2015, 43, 377–383. [Google Scholar] [CrossRef]
- Booth, M.E.; Treanor, D.; Roberts, N.; Magee, D.R.; Speirs, V.; Hanby, A.M. Three-dimensional reconstruction of ductal carcinoma in situ with virtual slides. Histopathology 2015, 66, 966–973. [Google Scholar] [CrossRef]
- Graff, C.G. A new, open-source, multi-modality digital breast phantom. In Medical Imaging 2016: Physics of Medical Imaging; San Diego, CA, USA, 2016; Volume 9783, p. 978309. Available online: https://www.spiedigitallibrary.org/conference-proceedings-of-spie/9783/978309/A-new-open-source-multi-modality-digital-breast-phantom/10.1117/12.2216312.short?SSO=1 (accessed on 7 December 2019). [CrossRef]
- Widmer, A.; Hu, Y. A viscoelastic model of a breast phantom for real-time palpation. In Proceedings of the 2011 Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Boston, MA, USA, 30 August–3 September 2011; pp. 4546–4549. [Google Scholar] [CrossRef]
- Celi, S.; Di Puccio, F.; Forte, P. Advances in finite element simulations of elastosonography for breast lesion detection. J. Biomech. Eng. 2011, 133, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Ikejimba, L.C.; Graff, C.G.; Rosenthal, S.; Badal, A.; Ghammraoui, B.; Lo, J.Y.; Glick, S.J. A novel physical anthropomorphic breast phantom for 2D and 3D X-ray imaging. Med. Phys. 2017, 44, 407–416. [Google Scholar] [CrossRef] [PubMed]
- Kiarashi, N.; Nolte, A.C.; Sturgeon, G.M.; Segars, W.P.; Ghate, S.V.; Nolte, L.W.; Samei, E.; Lo, J.Y. Development of realistic physical breast phantoms matched to virtual breast phantoms based on human subject data. Med. Phys. 2015, 42, 4116–4126. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.T.; Samsuzzaman, M.; Kibria, S.; Islam, M.T. Experimental breast phantoms for estimation of breast tumor using microwave imaging systems. IEEE Access 2018, 6, 78587–78597. [Google Scholar] [CrossRef]
- Joachimowicz, N.; Duchêne, B.; Conessa, C.; Meyer, O. Easy-to-produce adjustable realistic breast phantoms for microwave imaging. In Proceedings of the 2016 10th European Conference Antennas Propagation (EuCAP 2016), Davos, Switzerland, 10–15 April 2016; pp. 3–6. [Google Scholar] [CrossRef]
- Truong, B.C.Q.; Fitzgerald, A.J.; Fan, S.; Wallace, V.P. Concentration analysis of breast tissue phantoms with terahertz spectroscopy. Biomed. Opt. Express 2018, 9, 1334. [Google Scholar] [CrossRef] [PubMed]
- Behmadi, M.; Gholamhosseinian, H.; Mohammadi, M.; Naseri, S.H.; Momennezhad, M.; Bayani, S.H.; Bahreyni Tossi, M.T. Evaluation of breast cancer radiation therapy techniques in outfield organs of rando phantom with thermoluminescence dosimeter. J. Biomed. Phys. Eng. 2019, 9, 179–188. [Google Scholar] [CrossRef]
- Quinn, A.; Holloway, L.; Metcalfe, P. Image guidance during breast radiotherapy: A phantom dosimetry and radiation-induced second cancer risk study. J. Phys. Conf. Ser. 2013, 444. [Google Scholar] [CrossRef]
- Di Meo, S.; Pasotti, L.; Iliopoulos, I.; Pasian, M.; Ettorre, M.; Zhadobov, M.; Matrone, G. Tissue-mimicking materials for breast phantoms up to 50 GHz. Phys. Med. Biol. 2019, 64. [Google Scholar] [CrossRef]
- Sabhachandani, P.; Motwani, V.; Cohen, N.; Sarkar, S.; Torchilin, V.; Konry, T.; Fenway, T.; Avenue, H. Fluorescent Aliphatic Hyperbranched Polyether: Chromophores-free and without any N and P Atoms. Phys. Chem. Chem. Phys. 2016, 16, 497–505. [Google Scholar] [CrossRef]
- Hwang, H.; Park, J.; Shin, C.; Do, Y.; Cho, Y.K. Three dimensional multicellular co-cultures and anti-cancer drug assays in rapid prototyped multilevel microfluidic devices. Biomed. Microdevices 2013, 15, 627–634. [Google Scholar] [CrossRef]
- Yildiz-Ozturk, E.; Gulce-Iz, S.; Anil, M.; Yesil-Celiktas, O. Cytotoxic responses of carnosic acid and doxorubicin on breast cancer cells in butterfly-shaped microchips in comparison to 2D and 3D culture. Cytotechnology 2017, 69, 337–347. [Google Scholar] [CrossRef] [PubMed]
- Grafton, M.M.G.; Wang, L.; Vidi, P.A.; Leary, J.; Lelièvre, S.A. Breast on-a-chip: Mimicry of the channeling system of the breast for development of theranostics. Integr. Biol. 2011, 3, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Yang, X.; Zou, J.; Jia, C.; Hu, Y.; Du, H.; Wang, H. Evaluation of photodynamic therapy efficiency using an in vitro three-dimensional microfluidic breast cancer tissue model. Lab Chip 2015, 15, 735–744. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.; Seo, H.I.; Bae, J.H.; Chung, B.G. Hydrogel microfluidic co-culture device for photothermal therapy and cancer migration. Electrophoresis 2017, 38, 1318–1324. [Google Scholar] [CrossRef]
- Sung, K.E.; Yang, N.; Pehlke, C.; Keely, P.J.; Eliceiri, K.W.; Friedl, A.; Beebe, D.J. Transition to invasion in breast cancer: A microfluidic in vitro model enables examination of spatial and temporal effects. Integr. Biol. (Camb.) 2011, 3, 439–450. [Google Scholar] [CrossRef]
- Bischel, L.L.; Beebe, D.J.; Sung, K.E. Microfluidic model of ductal carcinoma in situ with 3D, organotypic structure. BMC Cancer 2015, 15, 1–10. [Google Scholar] [CrossRef]
- Truong, D.; Puleo, J.; Llave, A.; Mouneimne, G.; Kamm, R.D.; Nikkhah, M. Breast cancer cell invasion into a three dimensional tumor-stroma microenvironment. Sci. Rep. 2016, 6, 34094. [Google Scholar] [CrossRef]
- Kwak, B.; Ozcelikkale, A.; Shin, C.S.; Park, K.; Han, B. Simulation of complex transport of nanoparticles around a tumor using tumor-microenvironment-on-chip. J. Control. Release 2014, 194, 157–167. [Google Scholar] [CrossRef]
- Bersini, S.; Jeon, J.S.; Dubini, G.; Arrigoni, C.; Chung, S.; Charest, J.L.; Moretti, M.; Kamm, R.D. A microfluidic 3D in vitro model for specificity of breast cancer metastasis to bone. Biomaterials 2014, 35, 2454–2461. [Google Scholar] [CrossRef]
- Song, J.W.; Cavnar, S.P.; Walker, A.C.; Luker, K.E.; Gupta, M.; Tung, Y.C.; Luker, G.D.; Takayama, S. Microfluidic endothelium for studying the intravascular adhesion of metastatic breast cancer cells. PLoS ONE 2009, 4. [Google Scholar] [CrossRef]
| ER | PR | HER2 | Ki-67 | Notes | |
|---|---|---|---|---|---|
| Luminal A | + +/− | +/− + | − | Low |
|
| Luminal B | + +/− | +/− + | + | Any |
|
| − | High | ||||
| HER2 Enriched | − | − | + | High |
|
| Triple-Negative A ≡ Basal-like | − | − | − | High |
|
| Triple-Negative B ≡ Normal-like | − | − | − | Low |
|
| Cell Line | ER | PR | AR | HER2 | Tumor Classification | Notes | Ref. | |
|---|---|---|---|---|---|---|---|---|
| BT-483 | + | +/− | + | − | LA | IDC | Medium: RPMI | [15,16,68,69,70,71,72,73] |
| HCC-712 | + | +/− | NA | − | DC | Medium: RPMI | [15,71] | |
| KPL-1 | + | − | NA | − | IDC | Medium: RPMI | [15,72,74] | |
| MCF-7 | + | + | + | − | IDC | Medium: RPMI, DMEM Ki67 low | [11,15,16,19,68,69,70,71,73] | |
| MDA-MB-415 | + | +/− | + | − | AC | Medium: DMEM | [15,68,69,70,73] | |
| T-47D | + | + | + | − | IDC | Medium: RPMI Ki67 low | [11,15,19,68,69,70,71,72,73] | |
| BT-474 | +/− | + | + | + | LB | IDC | Medium: RPMI Ki67 high | [15,16,19,68,69,70,71,72,73] |
| EFM-192A | + | + | NA | + | AC | Medium: RPMI | [15,71] | |
| IBEP-1 | - | + | NA | + | IDC | Medium: DMEM | [15,72] | |
| MDA-MB-330 | +/- | − | NA | + | ILC | Medium: RPMI | [15,68,70,72] | |
| UACC-812 | +/- | − | + | + | IDC | Medium: RPMI, DMEM | [15,68,69,70,71,72,73] | |
| ZR-75-30 | + | − | NA | + | IDC | Medium: RPMI | [15,68,69,70,71,72] | |
| 21-PT | − | +/− | NA | + | H | IDC | Medium: α-MEM/DFC1 | [15,62] |
| HCC-1569 | − | − | − | + | MC | Medium: RPMI | [15,68,69,71,73] | |
| MDA-MB-453 | − | − | + | + | AC | Medium: RPMI, DMEM Ki67 high | [11,15,16,19,66,67,68,69,70,71,72,73] | |
| SK-BR-3 | − | − | + | + | AC | Medium: RPMI, McCoys Ki67 high | [15,19,68,69,70,71,72,73] | |
| SUM-190PT | − | − | NA | + | InfC | Medium: Ham’s F12 | [15,68,69,70,71] | |
| SUM-225CWN | − | − | NA | + | IDC | Medium: Ham’s F12 | [15,68,69,70] | |
| DU-4475 | − | − | NA | − | TNA | IDC | Medium: RPMI | [15,68,70,72] |
| HCC-1806 | − | − | +/- | − | SqC | Medium: RPMI | [15,67,68,71,73] | |
| HCC-70 | − | − | +/- | − | DC | Medium: RPMI | [15,68,69,71,73] | |
| HMT-3522 | − | − | NA | − | B | Medium: DMEM, Ham’s F12 | [15,75] | |
| MA-11 | − | − | NA | − | ILC | Medium: DMEM | [15,72,76,77] | |
| MDA-MB-157 | − | − | - | − | TNB | MC | Medium: RPMI, DMEM | [15,68,69,70,71,72,73] |
| MDA-MB-231 | − | − | + | − | AC | Medium: RPMI, DMEM Ki67, E-cadherin, claudin-3, claudinin-4 and claudinin-7 low | [11,15,16,19,66,67,68,69,70,71,72,73] | |
| SUM-149PT | − | − | NA | − | InfDC | Medium: Ham’s F12 | [15,68,69,70,71] | |
| SUM-159PT | − | − | + | − | AC | Medium: Ham’s F12 | [15,67,68,69,70] | |
| Model | Implantation Site | Mice Strain | Cell Line | Tumor Classification |
|---|---|---|---|---|
| CDX | Subcutaneous (Heterotopic model) | BALB/c, Nude | MDA-MB-231 | TN |
| MDA-MB-435 | TN | |||
| BT474 | LB | |||
| Mammary fat pad (Orthotopic model) | NOD/SCID | MDA-MB-231 | TN | |
| MDA-MB4-35 | TN | |||
| SUM1315 | TN | |||
| MCF7 | LA | |||
| T47D | LA | |||
| Tail vein (Metastatic model) | NOD/SCID | MDA-MB-231 | TN | |
| SUM149 | TN | |||
| PDX | Subcutaneous | BALB/c, Nude | / | / |
| Mammary fat pad (Orthotopic model) | NOD/SCID NSG | / | / | |
| Humanized Mammary fat pad (Orthotopic model) | NOD/SCID | / | / | |
| Syngeneic | Mammary fat pad | BALB/c | 4T1 | / |
| Models | Pros | Cons | ||
|---|---|---|---|---|
| Xenograft | CDX | Subcutaneous administration |
|
|
| Orthotopic administration |
|
| ||
| PDX |
|
| ||
| Syngeneic models (allograft) |
|
| ||
| GEM |
|
| ||
| Tumor-inducted by | Chemicals and Hormones |
|
| |
| Radiation |
|
| ||
| Models | Pros | Cons |
|---|---|---|
| In vitro |
|
|
| In vivo |
|
|
| In silico |
|
|
| Physical Phantoms |
|
|
| 3D Microfluidic Models |
|
|
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Costa, E.; Ferreira-Gonçalves, T.; Chasqueira, G.; Cabrita, A.S.; Figueiredo, I.V.; Reis, C.P. Experimental Models as Refined Translational Tools for Breast Cancer Research. Sci. Pharm. 2020, 88, 32. https://doi.org/10.3390/scipharm88030032
Costa E, Ferreira-Gonçalves T, Chasqueira G, Cabrita AS, Figueiredo IV, Reis CP. Experimental Models as Refined Translational Tools for Breast Cancer Research. Scientia Pharmaceutica. 2020; 88(3):32. https://doi.org/10.3390/scipharm88030032
Chicago/Turabian StyleCosta, Eduardo, Tânia Ferreira-Gonçalves, Gonçalo Chasqueira, António S. Cabrita, Isabel V. Figueiredo, and Catarina Pinto Reis. 2020. "Experimental Models as Refined Translational Tools for Breast Cancer Research" Scientia Pharmaceutica 88, no. 3: 32. https://doi.org/10.3390/scipharm88030032
APA StyleCosta, E., Ferreira-Gonçalves, T., Chasqueira, G., Cabrita, A. S., Figueiredo, I. V., & Reis, C. P. (2020). Experimental Models as Refined Translational Tools for Breast Cancer Research. Scientia Pharmaceutica, 88(3), 32. https://doi.org/10.3390/scipharm88030032