StoolNet for Color Classification of Stool Medical Images
Abstract
:1. Introduction
- The current stool color examination heavily depends on the medical laboratorians’ professional skills. We first focus on this important factor and propose a lightweight, practical, and efficient automatic color classification method to remarkably alleviate burdens on medical laboratorians.
- The developed preprocessing method can automatically and accurately segment the effective stool region. The designed model, namely StoolNet, can converge well and automatically and accurately classify the stool color.
2. Proposed Method
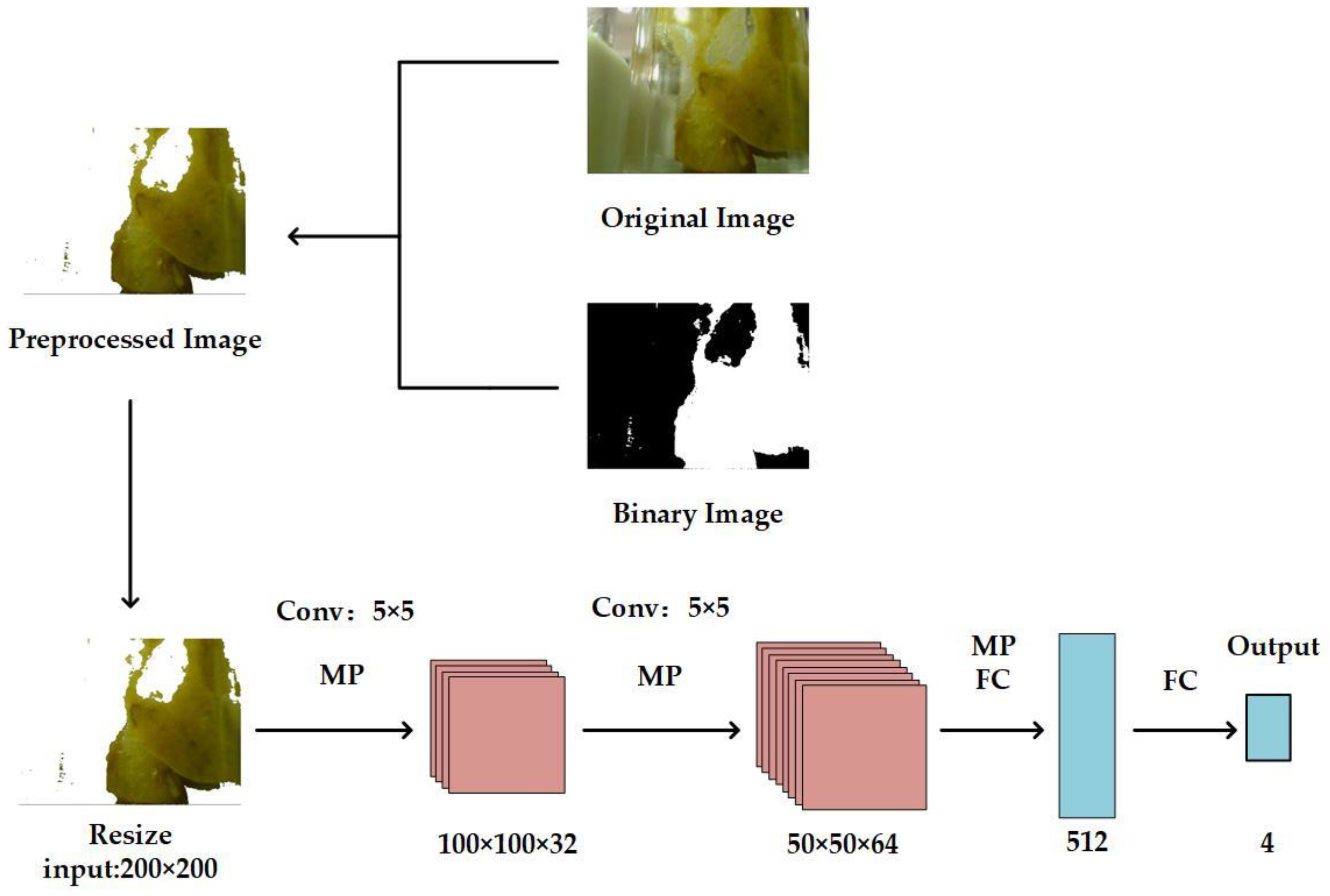
2.1. Overview of the Proposed Method
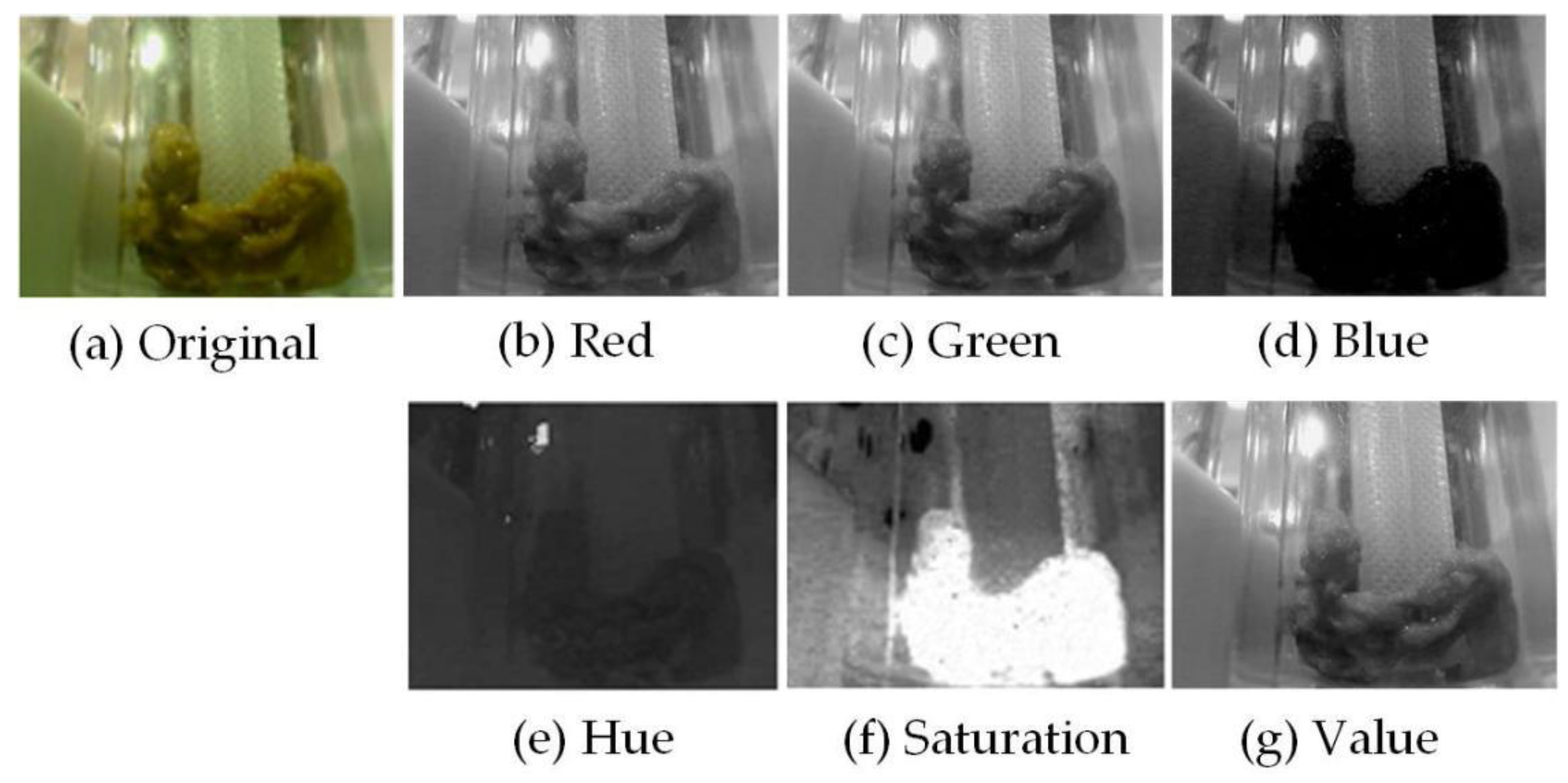
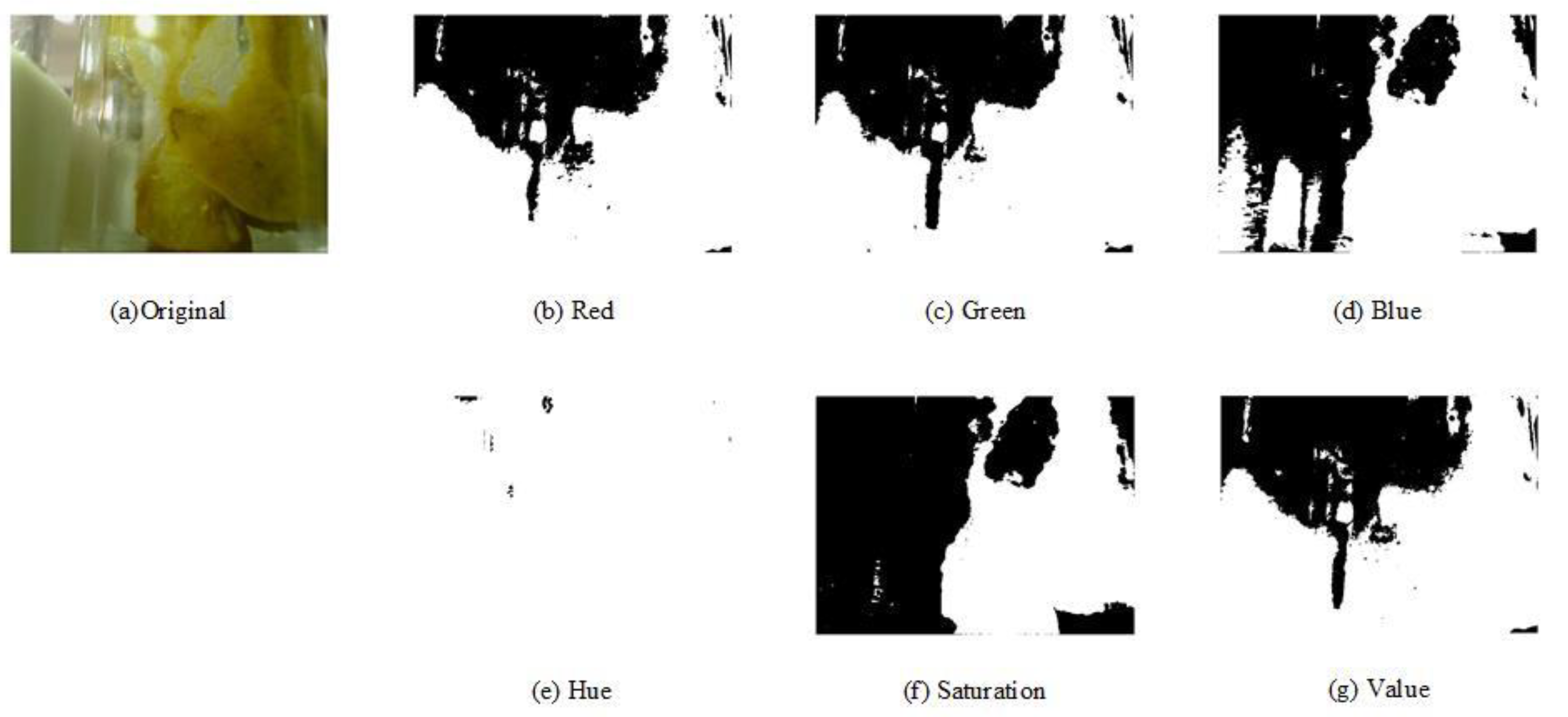
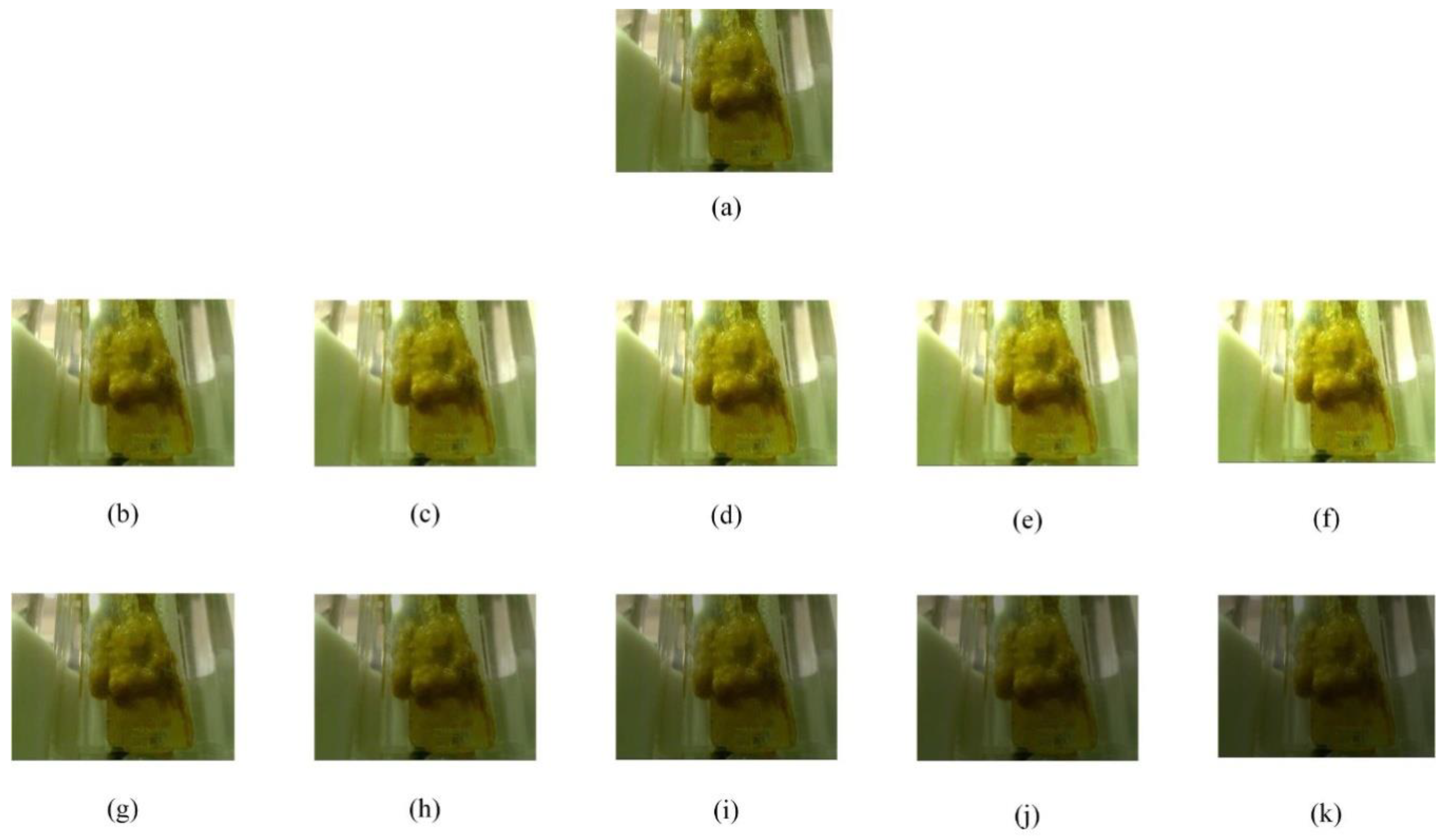
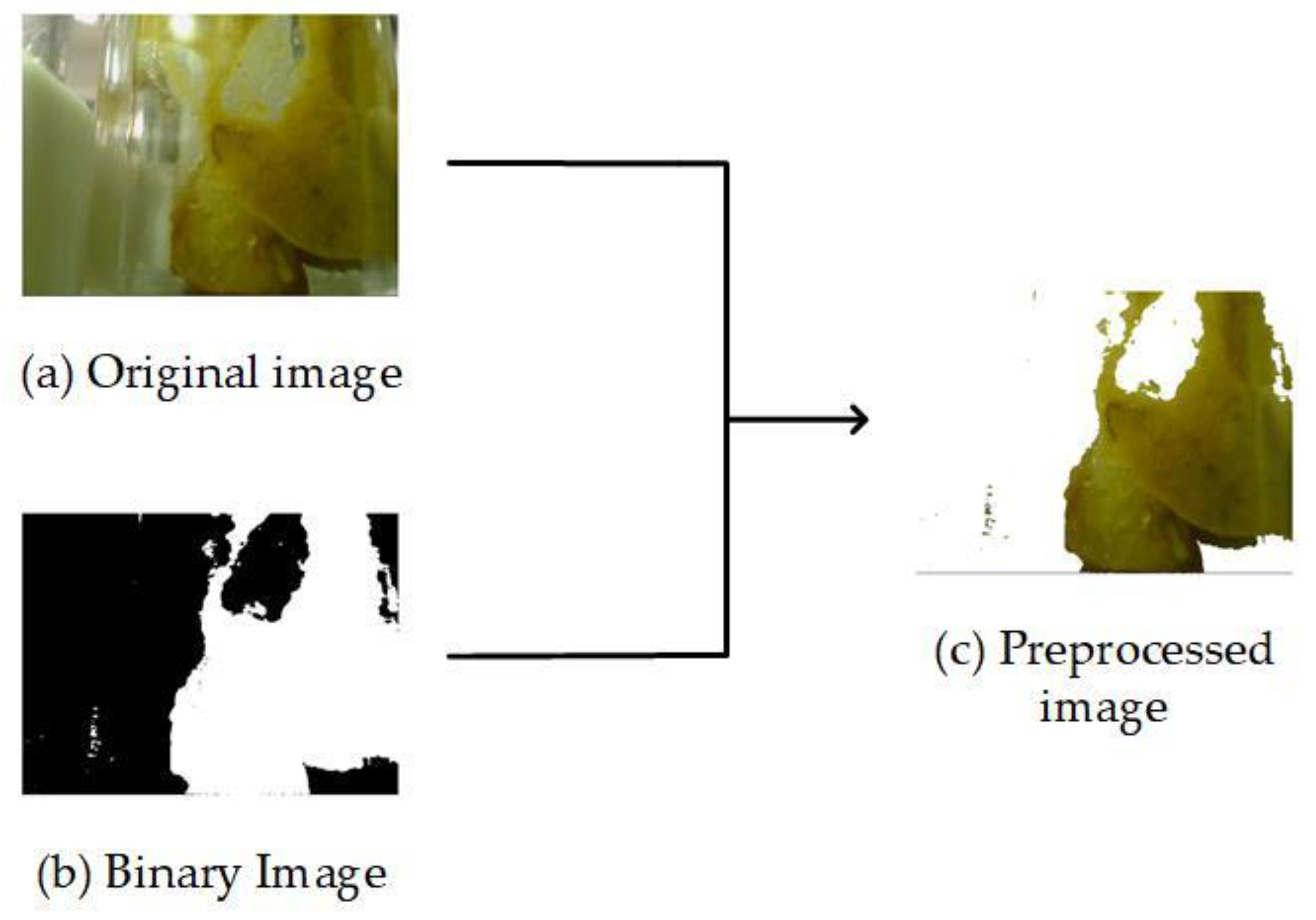
2.2. Preprocessing Stage
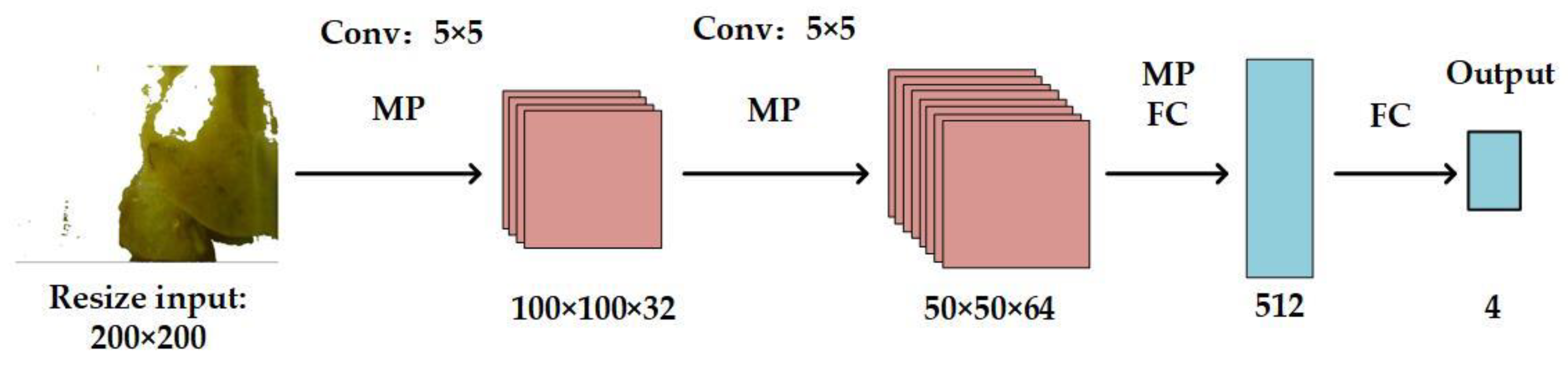
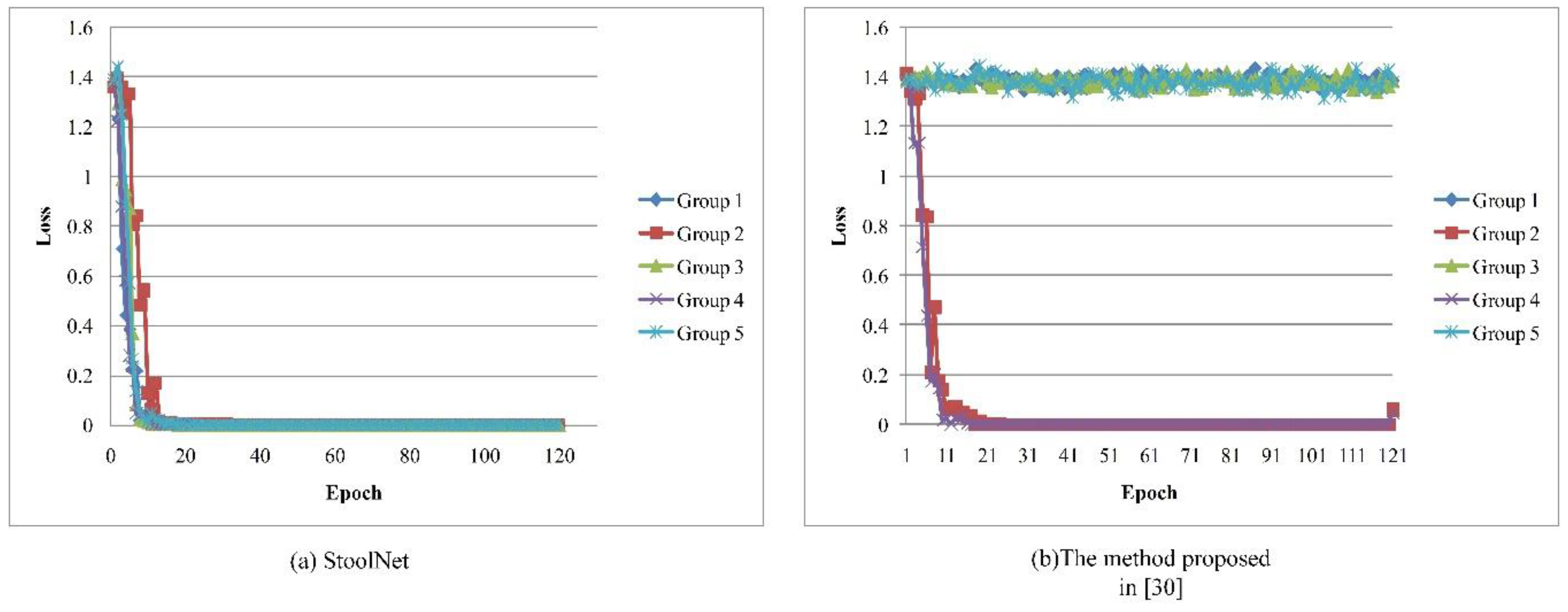
2.3. StoolNet
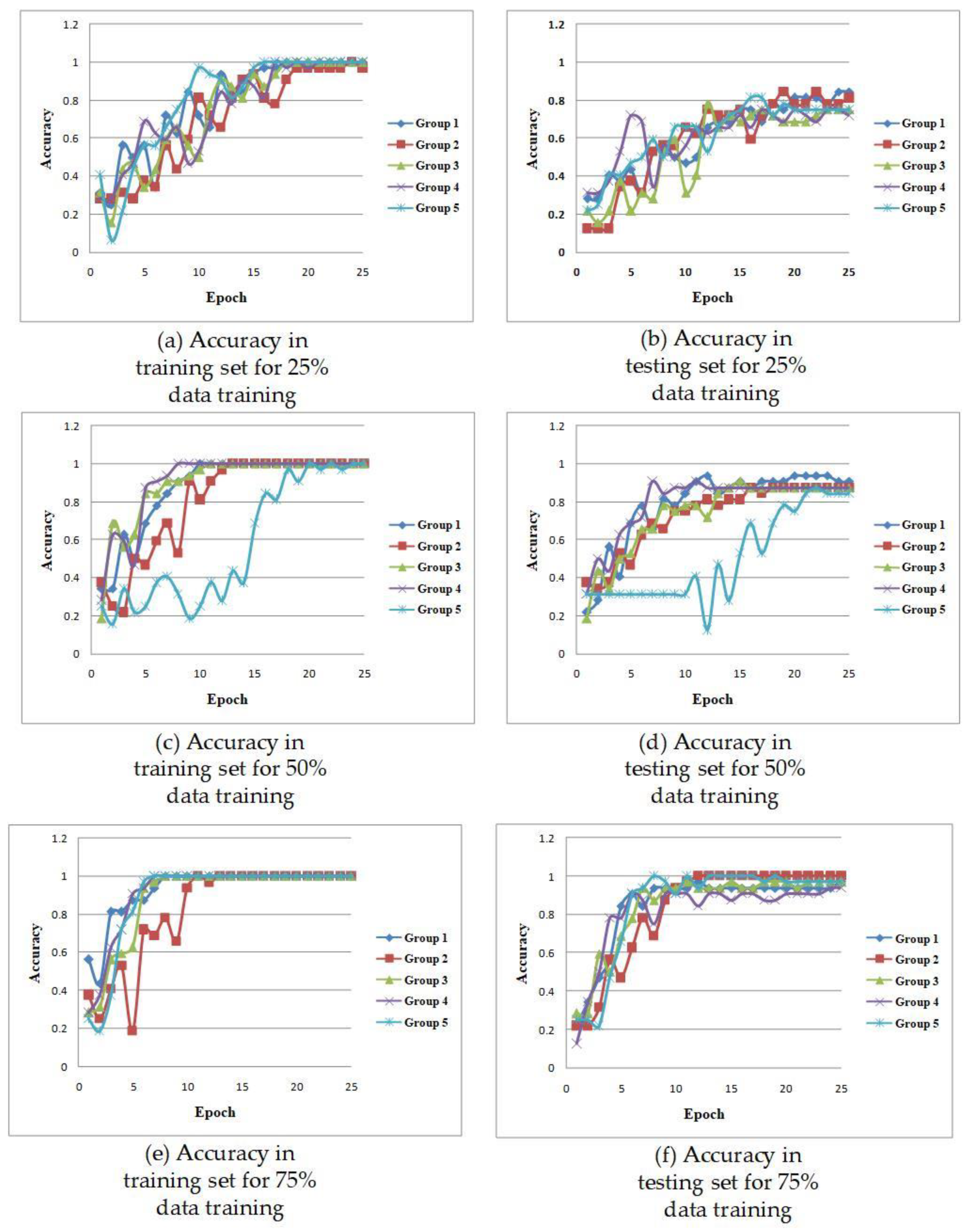
3. Experiments
4. Conclusions and Future Work
Author Contributions
Funding
Conflicts of Interest
References
- Marques, G.; Pitarma, R.; Garcia, N.M.; Pombo, N. Internet of Things Architectures, Technologies, Applications, Challenges, and Future Directions for Enhanced Living Environments and Healthcare Systems: A Review. Electronics 2019, 8, 1081. [Google Scholar] [CrossRef]
- Gil-Martin, M.; Montero, J.M.; San-Segundo, R. Parkinson’s Disease Detection from Drawing Movements Using Convolutional Neural Networks. Electronics 2019, 8, 907. [Google Scholar] [CrossRef]
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global Cancer Statistics 2018: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Casavant, E.P.; Dethlefsen, L.; Sankaran, K.; Sprockett, D.; Holmes, S.; Relman, D.A.; Elias, J. Strategies for understanding Dynamic, Personalized Profiles of Host-Derived Proteins and Microbes from Human Stool. bioRxiv 2019, 551143. [Google Scholar] [CrossRef]
- Ahad, A.; Tahir, M.; Yau, K.A. 5G-Based Smart Healthcare Network: Architecture, Taxonomy, Challenges and Future Research Directions. IEEE Access 2019, 7, 100747–100762. [Google Scholar] [CrossRef]
- Zhao, C.Y.; Wang, Z.Q.; Li, H.Y.; Wu, X.Y.; Qiao, S.; Sun, J.N. A New Approach for Medical Image Enhancement Based on Luminance-Level Modulation and Gradient Modulation. Biomedical Signal Process. Control 2019, 48, 189–196. [Google Scholar] [CrossRef]
- Shen, T.; Wang, Y. Medical Image Segmentation Based on Improved Watershed Algorithm. In Proceedings of the IEEE Advanced Information Technology, Electronic and Automation Control Conference (IAEAC), Chongqing, China, 12–14 October 2018; pp. 1695–1698. [Google Scholar]
- Huang, C.C.; Nguyen, M.H. X-Ray Enhancement Based on Component Attenuation, Contrast Adjustment, and Image Fusion. IEEE Trans. Image Process. 2019, 28, 127–141. [Google Scholar] [CrossRef]
- Higaki, T.; Nakamura, Y.; Tatsugami, F.; Nakaura, T.; Awai, K. Improvement of Image Quality at CT and MRI using Deep Learning. Jpn. J. Radiol. 2019, 37, 73–80. [Google Scholar] [CrossRef]
- Xue, Y.M.; Liu, P. Medical Ultrasonic Images Denoising and Enhancement. Investig. Clin. 2019, 60, 728–738. [Google Scholar]
- Sharp, S.E.; Suarez, C.A.; Duran, Y.; Poppiti, R.J. Evaluation of the Triage Micro Parasite Panel for Detection of Giardia Lamblia, Entamoeba Histolytica/Entamoeba Dispar, and Cryptosporidium Parvum in Patient Stool Specimens. J. Clin. Microbiol. 2001, 39, 332–334. [Google Scholar] [CrossRef]
- Tchiotsop, D.; Tchinda, B.S.; Tchinda, R.; Kenne, G. Edge Detection of Intestinal Parasites in Stool Microscopic Images Using Multi-scale Wavelet Transform. Signal Image Video Process. 2015, 9, 121–134. [Google Scholar] [CrossRef]
- Piekkala, M.; Alfthan, H.; Merras-Salmio, L.; PuustinenWikström, A.; Heiskanen, K.; Jaakkola, T.; Klemetti, P.; Färkkilä, M.; Kolho, K. Fecal Calprotectin Test Performed at Home: A Prospective Study of Pediatric Patients with Inflammatory Bowel Disease. J. Pediatric Gastroenterol. Nutr. 2018, 66, 926–931. [Google Scholar] [CrossRef] [PubMed]
- Rundo, F.; Conoci, S.; Ortis, A.; Battiato, S. An Advanced Bio-inspired PhotoPlethymoGraphy (PPG) and ECG Pattern Recognition System for Medical Assessment. Sensors 2018, 18, 405. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Pan, H.; Han, J.; Xie, X.; Zhang, Z.; Zhai, X. Corner Detection and Matching Methods for Brain Medical Image Classification. In Proceedings of the IEEE International Bioinformatics and Biomedicine (BIBM), Shenzhen, China, 15–18 December 2016; pp. 475–478. [Google Scholar]
- Chen, J.; You, H. Efficient Classification of Benign and Malignant Thyroid Tumors based on Characteristics of Medical Ultrasonic Images. In Proceedings of the IEEE Advanced Information Management, Communicates, Electronic and Automation Control Conference (IMCEC), Xi’an, China, 3–5 October 2016; pp. 950–954. [Google Scholar]
- Opbroek, A.V.; Achterberg, H.C.; Vernooij, M.W.; De Bruijne, M. Transfer Learning for Image Segmentation by Combining Image Weighting and Kernel Learning. IEEE Trans. Med. Imaging 2019, 38, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.J.; Lin, C.H.; Sun, C.C.; Wang, S.H. Evolutionary-Fuzzy-Integral-Based Convolutional Neural Networks for Facial Image Classification. Electronics 2019, 8, 997. [Google Scholar] [CrossRef]
- Cui, G.; Wang, S.; Wang, Y.; Liu, Z.; Yuan, Y.; Wang, Q. Preceding Vehicle Detection Using Faster R-CNN Based on Speed Classification Random Anchor and Q-Square Penalty Coefficient. Electronics 2019, 8, 1024. [Google Scholar] [CrossRef]
- Nonis, F.; Dagnes, N.; Marcolin, F.; Vezzetti, E. 3D Approaches and Challenges in Facial Expression Recognition Algorithms—A Literature Review. Appl. Sci. 2019, 9, 3904. [Google Scholar] [CrossRef]
- Olga, R.; Deng, J.; Su, H.; Krause, J.; Satheesh, S.; Ma, S.; Huang, Z.H.; Karpathy, A.; Khosla, A.; Bernstein, M.; et al. ImageNet Large Scale Visual Recognition Challenge. Int. J. Comput. Vis. 2015, 115, 211–252. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Neural Networks for Large-scale Image Recognition. arXiv 2014, arXiv:abs/1409.1556. Available online: https://arxiv.org/abs/1409.1556 (accessed on 9 November 2019).
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Iandola, F.; Moskewicz, M.; Karayev, S.; Girshick, R.; Darrell, T.; Keutzer, K. DenseNet: Implementing Efficient ConvNet Descriptor Pyramids. arXiv 2014, arXiv:abs/1404.1869. Available online: https://arxiv.org/abs/1404.1869 (accessed on 9 November 2019).
- Razzak, M.I.; Naz, S.; Zaib, A. Deep Learning for Medical Image Processing: Overview, Challenges and the Future. Classif. BioApps 2018, 26, 323–350. [Google Scholar]
- Lee, J.Y.; Choi, S.; Chung, J.W. Automated Classification of the Tympanic Membrane Using a Convolutional Neural Network. Appl. Sci. 2019, 9, 1827. [Google Scholar] [CrossRef]
- Sugimori, H.; Kawakami, M. Automatic Detection of a Standard Line for Brain Magnetic Resonance Image Using Deep Learning. Appl. Sci. 2019, 9, 3849. [Google Scholar] [CrossRef]
- Khawaldeh, S.; Pervaiz, U.; Rafiq, A.; Alkhawaldeh, R.S. Noninvasive Grading of Glioma Tumor Using Magnetic Resonance Imaging with Convolutional Neural Networks. Appl. Sci. 2018, 8, 27. [Google Scholar] [CrossRef]
- Oktay, O.; Ferrante, E.; Kamnitsas, K.; Heinrich, M.; Bai, W.; Caballero, J.; Cook, S.A.; De Marvao, A.; Dawes, T.; O‘Regan, D.P.; et al. Anatomically Constrained Neural Networks (ACNNs): Application to Cardiac Image Enhancement and Segmentation. IEEE Trans. Med Imaging 2018, 37, 384–395. [Google Scholar] [CrossRef]
- Hachuel, D.; Jha, A.; Estrin, D.; Martinez, A.; Staller, K.; Velez, C. Augmenting Gastrointestinal Health: A Deep Learning Approach to Human Stool Recognition and Characterization in Macroscopic Images. arXiv 2019, arXiv:Abs/1903.10578. Available online: https://arxiv.org/abs/1903.10578 (accessed on 9 November 2019). [CrossRef]
- Ostu, N. A Threshold Selection Method from Gray Level Histograms. IEEE Trans. Syst. Man Cybern. 1979, 9, 62–69. [Google Scholar]
- Abadi, M. TensorFlow: Learning Functions at Scale. ACM Sigplan Not. 2016, 51, 1. [Google Scholar] [CrossRef]



| Group 1 | Group 2 | Group 3 | Group 4 | Group 5 | Average | |
|---|---|---|---|---|---|---|
| 25% training | 48 | 18 | 18 | 26 | 27 | 27.4 |
| 25% training * | 18 | 26 | 18 | 21 | 16 | 19.8 |
| 50% training | 20 | 38 | 30 | 22 | 24 | 26.8 |
| 50% training * | 10 | 13 | 11 | 24 | 24 | 16.4 |
| 75% training | 26 | 25 | 13 | 28 | 41 | 26.6 |
| 75% training * | 8 | 13 | 8 | 7 | 7 | 8.6 |
| Group 1 | Group 2 | Group 3 | Group 4 | Group 5 | Average | |
|---|---|---|---|---|---|---|
| 25% training | 0.6250 | 0.7500 | 0.8750 | 0.6250 | 0.7500 | 0.7250 |
| 25%training * | 0.8125 | 0.8125 | 0.7188 | 0.7500 | 0.7188 | 0.7625 |
| 50% training | 0.9375 | 0.9375 | 0.8125 | 0.8750 | 0.9375 | 0.9000 |
| 50%training * | 0.9375 | 0.8438 | 0.8125 | 0.8750 | 0.8438 | 0.8625 |
| 75% training | 0.8750 | 1.0000 | 0.9375 | 0.8750 | 0.8125 | 0.9000 |
| 75%training * | 0.9688 | 1.0000 | 0.9688 | 0.9375 | 1.0000 | 0.9750 |
| Illumination Scale | Yellow | Brown | Black | Green |
|---|---|---|---|---|
| 0.5 | 0.0043 | 0.0072 | 0.0298 | 0.0072 |
| 0.6 | 0.0035 | 0.0060 | 0.0310 | 0.0058 |
| 0.7 | 0.0033 | 0.0054 | 0.0194 | 0.0052 |
| 0.8 | 0.0026 | 0.0047 | 0.0212 | 0.0045 |
| 0.9 | 0.0029 | 0.0048 | 0.0139 | 0.0047 |
| 1.1 | 0.0026 | 0.0040 | 0.0112 | 0.0040 |
| 1.2 | 0.0025 | 0.0035 | 0.0086 | 0.0040 |
| 1.3 | 0.0039 | 0.0050 | 0.0159 | 0.0055 |
| 1.4 | 0.0067 | 0.0079 | 0.0198 | 0.0080 |
| 1.5 | 0.0131 | 0.0156 | 0.0410 | 0.0189 |
| Illumination Scale | Group 1 | Group 2 | Group 3 | Group 4 | Group 5 | Average |
|---|---|---|---|---|---|---|
| 0.5 | 90.63 | 100.00 | 93.75 | 93.75 | 96.88 | 95.00 |
| 0.6 | 96.88 | 96.88 | 100.00 | 93.75 | 93.75 | 96.25 |
| 0.7 | 100.00 | 93.75 | 100.00 | 93.75 | 100.00 | 97.50 |
| 0.8 | 96.88 | 93.75 | 96.88 | 90.63 | 96.88 | 95.00 |
| 0.9 | 93.75 | 96.88 | 93.75 | 96.88 | 94.79 | 95.21 |
| 1.1 | 94.27 | 93.75 | 96.88 | 96.88 | 96.88 | 95.73 |
| 1.2 | 96.88 | 94.27 | 95.31 | 93.75 | 100.00 | 96.04 |
| 1.3 | 100.00 | 100.00 | 90.63 | 96.88 | 96.88 | 96.88 |
| 1.4 | 93.75 | 96.88 | 100.00 | 93.75 | 93.75 | 95.63 |
| 1.5 | 96.88 | 100.00 | 100.00 | 100.00 | 87.50 | 96.88 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Z.; Leng, L.; Kim, B.-G. StoolNet for Color Classification of Stool Medical Images. Electronics 2019, 8, 1464. https://doi.org/10.3390/electronics8121464
Yang Z, Leng L, Kim B-G. StoolNet for Color Classification of Stool Medical Images. Electronics. 2019; 8(12):1464. https://doi.org/10.3390/electronics8121464
Chicago/Turabian StyleYang, Ziyuan, Lu Leng, and Byung-Gyu Kim. 2019. "StoolNet for Color Classification of Stool Medical Images" Electronics 8, no. 12: 1464. https://doi.org/10.3390/electronics8121464
APA StyleYang, Z., Leng, L., & Kim, B.-G. (2019). StoolNet for Color Classification of Stool Medical Images. Electronics, 8(12), 1464. https://doi.org/10.3390/electronics8121464







