Activity of Bacteriophage D29 Loaded on Nanoliposomes against Macrophages Infected with Mycobacterium tuberculosis
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemical Reagents
2.2. Preparation of Nanoliposomes
2.3. Characterization of Nanoliposomes
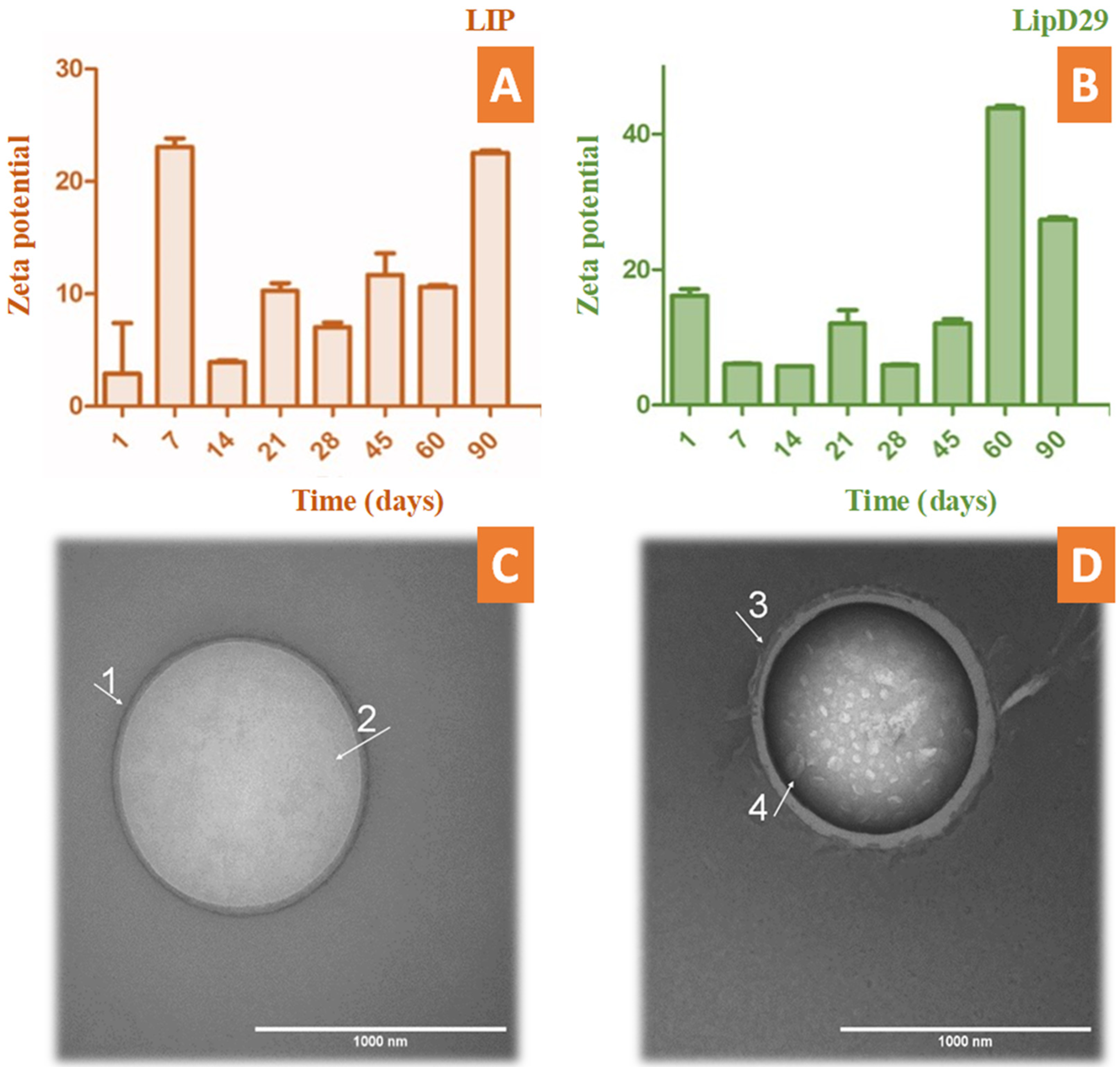
2.4. Nanostructure of Nanoliposomes
2.5. Mycobacterial Strains
2.6. Antimycobacterial Activity
2.7. Cytotoxic Activity
2.8. Fluorescence Microscopy
2.9. Time–Kill Studies
2.10. Intramacrophage Assay
2.11. Statistical Analysis
3. Results and Discussion
3.1. Physicochemical Results
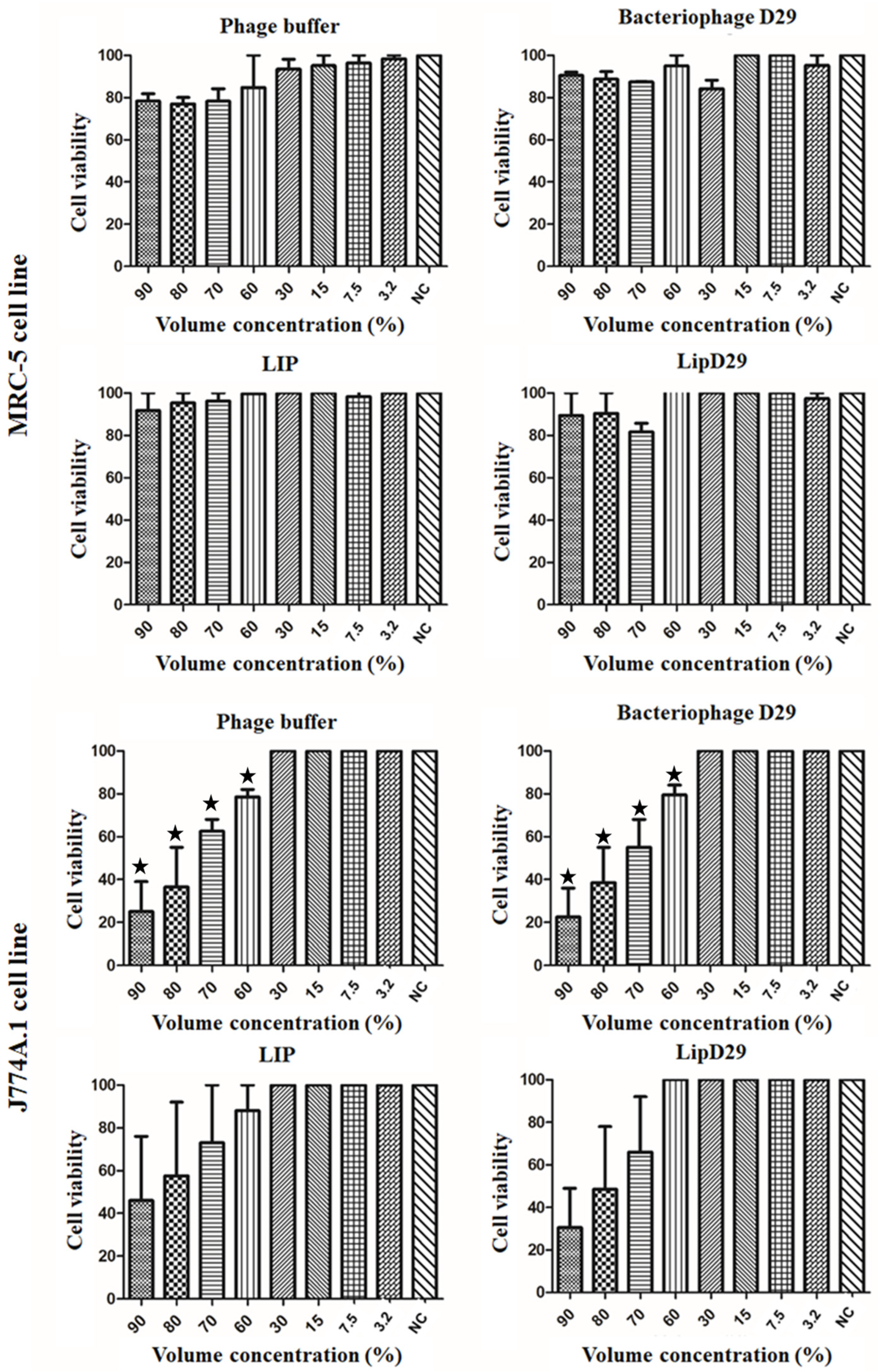
3.2. Cytotoxic Activity
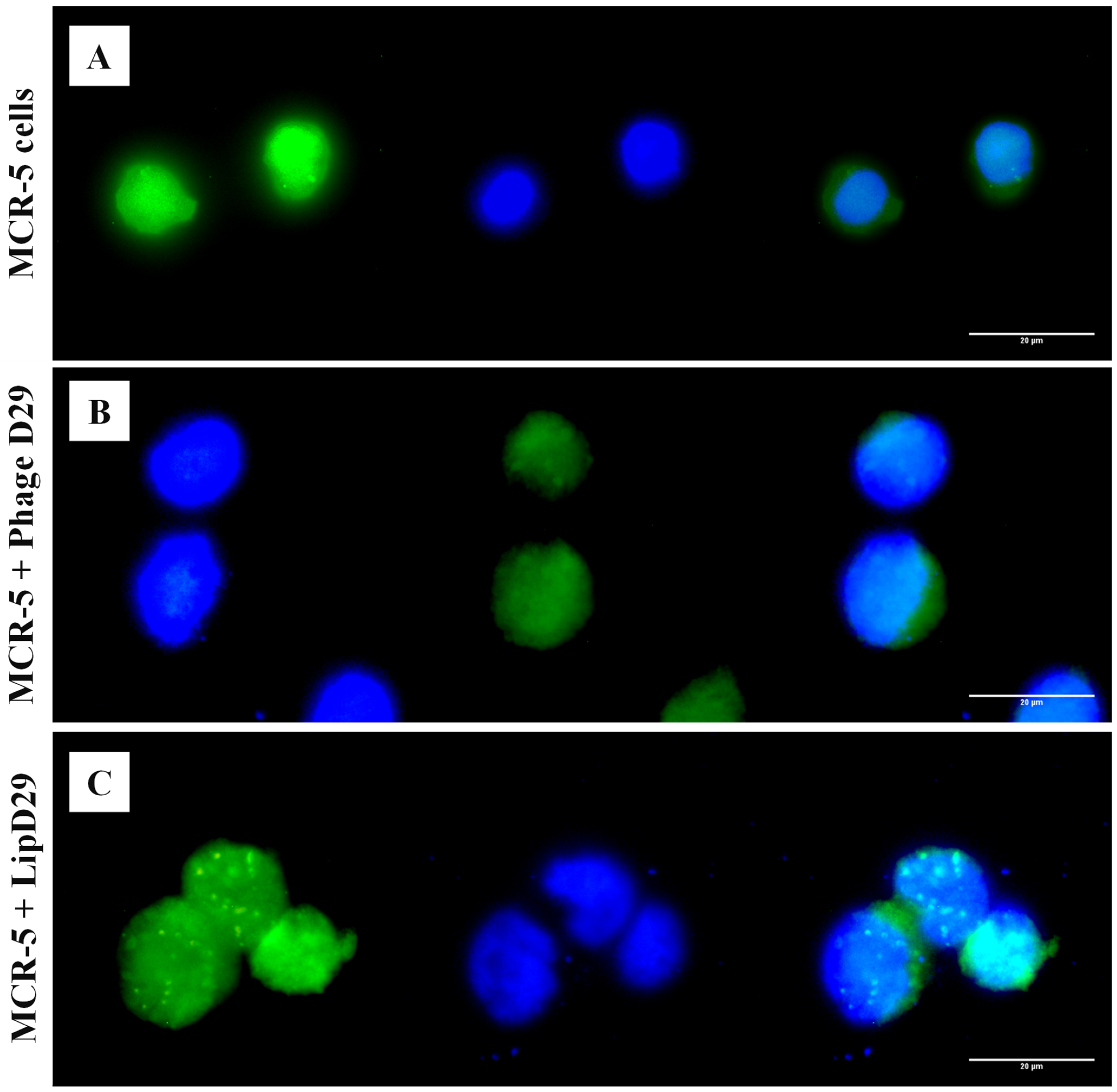
3.3. Fluorescence Microscopy
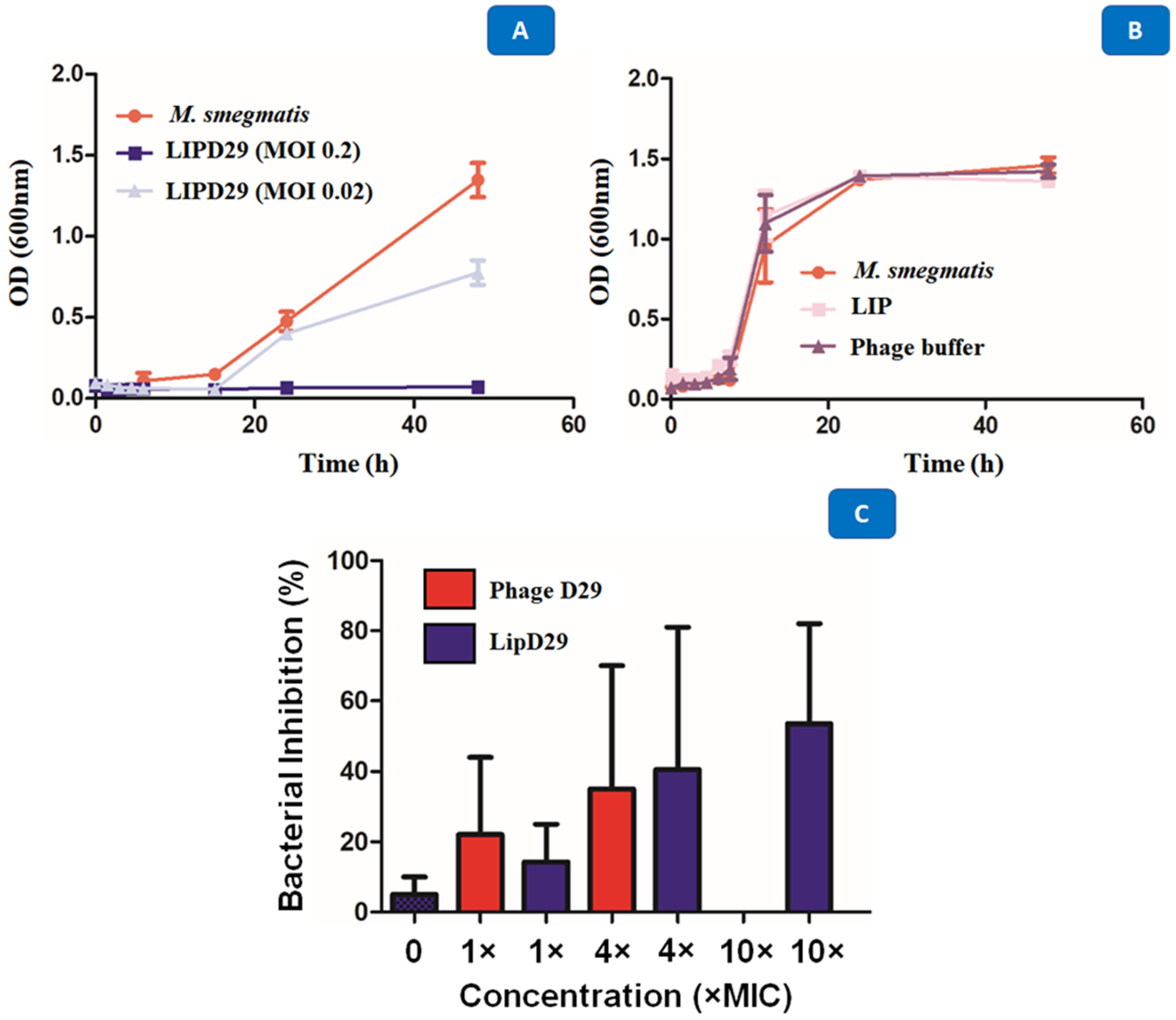
3.4. Antimycobacterial Results
3.5. Time–Kill Results
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- WHO. Global Tuberculosis Report 2022; WHO: Geneva, Switzerland, 2022; ISBN 9789240061729. [Google Scholar]
- Gordillo Altamirano, F.L.; Barr, J.J. Phage Therapy in the Postantibiotic Era. Clin. Microbiol. Rev. 2019, 32, e00066-18. [Google Scholar] [CrossRef] [PubMed]
- Souza, G.d.C.d.; Roque-Borda, C.A.; Pavan, F.R. Beta-lactam Resistance and the Effectiveness of Antimicrobial Peptides against KPC-Producing Bacteria. Drug Dev. Res. 2022, 83, 1534–1554. [Google Scholar] [CrossRef]
- Kelishomi, F.Z.; Khanjani, S.; Fardsanei, F.; Sarabi, H.S.; Nikkhahi, F.; Dehghani, B. Bacteriophages of Mycobacterium Tuberculosis, Their Diversity, and Potential Therapeutic Uses: A Review. BMC Infect. Dis. 2022, 22, 957. [Google Scholar] [CrossRef]
- Roque-Borda, C.A.; da Silva, B.P.; Rodrigues, M.C.; Azevedo, R.B.; Di Filippo, L.; Duarte, J.L.; Chorilli, M.; Festozo Vicente, E.; Pavan, F.R. Challenge in the Discovery of New Drugs: Antimicrobial Peptides against WHO-List of Critical and High-Priority Bacteria. Pharmaceutics 2021, 13, 773. [Google Scholar] [CrossRef] [PubMed]
- Vasava, M.S.; Bhoi, M.N.; Rathwa, S.K.; Borad, M.A.; Nair, S.G.; Patel, H.D. Drug Development against Tuberculosis: Past, Present and Future. Indian J. Tuberc. 2017, 64, 252–275. [Google Scholar]
- WHO. Tuberculosis. Available online: https://www.who.int/news-room/fact-sheets/detail/tuberculosis (accessed on 30 November 2020).
- Ahmad, S. Pathogenesis, Immunology, and Diagnosis of Latent Mycobacterium Tuberculosis Infection. Clin. Dev. Immunol. 2011, 2011, 814943. [Google Scholar] [CrossRef] [PubMed]
- Du Bruyn, E.; Ruzive, S.; Lindestam Arlehamn, C.S.; Sette, A.; Sher, A.; Barber, D.L.; Wilkinson, R.J.; Riou, C. Mycobacterium Tuberculosis-Specific CD4 T Cells Expressing CD153 Inversely Associate with Bacterial Load and Disease Severity in Human Tuberculosis. Mucosal Immunol. 2021, 14, 491–499. [Google Scholar] [CrossRef]
- Hoagland, D.T.; Liu, J.; Lee, R.B.; Lee, R.E. New Agents for the Treatment of Drug-Resistant Mycobacterium Tuberculosis. Adv. Drug Deliv. Rev. 2016, 102, 55–72. [Google Scholar] [CrossRef] [PubMed]
- Lapenkova, M.B.; Smirnova, N.S.; Rutkevich, P.N.; Vladimirsky, M.A. Evaluation of the Efficiency of Lytic Mycobacteriophage D29 on the Model of M. Tuberculosis-Infected Macrophage RAW 264 Cell Line. Bull. Exp. Biol. Med. 2018, 164, 344–346. [Google Scholar] [CrossRef]
- Gondhi, V.S.; Chhibber, S. Exploring Potential of Phage Therapy for Tuberculosis Using Model Organism. Biomed. Biotechnol. Res. J. 2018, 2, 9–15. [Google Scholar] [CrossRef]
- Carrigy, N.B.; Larsen, S.E.; Reese, V.; Pecor, T.; Harrison, M.; Kuehl, P.J.; Hatfull, G.F.; Sauvageau, D.; Baldwin, S.L.; Finlay, W.H.; et al. Prophylaxis of Mycobacterium Tuberculosis H37Rv Infection in a Preclinical Mouse Model via Inhalation of Nebulized Bacteriophage D29. Antimicrob. Agents Chemother. 2019, 63, e00871-19. [Google Scholar] [CrossRef]
- Carrigy, N.B.; Chang, R.Y.; Leung, S.S.Y.; Harrison, M.; Petrova, Z.; Pope, W.H.; Hatfull, G.F.; Britton, W.J.; Chan, H.-K.; Sauvageau, D.; et al. Anti-Tuberculosis Bacteriophage D29 Delivery with a Vibrating Mesh Nebulizer, Jet Nebulizer, and Soft Mist Inhaler. Pharm. Res. 2017, 34, 2084–2096. [Google Scholar] [CrossRef] [PubMed]
- Roque-Borda, C.A.; Bento da Silva, P.; Rodrigues, M.C.; Di Filippo, L.D.; Duarte, J.L.; Chorilli, M.; Vicente, E.F.; Garrido, S.S.; Rogério Pavan, F. Pharmaceutical Nanotechnology: Antimicrobial Peptides as Potential New Drugs against WHO List of Critical, High, and Medium Priority Bacteria. Eur. J. Med. Chem. 2022, 241, 114640. [Google Scholar] [CrossRef] [PubMed]
- Hatae, A.C.; Roque-Borda, C.A.; Pavan, F.R. Strategies for Lipid-Based Nanocomposites with Potential Activity against Mycobacterium Tuberculosis: Microbial Resistance Challenge and Drug Delivery Trends. OpenNano 2023, 13, 100171. [Google Scholar] [CrossRef]
- Roque-Borda, C.A.; Saraiva, M.d.M.S.; Monte, D.F.M.; Alves, L.B.R.; de Almeida, A.M.; Ferreira, T.S.; de Lima, T.S.; Benevides, V.P.; Cabrera, J.M.; Claire, S.; et al. HPMCAS-Coated Alginate Microparticles Loaded with Ctx(Ile 21)-Ha as a Promising Antimicrobial Agent against Salmonella Enteritidis in a Chicken Infection Model. ACS Infect. Dis. 2022, 8, 472–481. [Google Scholar] [CrossRef] [PubMed]
- Sheth, V.; Wang, L.; Bhattacharya, R.; Mukherjee, P.; Wilhelm, S. Strategies for Delivering Nanoparticles across Tumor Blood Vessels. Adv. Funct. Mater. 2021, 31, 2007363. [Google Scholar] [CrossRef]
- Di Filippo, L.D.; Duarte, J.L.; Azambuja, J.H.; Mancuso, R.I.; Luiz, M.T.; Araújo, V.H.S.; Figueiredo, I.D.; Barretto-De-Souza, L.; Sábio, R.M.; Sasso-Cerri, E.; et al. Glioblastoma Multiforme Targeted Delivery of Docetaxel Using Bevacizumab-Modified Nanostructured Lipid Carriers Impair in Vitro Cell Growth and in Vivo Tumor Progression. Int. J. Pharm. 2022, 618, 121682. [Google Scholar] [CrossRef]
- Roque-Borda, C.A.; Gualque, M.W.d.L.; da Fonseca, F.H.; Pavan, F.R.; Santos-Filho, N.A. Nanobiotechnology with Therapeutically Relevant Macromolecules from Animal Venoms: Venoms, Toxins, and Antimicrobial Peptides. Pharmaceutics 2022, 14, 891. [Google Scholar] [CrossRef]
- Bilal, M.; Qindeel, M.; Raza, A.; Mehmood, S.; Rahdar, A. Stimuli-Responsive Nanoliposomes as Prospective Nanocarriers for Targeted Drug Delivery. J. Drug Deliv. Sci. Technol. 2021, 66, 102916. [Google Scholar] [CrossRef]
- Aguilar-Pérez, K.M.; Avilés-Castrillo, J.I.; Medina, D.I.; Parra-Saldivar, R.; Iqbal, H.M.N. Insight Into Nanoliposomes as Smart Nanocarriers for Greening the Twenty-First Century Biomedical Settings. Front. Bioeng. Biotechnol. 2020, 8, 579536. [Google Scholar] [CrossRef]
- Nieth, A.; Verseux, C.; Barnert, S.; Süss, R.; Römer, W. A First Step toward Liposome-Mediated Intracellular Bacteriophage Therapy. Expert Opin. Drug Deliv. 2015, 12, 1411–1424. [Google Scholar] [CrossRef] [PubMed]
- Russell, D.A.; Hatfull, G.F. PhagesDB: The Actinobacteriophage Database. Bioinformatics 2017, 33, 784–786. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, J.L.; Piuri, M.; Broussard, G.; Marinelli, L.J.; Bastos, G.M.; Hirata, R.D.C.; Hatfull, G.F.; Hirata, M.H. Application of BRED Technology to Construct Recombinant D29 Reporter Phage Expressing EGFP. FEMS Microbiol. Lett. 2013, 344, 166–172. [Google Scholar] [CrossRef]
- Mady, M.M.; Darwish, M.M. Effect of Chitosan Coating on the Characteristics of DPPC Liposomes. J. Adv. Res. 2010, 1, 187–191. [Google Scholar] [CrossRef]
- Mady, M.M.; Darwish, M.M.; Khalil, S.; Khalil, W.M. Biophysical Studies on Chitosan-Coated Liposomes. Eur. Biophys. J. 2009, 38, 1127–1133. [Google Scholar] [CrossRef]
- Eloy, J.O.; Petrilli, R.; Topan, J.F.; Antonio, H.; Barcellos, J.P.A.; Chesca, D.L.; Serafini, L.N.; Tiezzi, D.G.; Lee, R.J.; Marchetti, J.M. Co-Loaded Paclitaxel/Rapamycin Liposomes: Development, Characterization and in Vitro and in Vivo Evaluation for Breast Cancer Therapy. Colloids Surf. B Biointerfaces 2016, 141, 74–82. [Google Scholar] [CrossRef]
- Performance Standards for Antimicrobial Susceptibility Testing, 28th ed.; CLSI: Malvern, PA, USA, 2018; Volume 1, ISBN 156238838X.
- Palomino, J.C.; Martin, A.; Camacho, M.; Guerra, H.; Swings, J.; Portaels, F. Resazurin Microtiter Assay Plate: Simple and Inexpensive Method for Detection of Drug Resistance in Mycobacterium Tuberculosis. Antimicrob. Agents Chemother. 2002, 46, 2720–2722. [Google Scholar] [CrossRef] [PubMed]
- Vipra, A.; Desai, S.N.; Junjappa, R.P.; Roy, P.; Poonacha, N.; Ravinder, P.; Sriram, B.; Padmanabhan, S. Determining the Minimum Inhibitory Concentration of Bacteriophages: Potential Advantages. Adv. Microbiol. 2013, 3, 181–190. [Google Scholar] [CrossRef]
- Cho, S.; Lee, H.S.; Franzblau, S. Microplate Alamar Blue Assay (MABA) and Low Oxygen Recovery Assay (LORA) for Mycobacterium Tuberculosis. In Mycobacteria Protocols. Methods in Molecular Biology; Humana Press: New York, NY, USA, 2015; Volume 1285, pp. 281–292. [Google Scholar]
- Silva, I.C.; Polaquini, C.R.; Regasini, L.O.; Ferreira, H.; Pavan, F.R. Evaluation of Cytotoxic, Apoptotic, Mutagenic, and Chemopreventive Activities of Semi-Synthetic Esters of Gallic Acid. Food Chem. Toxicol. 2017, 105, 300–307. [Google Scholar] [CrossRef]
- Wikler, M. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria That Grow Aerobically: Approved Standard. CLSI 2006, 26, M7-A7. [Google Scholar]
- Snewin, V.A.; Gares, M.-P.; Ó Gaora, P.; Hasan, Z.; Brown, I.N.; Young, D.B. Assessment of Immunity to Mycobacterial Infection with Luciferase Reporter Constructs. Infect. Immun. 1999, 67, 4586–4593. [Google Scholar] [CrossRef]
- Yuba, E.; Harada, A.; Sakanishi, Y.; Watarai, S.; Kono, K. A Liposome-Based Antigen Delivery System Using PH-Sensitive Fusogenic Polymers for Cancer Immunotherapy. Biomaterials 2013, 34, 3042–3052. [Google Scholar] [CrossRef] [PubMed]
- Moghimi, S.M.; Szebeni, J. Stealth Liposomes and Long Circulating Nanoparticles: Critical Issues in Pharmacokinetics, Opsonization and Protein-Binding Properties. Prog. Lipid Res. 2003, 42, 463–478. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.; Yuan, L.; Lin, L. Novel Chitosan Film Embedded with Liposome-Encapsulated Phage for Biocontrol of Escherichia Coli O157:H7 in Beef. Carbohydr. Polym. 2017, 177, 156–164. [Google Scholar] [CrossRef]
- Colom, J.; Cano-Sarabia, M.; Otero, J.; Cortés, P.; Maspoch, D.; Llagostera, M. Liposome-Encapsulated Bacteriophages for Enhanced Oral Phage Therapy against Salmonella spp. Appl. Environ. Microbiol. 2015, 81, 4841–4849. [Google Scholar] [CrossRef]
- Reimer, L.; Kohl, H. Transmission Electron Microscopy; Springer Series in Optical Sciences; Springer New York: New York, NY, USA, 2008; Volume 36, ISBN 978-0-387-40093-8. [Google Scholar]
- Schafer, R.; Huber, U.; Franklin, R.M. Chemical and Physical Properties of Mycobacteriophage D29. Eur. J. Biochem. 1977, 73, 239–246. [Google Scholar] [CrossRef] [PubMed]
- Sapra, P.; Allen, T.M. Ligand-Targeted Liposomal Anticancer Drugs. Prog. Lipid Res. 2003, 42, 439–462. [Google Scholar] [CrossRef]
- Monteiro, N.; Martins, A.; Reis, R.L.; Neves, N.M. Liposomes in Tissue Engineering and Regenerative Medicine. J. R. Soc. Interface 2014, 11, 20140459. [Google Scholar] [CrossRef]
- David, H.; Clavel, S.; Clement, F. Adsorption and Growth of the Bacteriophage D29 in Selected Mycobacteria. Ann. l’Institut Pasteur/Virol. 1980, 131, 167–184. [Google Scholar] [CrossRef]
- Guse, C.; Koennings, S.; Maschke, A.; Hacker, M.; Becker, C.; Schreiner, S.; Blunk, T.; Spruss, T.; Goepferich, A. Biocompatibility and Erosion Behavior of Implants Made of Triglycerides and Blends with Cholesterol and Phospholipids. Int. J. Pharm. 2006, 314, 153–160. [Google Scholar] [CrossRef]
- Ding, Y.; Cui, W.; Sun, D.; Wang, G.-L.; Hei, Y.; Meng, S.; Chen, J.; Xie, Y.; Wang, Z. In Vivo Study of Doxorubicin-Loaded Cell-Penetrating Peptide-Modified PH-Sensitive Liposomes: Biocompatibility, Bio-Distribution, and Pharmacodynamics in BALB/c Nude Mice Bearing Human Breast Tumors. Drug Des. Dev. Ther. 2017, 11, 3105–3117. [Google Scholar] [CrossRef] [PubMed]
- Barlow, S.T.; Figueroa, B.; Fu, D.; Zhang, B. Membrane Tension Modifies Redox Loading and Release in Single Liposome Electroanalysis. Anal. Chem. 2021, 93, 3876–3882. [Google Scholar] [CrossRef] [PubMed]
- Jorquera, D.; Galarce, N.; Borie, C. El Desafío de Controlar Las Enfermedades Transmitidas Por Alimentos: Bacteriófagos Como Una Nueva Herramienta Biotecnológica. Rev. Chil. Infectol. 2015, 32, 678–688. [Google Scholar] [CrossRef]
- Cooper, C.J.; Mirzaei, M.K.; Nilsson, A.S. Adapting Drug Approval Pathways for Bacteriophage-Based Therapeutics. Front. Microbiol. 2016, 7, 1209. [Google Scholar] [CrossRef]
- Guerrero-Bustamante, C.A.; Dedrick, R.M.; Garlena, R.A.; Russell, D.A.; Hatfull, G.F. Toward a Phage Cocktail for Tuberculosis: Susceptibility and Tuberculocidal Action of Mycobacteriophages against Diverse Mycobacterium Tuberculosis Strains. mBio 2021, 12, e00973-21. [Google Scholar] [CrossRef]
- Gordillo Altamirano, F.; Forsyth, J.H.; Patwa, R.; Kostoulias, X.; Trim, M.; Subedi, D.; Archer, S.K.; Morris, F.C.; Oliveira, C.; Kielty, L.; et al. Bacteriophage-Resistant Acinetobacter Baumannii Are Resensitized to Antimicrobials. Nat. Microbiol. 2021, 6, 157–161. [Google Scholar] [CrossRef] [PubMed]
- Bertozzi Silva, J.; Storms, Z.; Sauvageau, D. Host Receptors for Bacteriophage Adsorption. FEMS Microbiol. Lett. 2016, 363, fnw002. [Google Scholar] [CrossRef]
- Kabwe, M.; Dashper, S.; Bachrach, G.; Tucci, J. Bacteriophage Manipulation of the Microbiome Associated with Tumour Microenvironments-Can This Improve Cancer Therapeutic Response? FEMS Microbiol. Rev. 2021, 45. [Google Scholar] [CrossRef]
- Hodyra-Stefaniak, K.; Miernikiewicz, P.; Drapała, J.; Drab, M.; Jończyk-Matysiak, E.; Lecion, D.; Kaźmierczak, Z.; Beta, W.; Majewska, J.; Harhala, M.; et al. Mammalian Host-Versus-Phage Immune Response Determines Phage Fate In Vivo. Sci. Rep. 2015, 5, 14802. [Google Scholar] [CrossRef]
- Xiong, X.; Zhang, H.-M.; Wu, T.-T.; Xu, L.; Gan, Y.-L.; Jiang, L.-S.; Zhang, L.; Guo, S.-L. Titer Dynamic Analysis of D29 within MTB-Infected Macrophages and Effect on Immune Function of Macrophages. Exp. Lung Res. 2014, 40, 86–98. [Google Scholar] [CrossRef]
- Goldstein, B.P. Resistance to Rifampicin: A Review. J. Antibiot. 2014, 67, 625–630. [Google Scholar] [CrossRef]
- Dookie, N.; Rambaran, S.; Padayatchi, N.; Mahomed, S.; Naidoo, K. Evolution of Drug Resistance in Mycobacterium Tuberculosis: A Review on the Molecular Determinants of Resistance and Implications for Personalized Care. J. Antimicrob. Chemother. 2018, 73, 1138–1151. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Liu, H.; Jia, X.; Wang, X.; Xu, P. Mechanisms and Detection Methods of Mycobacterium Tuberculosis Rifampicin Resistance: The Phenomenon of Drug Resistance Is Complex. Tuberculosis 2021, 128, 102083. [Google Scholar] [CrossRef] [PubMed]
- Davoodi, S.; Daryaee, F.; Iuliano, J.N.; Tolentino Collado, J.; He, Y.; Pollard, A.C.; Gil, A.A.; Aramini, J.M.; Tonge, P.J. Evaluating the Impact of the Tyr158 pKa on the Mechanism and Inhibition of InhA, the Enoyl-ACP Reductase from Mycobacterium Tuberculosis. Biochemistry 2023, 62, 1943–1952. [Google Scholar] [CrossRef] [PubMed]
- Hsu, L.-Y.; Lai, L.-Y.; Hsieh, P.-F.; Lin, T.-L.; Lin, W.-H.; Tasi, H.-Y.; Lee, W.-T.; Jou, R.; Wang, J.-T. Two Novel KatG Mutations Conferring Isoniazid Resistance in Mycobacterium Tuberculosis. Front. Microbiol. 2020, 11, 1644. [Google Scholar] [CrossRef]
- Allué-Guardia, A.; Saranathan, R.; Chan, J.; Torrelles, J.B. Mycobacteriophages as Potential Therapeutic Agents against Drug-Resistant Tuberculosis. Int. J. Mol. Sci. 2021, 22, 735. [Google Scholar] [CrossRef]
- Bavda, V.R.; Jain, V. Deciphering the Role of Holin in Mycobacteriophage D29 Physiology. Front. Microbiol. 2020, 11, 883. [Google Scholar] [CrossRef]
- Shleider Carnero Canales, C.; Marquez Cazorla, J.; Furtado Torres, A.H.; Monteiro Filardi, E.T.; Di Filippo, L.D.; Costa, P.I.; Roque-Borda, C.A.; Pavan, F.R. Advances in Diagnostics and Drug Discovery against Resistant and Latent Tuberculosis Infection. Pharmaceutics 2023, 15, 2409. [Google Scholar] [CrossRef]
- Primo, L.M.D.G.; Roque-Borda, C.A.; Carnero Canales, C.S.; Caruso, I.P.; de Lourenço, I.O.; Colturato, V.M.M.; Sábio, R.M.; de Melo, F.A.; Vicente, E.F.; Chorilli, M.; et al. Antimicrobial Peptides Grafted onto the Surface of N-Acetylcysteine-Chitosan Nanoparticles Can Revitalize Drugs against Clinical Isolates of Mycobacterium Tuberculosis. Carbohydr. Polym. 2024, 323, 121449. [Google Scholar] [CrossRef]
- Lapenkova, M.B.; Alyapkina, Y.S.; Vladimirsky, M.A. Bactericidal Activity of Liposomal Form of Lytic Mycobacteriophage D29 in Cell Models of Tuberculosis Infection In Vitro. Bull. Exp. Biol. Med. 2020, 169, 361–364. [Google Scholar] [CrossRef] [PubMed]
- Avdeev, V.V.; Kuzin, V.V.; Vladimirsky, M.A.; Vasilieva, I.A. Experimental Studies of the Liposomal Form of Lytic Mycobacteriophage D29 for the Treatment of Tuberculosis Infection. Microorganisms 2023, 11, 1214. [Google Scholar] [CrossRef] [PubMed]

| Sample | REMA | LORA | ||
|---|---|---|---|---|
| MIC90 (µg/mL) | MOI | MIC90 (µg/mL) | MOI | |
| Rifampicin | 0.06 ± 0.05 | - | 1.2 ± 0.81 | - |
| Isoniazid | 0.03 ± 0.02 | - | 152.4 ± 3.14 | - |
| PhageD29 | - | 0.001 | - | >2.7 |
| LipD29 | - | 0.001 ± 0.0008 | - | >2.7 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Silva, A.P.B.; Roque-Borda, C.A.; Carnero Canales, C.S.; Duran Gleriani Primo, L.M.; Silva, I.C.; Ribeiro, C.M.; Chorilli, M.; da Silva, P.B.; Silva, J.L.; Pavan, F.R. Activity of Bacteriophage D29 Loaded on Nanoliposomes against Macrophages Infected with Mycobacterium tuberculosis. Diseases 2023, 11, 150. https://doi.org/10.3390/diseases11040150
Silva APB, Roque-Borda CA, Carnero Canales CS, Duran Gleriani Primo LM, Silva IC, Ribeiro CM, Chorilli M, da Silva PB, Silva JL, Pavan FR. Activity of Bacteriophage D29 Loaded on Nanoliposomes against Macrophages Infected with Mycobacterium tuberculosis. Diseases. 2023; 11(4):150. https://doi.org/10.3390/diseases11040150
Chicago/Turabian StyleSilva, Ana P. B., Cesar Augusto Roque-Borda, Christian S. Carnero Canales, Laura Maria Duran Gleriani Primo, Isabel C. Silva, Camila M. Ribeiro, Marlus Chorilli, Patrícia Bento da Silva, Joás L. Silva, and Fernando Rogério Pavan. 2023. "Activity of Bacteriophage D29 Loaded on Nanoliposomes against Macrophages Infected with Mycobacterium tuberculosis" Diseases 11, no. 4: 150. https://doi.org/10.3390/diseases11040150
APA StyleSilva, A. P. B., Roque-Borda, C. A., Carnero Canales, C. S., Duran Gleriani Primo, L. M., Silva, I. C., Ribeiro, C. M., Chorilli, M., da Silva, P. B., Silva, J. L., & Pavan, F. R. (2023). Activity of Bacteriophage D29 Loaded on Nanoliposomes against Macrophages Infected with Mycobacterium tuberculosis. Diseases, 11(4), 150. https://doi.org/10.3390/diseases11040150










