Molecular Aggregation in Immune System Activation Studied by Dynamic Light Scattering
Abstract
1. Introduction
2. Materials and Methods
2.1. Dynamic Light Scattering Spectrometer
2.2. Experimental Samples
3. Results
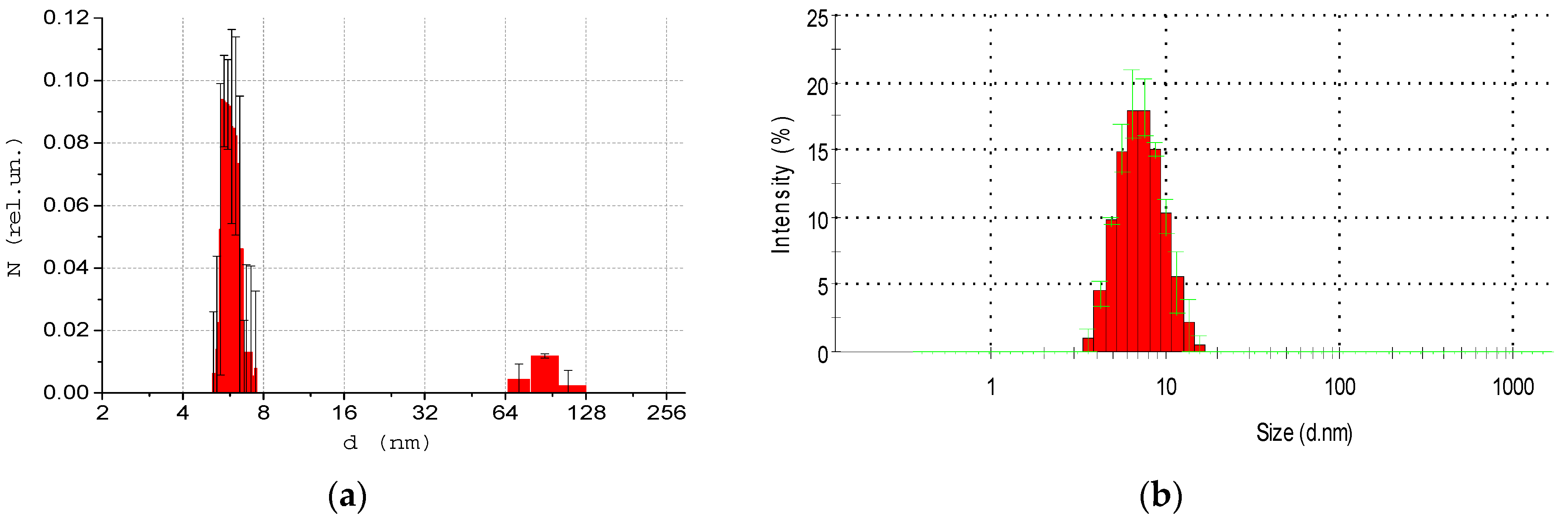
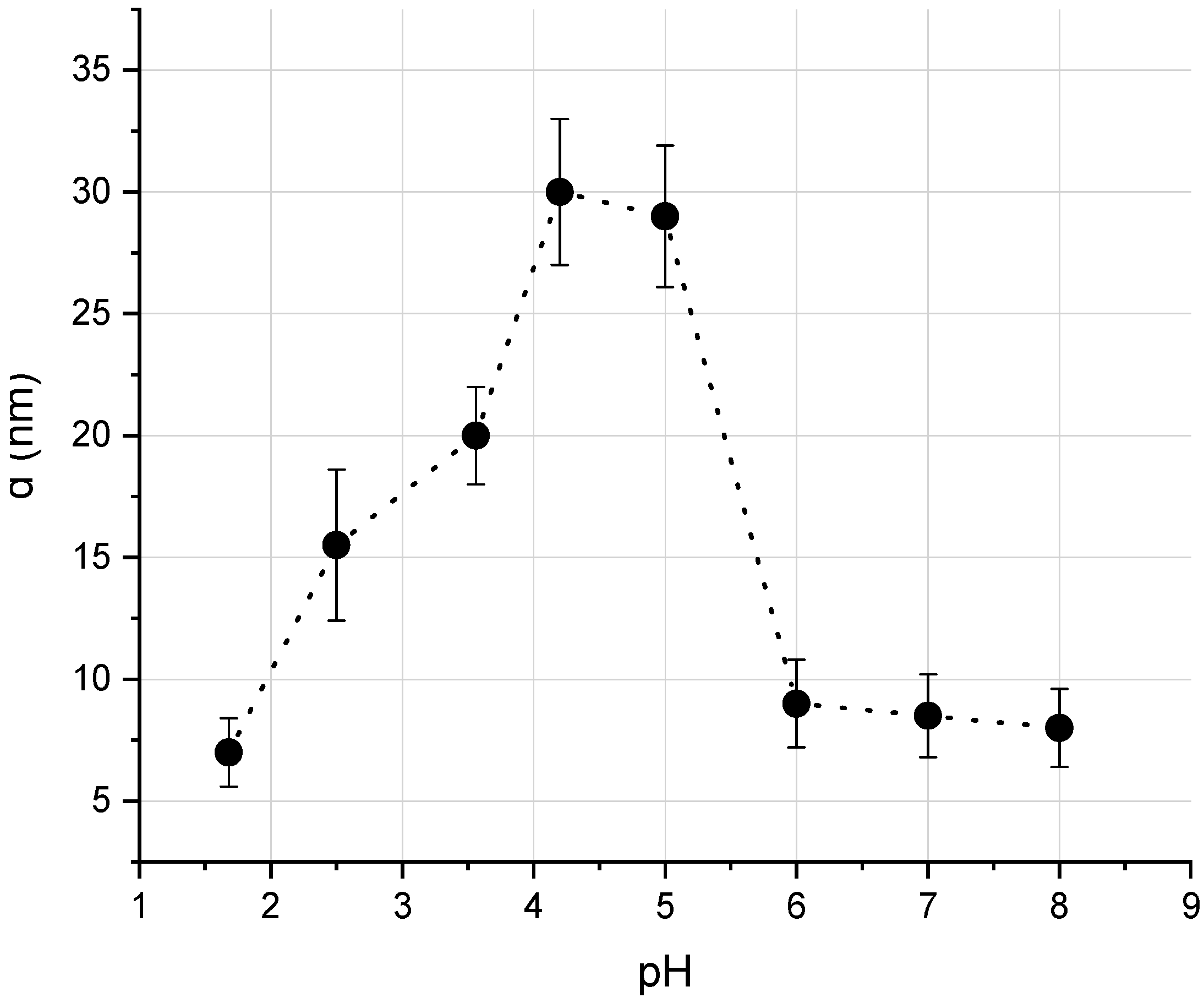
3.1. Solutions of Albumin
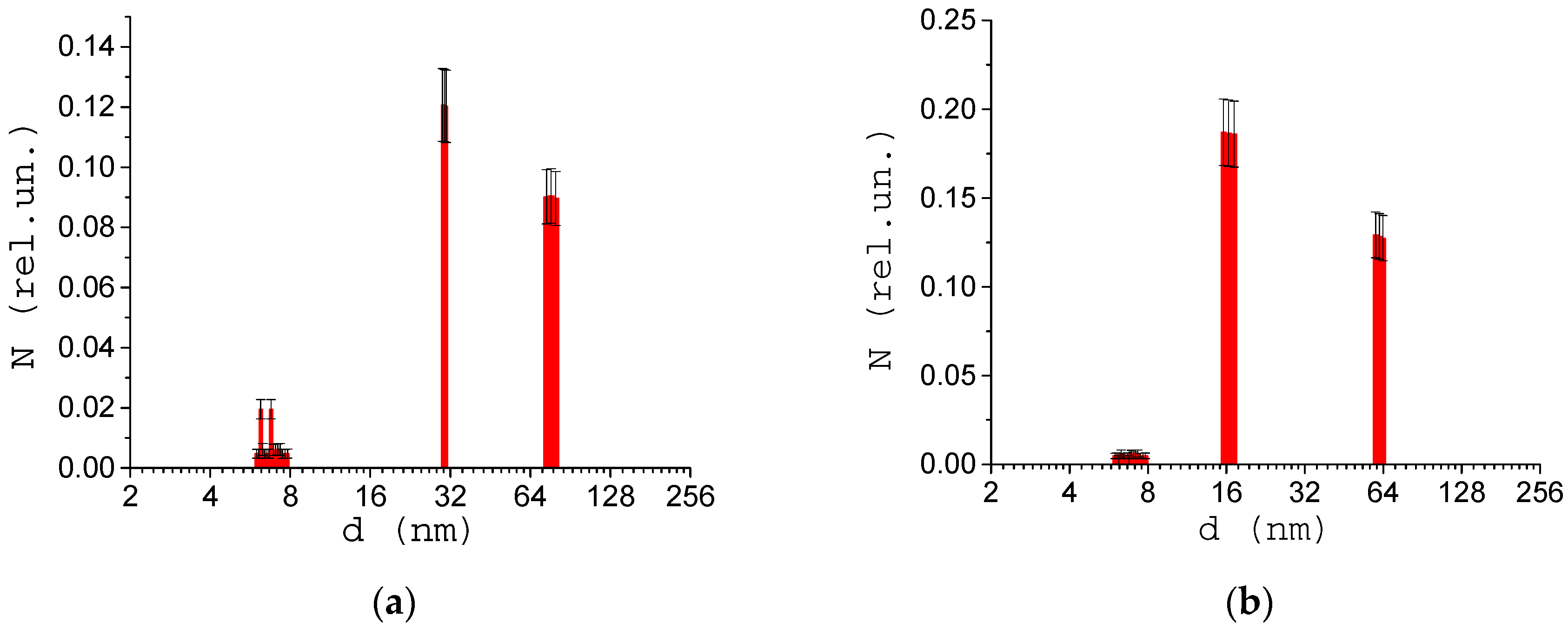
3.2. Size of Structures in the Blood Serum
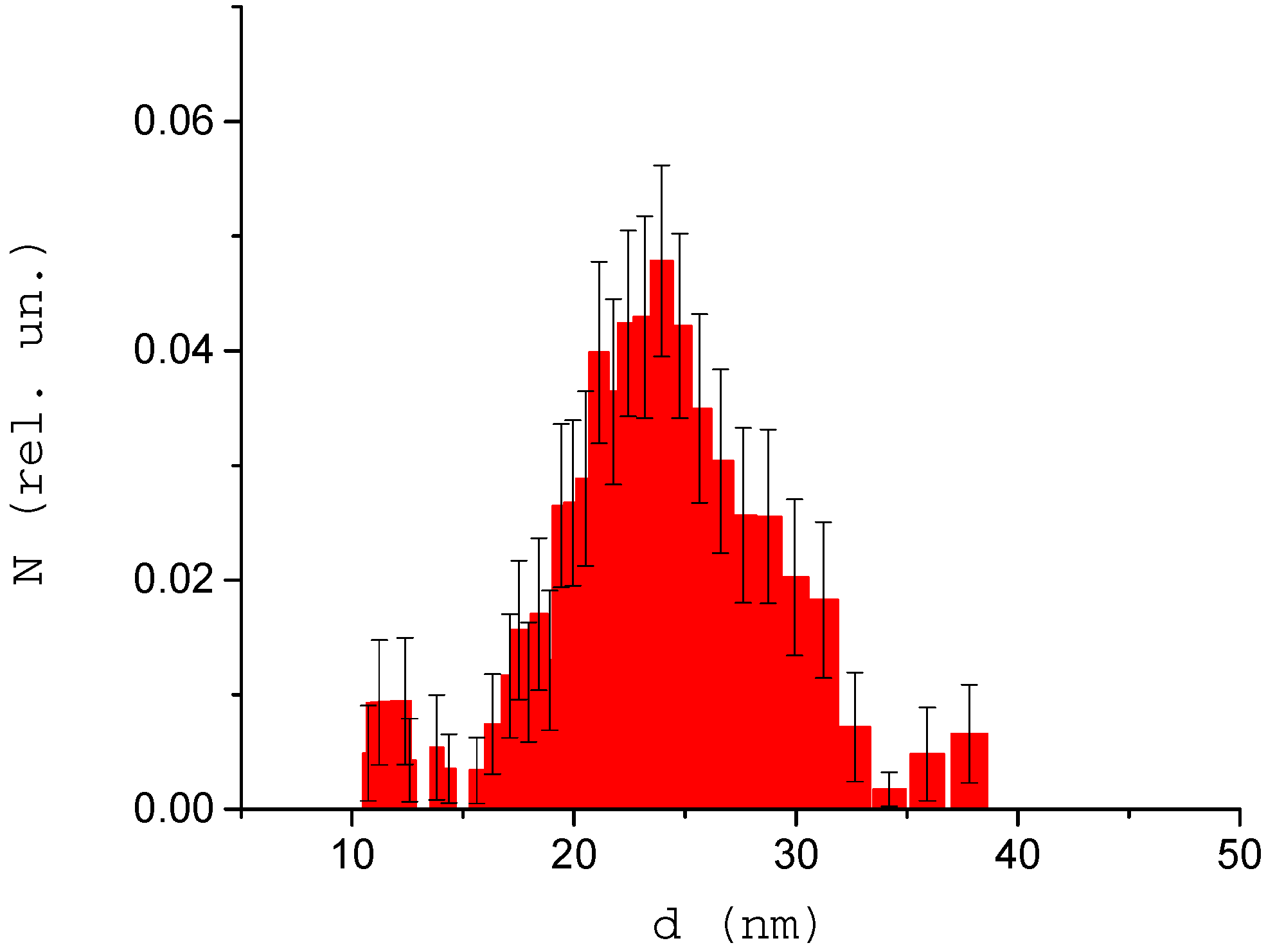
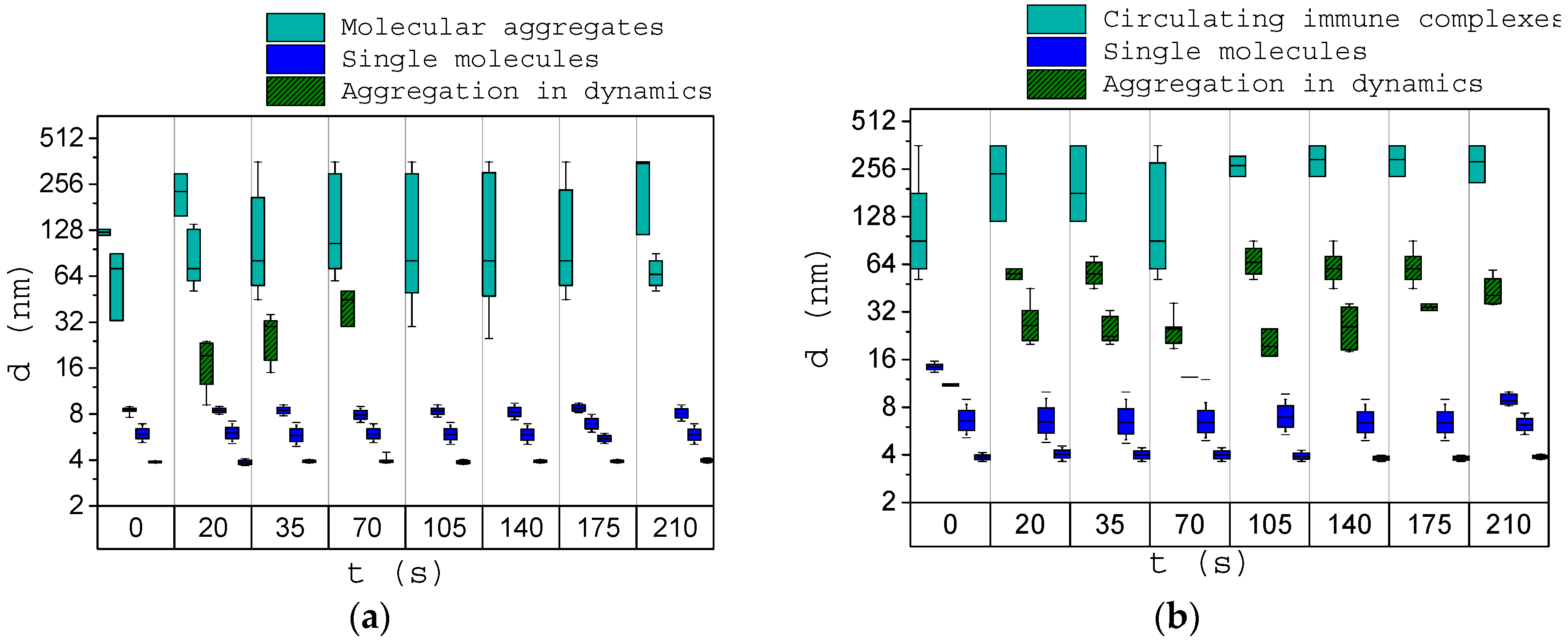
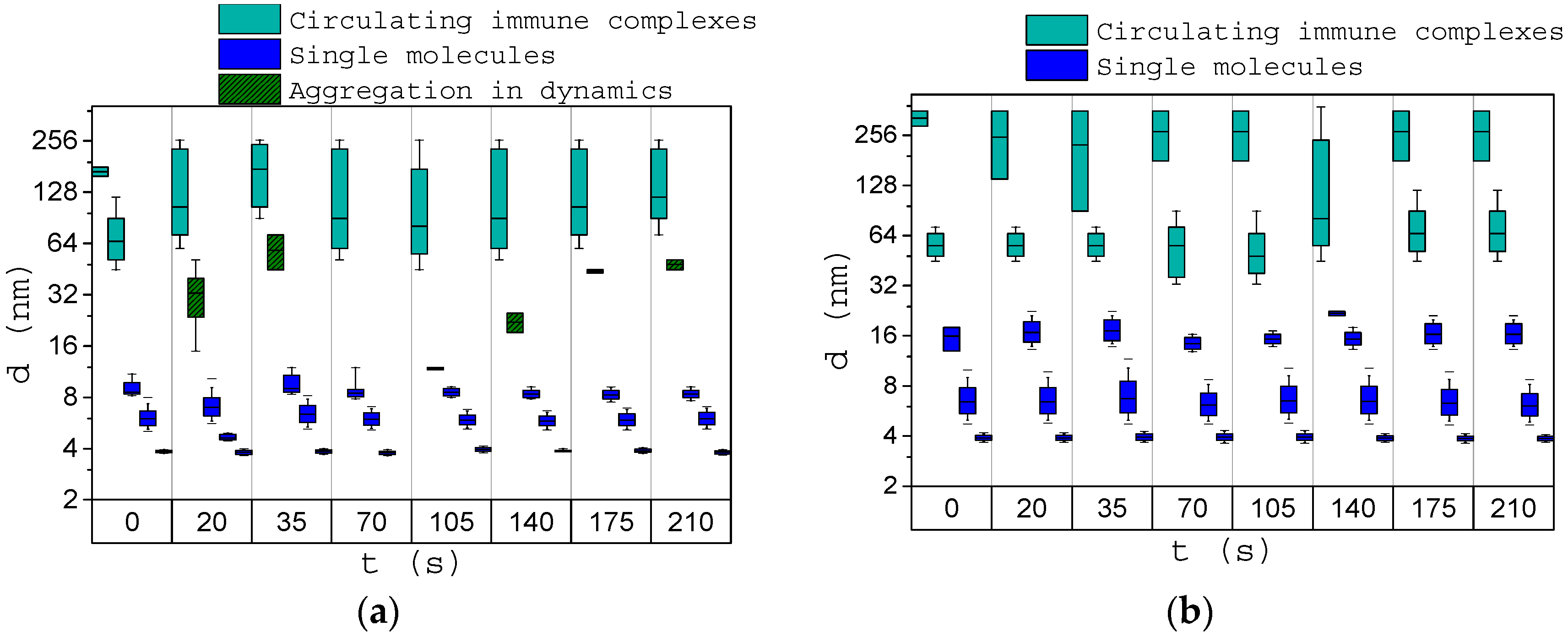
3.3. Analysis of the Dynamics of Immune Activation
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gopalakrishnan, V.; Helmink, B.A.; Spencer, C.N.; Reuben, A.; Wargo, J.A. The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 2018, 33, 570–580. [Google Scholar] [CrossRef] [PubMed]
- Finn, O.J. Immuno-oncology: Understanding the function and dysfunction of the immune system in cancer. Ann. Oncol. 2012, 23 (Suppl. 8), viii6–viii9. [Google Scholar] [CrossRef] [PubMed]
- Brodin, P.; Davis, M.M. Human immune system variation. Nat. Rev. Immunol. 2017, 17, 21–29. [Google Scholar] [CrossRef]
- Nuriddinovna, S.K.; Nigmatovna, A.M. Formation of circulating immune complexes (CIC) and immunoglobulins in intrauterine infection (IUI). Eur. Sci. Rev. 2016, 9–10, 134–135. [Google Scholar]
- Barnett, E.V.; Knutson, D.W.; Abrass, C.K.; Chia, D.S.; Young, L.S.; Liebling, M.R. Circulating immune complexes: Their immunochemistry, detection, and importance. Ann. Intern. Med. 1979, 91, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Tsvilihovskyy, V.; Vovk, N.; Gavrilova, T. Method of immunoferment analysis for determination of lysozyme of mucus and fish tissues. Ukr. J. Ecol. 2017, 7, 210–219. [Google Scholar] [CrossRef][Green Version]
- Li, J.; Cheng, B.; Yang, L.; Zhao, Y.; Pan, M.; Zheng, G.; Cai, Y. Development and implementation of autoverification rules for ELISA results of HBV serological markers. J. Lab. Autom. 2016, 21, 642–651. [Google Scholar] [CrossRef]
- Guo, Y.; Huo, Z.; Wang, Z.; Liu, B. Analysis of three serological tests results of hemolytic disease of newborn in 1 350 cases. Chin. J. Immunol. 2016, 32, 1357–1359. [Google Scholar]
- Atkins, C.G.; Buckley, K.; Blades, M.W.; Turner, R.F.B. Raman spectroscopy of blood and blood components. Appl. Spectrosc. 2017, 71, 767–793. [Google Scholar] [CrossRef]
- Alekseev, S.G.; Ivanov, A.V.; Sviridov, S.V.; Petrova, G.P.; Petrusevich, Y.M.; Boiko, A.V.; Ten, D.I. Multiparametric testing of blood protein solutions with diagnostic purpose. In Current Research on Laser Use in Oncology: 2000–2004; International Society for Optics and Photonics: Bellingham, WA, USA, 2006; Volume 5973, p. 597301. [Google Scholar]
- Niedziela, J.T.; Hudzik, B.; Szygula-Jurkiewicz, B.; Nowak, J.U.; Polonski, L.; Gasior, M.; Rozentryt, P. Albumin-to-globulin ratio as an independent predictor of mortality in chronic heart failure. Biomark. Med. 2018, 12, 749–757. [Google Scholar] [CrossRef]
- Wu, P.P.; Hsieh, Y.P.; Kor, C.T.; Chiu, P.F. Association between Albumin–Globulin Ratio and Mortality in Patients with Chronic Kidney Disease. J. Clin. Med. 2019, 8, 1991. [Google Scholar] [CrossRef] [PubMed]
- Petrova, G.P.; Boiko, A.V.; Fedorova, K.V.; Sergeeva, I.A.; Sokol, N.V.; Tichonova, T.N. Optical properties of solutions consisting of albumin and γ-globulin molecules in different ratio modeling blood serum. Laser Phys. 2009, 19, 1303–1307. [Google Scholar] [CrossRef]
- Kirichenko, M.N.; Bulychev, N.A.; Chaikov, L.L.; Kazaryan, M.A.; Masalov, A.V. Effect of iron oxide nanoparticles on the blood coagulation according to light scattering data. Int. Conf. Mol. Pulsed Lasers Xiii 2018, 10614, 106142C. [Google Scholar]
- Kirichenko, M.N.; Masalov, A.V.; Chaikov, L.L.; Zaritskii, A.R. Relation between particle sizes and concentration in undiluted and diluted blood plasma according to light scattering data. Bull. Lebedev Phys. Inst. 2015, 42, 33–36. [Google Scholar] [CrossRef]
- Lawrie, A.S.; Albanyan, A.; Cardigan, R.A.; Mackie, I.J.; Harrison, P. Microparticle sizing by dynamic light scattering in fresh-frozen plasma. Vox Sang. 2009, i96, 206–212. [Google Scholar] [CrossRef] [PubMed]
- Berne, B.J.; Pecora, R. Dynamic Light Scattering; Wdey: New York, NY, USA, 1976. [Google Scholar]
- Pusey, P.N.; Schaefer, D.W.; Koppel, D.E.; Camerini-Otero, R.D.; Franklin, R.M. A Study of the Diffusion Properties of R 17 Virus by Time-Dependent Light Scattering. Le J. De Phys. Colloq. 1972, 33, 1–163. [Google Scholar] [CrossRef]
- Stetefeld, J.; McKenna, S.A.; Patel, T.R. Dynamic light scattering: A practical guide and applications in biomedical sciences. Biophys. Rev. 2016, 8, 409–427. [Google Scholar] [CrossRef]
- Baranov, A.N.; Vlasova, I.M.; Saletskii, A.M. Investigation of serum-albumin aggregation. J. Appl. Spectrosc. 2004, 71, 222–226. [Google Scholar] [CrossRef]
- Militello, V.; Casarino, C.; Emanuele, A.; Giostra, A.; Pullara, F.; Leone, M. Aggregation kinetics of bovine serum albumin studied by FTIR spectroscopy and light scattering. Biophys. Chem. 2004, 107, 175–187. [Google Scholar] [CrossRef]
- Papok, I.M.; Petrova, G.P.; Anenkova, K.A.; Papish, E.A. Using the dynamic light-scattering method for the analysis of a blood-serum model solution. Mosc. Univ. Phys. Bull. 2012, 67, 452–456. [Google Scholar] [CrossRef]
- Mishra, I.; Patel, V.; Robinson, M.D.; Gordon, K.; Deodhar, S.; Cistola, D.P. Dynamics light scattering as a tool for assessing health status and disease risk. Biophys. J. 2016, 110, 476a. [Google Scholar] [CrossRef]
- Chaikov, L.; Kirichenko, M.N.; Krivokhizha, S.V.; Zaritskiy, A.R. Dynamics of statistically confident particle sizes and concentrations in blood plasma obtained by the dynamic light scattering method. J. Biomed. Opt. 2015, 20, 057003. [Google Scholar] [CrossRef] [PubMed]
- Fischer, K.; Schmidt, M. Pitfalls and novel applications of particle sizing by dynamic light scattering. Biomaterials 2016, 98, 79–91. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Ma, J.; Fan, Q.; Tao, Y.; Li, G. Dynamic light scattering (DLS)-based immunoassay for ultra-sensitive detection of tumor marker protein. Chem. Commun. 2016, 52, 7850–7853. [Google Scholar] [CrossRef]
- Huo, Q.; Liu, X.; Dai, Q. Detection of analtyes using metal nanoparticle probes and dynamic light scattering. U.S. Patent 8,883,094 B2, 11 November 2014. [Google Scholar]
- Nepomnyashchaya, E.K. On approximation and experimental accuracy in dynamic light scattering. J. Phys. Conf. Ser. 2019, 1236, 012041. [Google Scholar] [CrossRef]
- Nepomnyashchaya, E.K.; Velichko, E.N. Modification of laser correlation spectroscopy method for analyzing polydisperse nanoparticle suspensions. St. Petersburg Polytech. State Univ. J. Phys. Math. 2019, 12, 73–87. [Google Scholar]
- Zakharov, P.; Scheffold, F. Advances in dynamic light scattering techniques. Light Scatt. Rev. 2009, 4, 433–467. [Google Scholar]
- Kostromitin, A.O.; Skliarov, P.V.; Liokumovich, L.B.; Ushakov, N.A. Laser Frequency Noise Measurement by Forming an Interference Signal with Subcarrier Frequency. In Proceedings of the 2019 IEEE International Conference on Electrical Engineering and Photonics (EExPolytech), St. Petersburg, Russia, 17–18 October 2019. [Google Scholar]
- Zhu, W.; Zhao, Z.; Guo, X.; Hong, X. Study on Hyperlipidemia Serum Ultraviolet Visible Absorption Spectra Based on Wavelet Transform. In Proceedings of the 2009 2nd International Congress on Image and Signal Processing, Tianjin, China, 17–19 October 2009. [Google Scholar]
- Gunasekaran, S.; Uthra, D. FTIR and UV-Visible spectral study on normal and jaundice blood samples. Asian J. Chem. 2008, 20, 5695. [Google Scholar]
- Kotov, O.; Chapalo, I. Signal-to-noise ratio for mode-mode fiber interferometer. In Proceedings of the Optical Measurement Systems for Industrial Inspection X, Munich, Germany, 26 June 2017. [Google Scholar]
- Liokumovich, L.B.; Kostromitin, A.O.; Ushakov, N.A.; Kudryashov, A.V. Method for Measuring Laser Frequency Noise. J. Appl. Spectrosc. 2020, 86, 1106–1112. [Google Scholar] [CrossRef]
- Nepomnyashchaya, E.; Velichko, E.; Kotov, O. Determination of Noise Components in Laser Correlation Spectroscopic Devices for Signal-to-Noise Ratio Estimation. In Proceedings of the 2019 IEEE International Conference on Electrical Engineering and Photonics (EExPolytech), St. Petersburg, Russia, 17–18 October 2019. [Google Scholar]
- Davydov, R.V.; Antonov, V.I.; Yushkova, V.V.; Grebenikova, N.M.; Dudkin, V.I. A new algorithm for processing the absorption and scattering signals of laser radiation on a blood vessel and human tissues. J. Phys. Conf. Ser. 2019, 1236, 012079. [Google Scholar] [CrossRef]
- Koppel, D.E. Analysis of macromolecular polydispersity in intensity correlation spectroscopy: The method of cumulants. J. Chem. Phys. 1972, 57, 4814–4820. [Google Scholar] [CrossRef]
- Provencher, S.W. A constrained regularization method for inverting data represented by linear algebraic or integral equations. Comput. Phys. Commun. 1982, 27, 213–227. [Google Scholar] [CrossRef]
- Nepomnyashchaya, E.; Antonova, E. Methods and algorithms for numerical calculations in dynamic light scattering problems. In Proceedings of the 2018 IEEE International Conference on Electrical Engineering and Photonics (EExPolytech), St. Petersburg, Russia, 22–23 October 2018. [Google Scholar]
- Men’shikov, V.V. Laboratory methods of research in the clinic. Moscow. Medicine 1987. [Google Scholar]
- Nepomnyashchaya, E.; Velichko, E.; Bogomaz, T. Diagnostic possibilities of dynamic light scattering technique. In Proceedings of the Nanoscale Imaging, Sensing, and Actuation for Biomedical Applications XVI, San Francisco, CA, USA, 5 March 2019. [Google Scholar]
- Abrass, C.K.; Heber, D.; Lieberman, J. Circulating immune complexes in patients with diabetes mellitus. Clin. Exp. Immunol. 1983, 52, 164. [Google Scholar] [PubMed]



© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Velichko, E.; Makarov, S.; Nepomnyashchaya, E.; Dong, G. Molecular Aggregation in Immune System Activation Studied by Dynamic Light Scattering. Biology 2020, 9, 123. https://doi.org/10.3390/biology9060123
Velichko E, Makarov S, Nepomnyashchaya E, Dong G. Molecular Aggregation in Immune System Activation Studied by Dynamic Light Scattering. Biology. 2020; 9(6):123. https://doi.org/10.3390/biology9060123
Chicago/Turabian StyleVelichko, Elena, Sergey Makarov, Elina Nepomnyashchaya, and Ge Dong. 2020. "Molecular Aggregation in Immune System Activation Studied by Dynamic Light Scattering" Biology 9, no. 6: 123. https://doi.org/10.3390/biology9060123
APA StyleVelichko, E., Makarov, S., Nepomnyashchaya, E., & Dong, G. (2020). Molecular Aggregation in Immune System Activation Studied by Dynamic Light Scattering. Biology, 9(6), 123. https://doi.org/10.3390/biology9060123





