1. Introduction
Microscopy plays a crucial role in the evaluation and diagnosis required by many medical disciplines, including microbiology, parasitology, or pathology. Even though other types of microscope, e.g., transmission electron microscope and fluorescent microscope, are widely available in the market, brightfield microscopy remains the gold standard for image analysis and the diagnosis of many medical specialities.
The technology used in optical microscopes has remained the same for the last decades. Most pathologists and microbiologists continue to use the same technology they were using fifty years ago and the workflows and protocols to follow are almost the same. The main difference is that the population has increased, and therefore the number of samples has increased along with the complexity of the diagnostic criteria. A serious limitation in low- and middle-income countries is the number of specialists, and the time they have to perform in-situ diagnosis is limited [
1].
The increasing need for trained microscopists, their increasing workload, and the accessibility and affordability of the technology can represent insurmountable constraints to deliver high quality and rapid healthcare attention in resource-constrained settings. However, new low-cost portable digital microscopes that leverage ever-improving quality and affordability of manufacturing processes; communications, mobile phones, and sensors, may be the way forward to point of care (PoC) diagnosis in remote locations, avoiding misleading results and delays. Thus, systems that could digitize microscope slides in a rapid, simple, and cheap manner could help bypass the dearth of microscopes, and more importantly, the scarcity of trained microscopists, as well as allow for the more specialized or even automated (artificial intelligence based) evaluation of the slides requiring microscopic evaluation.
It was seventy years ago when the concept of portable microscopy was firstly conceived by the hand of McArthur [
2], guiding the portable microscopy industry path for several decades with many iterations of portable microscopes commercialized [
2,
3]. With this approach, remote areas where high-risk patients reside could be reached, but there was still the extreme need for highly trained field microscopists. In 1985, the first digital microscope was invented by Hirox Co Ltd., a Japanese lens company [
4]. In the same decade, with the advances of moldable plastic technology and computer design in the late 1980s, low-cost portable microscopes were conceived [
5]. It was not until the end of the first decade of the twenty-first century when an explosion within this field occurred. With the collective technological advances of LEDs [
6], smartphones, sensors, and communications, together with the expiration in 2009 of the FDM 3D printing technique patent and the RepRap Project [
7], a new concept of portable, low-cost, mobile-based digital microscopes was born [
8,
9,
10,
11].
The penetration rate of the smartphone’s market is expected to cover approximately 80% of the world’s population by 2023 [
12]. In addition, smartphone technology is advancing by leaps and bounds, incorporating new improvements in terms of sensors, software, electronics, and connectivity, thereby growing the computing power exponentially and reducing its cost every year. Thus, smartphones used together with networking and cloud SaaS (software as a service) solutions open the door to the possibility for effective and rapid digitalization, allowing for not only remote diagnosis, but also slide archives, quality control, opinion sharing, and importantly, training purposes.
As a result of this widespread use of smartphones and their capabilities, mobile health diagnostics solutions are becoming a game-changing technology that could offer primary attention in low resource settings delivering PoC diagnostics, therefore decentralizing, democratizing, and digitizing medical diagnostic care. Moreover, the use of smartphones combined with 3D printing technology allows for extending the smartphone capabilities to additional digitization applications.
Several studies proposed low-cost microscopes built with 3D printing and mobile technology for low-resource areas [
5,
10,
13,
14,
15,
16,
17]. For instance, Switz et al. presented a mobile phone microscope that achieved a resolution of smaller than 5 µm and a field of view of 10 mm
2. They demonstrated its use in the identification of blood cells in blood smears and soil-transmitted helminth eggs in stool samples [
10]. Skandarajah evaluated CellScope scanner, which uses a 20× objective lens, for oral cancer screening [
16]. Vasiman et al. reviewed existing mobile phone and handheld microscopes and their application in malaria and neglected tropical diseases, finding that some of them have high sensitivity and specificity compared with conventional microscopes [
5]. Even so, these solutions yield a set of images from different single fields from the sample, which in some cases are not enough to render a proper examination of the sample. As an example, pathologists have to see the whole slide to determine if the patient is healthy or the slide contains some malignant cells. In order to address this problem, whole-slide scanners are used.
Currently, commercial slide scanning technology that uses precise sub-micron motor stages to capture image tiles at a magnification and stitch them together entails a cost in the order of tens of thousands dollars [
18,
19,
20]. An alternative is to use more affordable hardware with less precise position information and more sophisticated computer vision methods to stitch the slide together [
21,
22,
23]. Li et al. presented Octopi, a scanner that is capable of automated slide scanning and multi-modal imaging. Instead of using nanometer precision motors, they used low-cost lead screw linear actuators. The device could be used for malaria and tuberculosis diagnostics and digital pathology [
24]. Similarly, Collins et al. presented OpenFlexure, a 3D printed microscope scanner using low-cost stepper motors [
11]. Nevertheless, implementation of those systems remains a difficult prospect for many institutions, especially those with stakeholders unfamiliar with these technologies.
Based on the previous considerations, this paper presents an affordable, 3D-printed, portable, robotic, mobile-based microscope scanner with an interactive augmented reality environment, connected to a telemicroscopy platform. We present the use of the system in different clinical applications, such as pathology and infectious diseases. We first (1) present all components (mechanics, optics, and mobile and cloud features) of the proposed robotic microscope; (2) summarize the performance assessment of the device and its application in two clinical scenarios; and (3) discuss its utility and possible future work lines derived from its use.
2. Robotic Mobile-Based Microscope Scanner
2.1. System Architecture
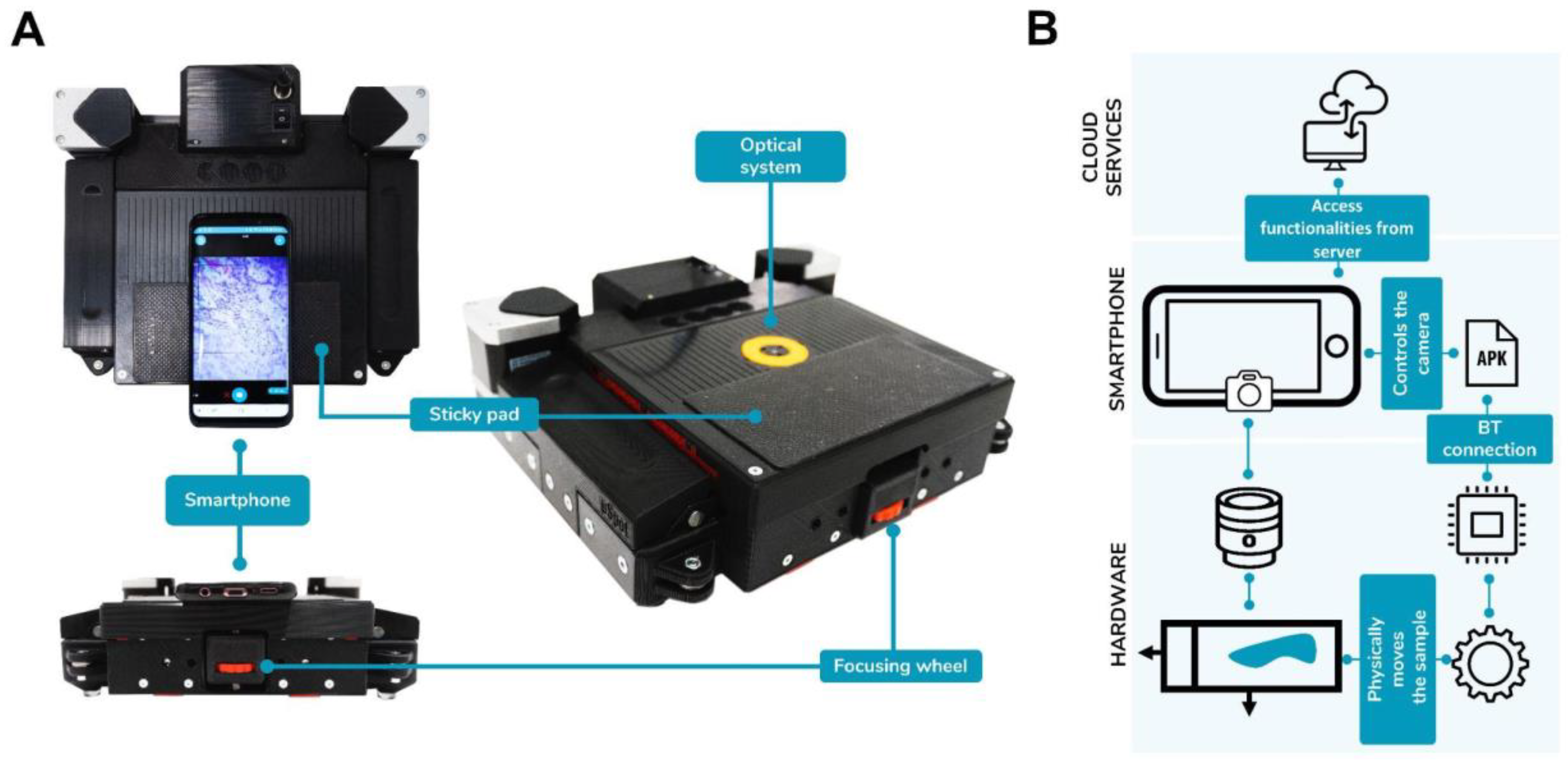
The system (
Figure 1) is the result of the expansion of the capabilities of a smartphone, by leveraging the infrastructures that the device itself supplies. Namely, the camera for image capture, the Bluetooth module for communicating nearby devices in an abstract manner, and the WiFi and broadband cellular network modem for connectivity with remote systems via the internet.
The smartphone is used in combination with an electromechanical device that implements the physical features needed in a microscopy scanner. Such electromechanical device is comprised of a mechanical cartesian XY stage powered by stepper motors that mounts a carriage capable of holding standard 78 × 25 mm2 sample glasses. An electronics module comprising a power stage equipped with stepper drives, an ATMega4809 Microcontroller, and an ESP32 based SoC; and a miniaturized optic system designed to provide a usable digital image of a microscopy sample when coupled with the camera optics.
Higher level functionalities such as user interface, machine routines, asset organization, and client-side interaction with the servers are implemented on the App. Servers provide the infrastructure for tele-medicine features as well as any computationally heavy task.
2.2. Mechanical Structure
The mechanical stage of the proposed device has been developed with aim to produce a compact, cost-effective design, enabled by the production of 3D printed parts. This manufacturing method has been selected because, compared to other methods such as moulding, it is very flexible. It allows for rapid iteration on the designing phase, and on demand and distributed production, which is optimal to reduce CO2 footprint due to transportation.
Regarding compactness, the target size has been selected in such a way that the device can fit in a regular size backpack. A maximum boundary box of 250 × 250 × 150 mm3 was taken into consideration, for the design not to exceed in any of these dimensions. The design targets a low production cost that could only be achieved by minimizing the customization of parts, i.e., composing the assembled product with the most customer off-the-shelf parts as possible; preferably standard.
The custom needs of the design have been addressed by features in 3D printed parts which become the nodes that connect the hardware commercial off-the-shelf parts of the design and provide the special features that the requisites mandate. The whole design has been optimized for this manufacturing technique and organized in functionally distinct modules that, while not completely interchangeable nor easily disassembled by the user, allowed for decoupled development through iterations with different versions during development. The machine used for 3D printing was a Raise3D Pro 3D printer and the filaments used were Raise 3D Premium PLA Filament for all the parts except for the ring that encloses the optical system, which was 3D printed with Recreus Filaflex.
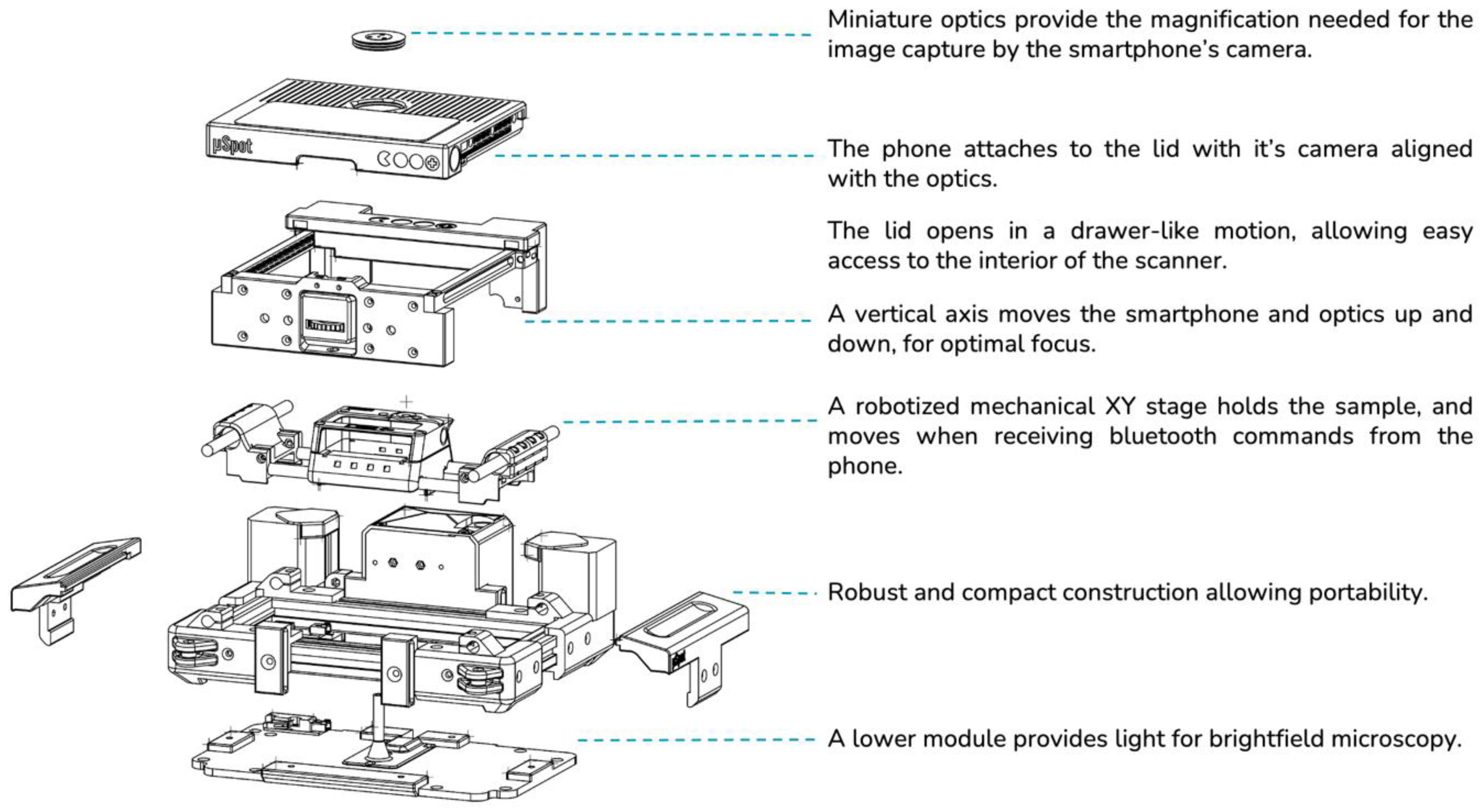
Every mechanical component (see
Figure 2) is mounted onto the Main Frame, directly or indirectly, effectively making it a chassis. The frame structural composition stems from the union of two pairs of aluminium profiles forming a rectangle. Instead of using off-the-shelf corner brackets or gussets, 3D printed corner parts have been used, which provide support for other modules, or actively participate in the other modules’ functionalities, such as mounting the stepper motors, the
Y-axis rods, and holding pulleys necessary for transmission in place. With the objective of minimizing the moving parts in the design, the actuation of such axes is laid out in a H-bot configuration. This allows for having the two stepper motors fixed to the chassis, reducing the number of moving parts.
In order to achieve focusing capabilities, a third vertical axis is needed in order to place the sample at the proper distance from the optics. While traditional microscopes move the sample stage towards the optics, our device moves the optics, along with the phone, closer to the sample by descending onto it. The vertical axis consists of a sliding platform anchored to the main rail, with vertical 3D printed linear rails to allow for vertical movement. Because friction between 3D printed parts is undesirable, small plastic spheres are used as rolling elements for these vertical rails. In order to actuate the linear motion, the front panel has a short spindle attached, whose nut is accessible from the front. The nut can be spun by the user using their thumb, causing a vertical movement on the spindle.
Mounted onto the cartesian stage’s carriage is a magnetically attached tray designed to hold the sample glass. The tray has flat fixings akin to the ones present in a classic microscope stage, designed to not stand above the glass, avoiding collisions with the optics.
The mechanical stage is enclosed on its top side by an upper lid, that additionally hosts the optics module and provides means of holding the smartphone in place while device operation takes place. The holding means being a sticky pad, commercially used to attach smartphones onto car dashboards. The lid is mounted onto sliding rails using the same 3D printed rails used on the vertical axis, allowing to uncover the mechanical stage by sliding the lid to the front out of the way, allowing to place the sample onto the scan tray.
The optics have been designed with miniaturization as a priority. In order to reduce the optical path in length as much as possible, miniature lenses have been used, resulting in a “disk-like” design. The optics mount has an exterior ring, 3D printed in flexible material, designed to fit through a hole in the upper lid. The flexible coupling makes it easy to replace the optics in case different magnifications are needed, and it also provides a damping mechanism in the event of the optics crashing against the sample upon focusing, as many commercially available microscope objectives have.
Electronic Layout and Firmware
The device’s electromechanical architecture shares the responsibility of operations between two microcontrollers: a mechanical microcontroller widely used on homemade CNC machining, and a Bluetooth enabled coordination microcontroller running upper-level Bluetooth communication firmware, responsible for communicating with the smartphone app and making upper level decisions based on the state of the machine. Additionally, means for actuating the axes are put in place, as well as illumination functionalities. Each axis is actuated by a stepper motor, equipped with a stepper driver, and an endstop. The endstops and the stepper drivers are connected to the mechanical microcontroller, the steppers being controlled via STEP/DIR signal pairs. A PWM line connects the mechanical microcontroller to an LED driver, which connects to a 3 W white light LED diode. The coordination microcontroller communicates the mechanical microcontroller via RX/TX serial lines, at 115,200 baud. The coordination microcontroller receives commands from and sends responses to the smartphone via bluetooth, and if concluded by some basic decision making, sends low-level movement commands to the mechanical microcontroller via the serial connection. The mechanical microcontroller reports back low-level status, e.g., pin states and axis position.
The mechanical microcontroller is an ATMega4809 running a CNC router firmware. The firmware implements a GCODE command interpreter, with step signal control for stepper drivers and endstop detection, governed by the states of a state machine. The firmware interprets GCODE commands received from the serial connection and coordinates the step signals to perform the requested movement. The state machine provides a readiness level that establishes a conservative approach to movement, to minimize the risk of mechanical damage. (e.g., no movement is performed unless the position of the stepper motors is known by the firmware via homing).
The coordination microcontroller is an ESP32 processor encapsulated within a NiNaW102 module, which provides antennas for WiFi and Bluetooth capabilities. The coordination microcontroller runs a firmware that connects to the mechanical microcontroller via serial connection and tracks its state. It establishes a Bluetooth connection with the app, acting as intermediary between the mechanical stage and the smartphone. Additionally, it provides construction information upon request, such as maximum and minimum coordinates, the characteristics of the lenses available in the device, as well as the illumination options. This information is later used to present magnification information to the user, as well as correct positioning.
Despite being carried over Bluetooth, communications with the smartphone are encoded with a REST API format for the requests, and a JSON format for the responses, to take advantage of the multiple smartphone app libraries available for parsing JSON-formatted strings.
The motors in charge of making the movement effective are bipolar stepper motors. The chosen motors are of the NEMA 17 family, which are over dimensioned in load but that allows for a steady, cold operation. Moreover, material costs are driven down as these motors have been widely used in 3D printing for the last decade, which resulted in more availability for sourcing. The motors are driven by Pololu’s A4988 stepper drivers, powered at 12 V, and with their STEP/DIR pin pairs connected to the mechanical microcontroller’s GPIOs. The motors are paired with 3D printed custom-made pulleys, and driven on 1/16 microstep configuration they yield a mathematical XY resolution of 5.5 µm.
2.3. Optics
In order to make the image taken by the smartphone uniform among smartphones, the optics have been designed to be infinity-corrected. That way, as we will discuss later, it is possible to estimate the total magnification of the image projected onto the sensor, when the focal length of the camera lens is known. In order to reduce component costs, the optics consist of two customer off-the-shelf lenses, chosen from commercial catalogs (e.g., Edmund Optics and Thorlabs) and evaluated via simulation to provide the best compromise between field of view size and resolution on the center.
As a reference for simulations, the camera of a Samsung S9 was considered, which reportedly provides a camera with a 4.3 mm focal length lens, and a 4032 × 3024 pixel sensor, measuring 5648 × 4236 mm2 in physical size. Simulations showed a total magnification of 2× (understanding this magnification as the relation of sizes between the object and the image projected onto the sensor), a numerical aperture of 0.13, and a working distance of 0.92 mm. Upon comparing these results with the ones obtained in a standard microscope, the resolution of images offered by this lens would be equivalent to a 10× microscope objective. Digital zoom is calculated and applied so that the field of view of digitized images is equivalent to the one obtained by a microscope objective of 40× magnification, with an eyepiece of field number 20, for a horizontal FOV of 0.5 mm.
In order to make brightfield microscopy effective, the sample is backlighted by the aforementioned 3 W LED. The LED is enclosed in the lower lid of the main frame, and a tube directs the light up to the very bottom of the sample glass. In order to increase the numerical aperture of the lighting and prevent lens flare, a light-diffusing sheet is glued to the tip of the tube.
2.4. Mobile Control App Features
The frontend of the device is presented to the user through a smartphone application, which provides the user interface and provides the upper-level automated operation routines, such as automatic scanning. The minimum requirement for the smartphone is that the operating system is over Android 6 and the camera pixel resolution is greater than 1440 × 1080, after the digital zoom is applied. In cases of the resolution of the camera being lower than this requirement, the fidelity of the image will be affected by pixel interpolation algorithms.
2.4.1. Augmented Reality Environment on Specimen
The application is designed to orient the user throughout the exploration of the sample, through an augmented reality (AR) environment projected onto the display of the sample. For that, it keeps track of the position in which the carriage is placed, as well as the area of the sample that is visible on the screen at any given time.
Since API 21, Android has provided reports on a series of physical properties of the camera modules, such as those described above. Those properties, along with the focal length of the additional lens that the device provides, and the digital zoom selected by the use, can be used to calculate an estimate of the area that is visible on the phone screen. Moreover, the application monitors the position of the carriage within the device, and therefore the position of the center of the visible field of the sample is known at all times.
That information is used to provide the user with a layout containing useful information that is understandable and meaningful to the professionals in the field, namely a virtual grid displayed on the screen, on top of the sample display. The size of the cells of such a grid are calculated using the data described above, in such a way that each cell encloses an area of the displayed sample that would correspond to a 0.1 × 0.1 mm2 area onto the real sample. Furthermore, the grid cells are numbered in such a way that the user is always oriented in relation to their position on the sample, as well as the actual size of the displayed objects.
This layout element, being responsive to the area of the sample displayed on the screen, effectively provides an AR environment that enhances the exploration of the sample, providing metrics that enhance the orientation of the user upon exploring the sample.
2.4.2. Mechanical Control
The application constitutes the upper command level of the device as a whole. That includes control over the mechanics as well as the illumination. By conception, the device attempts to provide a familiar and user-friendly interface and operation to the user, and the mechanical control is designed according to such a concept.
The application effectively transforms the three well-known gestures (panning, flinging, and pinching) into movements of the microscope, as well as zooming onto the sample. In the case of panning and flinging, the movement of the finger across the screen is translated into micrometric movements of the carriage, that correspond to the movement of the finger scaled for magnification, in such a way that the finger is always pointing to the same zone on the sample. Pinching is used for zoom, translating the movement of the fingers into digital zoom, adjusting the magnification at the user’s will.
The aforementioned virtual grid is updated in real time upon user interaction, in such a way that it always corresponds to the area of the sample that is displayed on the screen.
In this way, the application brings the smartphone control over the mechanics of the device, as well as collecting imaging information from the examined sample, via the camera feed. This effectively creates a closed control loop that can be used to implement fully automated routines such as a scan procedure, in which an area of the sample is swept across, step by step, making evenly spaced photos distributed in a grid. The distance between each photo’s coordinates is calculated using the sample area visible to the camera, in such a way that an overlap exists between the photos, allowing for a later stitching into a big photo map of the whole area.
The device can execute this task unattended, which allows for non-specialized operators to perform the scanning process. The scan photos can be uploaded onto a cloud server for specialized personnel to analyze.
2.5. Remote Telemedicine Platform
All acquired images using the mobile app are transferred via the mobile network to an Amazon Simple Storage Service (Amazon S3, Seattle, WA, USA) bucket, which is encrypted with the industry-standard AES-256 encryption algorithm. In addition, data security in transit is ensured through enforcing HTTPS (Secure HTTPS) through TLS (Transport Layer Security).
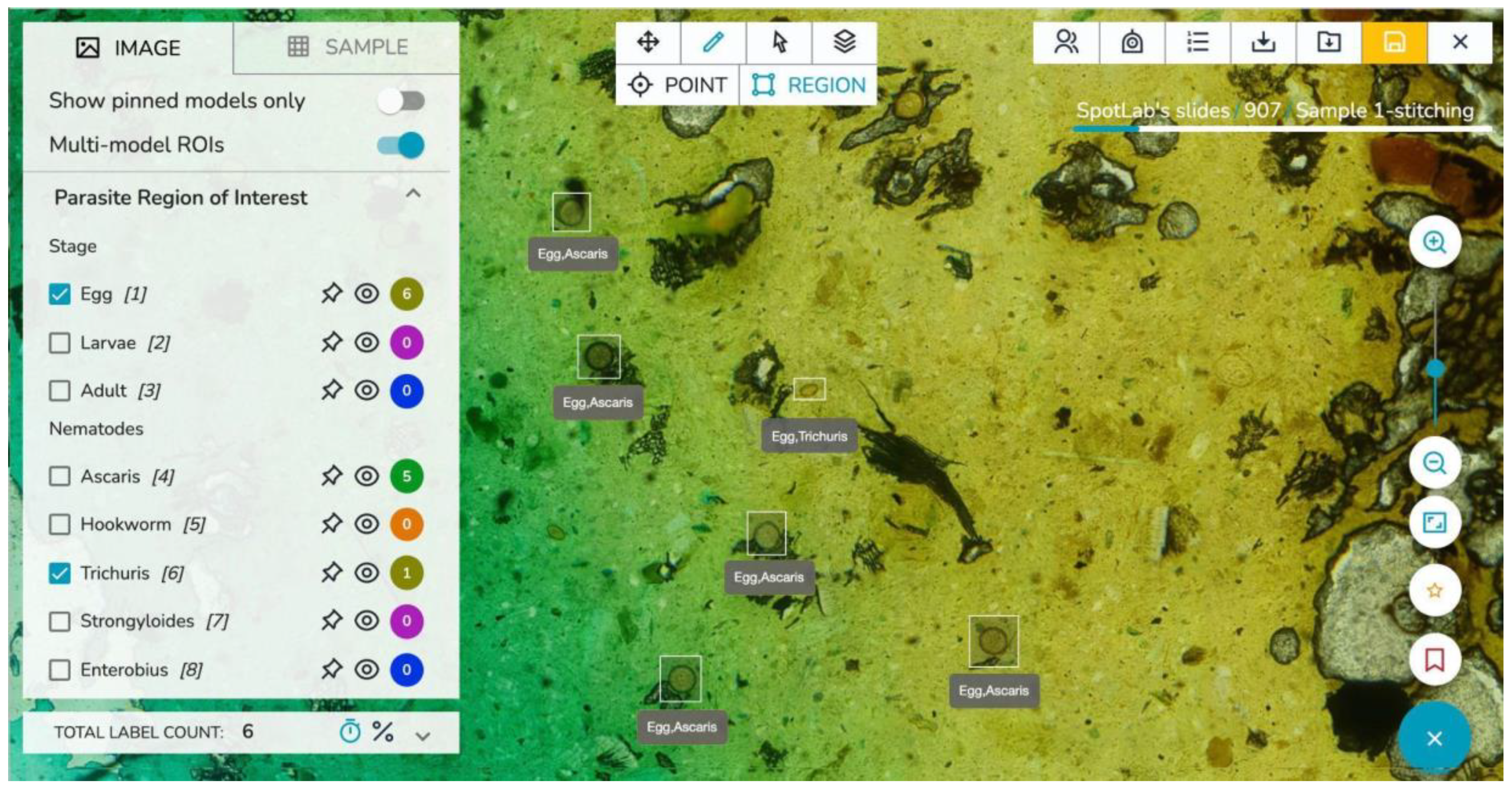
The images can be visualized through a remote telemedicine platform entirely developed within SpotLab, where images are presented in an easy-to-use dashboard that allows their management, sharing, analysis and reporting [
25]. The remote platform, which is accessible from all desktops, laptops, tablets, and mobile phones, translates standard clinical diagnostic protocols into digital tasks which are adapted to each case under study.
Figure 3 illustrates a screenshot of the telemedicine platform.
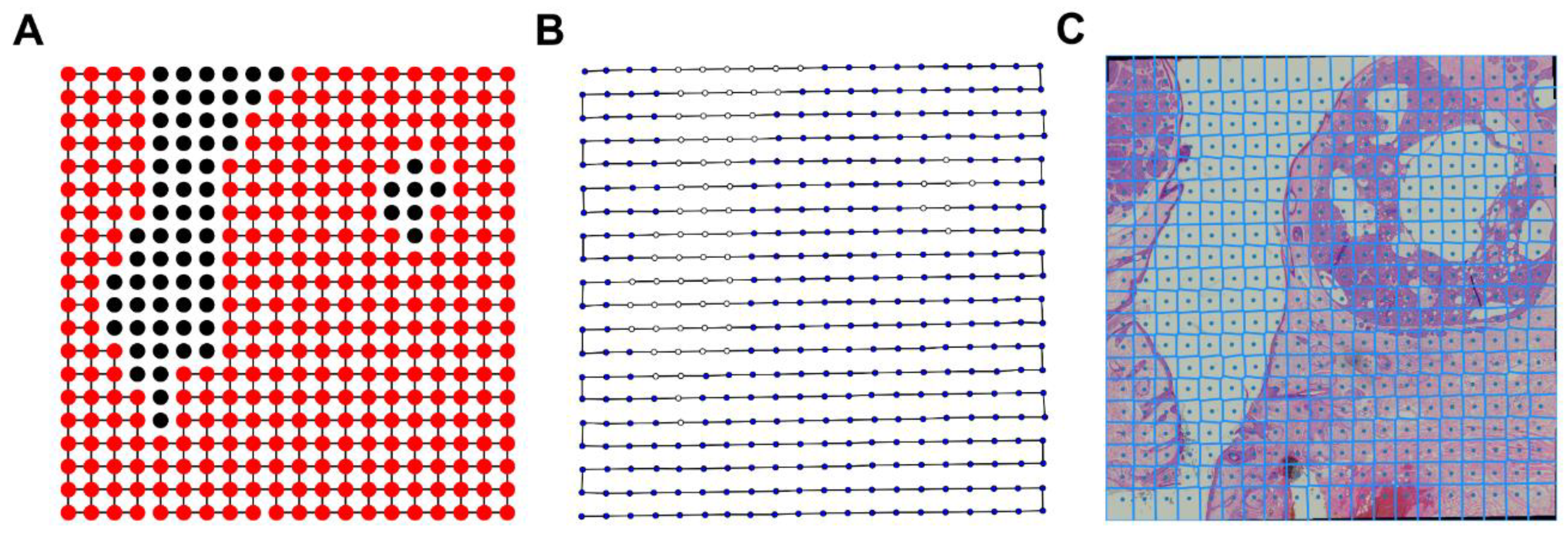
As previously mentioned, a scan routine is implemented to acquire the necessary captures to produce whole slide images so that once all individual images are uploaded, an image processing pipeline is automatically triggered to stitch and generate the final whole slide image. This processing pipeline includes several steps such as vignetting correction without reference or background image, integrated distortion correction, pairwise registration between adjacent scan images and fusion using Voronoi cells and Laplacian blending. A detailed description of the stitching algorithm as well as its performance are presented elsewhere [
26]. This stitching algorithm is deployed in Amazon Web Services using a docker container. Resulting stitched images are also uploaded to the Amazon S3 container and can be visualized through the telemedicine platform.
3. Performance Assessment
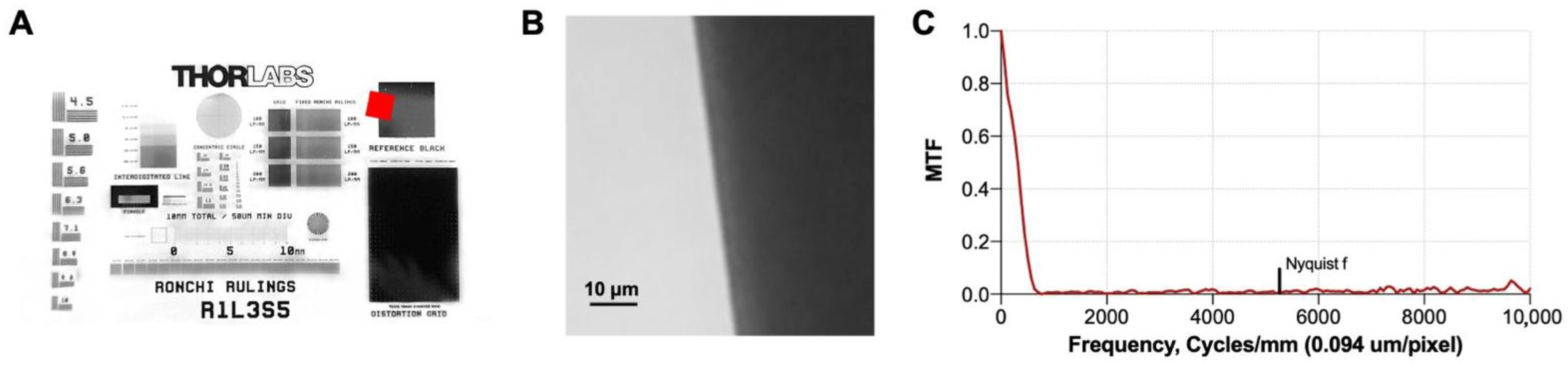
In order to evaluate the performance of the optical system, the slanted edge test [
27] was used to determine the optical resolution of the optical system. We imaged a slanted edge from a R1L3S5P resolution target (Thorlabs, Inc., Newton, NJ, USA) (
Figure 4A,B), computed the edge spread function, derived it, and obtained the modulation transfer function (MTF) (
Figure 4C). The MTF describes the ability of an optical system to transfer the object contrast to the acquired image as a function of frequency and can be used to determine the system resolution.
Table 1 reports the spatial resolution of the system quantified as the spatial frequency at different levels of the MTF. Note that spatial frequencies between 10% and 20% MTF (MTF10, MTF20) correspond to the classic vanishing resolution, while MTF50 ensures resolution where objects can be imaged with sharpness and detail. As derived from
Table 1, the proposed system is able to distinguish objects as small as approximately 1 micrometer (µm).
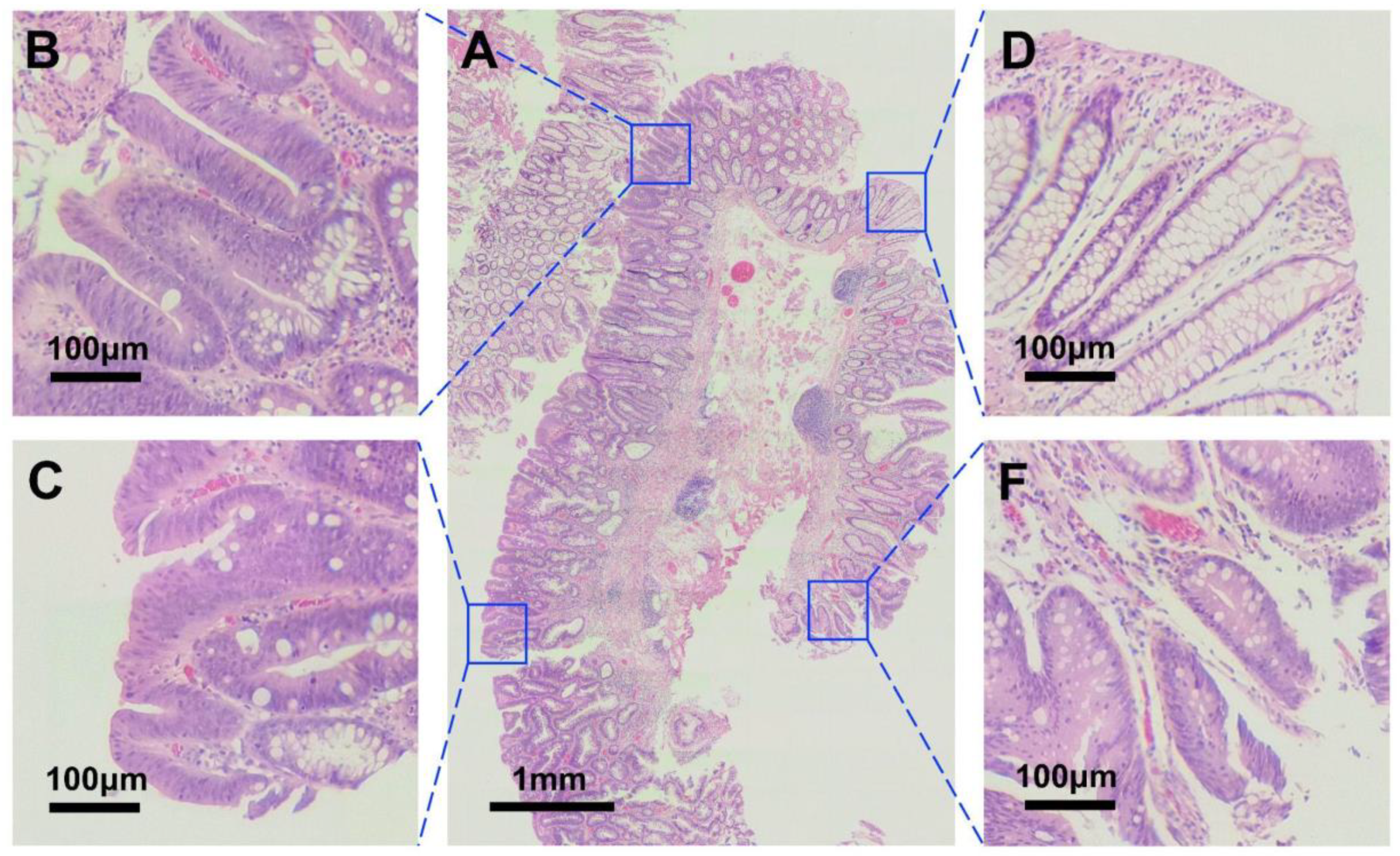
To compare the quality of the images obtained through our proposed robotic microscope to the one of the images obtained with a conventional microscope, we have digitized the same sample with both the proposed device and a standard microscope (Leica DM-200, 40× objective, field number 22). The image corresponding to the conventional microscope was obtained by coupling a smartphone to the microscope’s eyepiece. Both images were captured with the same device (Samsung S9). As shown in
Figure 5, the quality of the images is similar and all objects of interest that can be visualized using the conventional microscope can also be correctly identified using our proposed device.
Regarding the performance of the scan procedure and the image processing pipeline, it should be noted that a typical 20 × 20 grid scan at a magnification of 40× that covers a field of view of 7.2 × 7.2 mm
2 can be obtained in under 5 min.
Figure 6 shows an example of a stitched scan composed by 20 × 20 independent acquisitions.
The uploading time for such a scan is about 7 min on a 2 Mb/s connection, and the stitching process is performed in the cloud within 8 min. For a field of view of 7.2 × 7.2 mm
2, an image of approximately 15,000 × 15,000 pixels is created, with a resulting pixel size of 0.48 µm/mm and a size of around 150 MB. These specifications fall within the typical range of commercial scanners [
20].
5. Discussion and Conclusions
At the present time, there is an urgent need, especially in low resource settings, for quality and low-cost point of care microscopy scanner diagnosis devices. In this work, we have presented an affordable, 3D-printed, portable, robotic, mobile-based microscope scanner, fully integrated with a telemedicine platform. It presents a combination of a simple 3D printed automatic stage with the increased computing and image-capturing power of smartphones, which allows for a completely automated, cost-effective, and mobile-based solution to microscope sample scanning.
The device can be built in non-specialized manufacturing environments on demand and made of less than 1.5 kg of 3D printed plastic parts, common hardware parts (such as normalized bolts and nuts, aluminum profiles, and smooth rods), off-the-shelf optics, and widely adopted electromechanic components, making it at least as affordable as other entry level scanners. The resulting weight of the device is around 3 kg.
Its compact design allows for physical portability, while its conceptualization around the smartphone hardware allows for connectivity, portability, and ease of use. Both the phone and device are combined in a compact package that is convenient for its operation in remote areas, by non-specialized personnel. Additionally, the proposed robotic microscope scanner is also connected to a telemedicine platform enabling remote diagnosis and analysis of digitized samples.
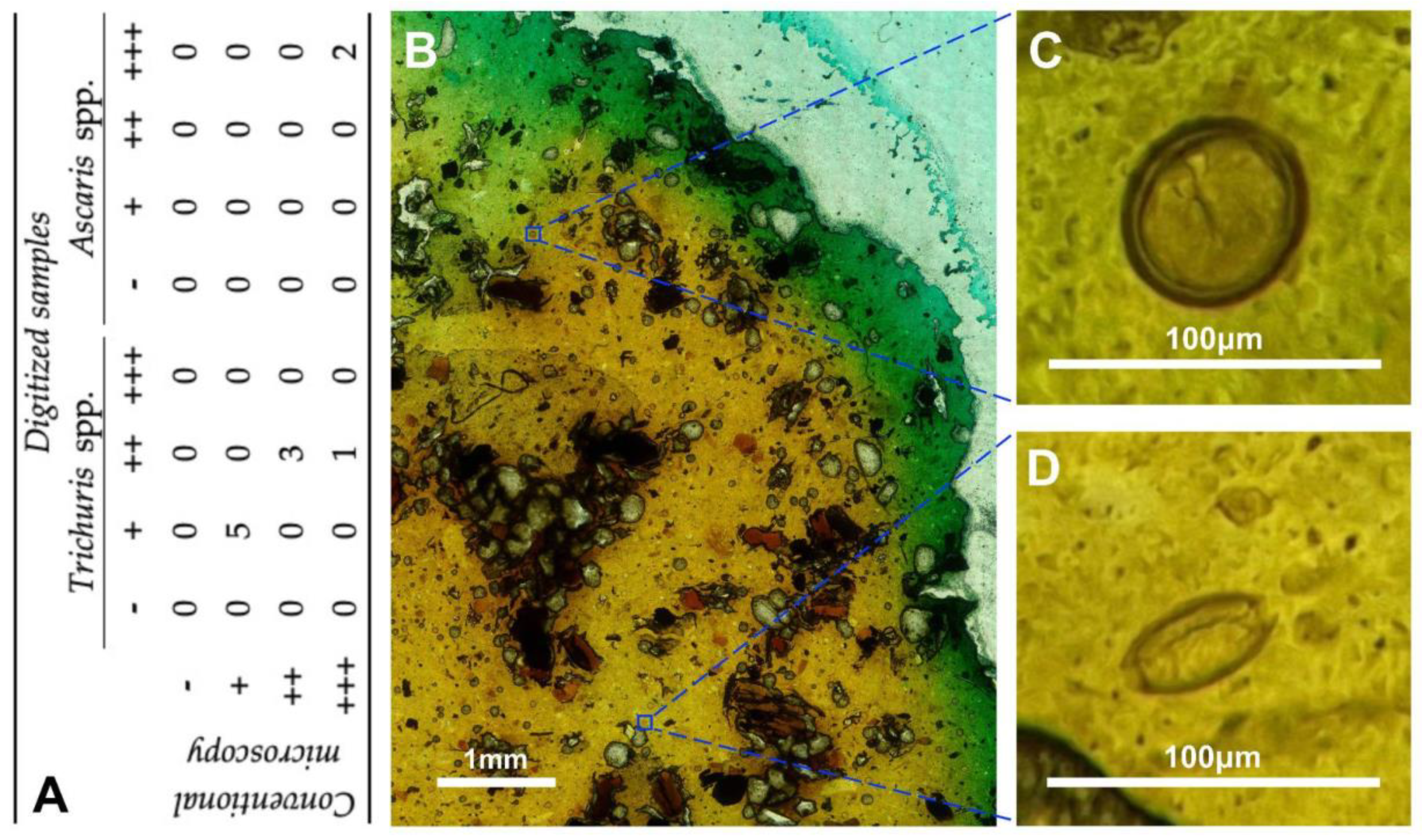
The proposed device has been assessed in two clinical scenarios, including histopathology (different tissue pathologies) and infectious diseases (soil transmitted helminthiasis). The results support its adequate usability and sufficient performance quality to allow diagnosis. Therefore, the quality of the digitized microscopy images generated by the proposed robotic microscope appear suitable for distinguishing and identifying all structures of interest for performing a proper diagnosis.
Future lines of work may benefit from the fact that the smartphone is responsible for capturing and ultimately storing the images as the media files used for later analysis, e.g., enabling the use of edge-detecting image algorithms. The use of AI algorithms to digest the camera feed allows for different in-situ automated and semi-automated routines, such as locking on to parasites or features detected by the AI algorithm when exploring the sample, or auto-searching specific features in a sample, making photos of areas of interest in a completely automated way. Two more example routines that could improve the user experience through the information received via the camera feed can be implemented, namely (1) a further enhancement of the augmented reality environment where features of the sample detected by artificial intelligence algorithms can be highlighted, and (2) an autofocus routine, with a future version of the device that provides a motorized Z-axis, to be operated according to focus-detecting measurements obtained from the camera feed. It should be noted that AI algorithms may also be integrated in the telemedicine platform for the automatic analysis of uploaded images, thus assisting experts in the diagnosis of different pathologies. Additionally, future lines of work will consider the validation of the proposed robotic microscope for the digitalization of other microscopic images and diagnosis of different pathologies. An cost analysis of the proposed device and the potential impact in low-resource settings will also be considered.
The development of cost-effective solutions affordable for physicians is key in the democratization of diagnosis and for the general improvement of the quality of care offered. The proposed robotic microscope digitization device aims to provide access to democratized and decentralized diagnosis, contributing to the achievement of universal health care.