BU-Net: Brain Tumor Segmentation Using Modified U-Net Architecture
Abstract
1. Introduction
- Both new modules in BU-Net help to get the contextual information along with aggregation in the global features.
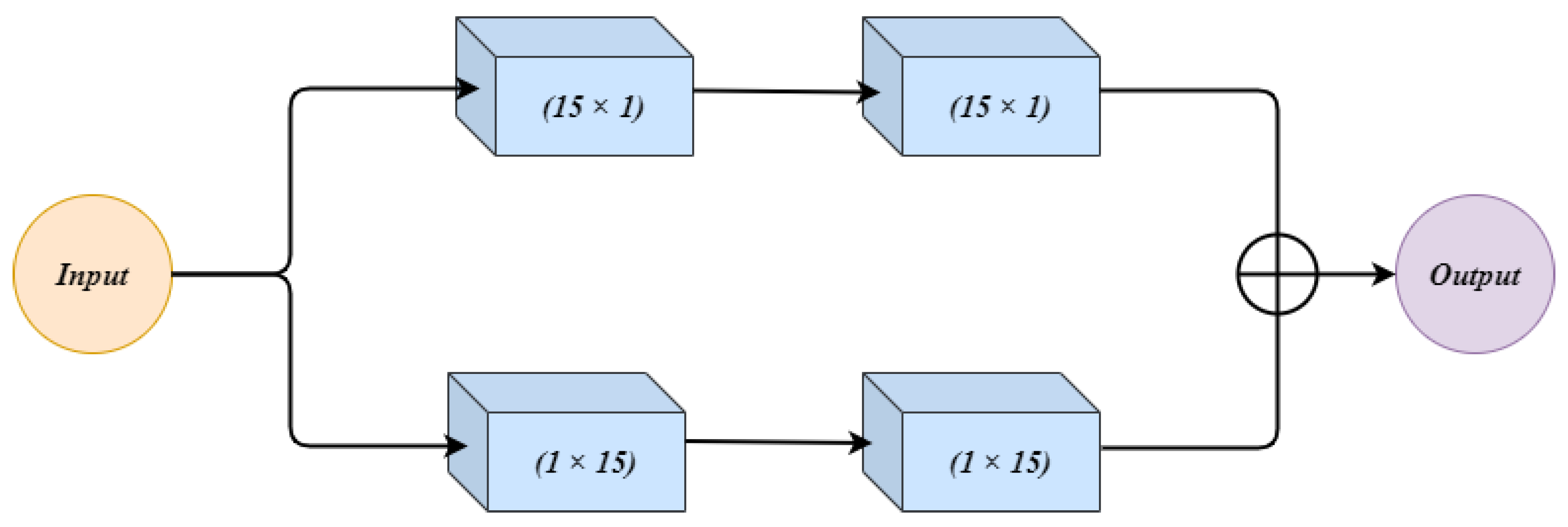
- Residual extended skip (RES) converts the low-level features to middle-level features.
- It is useful when scale-invariant features are used, which is important in the case of brain tumor segmentation, as the cancer regions vary from case to case.
- The RES module increases the valid receptive field, which remains a problem in previous techniques, as in those techniques the theoretical receptive field is always dominant.
- Two combined loss functions are used to tackle the problem associated with a huge difference in the percentage of pixels occupied by each class.
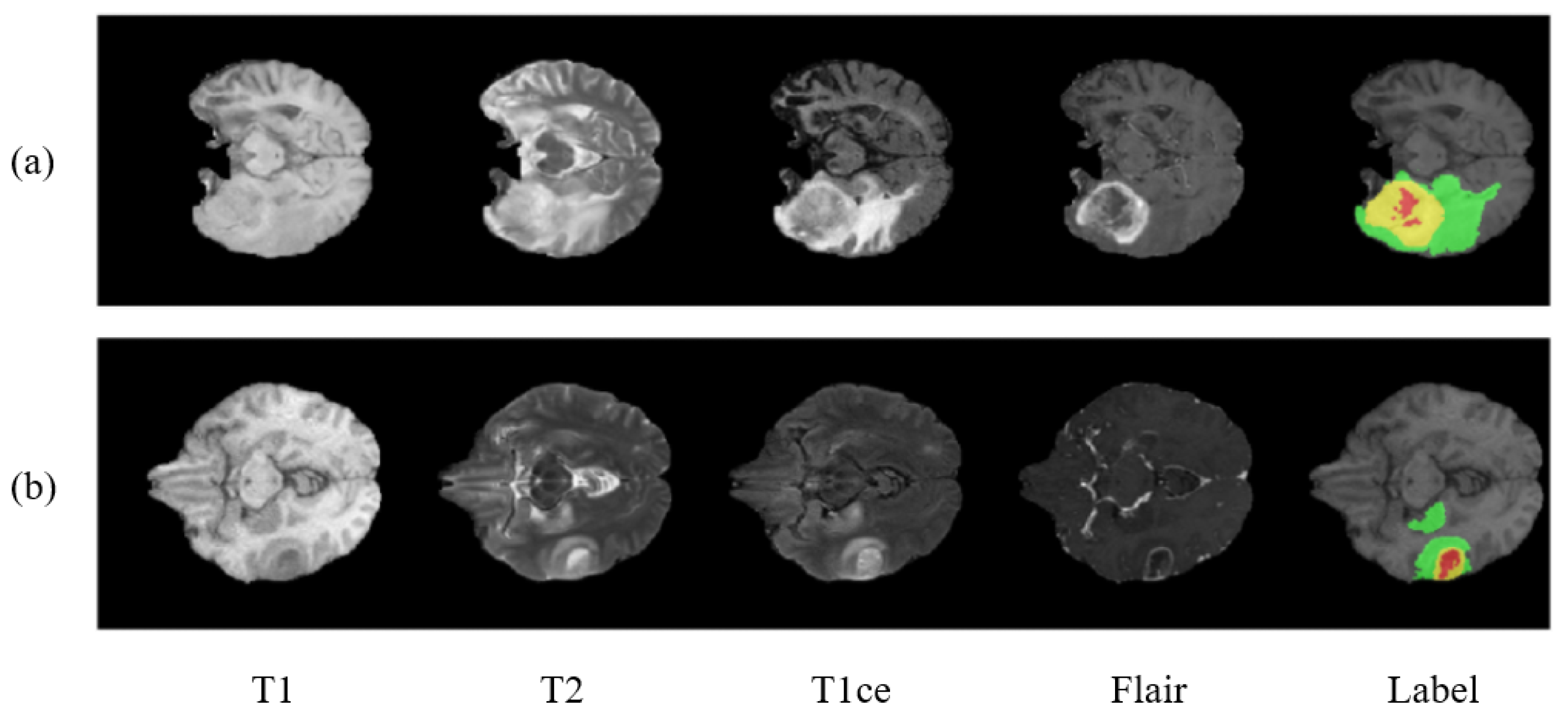
2. Datasets
- Enhancing tumor.
- Necrosis and non-enhancing tumor.
- Edema.
- Healthy tissue.
3. Methodology
3.1. Image Preprocessing
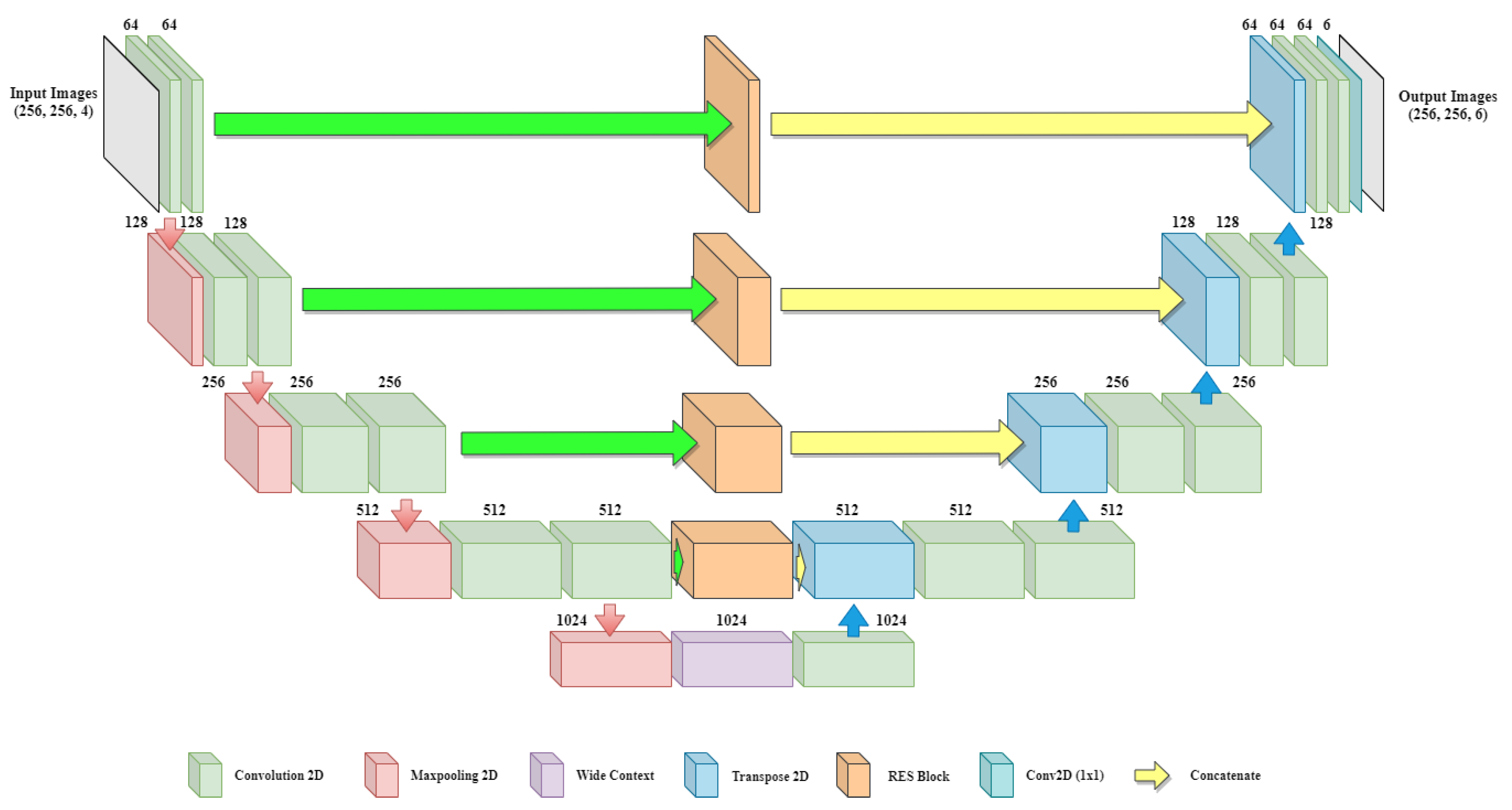
3.2. Proposed BU-Net
3.2.1. Residual Extended Skip (RES)
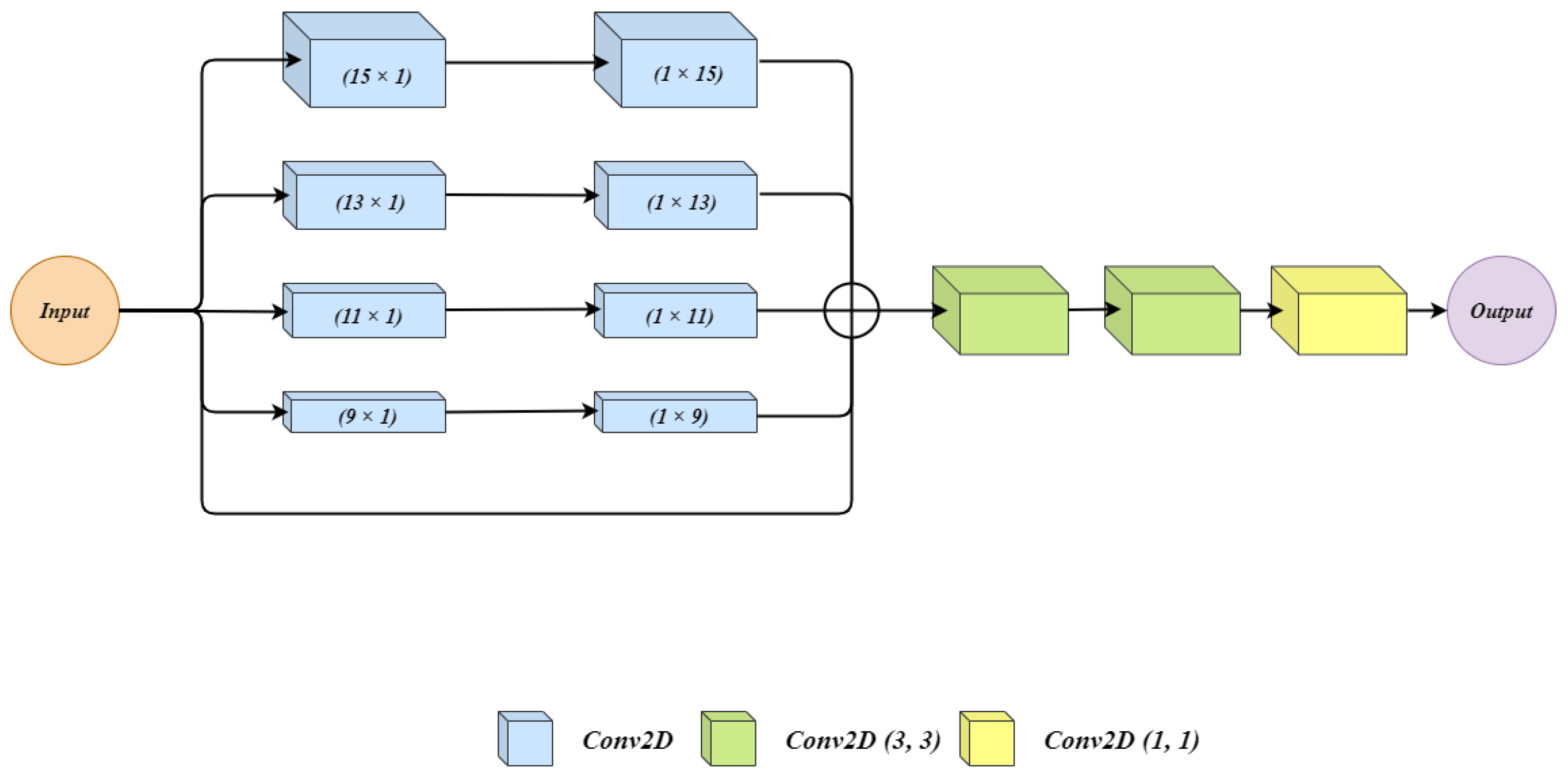
3.2.2. Wide Context (WC)
3.2.3. Customized Loss Function
4. Results and Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Işın, A.; Direkoğlu, C.; Şah, M. Review of MRI-based brain tumor image segmentation using deep learning methods. Procedia Comput. Sci. 2016, 102, 317–324. [Google Scholar] [CrossRef]
- Bakas, S.; Akbari, H.; Sotiras, A.; Bilello, M.; Rozycki, M.; Kirby, J.S.; Freymann, J.B.; Farahani, K.; Davatzikos, C. Advancing the cancer genome atlas glioma MRI collections with expert segmentation labels and radiomic features. Sci. Data 2017, 4, 170117. [Google Scholar] [CrossRef] [PubMed]
- Bakas, S.; Reyes, M.; Jakab, A.; Bauer, S.; Rempfler, M.; Crimi, A.; Shinohara, R.T.; Berger, C.; Ha, S.M.; Rozycki, M.; et al. Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv 2018, arXiv:1811.02629. [Google Scholar]
- Saut, O.; Lagaert, J.B.; Colin, T.; Fathallah-Shaykh, H.M. A multilayer grow-or-go model for GBM: Effects of invasive cells and anti-angiogenesis on growth. Bull. Math. Biol. 2014, 76, 2306–2333. [Google Scholar] [CrossRef]
- Goceri, E.; Goceri, N. Deep learning in medical image analysis: Recent advances and future trends. In Proceedings of the International Conferences on Computer Graphics, Visualization, Computer Vision and Image Processing 2017 and Big Data Analytics, Data Mining and Computational Intelligence 2017—Part of the Multi Conference on Computer Science and Information Systems 2017, Lisbon, Portugal, 21–23 November 2017. [Google Scholar]
- Rehman, M.U.; Chong, K.T. DNA6mA-MINT: DNA-6mA modification identification neural tool. Genes 2020, 11, 898. [Google Scholar] [CrossRef]
- Abbas, Z.; Tayara, H.; Chong, K.T. SpineNet-6mA: A Novel Deep Learning Tool for Predicting DNA N6-Methyladenine Sites in Genomes. IEEE Access 2020, 8, 201450–201457. [Google Scholar] [CrossRef]
- Alam, W.; Ali, S.D.; Tayara, H.; to Chong, K. A CNN-Based RNA N6-Methyladenosine Site Predictor for Multiple Species Using Heterogeneous Features Representation. IEEE Access 2020, 8, 138203–138209. [Google Scholar] [CrossRef]
- Ur Rehman, M.; Khan, S.H.; Rizvi, S.D.; Abbas, Z.; Zafar, A. Classification of skin lesion by interference of segmentation and convolotion neural network. In Proceedings of the 2018 2nd International Conference on Engineering Innovation (ICEI), Bangkok, Thailand, 5–6 July 2018; pp. 81–85. [Google Scholar]
- Khan, S.H.; Abbas, Z.; Rizvi, S.D.; Rizvi, S.M.D. Classification of Diabetic Retinopathy Images Based on Customised CNN Architecture. In Proceedings of the 2019 Amity International Conference on Artificial Intelligence (AICAI), Dubai, UAE, 4–6 February 2019; pp. 244–248. [Google Scholar]
- Banan, A.; Nasiri, A.; Taheri-Garavand, A. Deep learning-based appearance features extraction for automated carp species identification. Aquac. Eng. 2020, 89, 102053. [Google Scholar] [CrossRef]
- Fan, Y.; Xu, K.; Wu, H.; Zheng, Y.; Tao, B. Spatiotemporal Modeling for Nonlinear Distributed Thermal Processes Based on KL Decomposition, MLP and LSTM Network. IEEE Access 2020, 8, 25111–25121. [Google Scholar] [CrossRef]
- Shamshirband, S.; Rabczuk, T.; Chau, K.W. A survey of deep learning techniques: Application in wind and solar energy resources. IEEE Access 2019, 7, 164650–164666. [Google Scholar] [CrossRef]
- Faizollahzadeh Ardabili, S.; Najafi, B.; Shamshirband, S.; Minaei Bidgoli, B.; Deo, R.C.; Chau, K.W. Computational intelligence approach for modeling hydrogen production: A review. Eng. Appl. Comput. Fluid Mech. 2018, 12, 438–458. [Google Scholar] [CrossRef]
- Taormina, R.; Chau, K.W. ANN-based interval forecasting of streamflow discharges using the LUBE method and MOFIPS. Eng. Appl. Artif. Intell. 2015, 45, 429–440. [Google Scholar] [CrossRef]
- Wu, C.; Chau, K.W. Prediction of rainfall time series using modular soft computingmethods. Eng. Appl. Artif. Intell. 2013, 26, 997–1007. [Google Scholar] [CrossRef]
- Ghani, A. Healthcare electronics—A step closer to future smart cities. ICT Express 2019, 5, 256–260. [Google Scholar] [CrossRef]
- Ghani, A.; See, C.H.; Sudhakaran, V.; Ahmad, J.; Abd-Alhameed, R. Accelerating Retinal Fundus Image Classification Using Artificial Neural Networks (ANNs) and Reconfigurable Hardware (FPGA). Electronics 2019, 8, 1522. [Google Scholar] [CrossRef]
- Ilyas, T.; Khan, A.; Umraiz, M.; Kim, H. SEEK: A Framework of Superpixel Learning with CNN Features for Unsupervised Segmentation. Electronics 2020, 9, 383. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; pp. 234–241. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Kamnitsas, K.; Ledig, C.; Newcombe, V.F.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Rueckert, D.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 2017, 36, 61–78. [Google Scholar] [CrossRef]
- Pereira, S.; Pinto, A.; Alves, V.; Silva, C.A. Deep convolutional neural networks for the segmentation of gliomas in multi-sequence MRI. In Proceedings of the BrainLes 2015, Munich, Germany, 5 October 2015; pp. 131–143. [Google Scholar]
- Liu, L.; Zheng, G.; Bastian, J.D.; Keel, M.J.B.; Nolte, L.P.; Siebenrock, K.A.; Ecker, T.M. Periacetabular osteotomy through the pararectus approach: Technical feasibility and control of fragment mobility by a validated surgical navigation system in a cadaver experiment. Int. Orthop. 2016, 40, 1389–1396. [Google Scholar] [CrossRef]
- Lloyd, C.T.; Sorichetta, A.; Tatem, A.J. High resolution global gridded data for use in population studies. Sci. Data 2017, 4, 1–17. [Google Scholar] [CrossRef]
- Havaei, M.; Davy, A.; Warde-Farley, D.; Biard, A.; Courville, A.; Bengio, Y.; Pal, C.; Jodoin, P.M.; Larochelle, H. Brain tumor segmentation with deep neural networks. Med. Image Anal. 2017, 35, 18–31. [Google Scholar] [CrossRef]
- Shen, H.; Wang, R.; Zhang, J.; McKenna, S.J. Boundary-aware fully convolutional network for brain tumor segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Quebec City, QC, Canada, 10–14 September 2017; pp. 433–441. [Google Scholar]
- Meng, Z.; Fan, Z.; Zhao, Z.; Su, F. ENS-Unet: End-to-end noise suppression U-Net for brain tumor segmentation. In Proceedings of the 2018 40th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Honolulu, HI, USA, 18–21 July 2018; pp. 5886–5889. [Google Scholar]
- Wang, X.; Girshick, R.; Gupta, A.; He, K. Non-local neural networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 7794–7803. [Google Scholar]
- Zhao, H.; Zhang, Y.; Liu, S.; Shi, J.; Change Loy, C.; Lin, D.; Jia, J. Psanet: Point-wise spatial attention network for scene parsing. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 267–283. [Google Scholar]
- Xia, X.; Kulis, B. W-net: A deep model for fully unsupervised image segmentation. arXiv 2017, arXiv:1711.08506. [Google Scholar]
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A. Inception-v4, inception-resnet and the impact of residual connections on learning. arXiv 2016, arXiv:1602.07261. [Google Scholar]
- Chen, L.C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. Deeplab: Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 40, 834–848. [Google Scholar] [CrossRef] [PubMed]
- Tustison, N.J.; Avants, B.B.; Cook, P.A.; Zheng, Y.; Egan, A.; Yushkevich, P.A.; Gee, J.C. N4ITK: Improved N3 bias correction. IEEE Trans. Med. Imaging 2010, 29, 1310–1320. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Wu, Y.; Song, G.; Li, Z.; Zhang, Y.; Fan, Y. A deep learning model integrating FCNNs and CRFs for brain tumor segmentation. Med. Image Anal. 2018, 43, 98–111. [Google Scholar] [CrossRef]
- Devunooru, S.; Alsadoon, A.; Chandana, P.; Beg, A. Deep learning neural networks for medical image segmentation of brain tumours for diagnosis: A recent review and taxonomy. J. Ambient Intell. Human. Comput. 2020. [Google Scholar] [CrossRef]
- Chollet, F. Keras: Deep Learning Library for Theano and Tensorflow. 2015, Volume 7, p. T1. Available online: Https://keras.Io (accessed on 22 July 2020).
- Pereira, S.; Pinto, A.; Alves, V.; Silva, C.A. Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans. Med. Imaging 2016, 35, 1240–1251. [Google Scholar] [CrossRef]
- Dong, H.; Yang, G.; Liu, F.; Mo, Y.; Guo, Y. Automatic brain tumor detection and segmentation using U-Net based fully convolutional networks. In Proceedings of the Annual Conference on Medical Image Understanding and Analysis, Southampton, UK, 9–11 July 2017; pp. 506–517. [Google Scholar]
- Chen, L.; Wu, Y.; DSouza, A.M.; Abidin, A.Z.; Wismüller, A.; Xu, C. MRI tumor segmentation with densely connected 3D CNN. In Proceedings of the Medical Imaging 2018: Image Processing, Houston, TX, USA, 10–15 February 2018; Volume 10574. [Google Scholar]
- Kermi, A.; Mahmoudi, I.; Khadir, M.T. Deep convolutional neural networks using U-Net for automatic brain tumor segmentation in multimodal MRI volumes. In Proceedings of the International MICCAI Brainlesion Workshop, Granada, Spain, 16 September 2018; pp. 37–48. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Li, H.; Li, A.; Wang, M. A novel end-to-end brain tumor segmentation method using improved fully convolutional networks. Comput. Biol. Med. 2019, 108, 150–160. [Google Scholar] [CrossRef]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: Learning dense volumetric segmentation from sparse annotation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Athens, Greece, 17–21 October 2016; pp. 424–432. [Google Scholar]
- Albiol, A.; Albiol, A.; Albiol, F. Extending 2D deep learning architectures to 3D image segmentation problems. In Proceedings of the International MICCAI Brainlesion Workshop, Granada, Spain, 16 September 2018; pp. 73–82. [Google Scholar]
- Wang, G.; Li, W.; Ourselin, S.; Vercauteren, T. Automatic brain tumor segmentation using convolutional neural networks with test-time augmentation. In Proceedings of the International MICCAI Brainlesion Workshop, Granada, Spain, 16 September 2018; pp. 61–72. [Google Scholar]
- Chen, W.; Liu, B.; Peng, S.; Sun, J.; Qiao, X. S3D-UNet: Separable 3D U-Net for brain tumor segmentation. In Proceedings of the International MICCAI Brainlesion Workshop, Granada, Spain, 16 September 2018; pp. 358–368. [Google Scholar]
- Hu, K.; Gan, Q.; Zhang, Y.; Deng, S.; Xiao, F.; Huang, W.; Cao, C.; Gao, X. Brain tumor segmentation using multi-cascaded convolutional neural networks and conditional random field. IEEE Access 2019, 7, 92615–92629. [Google Scholar] [CrossRef]
| Class | Area Covered in % |
|---|---|
| Healthy Tissue | 98.46 |
| Edema | 1.02 |
| Enhancing Tumor | 0.29 |
| Non-Enhancing Tumor | 0.23 |
| Architecture | Whole | Core | Enhancing |
|---|---|---|---|
| CNN [38] | 0.840 | 0.720 | 0.620 |
| U-Net [39] | 0.831 | 0.801 | 0.750 |
| Densely CNN [40] | 0.720 | 0.830 | 0.810 |
| ResU-Net [41] | 0.88 | 0.850 | 0.750 |
| FCNN [35] | 0.865 | 0.864 | 0.816 |
| Proposed BU-Net | 0.901 | 0.867 | 0.835 |
| Architecture | Whole | Core | Enhancing |
|---|---|---|---|
| Seg-Net [42] | 0.833 | 0.703 | 0.496 |
| U-Net [39] | 0.870 | 0.762 | 0.700 |
| ResU-Net [41] | 0.873 | 0.768 | 0.716 |
| PSPNet [43] | 0.809 | 0.701 | 0.554 |
| NovelNet [43] | 0.876 | 0.763 | 0.642 |
| Proposed BU-Net | 0.892 | 0.783 | 0.736 |
| Architecture | Whole | Core | Enhancing |
|---|---|---|---|
| U-Net [39] | 0.860 | 0.790 | 0.767 |
| 3DU-Net [44] | 0.885 | 0.718 | 0.760 |
| ResU-Net [41] | 0.867 | 0.803 | 0.768 |
| Ensemble Net [45] | 0.881 | 0.777 | 0.773 |
| TTA [46] | 0.873 | 0.783 | 0.754 |
| S3DU-Net [47] | 0.894 | 0.831 | 0.749 |
| MCC [48] | 0.882 | 0.748 | 0.718 |
| Proposed BU-Net | 0.901 | 0.837 | 0.788 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rehman, M.U.; Cho, S.; Kim, J.H.; Chong, K.T. BU-Net: Brain Tumor Segmentation Using Modified U-Net Architecture. Electronics 2020, 9, 2203. https://doi.org/10.3390/electronics9122203
Rehman MU, Cho S, Kim JH, Chong KT. BU-Net: Brain Tumor Segmentation Using Modified U-Net Architecture. Electronics. 2020; 9(12):2203. https://doi.org/10.3390/electronics9122203
Chicago/Turabian StyleRehman, Mobeen Ur, SeungBin Cho, Jee Hong Kim, and Kil To Chong. 2020. "BU-Net: Brain Tumor Segmentation Using Modified U-Net Architecture" Electronics 9, no. 12: 2203. https://doi.org/10.3390/electronics9122203
APA StyleRehman, M. U., Cho, S., Kim, J. H., & Chong, K. T. (2020). BU-Net: Brain Tumor Segmentation Using Modified U-Net Architecture. Electronics, 9(12), 2203. https://doi.org/10.3390/electronics9122203








