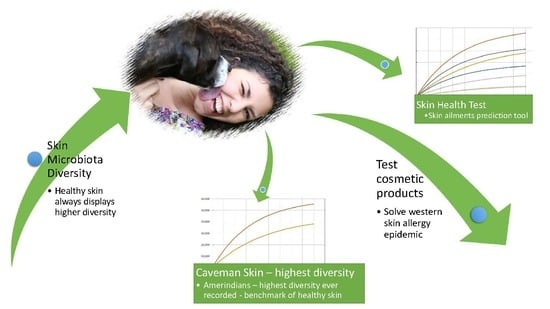
Meta Analysis of Skin Microbiome: New Link between Skin Microbiota Diversity and Skin Health with Proposal to Use This as a Future Mechanism to Determine Whether Cosmetic Products Damage the Skin
Abstract
:1. Skin Microbiota
1.1. Skin Immune System
1.2. The Rise of Skin Problems in the Developed World
1.3. Skin Microbiota and Possible Links to Ill Health
1.3.1. The Importance of (Bio)diversity
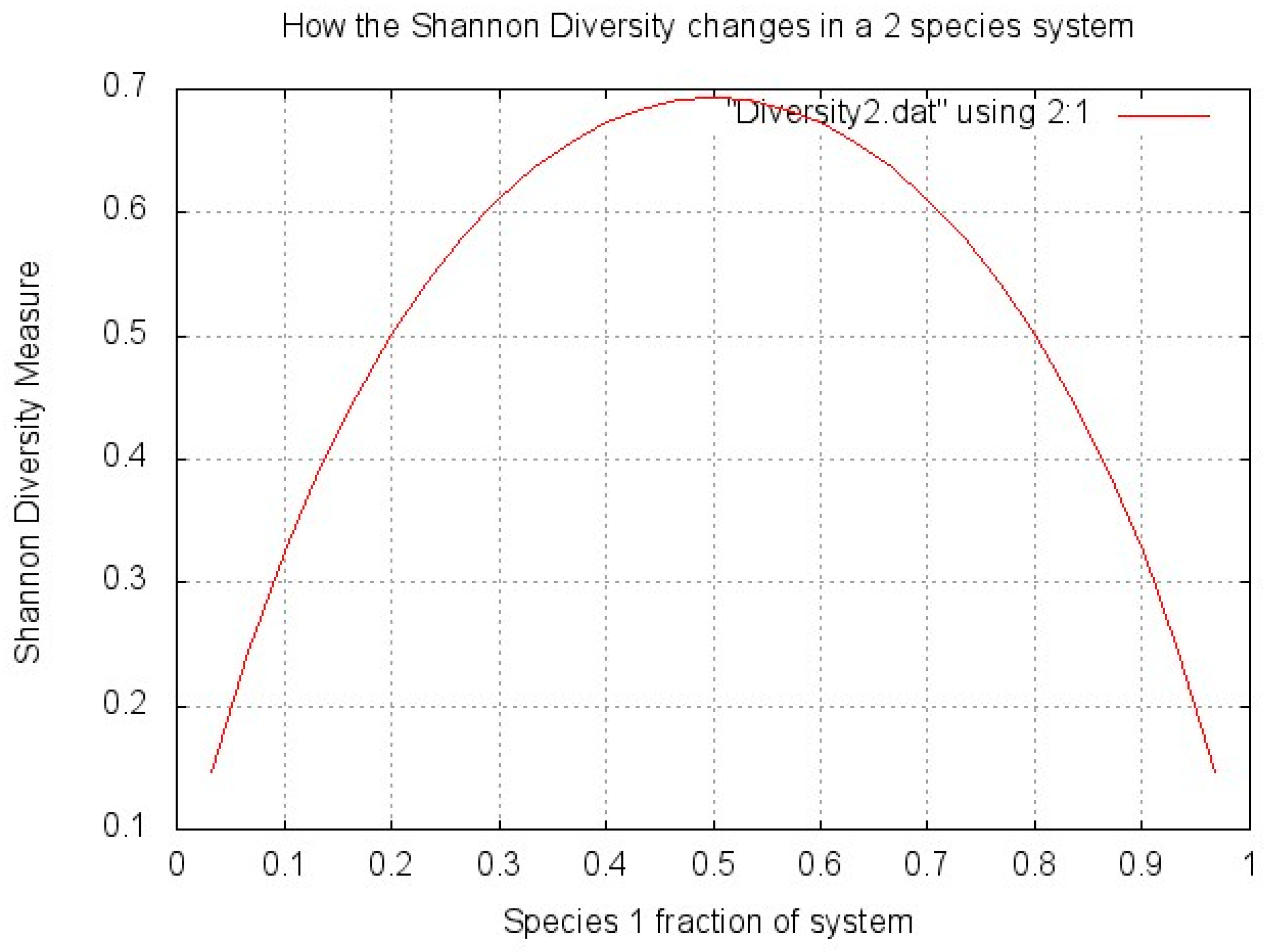
1.3.2. Measuring Diversity
- H’ = biodiversity index
- i = species
- pi = ni/N
- ni = total number of organisms of a particular species
- N = total number of organisms of all species.
- Sest = estimated number of species
- Sobs = observed number of species
- f1 = number of singleton taxa (taxa represented by a single read in that community)
- f2 = number of doubleton taxa
- OTU = Operational Taxonomic Unit (the most commonly used units of microbial diversity, refers to clusters of (uncultivated or unknown) organisms, grouped by DNA sequence similarity of a specific taxonomic marker gene)
2. Field Data and Key Results
2.1. Setting a Baseline for Determining “Healthy” vs. “Damaged” Skin
2.1.1. Previously Uncontacted Yanomami People
2.1.2. Rural Guahibo Settlement
2.1.3. Agricultural Community in Brazil
2.1.4. Healthy vs. Diseased or Damaged Skin
2.1.5. Summary of Results
2.2. Using the Data
2.3. Comparing the Data
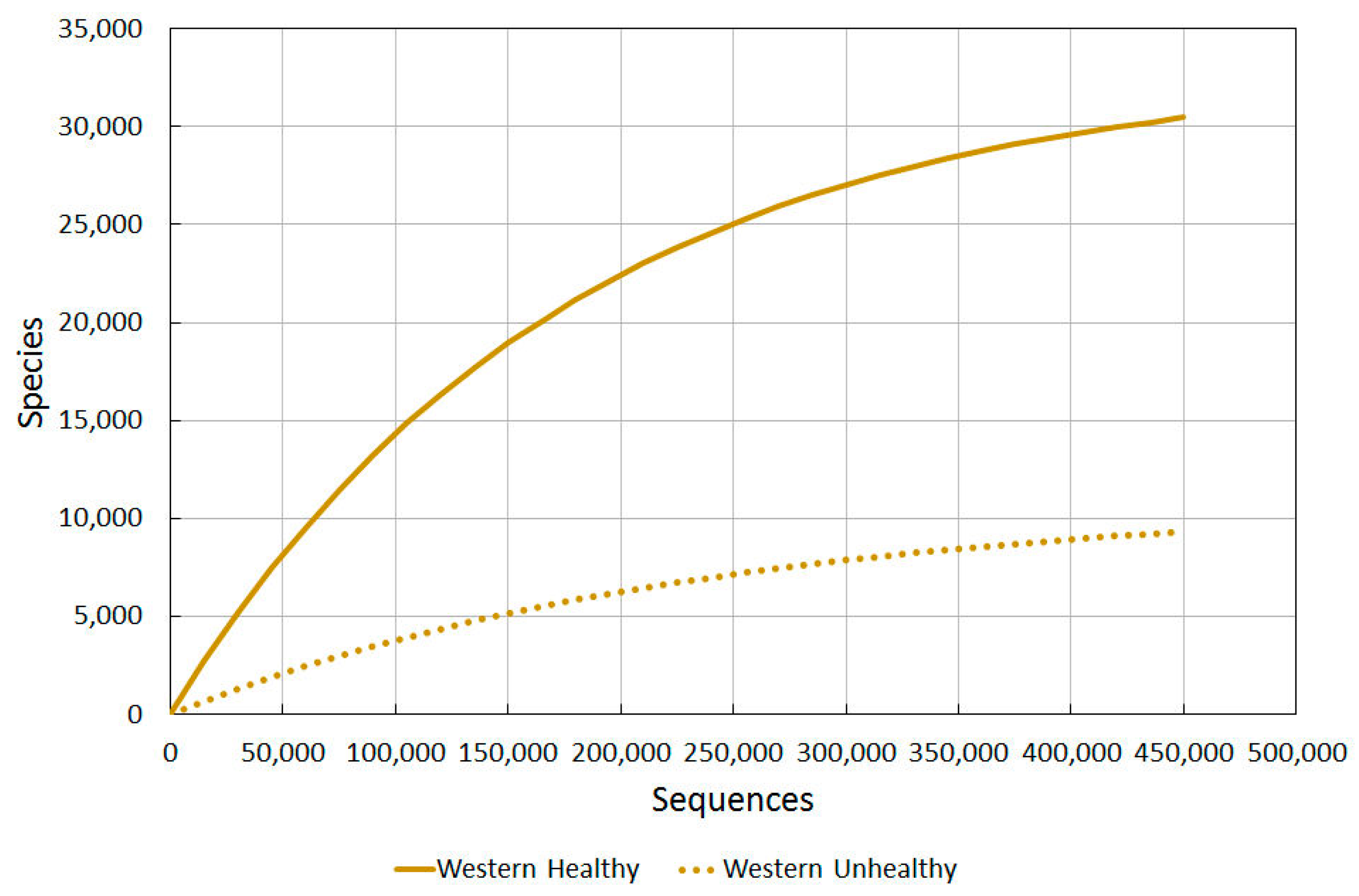
2.3.1. Western Healthy vs. Western Unhealthy
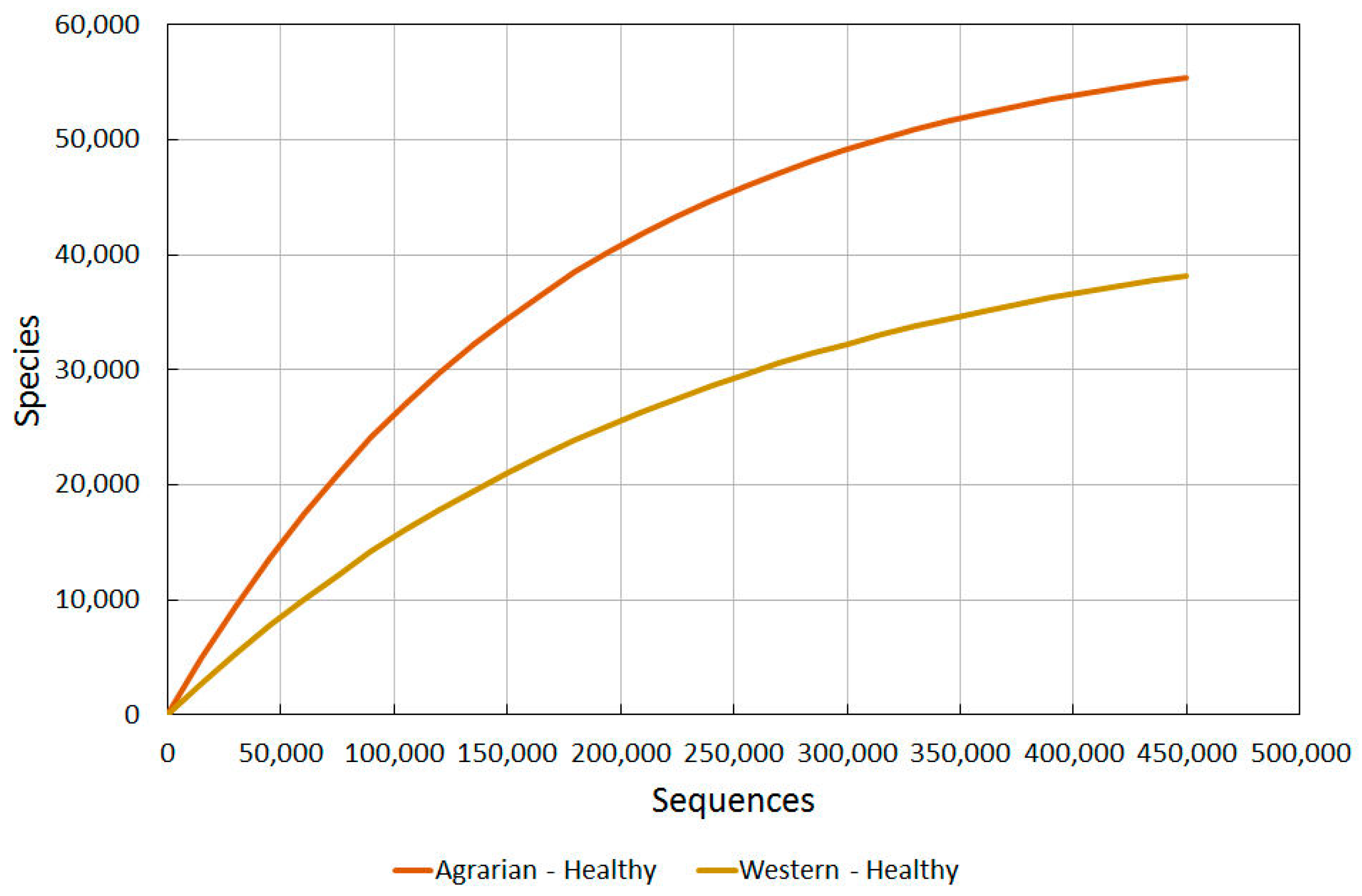
2.3.2. Agrarian Healthy vs. Western Healthy Skin
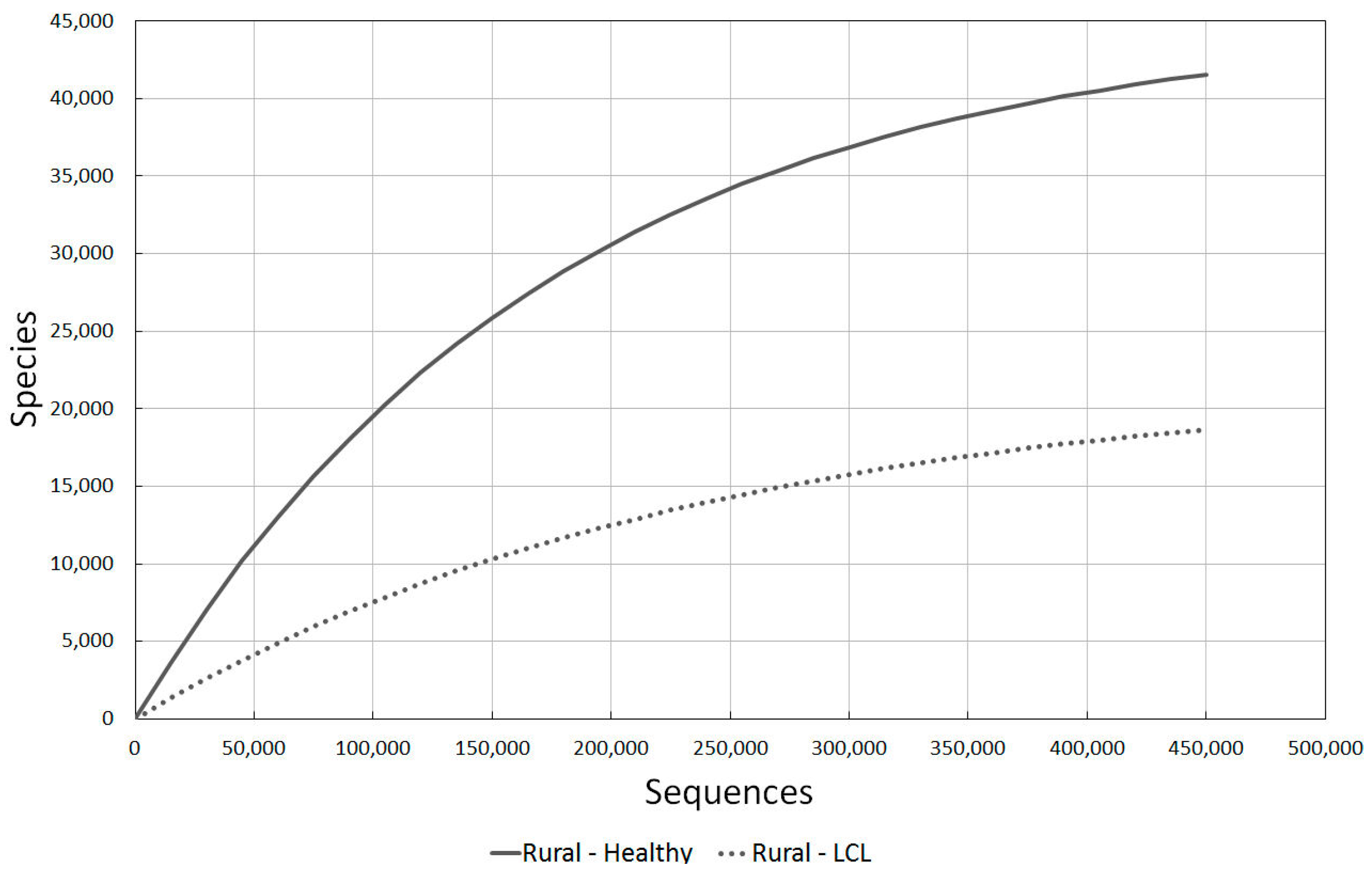
2.3.3. Rural Healthy vs. LCL Lesion Skin
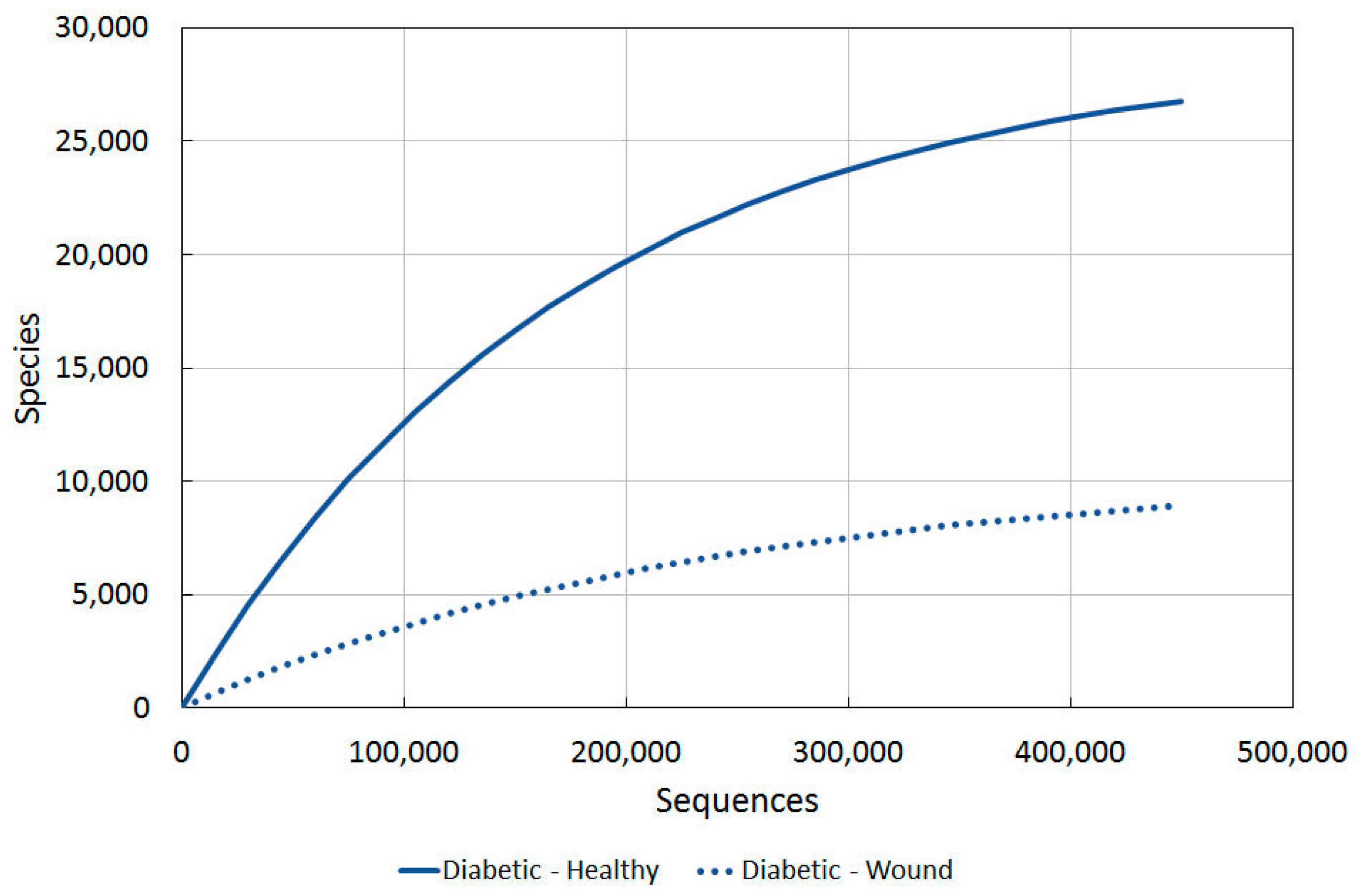
2.3.4. Diabetic vs. Intact Skin
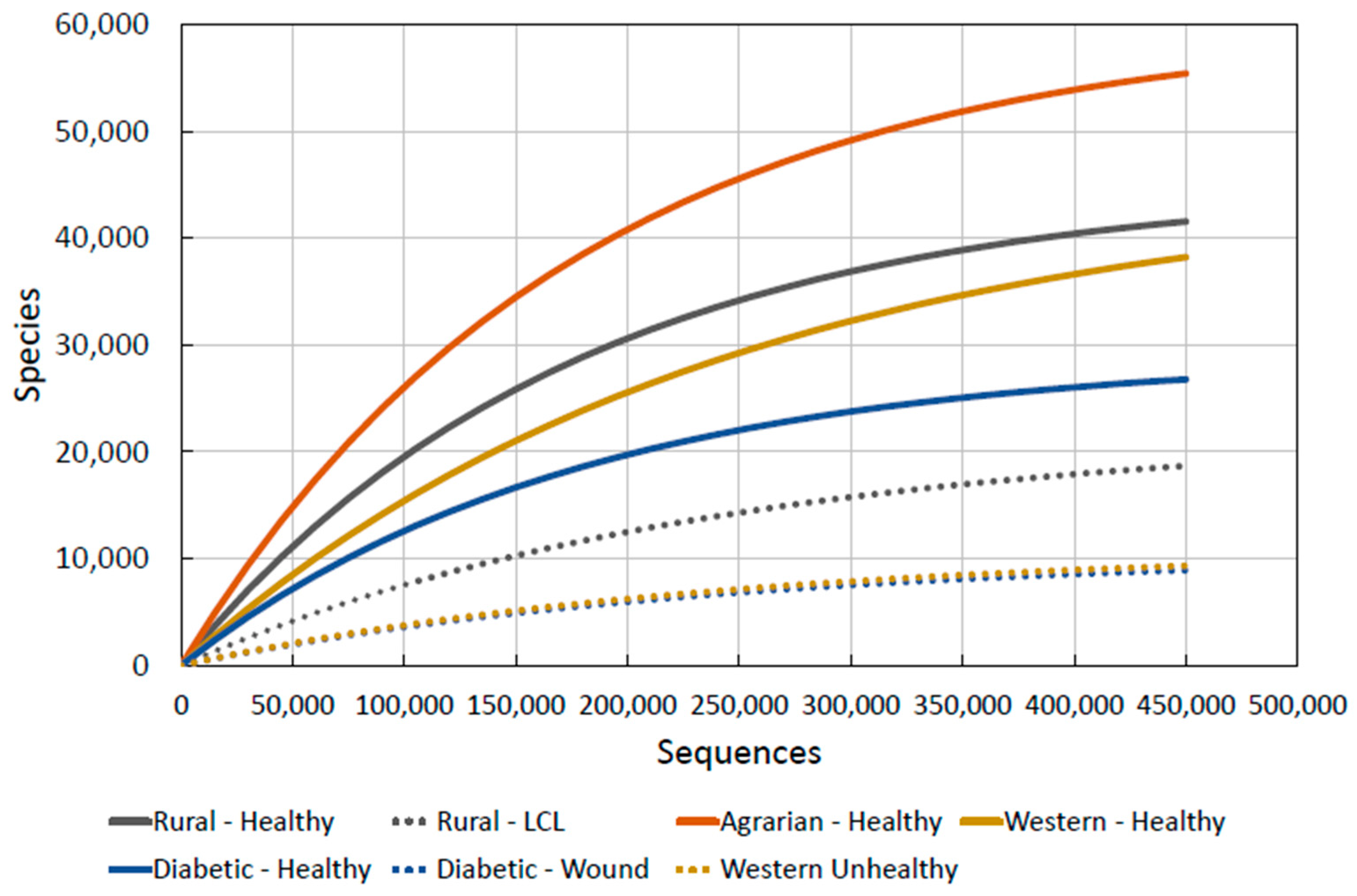
2.3.5. Combining the Data
2.4. Proposal for Using Skin Microbiota Diversity
2.4.1 Improving Our Mechanism in Future Work
3. Discussion: Solving the Skin Allergy Epidemic
3.1. Comparisons with Gut Microbiota Studies
3.1.1. Why Microbial Diversity Is Key to Health
3.1.2. Using Probiotics
3.2. Why Agrarian Cultures Have a Greater Skin Microbial Diversity
3.3. Testing for Skin Health: A New Mechanism Revealed
3.4. Are Cosmetics a Major Contributor to the Skin Allergy Epidemic?
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Eckburg, P.B.; Bik, E.M.; Bernstein, C.N.; Purdom, E.; Dethlefsen, L.; Sargent, M.; Gill, S.R.; Nelson, K.E.; Relman, D.A. Diversity of the human intestinal microbial flora. Science 2005, 308, 1635–1638. [Google Scholar] [CrossRef] [PubMed]
- O’Hara, A.M.; Shanahan, F. The gut flora as a forgotten organ. EMBO Rep. 2006, 7, 688–693. [Google Scholar] [CrossRef] [PubMed]
- Kelly, D.; Conway, S.; Aminov, R. Commensal gut bacteria: Mechanisms of immune modulation. Trends Immunol. 2005, 26, 326–333. [Google Scholar] [CrossRef] [PubMed]
- Finlay, B.B.; Arrieta, M.-C. Let Them Eat Dirt. Saving Your Child from an Oversanitized World; Windmill Books: London, UK, 2016. [Google Scholar]
- Marrs, T.; Flohr, C. The role of skin and gut microbiota in the development of atopic eczema. Br. J. Dermatol. 2016, 175, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Martínez, I.; Stegen, J.C.; Maldonado-Gómez, M.X.; Eren, A.M.; Siba, P.M.; Greenhill, A.R.; Walter, J. The Gut Microbiota of Rural Papua New Guineans: Composition, Diversity Patterns, and Ecological Processes. Cell Rep. 2015, 11, 527–538. [Google Scholar] [CrossRef] [PubMed]
- Taylor, B.; Wadsworth, M.; Wadsworth, J.; Peckham, C. Changes in the reported prevalence of childhood eczema since the 1939–45 war. Lancet 1984, 324, 1255–1257. [Google Scholar] [CrossRef]
- Ribeiro, A.; Estanqueiro, M.; Oliveira, M.; Sousa Lobo, J. Main Benefits and Applicability of Plant Extracts in Skin Care Products. Cosmetics 2015, 2, 48–65. [Google Scholar] [CrossRef]
- Grice, E.A.; Segre, J.A. The skin microbiome. Nat. Rev. Microbiol. 2011, 9, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Findley, K.; Grice, E.A. The skin microbiome: A focus on pathogens and their association with skin disease. PLoS Pathog. 2014, 10, e1004436. [Google Scholar] [CrossRef] [PubMed]
- Cogen, A.L.; Nizet, V.; Gallo, R.L. Skin microbiota: A source of disease or defence? Br. J. Dermatol. 2009, 158, 442–455. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Snitkin, E.S.; Yockey, L.J.; Bermudez, D.M.; NISC Comparative Sequencing Program; Liechty, K.W.; Segre, J.A. Longitudinal shift in diabetic wound microbiota correlates with prolonged skin defense response. Proc. Natl. Acad. Sci. USA 2010, 33107, 14799–14804. [Google Scholar] [CrossRef] [PubMed]
- Kong, H.H.; Segre, J.A. Skin microbiome: Looking back to move forward. J. Investig. Dermatol. 2012, 132, 933–939. [Google Scholar] [CrossRef] [PubMed]
- Blaser, M.J.; Falkow, S. What are the consequences of the disappearing human microbiota? Nat. Rev. Microbiol. 2009, 7, 887–894. [Google Scholar] [CrossRef] [PubMed]
- Noble, W.C. Staphylococci on the skin. In The Skin Microflora and Microbial Skin Disease; Noble, W.C., Ed.; Cambridge University Press: London, UK, 2004; pp. 135–152. [Google Scholar]
- Holland, K.T.; Bojar, R.A. Cosmetics: What is their influence on the skin microflora? Am. J. Clin. Dermatol. 2002, 3, 445–449. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Kong, H.H.; Renaud, G.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Wolfsberg, T.G.; Turner, M.L.; Segre, J.A.; NISC, C.S.P. A diversity profile of the human skin microbiota. Genome Res. 2008, 18, 1043–1050. [Google Scholar] [CrossRef] [PubMed]
- Katsuyama, M.; Ichikawa, H.; Ogawa, S.; Ikezawa, Z. A novel method to control the balance of skin microflora: Part 1. Attack on biofilm of Staphylococcus aureus without antibiotics. J. Dermatol. Sci. 2005, 38, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Lambers, H.; Piessens, S.; Bloem, A.; Pronk, H.; Finkel, P. Natural skin surface pH is on average below 5, which is beneficial for its resident flora. Int. J. Cosmet. Sci. 2006, 28, 359–370. [Google Scholar] [CrossRef] [PubMed]
- Brogden, N.K.; Mehalick, L.; Fischer, C.L.; Wertz, P.W.; Brogden, K.A. The emerging role of peptides and lipids as antimicrobial epidermal barriers and modulators of local inflammation. Skin Pharmacol. Physiol. 2012, 25, 167–181. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Kong, H.H.; Conlan, S.; Deming, C.B.; Davis, J.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Murray, P.R.; Green, E.D. Topographical and temporal diversity of the human skin microbiome. Science 2009, 324, 1190–1192. [Google Scholar] [CrossRef] [PubMed]
- Rinne, M.; Kalliomäki, M.; Salminen, S.; Isolauri, E. Probiotic intervention in the first months of life: Short-term effects on gastrointestinal symptoms and long-term effects on gut microbiota. J. Pediatr. Gastroenterol. Nutr. 2006, 43, 200–205. [Google Scholar] [CrossRef] [PubMed]
- Noverr, M.C.; Huffnagle, G.B. The “microflora hypothesis” of allergic diseases. Clin. Exp. Allergy 2005, 35, 1511–1520. [Google Scholar] [CrossRef] [PubMed]
- Simpson, C.R.; Newton, J.; Hippisley-Cox, J.; Sheikh, A. Trends in the epidemiology and prescribing of medication for eczema in England. J. R. Soc. Med. 2009, 102, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Burd, R.M. Psoriasis: A general overview. Br. J. Hosp. Med. 2006, 67, 259–262. [Google Scholar] [CrossRef] [PubMed]
- Prescott, S.L.; Tang, M.L.K. The Australasian Society of Clinical Immunology and Allergy position statement: Summary of allergy prevention in children. Med. J. Aust. 2005, 182, 464–467. [Google Scholar] [PubMed]
- Srinivas, G.; Möller, S.; Wang, J.; Künzel, S.; Zillikens, D.; Baines, J.F.; Ibrahim, S.M. Genome-wide mapping of gene–microbiota interactions in susceptibility to autoimmune skin blistering. Nat. Commun. 2013, 4, 2462. [Google Scholar] [CrossRef] [PubMed]
- Asher, M.I.; Montefort, S.; Björkstén, B.; Lai, C.K.; Strachan, D.P.; Weiland, S.K.; Williams, H.; ISAAC Phase Three Study Group. Worldwide time trends in the prevalence of symptoms of asthma, allergic rhinoconjunctivitis, and eczema in childhood: ISAAC Phases One and Three repeat multicountry cross-sectional surveys. Lancet 2006, 368, 733–743. [Google Scholar] [CrossRef]
- Shaw, T.E.; Currie, G.P.; Koudelka, C.W.; Simpson, E.L. Eczema Prevalence in the United States: Data from the 2003 National Survey of Children’s Health. J. Investig. Dermatol. 2011, 131, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Bickers, D.R.; Lim, H.W.; Margolis, D.; Weinstock, M.A.; Goodman, C.; Faulkner, E.; Gould, C.; Gemmen, E.; Dall, T.; American Academy of Dermatology Association Society for Investigative Dermatology. The burden of skin diseases: 2004. J. Am. Acad. Dermatol. 2006, 55, 490–500. [Google Scholar] [CrossRef] [PubMed]
- Bjarnsholt, T.; Kirketerp-Møller, K.; Jensen, P.Ø.; Madsen, K.G.; Phipps, R.; Krogfelt, K.; Høiby, N.; Givskov, M. Why chronic wounds will not heal: A novel hypothesis. Wound Repair Regen 2008, 16, 2–10. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulos, N.G.; Agache, I.; Bavbek, S.; Bilo, B.M.; Braido, F.; Cardona, V.; Custovic, A.; Demoly, P.; Eigenmann, P.; Gayraud, J.; et al. Research needs in allergy: An EAACI position paper, in collaboration with EFA. Clin. Transl. Allergy 2012, 2, 21. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.; Kiyu, A.; Milman, H.M.; Jimenez, J. Improving health and building human capital through an effective primary care system. J. Urban Health 2007, 84 (Suppl. S3), i75–i85. [Google Scholar] [CrossRef] [PubMed]
- Dessinioti, C.; Katsambas, A.D. The role of Propionibacterium acnes in acne pathogenesis: Facts and controversies. Clin. Dermatol. 2010, 28, 2–7. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Lozupone, C.A.; Hamady, M.; Knight, R.; Gordon, J.I. Worlds within worlds: Evolution of the vertebrate gut microbiota. Nat. Rev. Microbiol. 2008, 6, 776–788. [Google Scholar] [CrossRef] [PubMed]
- Pei, Z.; Bini, E.J.; Yang, L.; Zhou, M.; Francois, F.; Blaser, M.J. Bacterial biota in the human distal esophagus. Proc. Natl. Acad. Sci. USA 2004, 101, 4250–4255. [Google Scholar] [CrossRef] [PubMed]
- Costello, E.K.; Lauber, C.L.; Hamady, M.; Fierer, N.; Gordon, J.I.; Knight, R. Bacterial Community Variation in Human Body Habitats Across Space and Time. Science 2009, 326, 1694–1697. [Google Scholar] [CrossRef] [PubMed]
- Van de Merwe, J.P.; Stegeman, J.H.; Hazenberg, M.P. The resident faecal flora is determined by genetic characteristics of the host. Implications for Crohn’s disease? Antonie Van Leeuwenhoek 1983, 49, 119–124. [Google Scholar] [CrossRef] [PubMed]
- De Filippo, C.; Cavalieri, D.; Di Paola, M.; Ramazzotti, M.; Poullet, J.B.; Massart, S.; Collini, S.; Pieraccini, G.; Lionetti, P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc. Natl. Acad. Sci. USA 2010, 107, 14691–14696. [Google Scholar] [CrossRef] [PubMed]
- Sonnenburg, E.D.; Sonnenburg, J.L. Starving our Microbial Self: The Deleterious Consequences of a Diet Deficient in Microbiota-Accessible Carbohydrates. Cell Metab. 2014, 20, 779–786. [Google Scholar] [CrossRef] [PubMed]
- Lefcheck, J.S.; Byrnes, J.E.K.; Isbell, F.; Gamfeldt, L.; Griffin, J.N.; Eisenhauer, N.; Hensel, M.J.; Hector, A.; Cardinale, B.J.; Duffy, J.E. Biodiversity enhances ecosystem multifunctionality across trophic levels and habitats. Nat. Commun. 2015, 6, 6936. [Google Scholar] [CrossRef] [PubMed]
- Balvanera, P.; Pfisterer, A.B.; Buchmann, N.; He, J.-S.; Nakashizuka, T.; Raffaelli, D.; Schmid, B. Quantifying the evidence for biodiversity effects on ecosystem functioning and services. Ecol. Lett. 2006, 9, 1146–1156. [Google Scholar] [CrossRef] [PubMed]
- Gamfeldt, L.; Hillebrand, H.; Jonsson, P.R. Multiple functions increase the importance of biodiversity for overall ecosystem functioning. Ecology 2008, 89, 1223–1231. [Google Scholar] [CrossRef] [PubMed]
- Thrupp, L.A. The importance of biodiversity in agroecosystems. J. Crop Improv. 2004, 12, 315–337. [Google Scholar] [CrossRef]
- Brookfield, H.; Padoch, C. Appreciating Agrodiversity: A Look at the Dynamism and Diversity of Indigenous Farming Practices. Environ. Sci. Policy Sustain. Dev. 1994, 36, 6–45. [Google Scholar] [CrossRef]
- Dudgeon, D.; Arthington, A.H.; Gessner, M.O.; Kawabata, Z.-I.; Knowler, D.J.; Lévêque, C.; Naiman, R.J.; Prieur-Richard, A.H.; Soto, D.; Stiassny, M.L. Freshwater biodiversity: Importance, threats, status and conservation challenges. Biol. Rev. 2006, 81, 163. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.U.; Chapin, F.S.; Ewel, J.J.; Hector, A.; Inchausti, P.; Lavorel, S.; Lawton, J.H.; Lodge, D.M.; Loreau, M.; Naeem, S.; et al. Effects of biodiversity on ecosystem functioning: A consensus of current knowledge. Ecol. Monogr. 2005, 75, 3–35. [Google Scholar] [CrossRef]
- Cardinale, B.J.; Srivastava, D.S.; Emmett Duffy, J.; Wright, J.P.; Downing, A.L.; Sankaran, M.; et al. Effects of biodiversity on the functioning of trophic groups and ecosystems. Nature 2006, 443, 989–992. [Google Scholar] [CrossRef] [PubMed]
- Worm, B.; Barbier, E.B.; Beaumont, N.; Duffy, J.E.; Folke, C.; Halpern, B.S.; Jackson, J.B.; Lotze, H.K.; Micheli, F.; Palumbi, S.R.; et al. Impacts of Biodiversity Loss on Ocean Ecosystem Services. Science 2006, 314, 787–790. [Google Scholar] [CrossRef] [PubMed]
- Costanza, R.; Mageau, M. What is a healthy ecosystem? In Aquatic Ecology; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1999; pp. 105–115. [Google Scholar]
- Ripple, W.J.; Beschta, R.L. Wolf reintroduction, predation risk, and cottonwood recovery in Yellowstone National Park. For. Ecol. Manag. 2003, 184, 299–313. [Google Scholar] [CrossRef]
- Zahniser, A.; Singh, A. Return of the wolves to Yellowstone National Park, USA: A model of ecosystem restoration. Biodiversity 2004, 5, 3–7. [Google Scholar] [CrossRef]
- Smith, D.W.; Peterson, R.O.; Houston, D.B. Yellowstone after wolves. Bioscience 2003, 53, 330–340. [Google Scholar] [CrossRef]
- Beschta, R.L.; Ripple, W.J. Berry-producing shrub characteristics following wolf reintroduction in Yellowstone National Park. For. Ecol. Manag. 2012, 276, 132–138. [Google Scholar] [CrossRef]
- Haines, A.L. The Yellowstone Story—A History of Our First National Park; University Press of Colorado: Boulder, CO, USA, 1996. [Google Scholar]
- Fischer, H. Wolf Wars: The Remarkable Inside Story of the Restoration of Wolves to Yellowstone; Falcon Press: Nashville, TN, USA, 1995; p. 183. [Google Scholar]
- Berger, K.M.; Gese, E.M.; Berger, J. Indirect effects and traditional trophic cascades: A test involving wolves, coyotes and pronghorn. Ecology 2008, 89, 818–828. [Google Scholar] [CrossRef] [PubMed]
- Stolzenburg, W. Where the Wild Things Were: Life, Death, and Ecological Wreckage in a Land of Vanishing Predators; Bloomsbury: London, UK, 2009; p. 291. ISBN 1596916249. [Google Scholar]
- Kay, C.E. Viewpoint: Ungulate herbivory, willows, and political ecology in yellowstone. J. Range Manag. 1997, 50, 139–145. [Google Scholar] [CrossRef]
- Platts, W.S.; Armour, C.L.; Booth, G.D.; Bryant, M.; Bufford, J.L.; Cuplin, P.; Jensen, S.; Lienkaemper, G.W.; Minshall, G.W.; Monsen, S.T.; et al. Methods for Evaluating Riparian Habitats with Applications to Management; General Technical Report INT-221; USDA Forest Service: Ogden, UT, USA; p. 177.
- Platts, W.S.; Megahan, W.F.; Minshall, G.W. Methods for Evaluating Stream, Riparian, and Biotic Conditions; General Technical Report INT-138; USDA Forest Service: Ogden, UT, USA; p. 70.
- Platts, W.S. Livestock grazing. Am. Fish Soc. Spec. Publ. 1991, 19, 389–424. [Google Scholar]
- Patten, D.T. Herbivore optimization and overcompensation: Does native herbivory on western rangelands support these theories? Ecol. Appl. 1993, 3, 35–36. [Google Scholar] [CrossRef] [PubMed]
- Rosgen, D.L. Stream Classification, Streambank Erosion, and Fluvial Interpretations for the Lamar River and Main Tributaries; Technical Report USDI Park Service; Yellowstone Natl Park: Pagosa Springs, CO, USA, 1993; p. 82. [Google Scholar]
- Kay, C.E. Yellowstone’s Northern Elk Herd: A Critical Evaluation of the “Natural Regulation” Paradigm. Ph.D. Dissertation, Utah State University, Logan, UT, USA, 1990. [Google Scholar]
- Kay, C.E. Browsing by native ungulates: Effects on shrub and seed production in the Greater Yellowstone Ecosystem. In Wild Land Shrub and Arid Land Restoration Symposium: Proceedings; Roundy, B.A., McArthur, E.D., Haley, J.S., Mann, D.K., Eds.; Forest Service General Technical Report INT-GTR-315; U.S. Department of Agriculture: Ogden, UT, USA, 1995; p. 384. [Google Scholar]
- Wagner, F.H.; Sax, J.L. Wildlife Policies in the U.S. National Parks; Island Press: Washington, DC, USA, 1995; p. 242. [Google Scholar]
- Beyer, H.L.; Merrill, E.H.; Varley, N.; Boyce, M.S. Willow on Yellowstone’s northern range: Evidence for a trophic cascade? Ecol. Appl. 2007, 17, 1563–1571. [Google Scholar] [CrossRef] [PubMed]
- Tyers, D.B. The Condition of the Northern Yellowstone Winter Range in Yellowstone National Park—A Discussion of the Controversy. Master’s Thesis, Montana State University, Bozeman, MT, USA, 1981. [Google Scholar]
- Kay, C.E. The impact of native ungulates and beaver on riparian communities in the Intermountain West. Nat. Resour. Environ. Issues 1994, 1, 23–44. [Google Scholar]
- Werndl, C.; Frigg, R. Entropy—A guide for the perplexed. In Probabilities in Phyics; Beisbart, C., Hartmann, S., Eds.; Oxford University Press: Oxford, UK, 2011. [Google Scholar] [CrossRef]
- Jost, L. Entropy and diversity. Oikos 2006, 113, 363–375. [Google Scholar] [CrossRef]
- Clemente, J.C.; Pehrsson, E.C.; Blaser, M.J.; Sandhu, K.; Gao, Z.; Wang, B.; Magris, M.; Hidalgo, G.; Contreras, M.; Noya-Alarcón, Ó.; et al. The microbiome of uncontacted Amerindians. Sci. Adv. 2015, 1, e1500183. [Google Scholar] [CrossRef] [PubMed]
- Blaser, M.J.; Dominguez-Bello, M.G.; Contreras, M.; Magris, M.; Hidalgo, G.; Estrada, I.; Gao, Z.; Clemente, J.C.; Costello, E.K.; Knight, R. Distinct cutaneous bacterial assemblages in a sampling of South American Amerindians and US residents. ISME J. 2013, 7, 85–95. [Google Scholar] [CrossRef] [PubMed]
- Salgado, V.R.; de Queiroz, A.T.L.; Sanabani, S.S.; de Oliveira, C.I.; Carvalho, E.M.; Costa, J.M.L.; Barral-Netto, M.; Barral, A. The microbiological signature of human cutaneous leishmaniasis lesions exhibits restricted bacterial diversity compared to healthy skin. Mem. Inst. Oswaldo Cruz 2016, 111, 241–251. [Google Scholar] [CrossRef] [PubMed]
- Gontcharova, V.; Youn, E.; Sun, Y.; Wolcott, R.D.; Dowd, S.E. A comparison of bacterial composition in diabetic ulcers and contralateral intact skin. Open Microbiol. J. 2010, 4, 8–19. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, A.R.; Patterson, A.P.; Diesel, A.; Lawhon, S.D.; Ly, H.J.; Stephenson, C.E.; Mansell, J.; Steiner, J.M.; Dowd, S.E.; Olivry, T.; et al. The skin microbiome in healthy and allergic dogs. PLoS ONE 2014, 9, e83197. [Google Scholar] [CrossRef]
- Nakatsuji, T.; Kao, M.C.; Zhang, L.; Zouboulis, C.C.; Gallo, R.L.; Huang, C.-M. Sebum free fatty acids enhance the innate immune defense of human sebocytes by upregulating β-defensin-2 expression. J. Investig. Dermatol. 2010, 130, 985–994. [Google Scholar] [CrossRef] [PubMed]
- Song, C.; Weichbrodt, C.; Salnikov, E.S.; Dynowski, M.; Forsberg, B.O.; Bechinger, B.; Steinem, C.; de Groot, B.L.; Zachariae, U.; Zeth, K. Crystal structure and functional mechanism of a human antimicrobial membrane channel. Proc. Natl. Acad. Sci. USA 2013, 110, 4586–4591. [Google Scholar] [CrossRef] [PubMed]
- Ghose, C.; Perez-Perez, G.I.; Dominguez-Bello, M.-G.; Pride, D.T.; Bravi, C.M.; Blaser, M.J. East Asian genotypes of Helicobacter pylori strains in Amerindians provide evidence for its ancient human carriage. Proc. Natl. Acad. Sci. USA 2002, 99, 15107–15111. [Google Scholar] [CrossRef] [PubMed]
- Marini, E.; Maldonado-Contreras, A.L.; Cabras, S.; Hidalgo, G.; Buffa, R.; Marin, A.; Floris, G.; Racugno, W.; Pericchi, L.R.; Castellanos, M.E.; et al. Helicobacter pylori and intestinal parasites are not detrimental to the nutritional status of Amerindians. Am. J. Trop. Med. Hyg. 2007, 76, 534–540. [Google Scholar] [CrossRef] [PubMed]
- Dominguez-Bello, M.G.; Costello, E.K.; Contreras, M.; Magris, M.; Hidalgo, G.; Fierer, N.; Knight, R. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc. Natl. Acad. Sci. USA 2010, 107, 11971–11975. [Google Scholar] [CrossRef] [PubMed]
- Kong, H.H.; Oh, J.; Deming, C.; Conlan, S.; Grice, E.A.; Beatson, M.A.; Nomicos, E.; Polley, E.C.; Komarow, H.D.; Murray, P.R.; Turner, M.L. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Res. 2012, 22, 850–859. [Google Scholar] [CrossRef] [PubMed]
- Dekio, I.; Hayashi, H.; Sakamoto, M.; Kitahara, M.; Nishikawa, T.; Suematsu, M.; Benno, Y. Detection of potentially novel bacterial components of the human skin microbiota using culture-independent molecular profiling. J. Med. Microbiol. 2005, 54, 1231–1238. [Google Scholar] [CrossRef] [PubMed]
- Bek-Thomsen, M.; Lomholt, H.B.; Kilian, M. Acne is not associated with yet-uncultured bacteria. J. Clin. Microbiol. 2008, 46, 3355–3360. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Tseng, C.; Pei, Z.; Blaser, M.J. Molecular analysis of human forearm superficial skin bacterial biota. Proc. Natl. Acad. Sci. USA 2007, 104, 2927–2932. [Google Scholar] [CrossRef] [PubMed]
- ScienceDaily. More than just Bacteria: Importance of Microbial Diversity in Gut Health, Disease. Available online: www.sciencedaily.com/releases/2014/03/140310090919.htm (accessed on 3 March 2017).
- Rowland, I.R.; Capurso, L.; Collins, K.; Cummings, J.; Delzenne, N.; Goulet, O.; Guarner, F.; Marteau, P.; Meier, R. Current level of consensus on probiotic science—Report of an expert meeting—London, 23 November 2009. Gut Microbes 2010, 1, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Grönlund, M.M.; Lehtonen, O.P.; Eerola, E.; Kero, P. Fecal microflora in healthy infants born by different methods of delivery: Permanent changes in intestinal flora after cesarean delivery. J. Pediatr. Gastroenterol. Nutr. 1999, 28, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Shono, Y.; Docampo, M.D.; Peled, J.U.; Perobelli, S.M.; Velardi, E.; Tsai, J.J.; Slingerland, A.E.; Smith, O.M.; Young, L.F.; Gupta, J.; et al. Increased GVHD-related mortality with broad-spectrum antibiotic use after allogeneic hematopoietic stem cell transplantation in human patients and mice. Sci. Transl. Med. 2016, 8, 339ra71. [Google Scholar] [CrossRef] [PubMed]
- Jernberg, C.; Lofmark, S.; Edlund, C.; Jansson, J.K. Long-term impacts of antibiotic exposure on the human intestinal microbiota. Microbiology 2010, 156, 3216–3223. [Google Scholar] [CrossRef] [PubMed]
- Jernberg, C.; Löfmark, S.; Edlund, C.; Jansson, J.K. Long-term ecological impacts of antibiotic administration on the human intestinal microbiota. ISME J. 2007, 1, 56–66. [Google Scholar] [CrossRef] [PubMed]
- Dethlefsen, L.; Huse, S.; Sogin, M.L.; Relman, D.A. The Pervasive Effects of an Antibiotic on the Human Gut Microbiota, as Revealed by Deep 16S rRNA Sequencing. PLoS Biol. 2008, 6, e280. [Google Scholar] [CrossRef] [PubMed]
- Fluhr, J.W.; Kao, J.; Ahn, S.K.; Feingold, K.R.; Elias, P.M.; Jain, M. Generation of free fatty acids from phospholipids regulates stratum corneum acidification and integrity. J. Investig. Dermatol. 2001, 117, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Pickford, M.; Senut, B. “Millennium Ancestor”, a 6-million-year-old bipedal hominid from Kenya. S. Afr. J. Sci. 2001, 97, 22. [Google Scholar]
- Kovarovic, K. Human evolution: A very short introduction. J. Hum. Evol. 2007, 52, 467. [Google Scholar] [CrossRef]
- Halliday, T.J.D.; Upchurch, P.; Goswami, A. Resolving the relationships of Paleocene placental mammals. Biol. Rev. 2017, 92, 521–550. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Hamady, M.; Lozupone, C.; Turnbaugh, P.J.; Ramey, R.R.; Bircher, J.S.; Schlegel, M.L.; Tucker, T.A.; Schrenzel, M.D.; Knight, R.; et al. Evolution of mammals and their gut microbes. Science 2008, 320, 1647–1651. [Google Scholar] [CrossRef] [PubMed]
- Goossens, A. Contact-allergic reactions to cosmetics. J. Allergy 2011, 2011, 467071. [Google Scholar] [CrossRef] [PubMed]
- Salverda, J.G.W.; Bragt, P.J.C.; de Wit-Bos, L.; Rustemeyer, T.; Coenraads, P.J.; Tupker, R.A.; Kunkeler, L.; Laheij-de Boer, A.M.; Stenveld, H.J.; van Ginkel, C.J.; et al. Results of a cosmetovigilance survey in The Netherlands. Contact Dermatitis 2013, 68, 139–148. [Google Scholar] [CrossRef] [PubMed]
- Heisterberg, M.V.; Menné, T.; Johansen, J.D. Contact allergy to the 26 specific fragrance ingredients to be declared on cosmetic products in accordance with the EU cosmetics directive. Contact Dermatitis 2011, 65, 266–275. [Google Scholar] [CrossRef] [PubMed]
- Warshaw, E.M.; Buchholz, H.J.; Belsito, D.V.; Maibach, H.I.; Fowler, J.F.; Rietschel, R.L.; Zug, K.A.; Mathias, C.T.; Pratt, M.D.; Sasseville, D.; et al. Allergic patch test reactions associated with cosmetics: Retrospective analysis of cross-sectional data from the North American Contact Dermatitis Group, 2001–2004. J. Am. Acad. Dermatol. 2009, 60, 23–38. [Google Scholar] [CrossRef] [PubMed]
- Berne, B.; Tammela, M.; Färm, G.; Inerot, A.; Lindberg, M. Can the reporting of adverse skin reactions to cosmetics be improved? A prospective clinical study using a structured protocol. Contact Dermatitis 2008, 58, 223–227. [Google Scholar] [CrossRef] [PubMed]
- Berne, B.; Boström, A.; Grahnén, A.F.; Tammela, M. Adverse effects of cosmetics and toiletries reported to the Swedish Medical Products Agency 1989–1994. Contact Dermatitis 1996, 34, 359–362. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wallen-Russell, C.; Wallen-Russell, S. Meta Analysis of Skin Microbiome: New Link between Skin Microbiota Diversity and Skin Health with Proposal to Use This as a Future Mechanism to Determine Whether Cosmetic Products Damage the Skin. Cosmetics 2017, 4, 14. https://doi.org/10.3390/cosmetics4020014
Wallen-Russell C, Wallen-Russell S. Meta Analysis of Skin Microbiome: New Link between Skin Microbiota Diversity and Skin Health with Proposal to Use This as a Future Mechanism to Determine Whether Cosmetic Products Damage the Skin. Cosmetics. 2017; 4(2):14. https://doi.org/10.3390/cosmetics4020014
Chicago/Turabian StyleWallen-Russell, Christopher, and Sam Wallen-Russell. 2017. "Meta Analysis of Skin Microbiome: New Link between Skin Microbiota Diversity and Skin Health with Proposal to Use This as a Future Mechanism to Determine Whether Cosmetic Products Damage the Skin" Cosmetics 4, no. 2: 14. https://doi.org/10.3390/cosmetics4020014
APA StyleWallen-Russell, C., & Wallen-Russell, S. (2017). Meta Analysis of Skin Microbiome: New Link between Skin Microbiota Diversity and Skin Health with Proposal to Use This as a Future Mechanism to Determine Whether Cosmetic Products Damage the Skin. Cosmetics, 4(2), 14. https://doi.org/10.3390/cosmetics4020014