A Hybrid Deep Learning-Based Approach for Brain Tumor Classification
Abstract
:1. Introduction
2. Related Works
3. Materials and Methods
3.1. Dataset and Preprocessing
3.2. DeepTumorNet
3.2.1. Image Input Layer
3.2.2. Convolutional Layer
3.2.3. Activation Function
3.2.4. Batch Normalization Layer
3.2.5. Pooling Layer
3.2.6. Fully Connected Layer
3.2.7. Softmax Layer
3.2.8. Classification Layer
3.2.9. Training Parameters
4. Results and Discussion
4.1. Performance Metrics (Accuracy, Precision, Recall, and F1 Score)
4.2. Comparison of DeepTumorNet with the Pretrained Transfer Learning Approaches
4.3. Comparative Results of DeepTumorNet with the State-of-the-Art Classification Approaches
5. Conclusions and Future Work
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| AI. | Artificial intelligence |
| ML | Machine learning |
| DL | Deep learning |
| TL | Transfer Learning |
| BT | Brain tumors |
| MRI. | Magnetic resonance imaging |
| K-NN | K-nearest neighbor |
| SVM | Support vector machine |
| CNN | Convolutional neural network |
| PCA. | Principal component analysis |
| LDA | Linear discriminant analysis |
References
- Zhao, L.; Jia, K. Multiscale CNNs for brain tumors segmentation and diagnosis. Comput. Math. Methods Med. 2016, 2016, 8356294. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saddique, M.; Kazmi, J.H.; Qureshi, K. A hybrid approach of using symmetry technique for brain tumors. Comput. Math. Methods Med. 2014, 2014, 712783. [Google Scholar] [CrossRef] [PubMed]
- Shree, N.V.; Kumar, T.N. Identification and classification of BTMRI images with feature extraction using DWT and probabilistic neural network. Brain Inform. 2018, 5, 23–30. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Komninos, J.; Vlassopoulou, V.; Protopapa, D.; Korfias, S.; Kontogeorgos, G.; Sakas, D.E.; Thalassinos, N.C. Tumors metastatic to the pituitary gland: Case report and literature review. J. Clin. Endocrinol. Metab. 2004, 2, 574–580. [Google Scholar] [CrossRef] [Green Version]
- DeAngelis, L.M. Brain tumors. N. Engl. J. Med. 2001, 344, 114–123. [Google Scholar] [CrossRef] [Green Version]
- Louis, D.N.; Perry, A.; Reifenberger, G.; Von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization classification of tumors of the central nervous system: A summary. Acta Neuropathol. 2016, 132, 803–820. [Google Scholar] [CrossRef] [Green Version]
- Chahal, P.K.; Pandey, S.; Goel, S. A survey on brain tumors detection techniques for MR images. Multimed. Tools Appl. 2020, 79, 21771–21814. [Google Scholar] [CrossRef]
- Sajjad, M.; Khan, S.; Muhammad, K.; Wu, W.; Ullah, A.; Baik, S.W. Multi-grade brain tumors classification using deep CNN with extensive data augmentation. J. Comput. Sci. 2019, 30, 174–182. [Google Scholar] [CrossRef]
- Rehman, A.; Naz, S.; Razzak, M.I.; Akram, F.; Imran, M. A deep learning-based framework for automatic brain tumors classification using transfer learning. Circuits Syst. Signal Process. 2020, 39, 757–775. [Google Scholar] [CrossRef]
- Wang, Y.; Zu, C.; Hu, G.; Luo, Y.; Ma, Z.; He, K.; Wu, X.; Zhou, J. Automatic tumor segmentation with deep convolutional neural networks for radiotherapy applications. Neural Process. Lett. 2018, 48, 1323–1334. [Google Scholar] [CrossRef]
- Jégou, S.; Drozdzal, M.; Vazquez, D.; Romero, A.; Bengio, Y. The one hundred layers tiramisu: Fully convolutional denseness for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition Workshops, Honolulu, HI, USA, 21–26 July 2017; pp. 11–19. [Google Scholar]
- Zhang, Q.; Cui, Z.; Niu, X.; Geng, S.; Qiao, Y. Image segmentation with pyramid dilated convolution based on ResNet and U-Net. In Proceedings of the International Conference on Neural Information Processing, Guangzhou, China, 14 November 2017; pp. 364–372. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 26 June–1 July 2016; pp. 2818–2826. [Google Scholar]
- Ding, Y.; Zhang, C.; Lan, T.; Qin, Z.; Zhang, X.; Wang, W. Classification of Alzheimer’s disease based on the combination of morphometric feature and texture feature. In Proceedings of the 2015 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Washington, DC, USA, 9–12 November 2015; pp. 409–412. [Google Scholar]
- Ijaz, A.; Ullah, I.; Khan, W.U.; Ur Rehman, A.; Adrees, M.S.; Saleem, M.Q.; Cheikhrouhou, O.; Hamam, H.; Shafiq, M. Efficient algorithms for E-healthcare to solve multiobject fuse detection problem. J. Healthc. Eng. 2021, 2021, 9500304. [Google Scholar]
- Ahmad, I.; Liu, Y.; Javeed, D.; Ahmad, S. A decision-making technique for solving order allocation problem using a genetic algorithm. IOP Conf. Ser. Mater. Sci. Eng. 2020, 853, 012054. [Google Scholar] [CrossRef]
- Binaghi, E.; Omodei, M.; Pedoia, V.; Balbi, S.; Lattanzi, D.; Monti, E. Automatic segmentation of MR brain tumors images using support vector machine in combination with graph cut. In Proceedings of the 6th International Joint Conference on Computational Intelligence (IJCCI), Rome, Italy, 22–24 October 2014; pp. 152–157. [Google Scholar]
- Bahadure, N.B.; Ray, A.K.; Thethi, H.P. The comparative approach of MRI-based brain tumors segmentation and classification using genetic algorithm. J. Digit. Imaging 2018, 31, 477–489. [Google Scholar] [CrossRef]
- Kaya, I.E.; Pehlivanlı, A.Ç.; Sekizkardeş, E.G.; Ibrikci, T. PCA based clustering for brain tumors segmentation of T1w MRI images. Comput. Methods Programs Biomed. 2017, 140, 19–28. [Google Scholar] [CrossRef]
- Zikic, D.; Glocker, B.; Konukoglu, E.; Criminisi, A.; Demiralp, C.; Shotton, J.; Thomas, O.M.; Das, T.; Jena, R.; Price, S.J. Forests for tissue-specific segmentation of high-grade gliomas in multi-channel MR. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Berlin, Heidelberg, 1 October 2012; pp. 369–376. [Google Scholar]
- Nikam, R.D.; Lee, J.; Choi, W.; Banerjee, W.; Kwak, M.; Yadav, M.; Hwang, H. Ionic sieving through one-atom-thick 2D material enables analog nonvolatile memory for neuromorphic computing. Adv. Electron. Mater. 2021, 17, 2100142. [Google Scholar] [CrossRef]
- Zhang, J.P.; Li, Z.W.; Yang, J. A parallel SVM training algorithm on large-scale classification problems. In Proceedings of the 2005 International Conference on Machine Learning and Cybernetics, Guangzhou, China, 18–21 August 2005; pp. 1637–1641. [Google Scholar]
- Ye, F.; Yang, J. A deep neural network model for speaker identification. Appl. Sci. 2021, 11, 3603. [Google Scholar] [CrossRef]
- Ijaz, A.; Liu, Y.; Javeed, D.; Shamshad, N.; Sarwr, D.; Ahmad, S. A review of artificial intelligence techniques for selection & evaluation. IOP Conf. Ser. Mater. Sci. Eng. 2020, 853, 012055. [Google Scholar]
- Akbari, H.; Khalighinejad, B.; Herrero, J.L.; Mehta, A.D.; Mesgarani, N. Towards reconstructing intelligible speech from the human auditory cortex. Sci. Rep. 2019, 9, 874. [Google Scholar] [CrossRef] [Green Version]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar]
- Harish, P.; Baskar, S. MRI based detection and classification of brain tumor using enhanced faster R-CNN and Alex Net model. Mater. Today Proc. 2020. [Google Scholar] [CrossRef]
- Iandola, F.N.; Han, S.; Moskewicz, M.W.; Ashraf, K.; Dally, W.J.; Keutzer, K. AlexNet-level accuracy with 50× fewer parameters and <0.5 MB model size. In Proceedings of the 5th International Conference Learning Representation, Toulon, France, 24–26 April 2016; pp. 1–8. [Google Scholar]
- Saleem, S.; Amin, J.; Sharif, M.; Anjum, M.A.; Iqbal, M.; Wang, S.H. A deep network designed for segmentation and classification of leukemia using fusion of the transfer learning models. Complex Intell. Syst. 2021, 7, 1–16. [Google Scholar] [CrossRef]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Zhang, X.; Zhou, X.; Lin, M.; Sun, J. Shufflenet: An extremely efficient convolutional neural network for mobile devices. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–22 June 2018; pp. 6848–6856. [Google Scholar]
- Gupta, A.; Verma, A.; Kaushik, D.; Garg, M. Applying deep learning approach for brain tumors detection. Mater. Today Proc. 2020, 50, 1–13. [Google Scholar] [CrossRef]
- Mehrotra, R.; Ansari, M.A.; Agrawal, R.; Anand, R.S. A transfer learning approach for AI-based classification of brain tumors. Mach. Learn. Appl. 2020, 2, 100003. [Google Scholar] [CrossRef]
- Ramzan, F.; Khan, M.U.G.; Rehmat, A.; Iqbal, S.; Saba, T.; Rehman, A.; Mehmood, Z. A deep learning approach for automated diagnosis and multi-class classification of Alzheimer’s disease stages using resting-state fMRI and residual neural networks. J. Med. Syst. 2020, 44, 37. [Google Scholar] [CrossRef]
- Selvapandian, A.; Manivannan, K. Fusion-based glioma brain tumors detection and segmentation using ANFIS classification. Comput. Methods Programs Biomed. 2018, 166, 33–38. [Google Scholar] [CrossRef]
- Sultan, H.H.; Salem, N.M.; Al-Atabany, W. Multi-classification of brain tumors images using deep neural Network. IEEE Access 2019, 7, 69215–69225. [Google Scholar] [CrossRef]
- Tufail, A.B.; Ijaz Ahmad, A. Diagnosis of diabetic retinopathy through retinal fundus images and 3D convolutional neural networks with limited number of samples. Wirel. Commun. Mob. Comput. 2021, 2021, 6013448. [Google Scholar] [CrossRef]
- Pitchai, R.; Supraja, P.; Victoria, A.H.; Madhavi, M. Brain tumors egmentation using deep learning and fuzzy K-means clustering for magnetic resonance images. Neural Processing Lett. 2021, 53, 2519–2532. [Google Scholar] [CrossRef]
- Amin, J.; Sharif, M.; Yasmin, M.; Fernandes, S.L. A distinctive approach in brain tumors detection and classification using MRI. Pattern Recognit. Lett. 2020, 139, 118–127. [Google Scholar] [CrossRef]
- Özyurt, F.; Sert, E.; Avci, E.; Dogantekin, E. Brain tumors detection based on Convolutional Neural Network with neutrosophic expert maximum fuzzy sure entropy. Measurement 2019, 147, 106830. [Google Scholar] [CrossRef]
- Kaplan, K.; Kaya, Y.; Kuncan, M.; Ertunç, H.M. Brain tumors classification using modified local binary patterns (LBP) feature extraction methods. Med. Hypotheses 2020, 139, 109696. [Google Scholar] [CrossRef] [PubMed]
- Çinar, A.; Yildirim, M. Detection of tumors on brain MRI images using the hybrid convolutional neural network architecture. Med. Hypotheses 2020, 139, 109684. [Google Scholar] [CrossRef] [PubMed]
- Mohsen, H.; El-Dahshan, E.S.; El-Horbaty, E.S.; Salem, A.B. Classification using deep learning neural networks for brain tumors. Future Comput. Inform. J. 2018, 3, 68–71. [Google Scholar] [CrossRef]
- Lather, M.; Singh, P. Investigating brain tumors segmentation and detection techniques. Procedia Comput. Sci. 2020, 167, 121–130. [Google Scholar] [CrossRef]
- Javeed, D.; Gao, T.; Khan, M.T.; Ahmad, I. A Hybrid Deep Learning-Driven SDN Enabled Mechanism for Secure Communication in Internet of Things (IoT). Sensors 2021, 21, 4884. [Google Scholar] [CrossRef]
- Marghalani, B.F.; Arif, M. Automatic classification of brain tumors and Alzheimer’s disease in MRI. Procedia Comput. Sci. 2019, 163, 78–84. [Google Scholar] [CrossRef]
- Swati, Z.N.K.; Zhao, Q.; Kabir, M.; Ali, F.; Ali, Z.; Ahmed, S.; Lu, J. Brain tumors classification for MR images using transfer learning and fine-tuning. Comput. Med. Imaging Graph. 2019, 75, 34–46. [Google Scholar] [CrossRef]
- Kumar, S.; Mankame, D.P. Optimization drove deep convolution neural network for brain tumors classification. Biocybern. Biomed. Eng. 2020, 40, 1190–1204. [Google Scholar] [CrossRef]
- Deepak, S.; Ameer, P.M. Brain tumors classification using in-depth CNN features via transfer learning. Comput. Biol. Med. 2019, 111, 103345. [Google Scholar] [CrossRef]
- Raja, P.S. Brain tumors classification using a hybrid deep autoencoder with Bayesian fuzzy clustering-based segmentation approach. Biocybern. Biomed. Eng. 2020, 40, 440–453. [Google Scholar] [CrossRef]
- Ramamurthy, D.; Mahesh, P.K. Whale Harris Hawks optimization-based deep learning classifier for brain tumors detection using MRI images. J. King Saud Univ. Comput. Inf. Sci. 2020, 32, 1–14. [Google Scholar] [CrossRef]
- Bahadure, N.B.; Ray, A.K.; Thethi, H.P. Image analysis for MRI-based brain tumors detection and feature extraction using biologically inspired BWT and SVM. Int. J. Biomed. Imaging 2017, 2017, 9749108. [Google Scholar] [CrossRef] [Green Version]
- Waghmare, V.K.; Kolekar, M.H. Brain tumors classification using deep learning. In The Internet of Things for Healthcare Technologies; Springer: Singapore, 2021; Volume 73, pp. 155–175. [Google Scholar]
- BT. Data Set. Available online: https://figshare.com/articles/dataset/brain_tumor_dataset/1512427 (accessed on 3 January 2022).
- Cheng, J.; Huang, W.; Cao, S.; Yang, R.; Yang, W.; Yun, Z.; Wang, Z.; Feng, Q. Enhanced performance of brain tumors classification via tumor region augmentation and partition. PLoS ONE 2015, 10, e0140381. [Google Scholar] [CrossRef]
- Resize Function. Available online: https://www.mathworks.com/products/matlab.html (accessed on 4 January 2022).
- Agarwal, M.; Gupta, S.; Biswas, K.K. A new conv2d model with modified relu activation function for identifying disease type and severity in cucumber plants. Sustain. Comput. Inform. Syst. 2021, 30, 100473. [Google Scholar]
- Botta, B.; Gattam, S.S.; Datta, A.K. Eggshell crack detection using deep convolutional neural networks. J. Food Eng. 2022, 315, 110798. [Google Scholar] [CrossRef]
- Thakur, S.; Kumar, A. X-ray and CT-scan-based automated detection and classification of COVID-19 using convolutional neural networks (CNN). Biomed. Signal Process. Control 2021, 69, 102920. [Google Scholar] [CrossRef]
- Liu, W.; Wen, Y.; Yu, Z.; Yang, M. Large-margin Softmax Loss for Convolutional Neural Networks. In Proceedings of the 33rd International Conference on Machine Learning, New York, NY, USA, 19 June 2016; pp. 1–7. [Google Scholar]
- Ruuska, S.; Hämäläinen, W.; Kajava, S.; Mughal, M.; Matilainen, P.; Mononen, J. Evaluation of the confusion matrix method invalidating an automated system for measuring feeding behavior of cattle. Behav. Process. 2018, 148, 56–62. [Google Scholar] [CrossRef] [Green Version]
- Anaraki, A.K.; Ayati, M.; Kazemi, F. Magnetic resonance imaging-based brain tumors grades classification and grading via convolutional neural networks and genetic algorithms. Biocybern. Biomed. Eng. 2019, 39, 63–74. [Google Scholar] [CrossRef]
- Abiwinanda, N.; Hanif, M.; Hesaputra, S.T.; Handayani, A.; Mengko, T.R. Brain tumors classification using convolutional neural network. In World Congress on Medical Physics and Biomedical Engineering; Springer: Singapore, 2018; pp. 183–189. [Google Scholar]
- Ullah, Z.; Farooq, M.U.; Lee, S.H.; An, D. A hybrid image enhancement-based brain MRI images classification technique. Med. Hypotheses 2020, 143, 109922. [Google Scholar] [CrossRef]
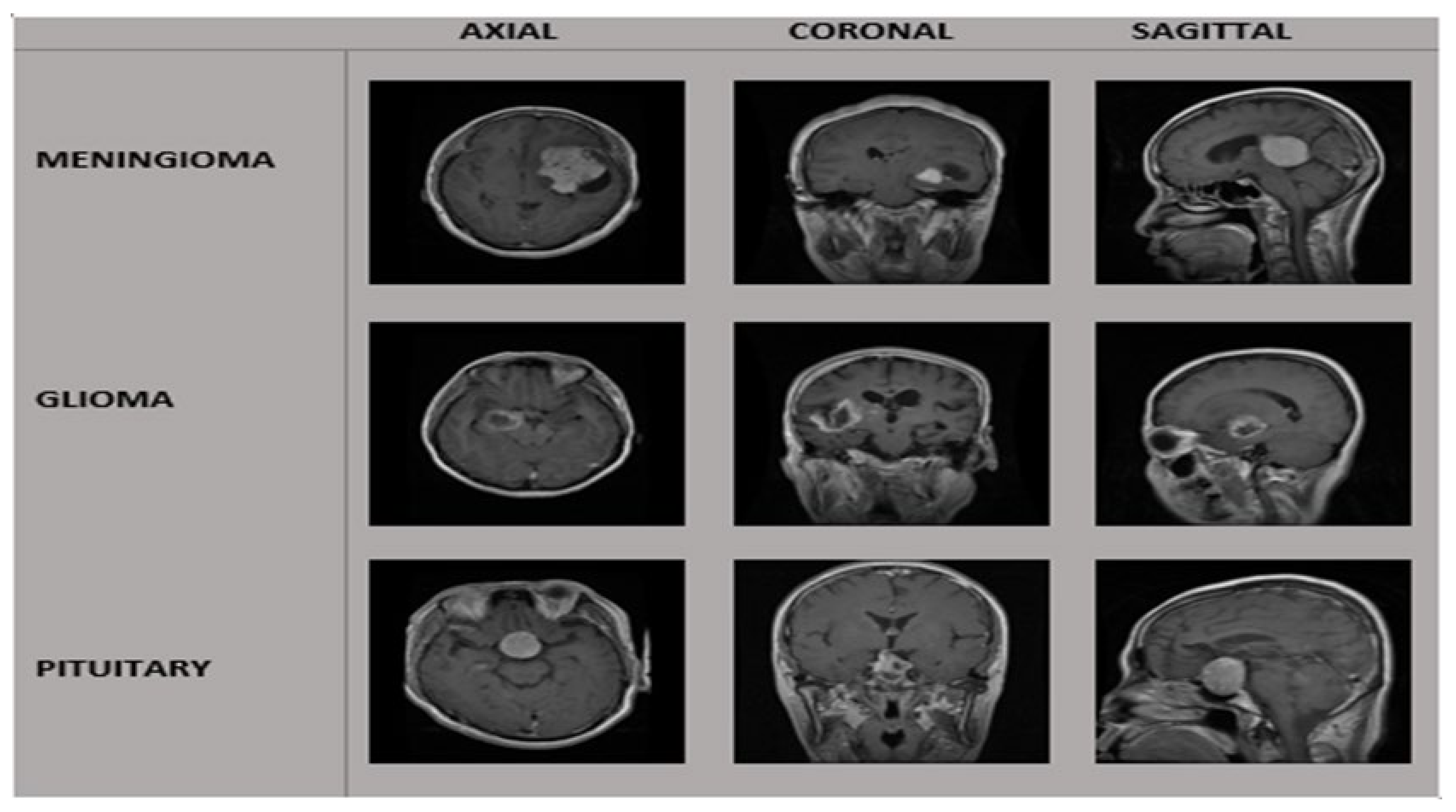

| Tumor Class | Patients | Images | View of MRI. | No. of MRI. Images |
|---|---|---|---|---|
| Meningioma | 82 | 708 | A * | 209 |
| C * | 268 | |||
| S * | 231 | |||
| Pituitary | 62 | 930 | A * | 291 |
| C * | 319 | |||
| S * | 320 | |||
| Glioma | 89 | 1426 | A * | 494 |
| C * | 437 | |||
| S * | 495 | |||
| Total | 233 | 3064 | 3064 |
| S.No | Layer Name | Type | No of Filter | Filter Size | Epsilon |
|---|---|---|---|---|---|
| 1 | block_16_expand | Conv | 960 | 1 × 1 | |
| 2 | block_16_expand_BN | Batch Norm | 0.001 | ||
| 3 | block_16_expand_relu | Clipped ReLU Layer | |||
| 4 | block_16_depthwise | Grouped Conv | 960 | 3 × 3 | |
| 5 | block_16_depthwise_BN | Batch Norm | 0.001 | ||
| 6 | block_16_depthwise_relu | Clipped ReLU Layer | 0.001 | ||
| 7 | block_16_project | Conv | 320 | 1 × 1 | |
| 8 | block_16_project_BN | Batch Norm | 0.001 | ||
| 9 | Conv_1 | Conv | 1280 | 1 × 1 | |
| 10 | Conv_1_bn | Batch Norm | 0.001 | ||
| 11 | out_relu | Clipped ReLU Layer | |||
| 12 | global_average_pooling2d_1 | Global Average Pooling | |||
| 13 | Logits | Fully Connected | |||
| 14 | Logits_softmax | Softmax | |||
| 15 | ClassificationLayer_Logits | Classification Layer |
| Name | SGDM |
|---|---|
| MiniBtachSize | 10 |
| Number of Epochs | 120 |
| Initial Learning Rate | 0.01 |
| Shuffle | every epoch |
| Validation Frequency | 50 |
| Predicated Classes | ||||
|---|---|---|---|---|
| Gliomas | Meningiomas | Pituitary | ||
| Actual Class | Gliomas | PGG | PMG | PPG |
| Meningiomas | PGM | PMM | PPM | |
| Pituitary | PGP | PMP | PPP | |
| Model | Accuracy | Precision | Recall | F1-Score |
|---|---|---|---|---|
| Proposed Model | 99.67% | 99.6% | 100% | 99.66% |
| AlexNet | 97.8% | 97.6% | 97.66% | 97.66% |
| GoogLeNet | 98.26% | 98% | 98.66% | 98.33% |
| Shufflenet | 98.37% | 98.33% | 98.66% | 98.33% |
| ResNet50 | 98.60% | 98.33% | 98.66% | 98.33% |
| MobileNet V2 | 99% | 99% | 99% | 99% |
| SqueezeNet | 97.91% | 97.66% | 98% | 97.66% |
| Darknet53 | 99.13% | 99% | 99.33% | 99% |
| Resnet101 | 98.91% | 98.66% | 99% | 98.66% |
| ExceptionNet | 98.69% | 98.33% | 98.33% | 98% |
| Predicated Classes | ||||
|---|---|---|---|---|
| Gliomas | Meningiomas | Pituitary | ||
| Actual Class | Gliomas | 426 | 01 | 01 |
| Meningiomas | 01 | 211 | 00 | |
| Pituitary | 00 | 00 | 278 | |
| Author | Technique | Classification Type | Dataset | Accuracy (%) |
|---|---|---|---|---|
| Mehrotra et al., (2020) [27] | PT-CNN: AlexNet | Binary Class | T1-weighed MRI (Benign = 224; malignant = 472) | 99.04% |
| Kaplan et al., (2020) [35] | LBP SVM KNN | Multi-Class | T1-weighed CE-MRI (meningiomas = 708; gliomas = 1426; pituitary = 930) | 95.56% |
| Sultan et al., (2019) [30] | CNN | Multi-Class | T1-weighed CE-MRI (meningiomas = 208; gliomas = 492; pituitary = 289) | 96% |
| Anaraki et al., (2019) [63] | GA-CNN | Multi-Class | T1-weighed CE-MRI (meningiomas = 708; gliomas = 1426; pituitary = 930) | 94.20% |
| Kumar et al., (2017) [31] | GWO+M-SVM | Multi-Class | T1-weighed CE-MRI (meningiomas = 248; gliomas = 12; pituitary = 55) | 95.23% |
| Bahadur et al., (2017) [64] | BWT+SVM | Binary | T2-weighted images (normal = 67; abnormal = 134) | 95% |
| Abiwinanda et al., (2019) | CNN | Multi-Class | T1-weighed CE-MRI (meningiomas = 708, gliomas = 1426; pituitary = 930) | 84% |
| Proposed method | Deep CNN | Multi-Class | T1-weighed CE-MRI (meningiomas = 708; gliomas = 1426; pituitary = 930) | 99.67% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raza, A.; Ayub, H.; Khan, J.A.; Ahmad, I.; S. Salama, A.; Daradkeh, Y.I.; Javeed, D.; Ur Rehman, A.; Hamam, H. A Hybrid Deep Learning-Based Approach for Brain Tumor Classification. Electronics 2022, 11, 1146. https://doi.org/10.3390/electronics11071146
Raza A, Ayub H, Khan JA, Ahmad I, S. Salama A, Daradkeh YI, Javeed D, Ur Rehman A, Hamam H. A Hybrid Deep Learning-Based Approach for Brain Tumor Classification. Electronics. 2022; 11(7):1146. https://doi.org/10.3390/electronics11071146
Chicago/Turabian StyleRaza, Asaf, Huma Ayub, Javed Ali Khan, Ijaz Ahmad, Ahmed S. Salama, Yousef Ibrahim Daradkeh, Danish Javeed, Ateeq Ur Rehman, and Habib Hamam. 2022. "A Hybrid Deep Learning-Based Approach for Brain Tumor Classification" Electronics 11, no. 7: 1146. https://doi.org/10.3390/electronics11071146
APA StyleRaza, A., Ayub, H., Khan, J. A., Ahmad, I., S. Salama, A., Daradkeh, Y. I., Javeed, D., Ur Rehman, A., & Hamam, H. (2022). A Hybrid Deep Learning-Based Approach for Brain Tumor Classification. Electronics, 11(7), 1146. https://doi.org/10.3390/electronics11071146









