A First Insight into North American Plant Pathogenic Fungi Armillaria sinapina Transcriptome
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Culture and Growth Conditions
2.2. Isolation of RNA
2.3. Transcriptome Sequencing, De Novo Assembly and Functional Annotation
2.4. Accession Numbers
2.5. Statistical Analysis
3. Results
3.1. Illumina Sequencing and De Novo Assembly
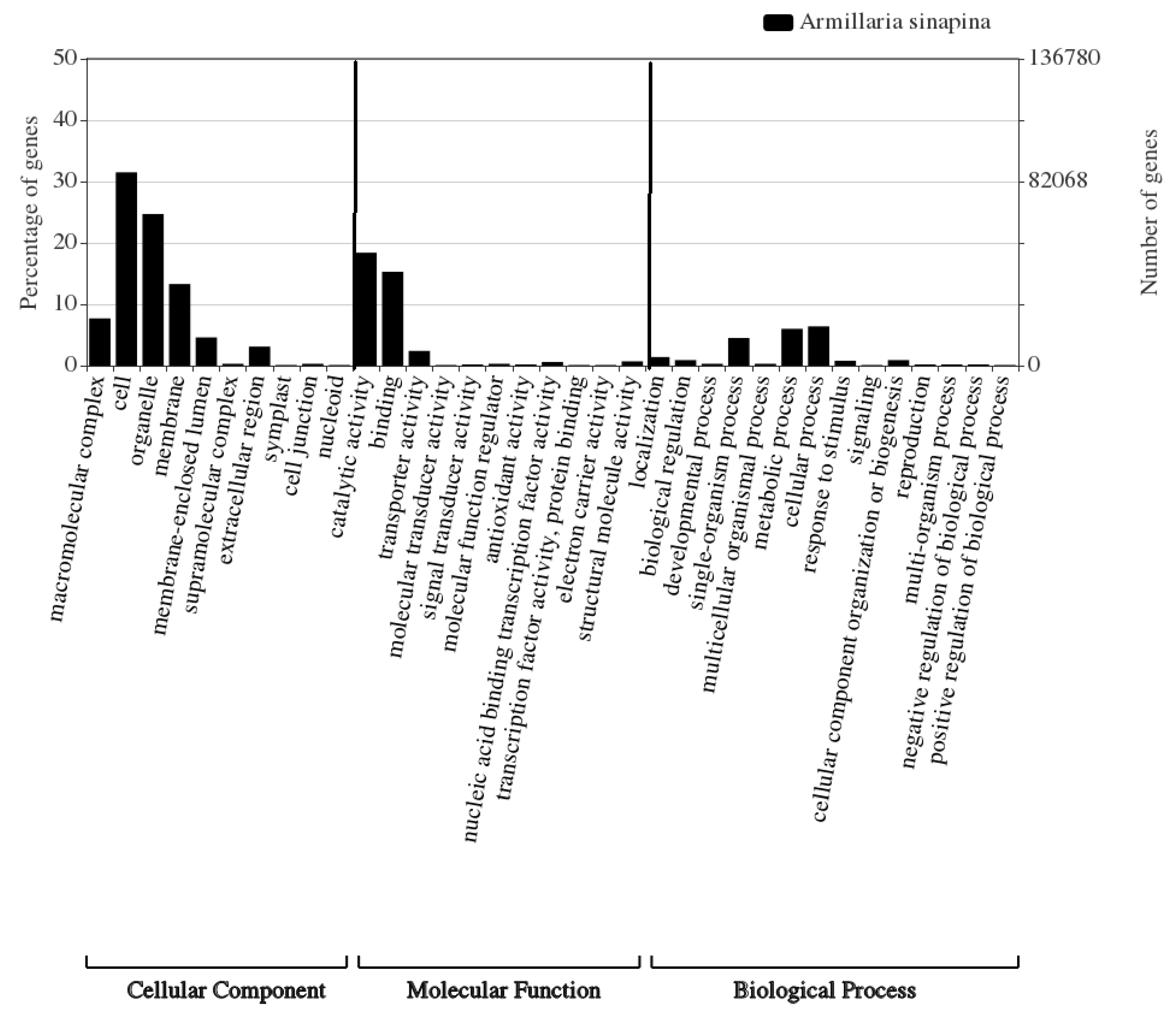
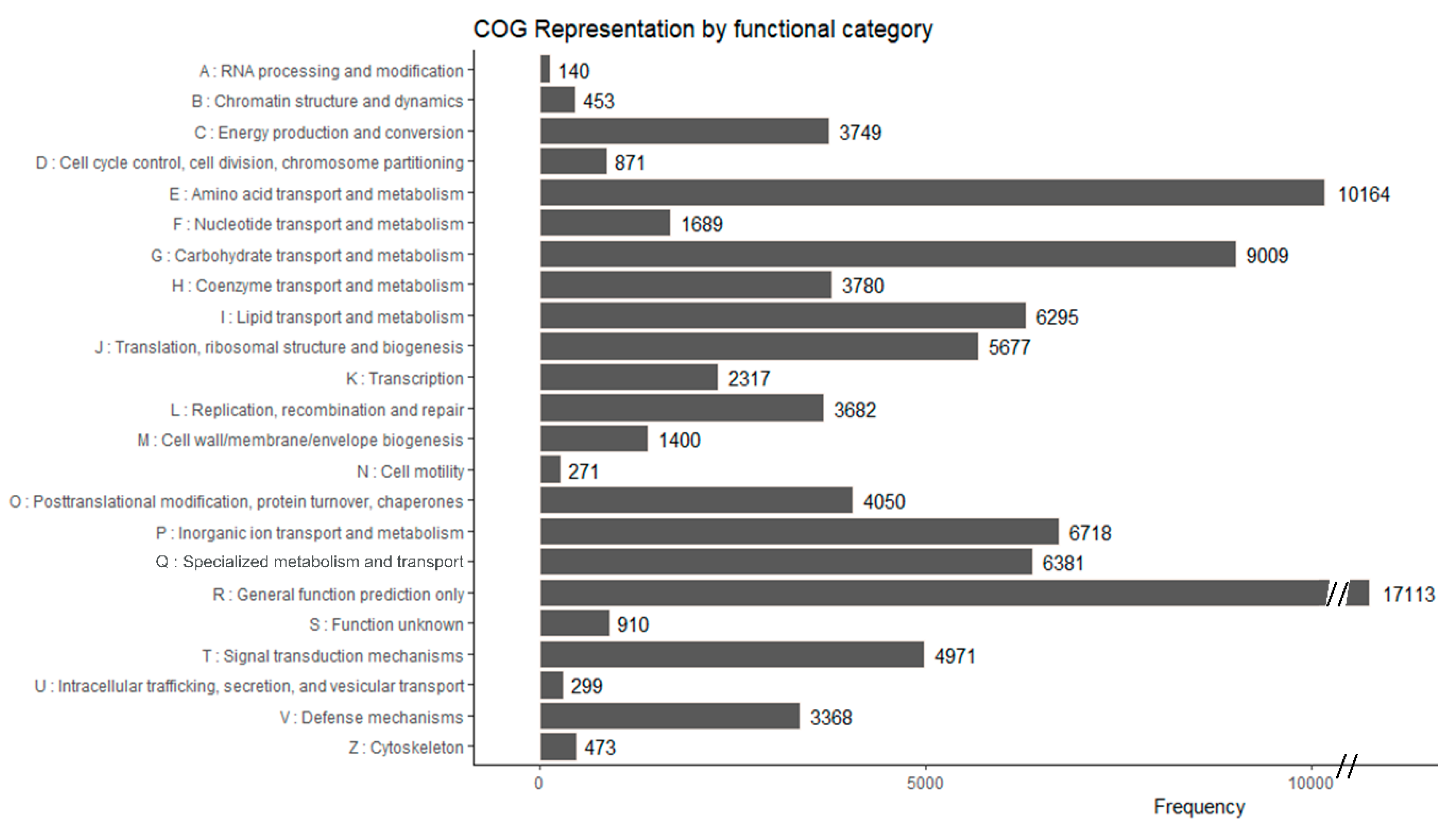
3.2. Functional Annotation of A. Sinapina Transcriptome
3.3. Differential Expression Analysis
3.4. Overview of Gene Expression with Biotechnological Relevance
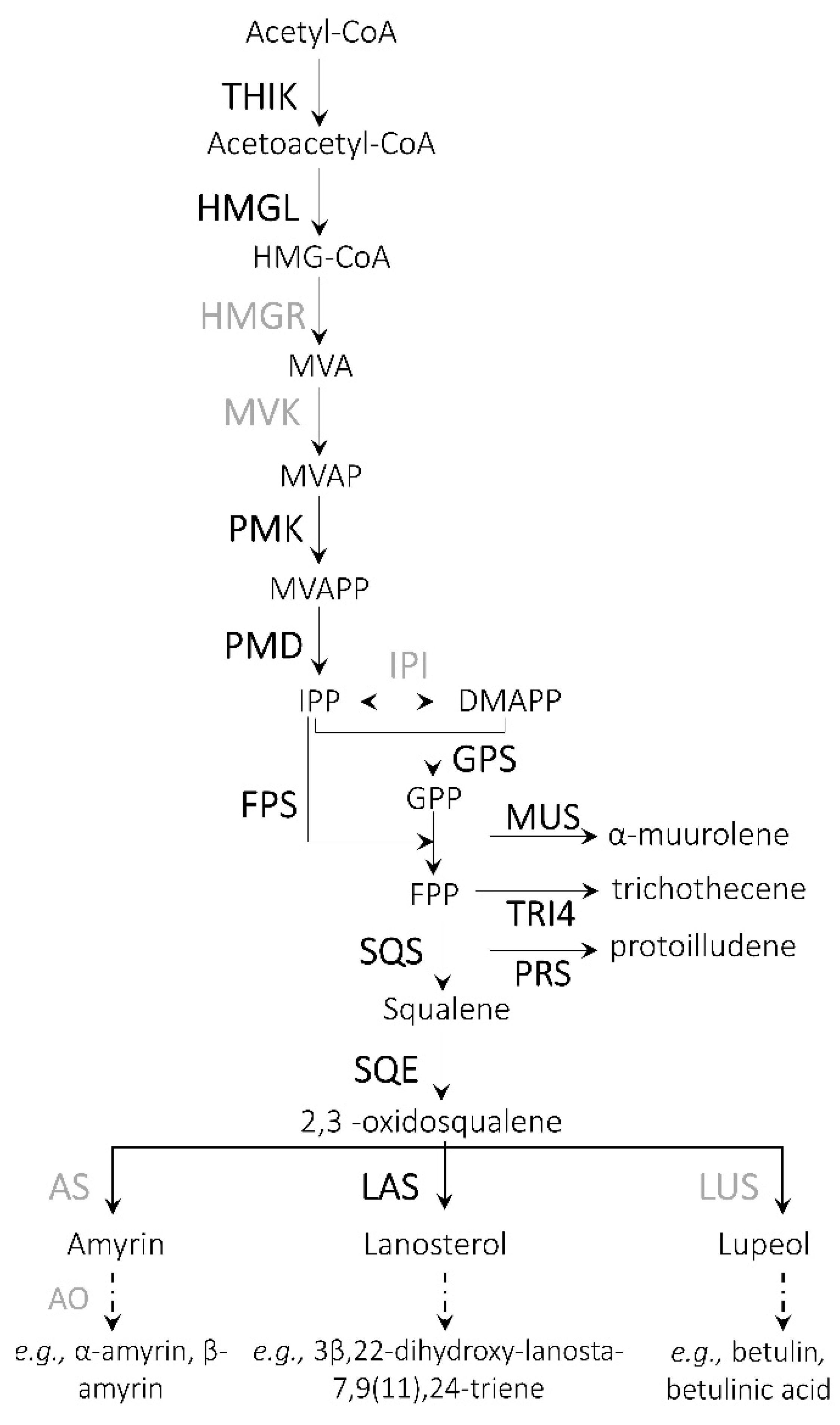
3.5. Terpenoids
3.6. Cytochrome P450s
3.7. Enzymes Involved in Plant Cell Wall Degradation
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Stewart, J.E.; Kim, M.-S.; Klopfenstein, N.B. Molecular Genetic Approaches Toward Understanding Forest-Associated Fungi and Their Interactive Roles Within Forest Ecosystems. Curr. For. Rep. 2018, 4, 72–84. [Google Scholar] [CrossRef]
- Araujo, R.; Sampaio-Maia, B. Fungal Genomes and Genotyping. In Advances in Applied Microbiology; Elsevier: Amsterdam, The Netherlands, 2018; Volume 102, pp. 37–81. [Google Scholar]
- Laperriere, G.; Desgagne-Penix, I.; Germain, H. DNA distribution pattern and metabolite profile of wild edible lobster mushroom (Hypomyces lactifluorum/Russula brevipes). Genome Natl. Res. Counc. Can. Genome Cons. Natl. Rech. Can. 2018, 61, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Genevieve, L.; Pierre-Luc, C.; Roxanne, G.-T.; Amélie, M.; Danny, B.; Vincent, M.; Hugo, G. Estimation of Fungal Diversity and Identification of Major Abiotic Drivers Influencing Fungal Richness and Communities in Northern Temperate and Boreal Quebec Forests. Forests 2019, 10, 1096. [Google Scholar] [CrossRef]
- Sipos, G.; Anderson, J.B.; Nagy, L.G. Armillaria. Curr. Biol. 2018, 28, R297–R298. [Google Scholar] [CrossRef] [PubMed]
- Heinzelmann, R.; Dutech, C.; Tsykun, T.; Labbé, F.; Soularue, J.-P.; Prospero, S. Latest advances and future perspectives in Armillaria research. Can. J. Plant Pathol. 2019, 41, 1–23. [Google Scholar] [CrossRef]
- Ross-Davis, A.; Stewart, J.; Hanna, J.; Kim, M.S.; Knaus, B.; Cronn, R.; Rai, H.; Richardson, B.; McDonald, G.; Klopfenstein, N. Transcriptome of an A rmillaria root disease pathogen reveals candidate genes involved in host substrate utilization at the host–pathogen interface. For. Pathol. 2013, 43, 468–477. [Google Scholar] [CrossRef]
- Schwartz, M.; Perrot, T.; Aubert, E.; Dumarçay, S.; Favier, F.; Gérardin, P.; Morel-Rouhier, M.; Mulliert, G.; Saiag, F.; Didierjean, C. Molecular recognition of wood polyphenols by phase II detoxification enzymes of the white rot Trametes versicolor. Sci. Rep. 2018, 8, 8472. [Google Scholar] [CrossRef]
- Mäkinen, M.A.; Risulainen, N.; Mattila, H.; Lundell, T.K. Transcription of lignocellulose-decomposition associated genes, enzyme activities and production of ethanol upon bioconversion of waste substrate by Phlebia radiata. Appl. Microbiol. Biotechnol. 2018, 102, 5657–5672. [Google Scholar] [CrossRef]
- Aranda, E. Promising approaches towards biotransformation of polycyclic aromatic hydrocarbons with Ascomycota fungi. Curr. Opin. Biotechnol. 2016, 38, 1–8. [Google Scholar] [CrossRef]
- Bérubé, J.; Dessureault, M. Morphological characterization of Armillaria ostoyae and Armillaria sinapina sp. nov. Can. J. Bot. 1988, 66, 2027–2034. [Google Scholar] [CrossRef]
- Klopfenstein, N.B.; Stewart, J.E.; Ota, Y.; Hanna, J.W.; Richardson, B.A.; Ross-Davis, A.L.; Elías-Román, R.D.; Korhonen, K.; Keča, N.; Iturritxa, E. Insights into the phylogeny of Northern Hemisphere Armillaria: Neighbor-net and Bayesian analyses of translation elongation factor 1-α gene sequences. Mycologia 2017, 109, 75–91. [Google Scholar] [CrossRef] [PubMed]
- McLaughlin, J. Distribution, hosts, and site relationships of Armillaria spp. in central and southern Ontario. Can. J. For. Res. 2001, 31, 1481–1490. [Google Scholar] [CrossRef]
- Mallett, K. Host range and geographic distribution of Armillaria root rot pathogens in the Canadian prairie provinces. Can. J. For. Res. 1990, 20, 1859–1863. [Google Scholar] [CrossRef]
- Dehelean, C.A.; Şoica, C.; Ledeţi, I.; Aluaş, M.; Zupko, I.; Gǎluşcan, A.; Cinta-Pinzaru, S.; Munteanu, M. Study of the betulin enriched birch bark extracts effects on human carcinoma cells and ear inflammation. Chem. Cent. J. 2012, 6, 137. [Google Scholar] [CrossRef]
- Joseph, A.; Kutty, N.G.; Moorkoth, S.; Alex, A.T. In vitro and In vivo Anticancer Activity of Semisynthetic Derivatives of Betulinic acid. Lat. Am. J. Pharm. 2019, 38, 1582–1590. [Google Scholar]
- Wang, X.; Yuan, Z.; Zhu, L.; Yi, X.; Ou, Z.; Li, R.; Tan, Z.; Pozniak, B.; Obminska-Mrukowicz, B.; Wu, J. Protective Effects of Betulinic Acid on Intestinal Mucosal Injury Induced by Cyclophosphamide in Mice. Pharmacol. Rep. 2019. [Google Scholar] [CrossRef]
- Huang, L.; Li, J.; Ye, H.; Li, C.; Wang, H.; Liu, B.; Zhang, Y. Molecular characterization of the pentacyclic triterpenoid biosynthetic pathway in Catharanthus roseus. Planta 2012, 236, 1571–1581. [Google Scholar] [CrossRef]
- Fukushima, E.O.; Seki, H.; Ohyama, K.; Ono, E.; Umemoto, N.; Mizutani, M.; Saito, K.; Muranaka, T. CYP716A Subfamily Members are Multifunctional Oxidases in Triterpenoid Biosynthesis. Plant Cell Physiol. 2011, 52, 2050–2061. [Google Scholar] [CrossRef]
- Liu, J.; Fu, M.L.; Chen, Q.H. Biotransformation optimization of betulin into betulinic acid production catalysed by cultured Armillaria luteo-virens Sacc ZJUQH100-6 cells. J. Appl. Microbiol. 2011, 110, 90–97. [Google Scholar] [CrossRef]
- Yin, J.; Ma, H.; Gong, Y.; Xiao, J.; Jiang, L.; Zhan, Y.; Li, C.; Ren, C.; Yang, Y. Effect of MeJA and Light on the Accumulation of Betulin and Oleanolic Acid in the Saplings of White Birch (Betula platyphylla Suk.). Am. J. Plant Sci. 2013, 4, 9. [Google Scholar] [CrossRef]
- Mandlik, A.; Livny, J.; Robins, W.P.; Ritchie, J.M.; Mekalanos, J.J.; Waldor, M.K. RNA-Seq-based monitoring of infection-linked changes in Vibrio cholerae gene expression. Cell Host Microbe 2011, 10, 165–174. [Google Scholar] [CrossRef] [PubMed]
- Cao, S.; Zhu, L.; Nie, H.; Yin, M.; Liu, G.; Yan, X. De novo assembly, gene annotation, and marker development using Illumina paired-end transcriptome sequencing in the Crassadoma gigantea. Gene 2018, 658, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Zhan, M.; Tian, M.; Wang, W.; Li, G.; Lu, X.; Cai, G.; Yang, H.; Du, G.; Huang, L. Draft genomic sequence of Armillaria gallica 012m: Insights into its symbiotic relationship with Gastrodia elata. Braz. J. Microbiol. 2020, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Collins, C. A Genomic and Proteomic Investigation of the Plant Pathogen Armillaria mellea: Buried Treasure or Hidden Threat? Ph.D. Thesis, National University of Ireland, Maynooth, Ireland, August 2013. [Google Scholar]
- Sipos, G.; Qi, W.; Künzli, M.; Okoniewski, M.; Rigling, D. 454 sequencing of transcriptomes for virulent and non-virulent Armillaria ostoyae strains and identification of their secretomes. In Proceedings of the XIII Conference”Root and Butt Rot of Forest Trees” IUFRO Working Party 7.02.01, Trento, Italy, 4–11 September 2011; p. 40. [Google Scholar]
- Ross-Davis, A.L.; Stewart, J.E.; Hanna, J.W.; Kim, M.-S.; Cronn, R.C.; Rai, H.S.; Richardson, B.A.; McDonald, G.I.; Klopfenstein, N.B. De novo assembly and transcriptome characterization of an Armillaria solidipes mycelial fan. In Proceedings of the 59th Annual Western International Forest Disease Work Conference, Leavenworth, WA, USA, 11–14 October 2011; Zeglen, S., Palacios, P., Eds.; Department of Agriculture, Forest Service, Forest Health Protection: Washington, DC, USA, 2011; pp. 165–168. [Google Scholar]
- Liu, M.-M.; Xing, Y.-M.; Zhang, D.-W.; Guo, S.-X. Transcriptome analysis of genes involved in defence response in Polyporus umbellatus with Armillaria mellea infection. Sci. Rep. 2015, 5, 16075. [Google Scholar] [CrossRef] [PubMed]
- Engels, B.; Heinig, U.; Grothe, T.; Stadler, M.; Jennewein, S. Cloning and characterization of an Armillaria gallica cDNA encoding protoilludene synthase, which catalyzes the first committed step in the synthesis of antimicrobial melleolides. J. Biol. Chem. 2011, 286, 6871–6878. [Google Scholar] [CrossRef] [PubMed]
- Ming-liang, F.; Jing, L.; Ya-chen, D.; Yu, F.; Ruo-si, F.; Qi-he, C.; Xiao-jie, L. Effect of ionic liquid-containing system on betulinic acid production from betulin biotransformation by cultured Armillaria luteo-virens Sacc cells. Eur. Food Res. Technol. 2011, 233, 507. [Google Scholar] [CrossRef]
- Zhang, H.-F.; He, G.-Q.; Liu, J.; Ruan, H.; Chen, Q.-H.; Zhang, Q.; Wang, J.-L.; Zhang, H.-B. Production of gastrodin through biotransformation of p-2-hydroxybenzyl alcohol by cultured cells of Armillaria luteo-virens Sacc. Enzyme Microbial Technol. 2008, 43, 25–30. [Google Scholar] [CrossRef]
- Fradj, N.; Gonçalves dos Santos, K.C.; de Montigny, N.; Awwad, F.; Boumghar, Y.; Germain, H.; Desgagné-Penix, I. RNA-Seq de Novo Assembly and Differential Transcriptome Analysis of Chaga (Inonotus obliquus) Cultured with Different Betulin Sources and the Regulation of Genes Involved in Terpenoid Biosynthesis. Int. J. Mol. Sci. 2019, 20, 4334. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Brown, C.T.; Howe, A.; Zhang, Q.; Pyrkosz, A.B.; Brom, T.H. A reference-free algorithm for computational normalization of shotgun sequencing data. arXiv 2012, arXiv:1203.4802. [Google Scholar]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Wang, T.; Li, B.; Nelson, C.E.; Nabavi, S. Comparative analysis of differential gene expression analysis tools for single-cell RNA sequencing data. BMC Bioinform. 2019, 20, 40. [Google Scholar] [CrossRef] [PubMed]
- Leinonen, R.; Sugawara, H.; Shumway, M.; International Nucleotide Sequence Database Collaboration. The sequence read archive. Nucleic Acids Res. 2011, 39, D19–D21. [Google Scholar] [CrossRef]
- Dewick, P.M. Medicinal Natural Products: A Biosynthetic Approach; John Wiley & Sons: Hoboken, NJ, USA, 2002. [Google Scholar]
- Thimmappa, R.; Geisler, K.; Louveau, T.; O’Maille, P.; Osbourn, A. Triterpene biosynthesis in plants. Annu. Rev. Plant Biol. 2014, 65, 225–257. [Google Scholar] [CrossRef] [PubMed]
- Cochrane, G.R.; Galperin, M.Y. The 2010 nucleic acids research database issue and online database collection: A community of data resources. Nucleic Acids Res. 2010, 38, D1–D4. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Attwood, T.K.; Babbitt, P.C.; Bateman, A.; Bork, P.; Bridge, A.J.; Chang, H.-Y.; Dosztányi, Z.; El-Gebali, S.; Fraser, M. InterPro in 2017—Beyond protein family and domain annotations. Nucleic Acids Res. 2017, 45, D190–D199. [Google Scholar] [CrossRef] [PubMed]
- Janocha, S.; Schmitz, D.; Bernhardt, R. Terpene hydroxylation with microbial cytochrome P450 monooxygenases. In Biotechnology of Isoprenoids; Springer: New York, NY, USA, 2015; pp. 215–250. [Google Scholar]
- Song, Y.; Yang, X.; Yang, S.; Wang, J. Transcriptome sequencing and functional analysis of Sedum lineare Thunb. Upon salt stress. Mol. Genet. Genom. 2019, 294, 1–13. [Google Scholar] [CrossRef]
- Prall, W.; Hendy, O.; Thornton, L.E. Utility of a phylogenetic perspective in structural analysis of CYP72A enzymes from flowering plants. PLoS ONE 2016, 11, e0163024. [Google Scholar] [CrossRef]
- Cao, J.; Wang, B.; Tan, X. Transcriptomic responses of the clam Meretrix meretrix to the organophosphorus pesticide (dimethoate). Ecotoxicology 2019, 28, 1–11. [Google Scholar] [CrossRef]
- Kuhad, R.C.; Deswal, D.; Sharma, S.; Bhattacharya, A.; Jain, K.K.; Kaur, A.; Pletschke, B.I.; Singh, A.; Karp, M. Revisiting cellulase production and redefining current strategies based on major challenges. Renew. Sustain. Energy Rev. 2016, 55, 249–272. [Google Scholar] [CrossRef]
- Jain, K.K.; Kumar, A.; Shankar, A.; Pandey, D.; Chaudhary, B.; Sharma, K.K. De novo transcriptome assembly and protein profiling of copper-induced lignocellulolytic fungus Ganoderma lucidum MDU-7 reveals genes involved in lignocellulose degradation and terpenoid biosynthetic pathways. Genomics 2019. [Google Scholar] [CrossRef] [PubMed]
- Bugg, T.D.; Ahmad, M.; Hardiman, E.M.; Rahmanpour, R. Pathways for degradation of lignin in bacteria and fungi. Nat. Prod. Rep. 2011, 28, 1883–1896. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Ma, F.; Zhao, H.; Zhang, S.; Wang, L.; Zhang, X.; Yu, H. A white-rot fungal lytic polysaccharide monooxygenase drives the degradation of lignin by a versatile peroxidase. Appl. Environ. Microbiol. 2019. [Google Scholar] [CrossRef]
- Dimarogona, M.; Topakas, E.; Olsson, L.; Christakopoulos, P. Lignin boosts the cellulase performance of a GH-61 enzyme from Sporotrichum thermophile. Bioresour. Technol. 2012, 110, 480–487. [Google Scholar] [CrossRef]
- Esquerré-Tugayé, M.-T.; Boudart, G.; Dumas, B. Cell wall degrading enzymes, inhibitory proteins, and oligosaccharides participate in the molecular dialogue between plants and pathogens. Plant Physiol. Biochem. 2000, 38, 157–163. [Google Scholar] [CrossRef]
- Miyara, I.; Shafran, H.; Kramer Haimovich, H.; Rollins, J.; Sherman, A.; Prusky, D. Multi-factor regulation of pectate lyase secretion by Colletotrichum gloeosporioides pathogenic on avocado fruits. Mol. Plant Pathol. 2008, 9, 281–291. [Google Scholar] [CrossRef]
- Drori, N.; Kramer-Haimovich, H.; Rollins, J.; Dinoor, A.; Okon, Y.; Pines, O.; Prusky, D. External pH and nitrogen source affect secretion of pectate lyase by Colletotrichum gloeosporioides. Appl. Environ. Microbiol. 2003, 69, 3258–3262. [Google Scholar] [CrossRef]
- Tardy, F.; Nasser, W.; Robert-Baudouy, J.; Hugouvieux-Cotte-Pattat, N. Comparative analysis of the five major Erwinia chrysanthemi pectate lyases: Enzyme characteristics and potential inhibitors. J. Bacteriol. 1997, 179, 2503–2511. [Google Scholar] [CrossRef]
- Tang, Y.; Wu, P.; Jiang, S.; Selvaraj, J.N.; Yang, S.; Zhang, G. A new cold-active and alkaline pectate lyase from Antarctic bacterium with high catalytic efficiency. Appl. Microbiol. Biotechnol. 2019, 103, 1–11. [Google Scholar] [CrossRef] [PubMed]

| Read Trimming and Clipping of Adapters | |
| Raw paired reads a | 170,592,464 |
| Surviving paired reads b | 166,006,772 |
| Surviving paired reads (%) c | 97.31 |
| Trinity De Novo Assembly | |
| Nb. Transcriptsd | 273,561 |
| Nb. Componentsd | 99,358 |
| Total Transcripts Length (bp) | 561,145,023 |
| Max. Transcript Length (bp) | 16,405 |
| Min. Transcript Length (bp) | 201 |
| Median Transcript Length (bp) | 1456 |
| Mean Transcript Length (bp) | 2051 |
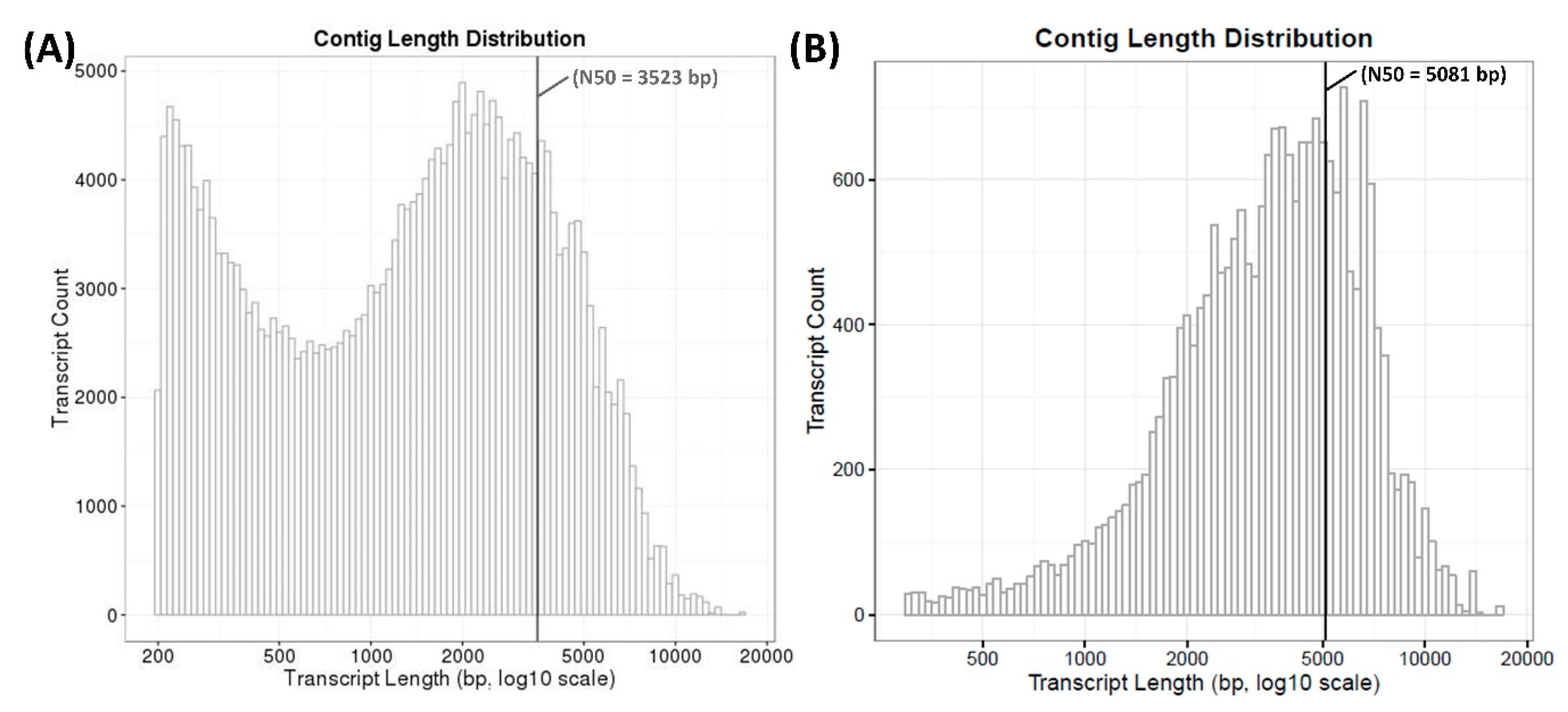
| N50 (bp) | 3523 |
| BLAST Annotation and Filtered Annotated Components | |
| Nb. Transcripts | 121,959 |
| Nb. Components | 13,544 |
| Total Transcripts Length (bp) | 288,576,284 |
| Max. Transcript Length (bp) | 16,405 |
| Min. Transcript Length (bp) | 298 |
| Median Transcript Length (bp) | 3661 |
| Mean Transcript Length (bp), | 4033 |
| N50 (bp) | 5081 |
| Public Database | Number of Transcripts | Percentage (%) |
|---|---|---|
| Blast-GO annotations | 103,267 | 37.75 |
| COG annotations | 61,272 | 23.53 |
| Pfam annotations | 45,683 | 16.69 |
| KEGG annotations | 105,195 | 40.39 |
| Total number of sequences not annotated | 164,622 | 60.18 |
| Total number of sequences annotated | 108,939 | 39.82 |
| Total of transcript sequences | 273,561 | 100.00 |
| Set | Differentially Expressed Transcripts | Up-Regulated | Down-Regulated |
|---|---|---|---|
| BET vs. Control | 3943 | 3863 | 80 |
| Rank | Log2 FC | Description | Species | E-Value | Accession |
|---|---|---|---|---|---|
| 1 | 10.96 | 3-ketoacyl-CoA thiolase 5 | Grifola fondosa | 0 | OBZ79350.1 |
| 2 | 10.92 | NAD-P-binding protein | Trametes versicolor | 0 | XP_008044855.1 |
| 3 | 10.82 | Hypothetical protein | Trametes versicolor | 0 | XP_008042994.1 |
| 4 | 10.73 | No hit | |||
| 5 | 10.68 | C4-methyl sterol oxidase | Trametes versicolor | 0 | XP_008044057.1 |
| 6 | 10.64 | Golgi apparatus membrane protein TVP38 | Trametes pubescens | 0 | OJT05825.1 |
| 7 | 10.56 | Hypothetical protein | Trametes versicolor | 0 | XP_008035434.1 |
| 8 | 10.55 | Hypothetical protein | Trametes versicolor | 0 | XP_008034798.1 |
| 9 | 10.51 | HSP20-like chaperone | Trametes versicolor | 0 | XP_008040595.1 |
| 10 | 10.51 | NADPH oxidase isoform 2 | Trametes versicolor | 0 | XP_008043698.1 |
| 11 | 10.44 | GroES-like protein | Trametes versicolor | 0 | XP_008042467.1 |
| 12 | 10.44 | Heat shock protein | Trametes versicolor | 0 | XP_008036836.1 |
| 13 | 10.33 | Phosphoglycerate kinase | Trametes versicolor | 0 | XP_008036919.1 |
| 14 | 10.32 | Heat shock protein 70 | Trametes versicolor | 0 | XP_008040622.1 |
| 15 | 10.31 | Hypothetical protein | Trametes versicolor | 0 | XP_008034620.1 |
| 16 | 10.29 | NAD-P-binding protein | Trametes versicolor | 0 | XP_008044855.1 |
| 17 | 10.28 | Hypothetical protein | Trametes versicolor | 0 | XP_008042390.1 |
| 18 | 10.27 | Hypothetical protein | Trametes versicolor | 0 | XP_008042803.1 |
| 19 | 10.25 | Hypothetical protein | Trametes versicolor | 0 | XP_008041395.1 |
| 20 | 10.23 | Calnexin-like protein | Trametes pubescens | 0 | OJT08454.1 |
| 21 | 10.21 | No hit | |||
| 22 | 10.21 | Heat shock protein 30 | Trametes versicolor | 0 | XP_008045263.1 |
| 23 | 10.20 | Pyruvate decarboxylase | Trametes versicolor | 0 | XP_008037637.1 |
| 24 | 10.17 | No hit | |||
| 25 | 10.15 | Hypothetical protein | Trametes versicolor | 0 | XP_008037020.1 |
| 26 | 10.08 | Hypothetical protein | Trametes versicolor | 0 | XP_008045579.1 |
| 27 | 10.08 | Fumarate reductase | Trametes pubescens | 0 | OJT15326.1 |
| 28 | 10.06 | Acid protease | Trametes versicolor | 0 | XP_008034004.1 |
| 29 | 10.06 | No hit | |||
| 30 | 10.06 | Glycoside hydrolase | Trametes versicolor | 0 | XP_008038270.1 |
| 31 | 10.06 | Hypothetical protein | Trametes versicolor | 0 | XP_008035168.1 |
| 32 | 10.05 | No hit | |||
| 33 | 10.04 | No hit | |||
| 34 | 10.01 | Phenylalanine-t-RNA synthethase | Trametes versicolor | 0 | XP_008035697.1 |
| 35 | 10.01 | No hit | |||
| 36 | 9.99 | No hit | |||
| 37 | 9.98 | Hypothetical protein | Trametes versicolor | 0 | XP_008045519.1 |
| 38 | 9.96 | Hypothetical protein | Trametes versicolor | 0 | XP_008040976.1 |
| 39 | 9.96 | No hit | |||
| 40 | 9.94 | Hypothetical protein | Trametes pubescens | 0 | OJT02899.1 |
| 41 | 9.93 | Aspartic peptidase A1 | Trametes versicolor | 0 | XP_008035722.1 |
| 42 | 9.93 | Heat shock protein 20-like chaperone | Trametes versicolor | 0 | XP_008037816.1 |
| 43 | 9.91 | Cytochrome P450 | Trametes versicolor | 0 | XP_008033952.1 |
| 44 | 9.91 | Hypothetical protein | Trametes versicolor | 0 | XP_008035218.1 |
| 45 | 9.88 | Delta 9-fatty acid desaturase | Trametes versicolor | 0 | XP_008041237.1 |
| 46 | 9.88 | No hit | |||
| 47 | 9.88 | Phosphopyruvate hydratase | Trametes versicolor | 0 | XP_008032334.1 |
| 48 | 9.88 | Ceramide very long chain fatty acid hydroxylase | Trametes pubescens | 0 | OJT03476.1 |
| 49 | 9.86 | No hit | |||
| 50 | 9.86 | No hit | |||
| Rank | Log2 FC | Description | Species | E-Value | Accession |
|---|---|---|---|---|---|
| 1 | −6.26 | AFG1-like ATPase | Armillaria gallica | 0 | PBL02956.1 |
| 2 | −6.23 | Pectate lyase | Armillaria gallica | 0 | PBL03123.1 |
| 3 | −5.82 | Putative O-fucosyltransferease-like protein | Moniliophtora roreri | 1 E-124 | KTB33430.1 |
| 4 | −5.45 | Hypothetical protein | Armillaria solidipes | 9 E-84 | PBK63479.1 |
| 5 | −5.38 | Uncharacterized protein | Armillaria ostoyae | 1 E-147 | SJL03005.1 |
| 6 | −5.32 | Uncharacterized protein | Armillaria ostoyae | 1 E-146 | SJL03005.1 |
| 7 | −5.08 | Hexose transporter | Armillaria gallica | 0 | PBK93947.1 |
| 8 | −4.88 | Glycosyltransferase | Armillaria solidipes | 0 | PBK58814.1 |
| 9 | −4.74 | Uncharacterized protein | Armillaria ostoyae | 0 | SJL10357.1 |
| 10 | −4.65 | Casein kinase II beta subunit | Armillaria gallica | 0 | PBL02672.1 |
| 11 | −3.47 | Hypothetical protein | Armillaria gallica | 0 | PBL00259.1 |
| 12 | −3.42 | No hit | |||
| 13 | −3.36 | Heavy metal translocatin | Armillaria solidipes | 0 | PBK63629.1 |
| 14 | −3.31 | Hypothetical protein | Armillaria solidipes | 0 | PBK76216.1 |
| 15 | −3.25 | WD40 repeat-like protein | Armillaria solidipes | 5 E-168 | PBK69270.1 |
| 16 | −3.16 | Kinase-like protein | Armillaria solidipes | 4 E-129 | PBK63656.1 |
| 17 | −3.02 | Hypothetical protein | Armillaria gallica | 0 | PBK93574.1 |
| 18 | −2.97 | Nicotinamide mononucleotide related permease | Armillaria ostoyae | 0 | SJL10502.1 |
| 19 | −2.97 | Alpha-aminoadipate reductase Lys1p | Armillaria gallica | 0 | PBK98805.1 |
| 20 | −2.91 | Hypothetical protein | Armillaria solidipes | 9 E-64 | PBK75291.1 |
| 21 | −2.84 | Hypothetical protein | Armillaria solidipes | 0 | PBK69763.1 |
| 22 | −2.66 | Cytochrome P450 | Armillaria gallica | 2 E-133 | PBK89827.1 |
| 23 | −2.63 | Kinase-like protein | Armillaria solidipes | 0 | PBK71971.1 |
| 24 | −2.59 | Hypothetical protein | Armillaria gallica | 0 | PBL01386.1 |
| 25 | −2.39 | Uncharacterized protein | Armillaria ostoyae | 0 | SJL04719.1 |
| 26 | −2.37 | Hypothetical protein | Armillaria gallica | 0 | PBK93754.1 |
| 27 | −2.26 | No hit | |||
| 28 | −2.23 | NAD-dependent deacylase | Armillaria ostoyae | 2 E-90 | SJL08753.1 |
| 29 | −2.20 | No hit | |||
| 30 | −2.16 | No hit | |||
| 31 | −2.12 | Hypothetical protein | Armillaria gallica | 3 E-70 | PBK96234.1 |
| 32 | −2.11 | Uncharacterized protein | Armillaria ostoyae | 1 E-149 | SJL09942.1 |
| 33 | −2.08 | No hit | |||
| 34 | −2.02 | No hit | |||
| 35 | −2.01 | No hit | |||
| 36 | −1.99 | No hit | |||
| 37 | −1.96 | HOOK-domain-containing protein | Armillaria solidipes | 0 | PBK77734.1 |
| 38 | −1.94 | DNA excision repair protein | Hypsizygus marmoreus | 0 | RDB19634.1 |
| 39 | −1.94 | Uncharacterized protein | Armillaria ostoyae | 0 | SJK99761.1 |
| 40 | −1.81 | BRO1-domain-containing protein | Armillaria gallica | 0 | PBK82405.1 |
| 41 | −1.78 | Ribonuclease subunit | Armillaria ostoyae | 0 | SJL08201.1 |
| 42 | −1.75 | Hypothetical protein | Armillaria solidipes | 0 | PBK63259.1 |
| 43 | −1.67 | No hit | |||
| 44 | −1.67 | Uncharacterized protein | Armillaria ostoyae | 0 | SJL12868.1 |
| 45 | −1.64 | BRO1-domain-containing protein | Armillaria gallica | 0 | PBK82405.1 |
| 46 | −1.62 | Uncharacterized protein | Armillaria ostoyae | 0 | SJL12868.1 |
| 47 | −1.61 | Uncharacterized protein | Armillaria ostoyae | 0 | SJL09034.1 |
| 48 | −1.60 | L-gulonolactone D-arabinono-1,4-lactone oxidase | Armillaria solidipes | 0 | PBK71727.1 |
| 49 | −1.59 | Vacuole import and degradation VID27 | Armillaria solidipes | 0 | PBK69736.1 |
| 50 | −1.58 | Uncharacterized protein | Armillaria ostoyae | 0 | SJL09949.1 |
| Transcript Name | Log2FC | Annotation |
|---|---|---|
| Terpenoids | ||
| Armillaria_TRINITY_DN29150_c0_g1_i1 | 10.68 | Methylsterol monooxygenase |
| Armillaria_TRINITY_DN22917_c0_g1_i3 | 9.00 | Lanosterol synthase |
| Armillaria_TRINITY_DN3215_c0_g1_i1 | 8.85 | C-5 sterol desaturase |
| Armillaria_TRINITY_DN31064_c0_g1_i13 | 8.70 | Lanosterol 14-alpha demethylase |
| Armillaria_TRINITY_DN6566_c0_g1_i2 | 8.27 | Sterol 24-C-methyltransferase |
| Armillaria_TRINITY_DN34616_c0_g1_i1 | 7.28 | Alpha-muurolene synthase |
| Armillaria_TRINITY_DN17473_c0_g1_i1 | 7.22 | Lanosterol 14-alpha demethylase |
| Armillaria_TRINITY_DN45244_c0_g1_i1 | 7.17 | Squalene epoxidase |
| Armillaria_TRINITY_DN18035_c0_g1_i1 | 7.13 | Delta(14)-sterol reductase |
| Armillaria_TRINITY_DN45179_c0_g1_i1 | 6.96 | Squalene synthase |
| Armillaria_TRINITY_DN54064_c0_g1_i1 | 6.94 | Sterol-4-alpha-carboxylate 3-dehydrogenase |
| Armillaria_TRINITY_DN13258_c0_g1_i1 | 6.93 | Sterol 14-demethylase |
| Armillaria_TRINITY_DN12757_c0_g1_i3 | 6.91 | Alpha-muurolene synthase |
| Armillaria_TRINITY_DN10601_c0_g1_i1 | 6.78 | Farnesyltransferase/geranylgeranyltransferase subunit alpha |
| Armillaria_TRINITY_DN10022_c0_g1_i1 | 6.78 | Isopentenyl-diphosphate Delta-isomerase |
| Armillaria_TRINITY_DN4217_c0_g1_i1 | 6.75 | Delta(24(24(1)))-sterol reductase |
| Armillaria_TRINITY_DN55954_c0_g1_i1 | 6.71 | C-8 sterol isomerase |
| Armillaria_TRINITY_DN9929_c0_g1_i1 | 6.66 | Diphosphomevalonate decarboxylase |
| CYP450s | ||
| Armillaria_TRINITY_DN20121_c0_g1_i1 | 9.91 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN8574_c0_g1_i1 | 8.67 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN61797_c0_g1_i1 | 8.50 | Cytochrome P450 4F5 |
| Armillaria_TRINITY_DN5903_c0_g1_i1 | 8.28 | Cytochrome P450 67 |
| Armillaria_TRINITY_DN4166_c0_g1_i1 | 7.74 | Cytochrome P450 61 |
| Armillaria_TRINITY_DN31064_c0_g1_i10 | 7.60 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN23798_c1_g2_i3 | 7.44 | Docosahexaenoic acid omega-hydroxylase CYP4F3 |
| Armillaria_TRINITY_DN31064_c0_g1_i5 | 7.21 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN23798_c1_g2_i1 | 7.16 | Docosahexaenoic acid omega-hydroxylase CYP4F3 |
| Armillaria_TRINITY_DN9123_c0_g1_i1 | 7.14 | Cytochrome P450 4F22 |
| Armillaria_TRINITY_DN23798_c1_g1_i1 | 7.00 | Cytochrome P450 3A9 |
| Armillaria_TRINITY_DN56547_c0_g1_i1 | 6.97 | Putative cytochrome P450 CYP13A10 |
| Armillaria_TRINITY_DN153_c0_g1_i1 | 6.95 | Cytochrome P450 3A24 |
| Armillaria_TRINITY_DN13346_c0_g1_i1 | 6.95 | Cytochrome P450 52E1 |
| Armillaria_TRINITY_DN7413_c0_g2_i1 | 6.90 | Cytochrome P450 3A24 |
| Armillaria_TRINITY_DN9340_c0_g1_i1 | 6.89 | Cytochrome P450 4A4 |
| Armillaria_TRINITY_DN31064_c0_g1_i27 | 6.89 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN14362_c0_g1_i1 | 6.86 | Taurochenodeoxycholic 6 alpha-hydroxylase |
| Armillaria_TRINITY_DN21389_c0_g1_i1 | 6.85 | Docosahexaenoic acid omega-hydroxylase CYP4F3 |
| Armillaria_TRINITY_DN31064_c0_g1_i3 | 6.71 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN6971_c0_g1_i1 | 6.65 | Putative cytochrome P450 CYP13A8 |
| Armillaria_TRINITY_DN3618_c0_g1_i1 | 6.58 | Cytochrome P450 72A15 |
| Armillaria_TRINITY_DN38009_c0_g1_i1 | 6.57 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN31064_c0_g1_i21 | 6.50 | Cytochrome P450 72A14 |
| Armillaria_TRINITY_DN30372_c0_g1_i50 | 4.07 | Cytochrome P450 98A1 |
| Armillaria_TRINITY_DN32870_c0_g1_i19 | 3.84 | Fumitremorgin C synthase |
| Armillaria_TRINITY_DN32870_c0_g1_i5 | 3.63 | O-methylsterigmatocystin oxidoreductase |
| Armillaria_TRINITY_DN32011_c0_g1_i15 | 1.69 | Docosahexaenoic acid omega-hydroxylase CYP4F3 |
| Armillaria_TRINITY_DN33108_c1_g2_i4 | 1.55 | Leukotriene-B4 omega-hydroxylase 3 |
| Armillaria_TRINITY_DN31587_c1_g1_i22 | 1.02 | Cytochrome P450 4F1 |
| Armillaria_TRINITY_DN33118_c0_g1_i13 | 0.99 | Cytochrome P450 52A6 |
| Armillaria_TRINITY_DN33118_c0_g1_i20 | 0.93 | Cytochrome P450 52A6 |
| Cell Wall Enzymes | ||
| Armillaria_TRINITY_DN19168_c0_g1_i1 | 9.19 | Probable feruloyl esterase A |
| Armillaria_TRINITY_DN31438_c0_g4_i1 | 8.93 | Laccase |
| Armillaria_TRINITY_DN22833_c0_g1_i1 | 8.87 | Glucan 1,3-beta-glucosidase D |
| Armillaria_TRINITY_DN20665_c0_g1_i1 | 8.76 | Beta-mannosidase A |
| Armillaria_TRINITY_DN8404_c0_g1_i1 | 8.75 | Probable glucan 1,3-beta-glucosidase D |
| Armillaria_TRINITY_DN21709_c0_g1_i2 | 8.68 | Polysaccharide monooxygenase |
| Armillaria_TRINITY_DN44356_c0_g1_i1 | 7.99 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase |
| Armillaria_TRINITY_DN15187_c0_g1_i1 | 7.52 | Glucosidase 2 subunit beta |
| Armillaria_TRINITY_DN2359_c0_g1_i1 | 7.42 | Invertase 2 |
| Armillaria_TRINITY_DN20388_c0_g1_i4 | 7.17 | Probable feruloyl esterase B-1 |
| Armillaria_TRINITY_DN31438_c0_g6_i2 | 7.09 | Laccase-1 |
| Armillaria_TRINITY_DN13563_c0_g1_i2 | 7.06 | Probable glucan endo-1,3-beta-glucosidase |
| Armillaria_TRINITY_DN16550_c0_g1_i1 | 7.05 | Glucosidase 2 subunit alpha |
| Armillaria_TRINITY_DN43871_c0_g1_i1 | 7.04 | Glucan endo-1,3-alpha-glucosidase agn1 |
| Armillaria_TRINITY_DN3408_c0_g1_i1 | 7.04 | Glycogen debranching enzyme |
| Armillaria_TRINITY_DN31438_c0_g3_i1 | 6.97 | Laccase-2 |
| Armillaria_TRINITY_DN71958_c0_g1_i1 | 6.94 | Probable glucan 1,3-beta-glucosidase D |
| Armillaria_TRINITY_DN11046_c0_g1_i2 | 6.91 | Glucan 1,3-beta-glucosidase |
| Armillaria_TRINITY_DN955_c0_g1_i1 | 6.83 | Alpha-mannosidase |
| Armillaria_TRINITY_DN55139_c0_g1_i1 | 6.82 | Probable alpha/beta-glucosidase |
| Armillaria_TRINITY_DN24913_c0_g1_i1 | 6.81 | Mannosyl-oligosaccharide alpha-1,2-mannosidase |
| Armillaria_TRINITY_DN33885_c0_g1_i1 | 6.78 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase |
| Armillaria_TRINITY_DN11477_c0_g1_i1 | 6.68 | Glucan 1,3-beta-glucosidase |
| Armillaria_TRINITY_DN38622_c0_g1_i1 | 6.63 | Uncharacterized family 31 glucosidase |
| Armillaria_TRINITY_DN12447_c0_g1_i1 | 6.55 | Probable endo-beta-1,4-glucanase D |
| Armillaria_TRINITY_DN13227_c0_g2_i1 | 6.55 | Probable glucan 1,3-beta-glucosidase D |
| Armillaria_TRINITY_DN13426_c0_g1_i1 | 6.55 | Versatile peroxidase VPL1 |
| Armillaria_TRINITY_DN13563_c0_g1_i1 | 6.54 | Probable glucan endo-1,3-beta-glucosidase |
| Armillaria_TRINITY_DN12447_c0_g1_i2 | 6.53 | Probable endo-beta-1,4-glucanase D |
| Armillaria_TRINITY_DN15833_c0_g1_i1 | 6.49 | Probable beta-glucosidase A |
| Armillaria_TRINITY_DN32420_c0_g1_i1 | 2.03 | Xyloglucanase |
| Armillaria_TRINITY_DN31740_c0_g1_i23 | −5.82 | Pectate lyase A |
| Armillaria_TRINITY_DN31740_c0_g1_i15 | −6.23 | Pectate lyase A |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fradj, N.; de Montigny, N.; Mérindol, N.; Awwad, F.; Boumghar, Y.; Germain, H.; Desgagné-Penix, I. A First Insight into North American Plant Pathogenic Fungi Armillaria sinapina Transcriptome. Biology 2020, 9, 153. https://doi.org/10.3390/biology9070153
Fradj N, de Montigny N, Mérindol N, Awwad F, Boumghar Y, Germain H, Desgagné-Penix I. A First Insight into North American Plant Pathogenic Fungi Armillaria sinapina Transcriptome. Biology. 2020; 9(7):153. https://doi.org/10.3390/biology9070153
Chicago/Turabian StyleFradj, Narimene, Nicolas de Montigny, Natacha Mérindol, Fatima Awwad, Yacine Boumghar, Hugo Germain, and Isabel Desgagné-Penix. 2020. "A First Insight into North American Plant Pathogenic Fungi Armillaria sinapina Transcriptome" Biology 9, no. 7: 153. https://doi.org/10.3390/biology9070153
APA StyleFradj, N., de Montigny, N., Mérindol, N., Awwad, F., Boumghar, Y., Germain, H., & Desgagné-Penix, I. (2020). A First Insight into North American Plant Pathogenic Fungi Armillaria sinapina Transcriptome. Biology, 9(7), 153. https://doi.org/10.3390/biology9070153





