Treatment of Leptothrix Cells with Ultrapure Water Poses a Threat to Their Viability
Abstract
:1. Introduction
2. Experimental Section
2.1. Strains, Medium, Culturing and Water
| Component | Amount (g/L) | Concentration (mM) |
|---|---|---|
| Glucose | 1.000 | 5.55 |
| Soy peptone | 1.000 | ND |
| Na2SiO3∙9H2O | 0.200 | 0.70 |
| CaCl2∙2H2O | 0.044 | 0.30 |
| MgSO4∙7H2O | 0.041 | 0.17 |
| Na2HPO4∙12H2O | 0.076 | 0.21 |
| KH2PO4∙2H2O | 0.020 | 0.15 |
| HEPES | 2.380 | 10.00 |
2.2. Drop- and Cfu-Tests to Examine Growth of Cells
2.3. LIVE/DEAD Stain to Examine Viability of Cells
2.4. Measurement of pH, Conductivity, Osmotic Pressure, and Ion Concentration in Suspension Agents or Cell Suspensions
2.5. Isolation of Genomic DNAs from OUMS1 Cells
2.6. Electron Microscopy
3. Results and Discussion
3.1. Autolysis of OUMS1 Cells Caused by Prolonged Treatment with UPW
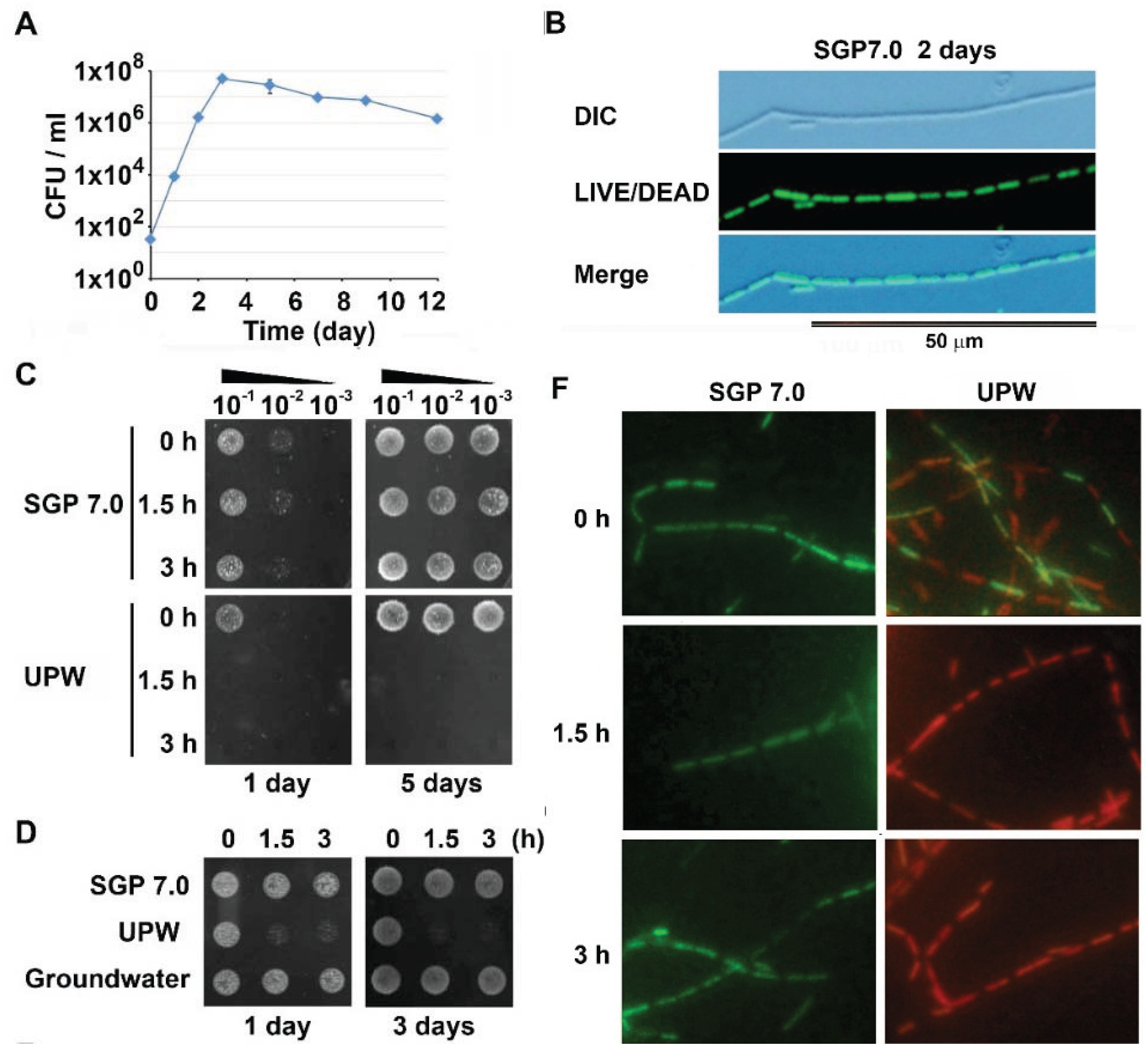
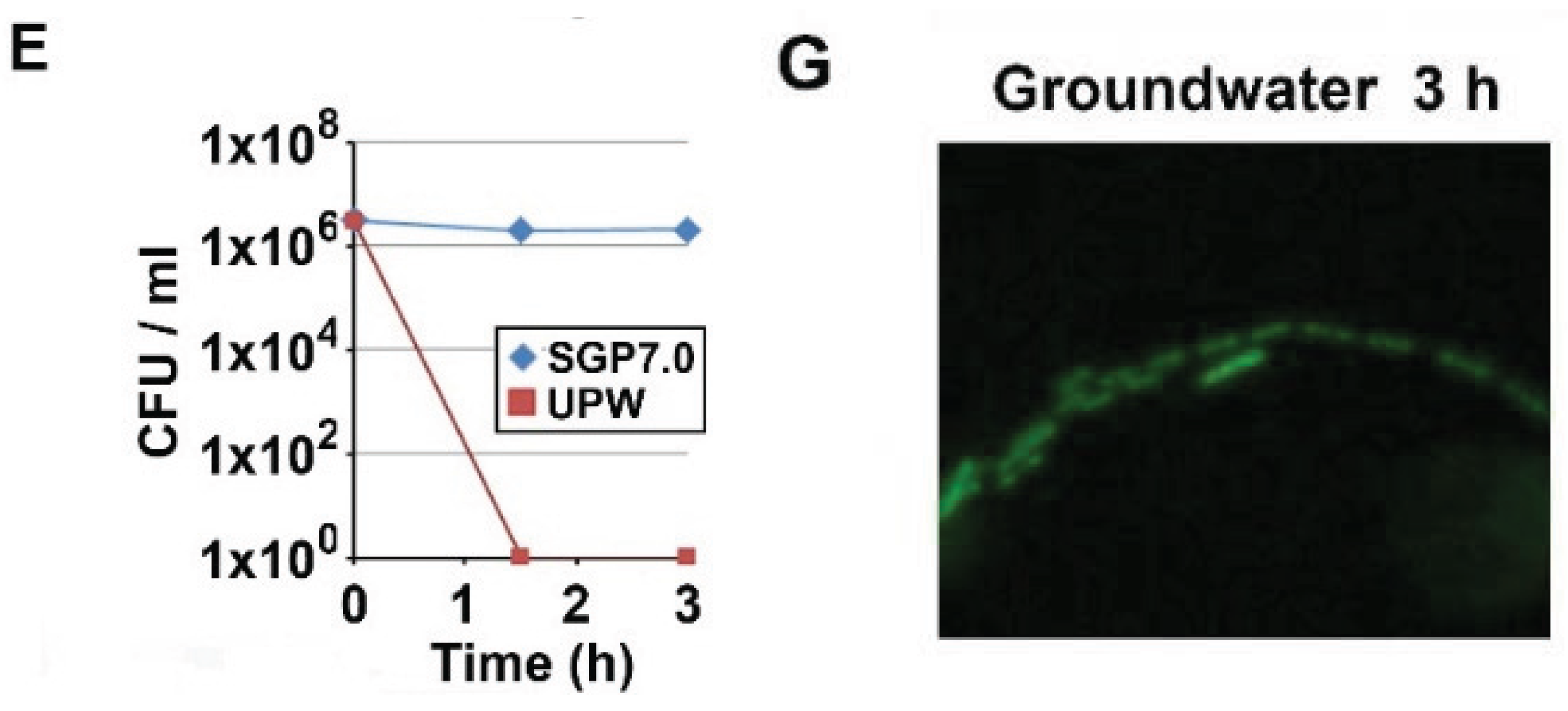
3.2. Acidic pH to Induce Autolysis of the Cells
| Solution | pH | Conductivity (mS/cm) | Osmotic pressure (mosmol/kg) |
|---|---|---|---|
| Tap water | 7.1 | 0.096 | 0.33 |
| UPW (autoclaved) | 5.7 | 0.002 | 0.00 |
| 10 mM Tris-HCl 6.0 | 6.0 | 0.850 | 18.33 |
| 10 mM Tris-HCl 6.5 | 6.4 | 0.890 | 18.00 |
| 10 mM Tris-HCl 7.0 | 6.9 | 0.870 | 18.00 |
| 10 mM HEPES 6.0 | 6.0 | 0.019 | 14.00 |
| 10 mM HEPES 6.5 | 6.5 | 0.054 | 14.00 |
| 10 mM HEPES 7.0 | 7.0 | 0.131 | 13.67 |
| Groundwater | 6.6 | 0.350 | 10.00 |
| Groundwater (filtrated) | 6.6 | 0.360 | 10.33 |
| SGP 6.0 | 6.0 | 0.510 | 31.00 |
| SGP 6.5 | 6.5 | 0.480 | 31.33 |
| SGP 7.0 | 7.0 | 0.510 | 31.33 |
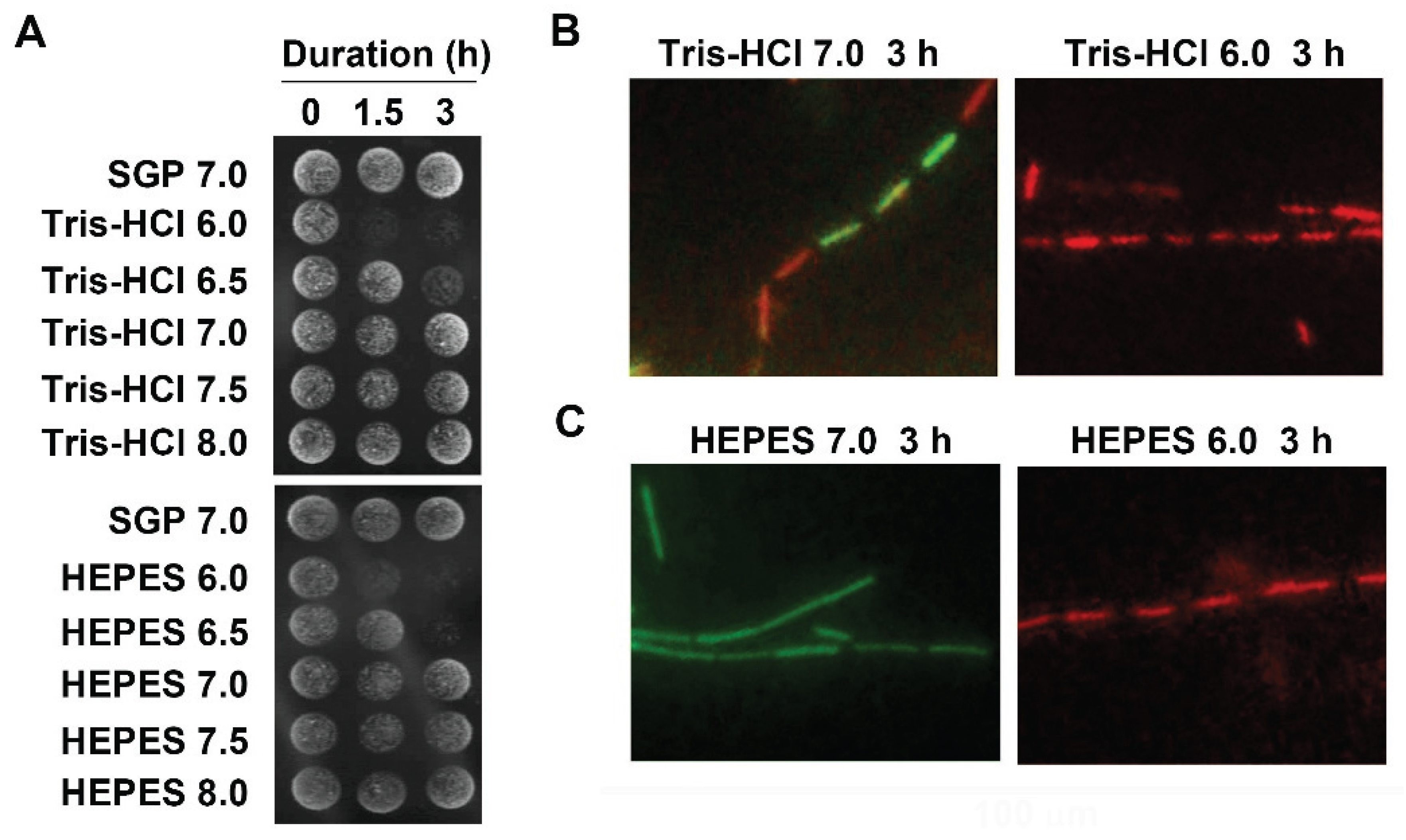
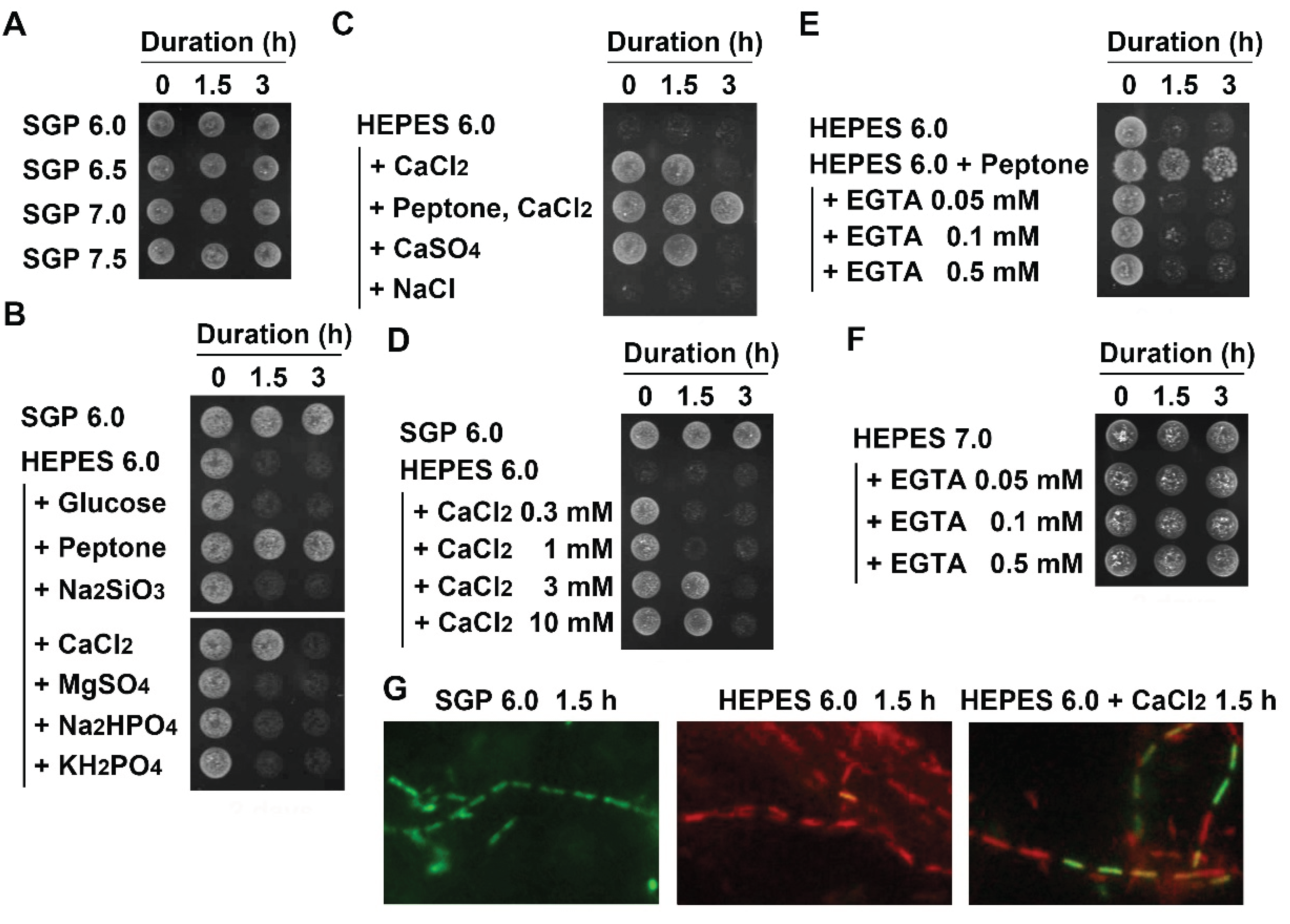
3.3. Suppression of Autolysis of OUMS1 Cells by Ca2+
3.4. Damage of Plasma Membrane and Cytoplasm in UPW- and HEPES6.0-Treated Cells
3.5. Relaxation of Genomic DNA and Leaking of Electrolytes Caused by Autolysis-Inducing Treatments
3.6. Lysozyme-Inducing Autolysis of OUMS1 Due to Damage of Peptidoglycan
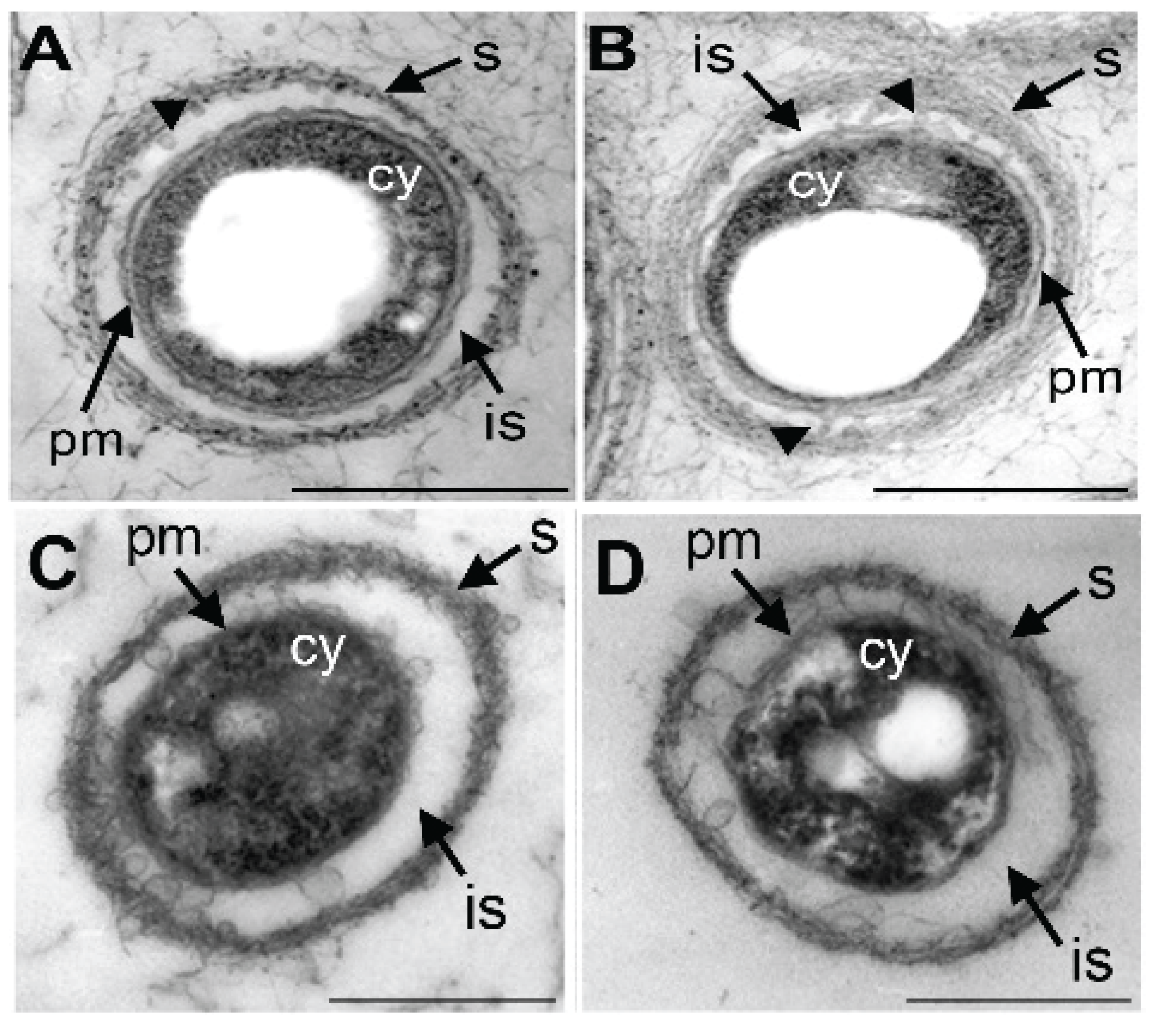
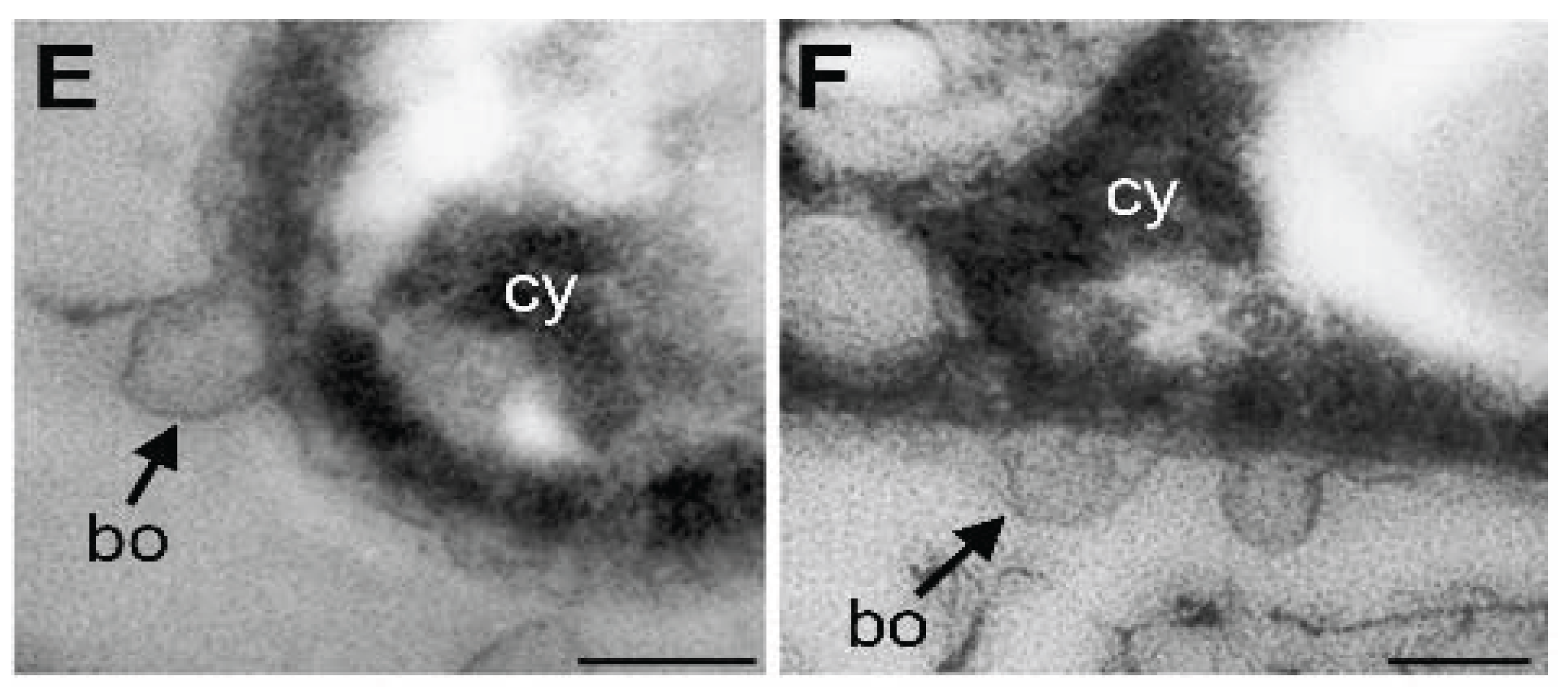
3.7. Lysozyme-inducing Autolysis of OUMS1 Due to Damage of Peptidoglycan

3.8. Autolysis of Leptothrix Discophora Strain SP-6 Cells Induced by the UPW Treatment
4. Conclusions
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Suzuki, T.; Ishihara, H.; Toyoda, K.; Shiraishi, T.; Kunoh, H.; Takada, J. Autolysis of bacterial cells leads to formation of empty sheaths by Leptothrix spp. Minerals 2013, 3, 247–257. [Google Scholar] [CrossRef]
- Spring, S. The general Leptothrix and Sphaerotilus. Prokaryotes 2006, 5, 758–777. [Google Scholar]
- Emerson, D.; Fleming, E.J.; McBeth, J.M. Iron-oxidizing bacteria: An environmental and genomic perspective. Annu. Rev. Microbiol. 2010, 64, 561–583. [Google Scholar] [CrossRef] [PubMed]
- Ghiorse, W.C. Biology of iron- and manganese-depositing bacteria. Annu. Rev. Microbiol. 1984, 38, 515–550. [Google Scholar] [CrossRef] [PubMed]
- Van Veen, W.L.; Mulder, E.G.; Deinema, M.H. The Sphaerotilus-Leptothrix group of bacteria. Microbiol. Rev. 1978, 42, 329–356. [Google Scholar] [PubMed]
- Sawayama, M.; Suzuki, T.; Hashimoto, H.; Kasai, T.; Furutani, M.; Miyata, N.; Kunoh, H.; Takada, J. Isolation of a Leptothrix strain, OUMS1, from ocherous deposits in groundwater. Curr. Microbiol. 2011, 63, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Rice, K.C.; Bayles, K.W. Molecular control of bacterial death and lysis. Microbiol. Mol. Biol. Rev. 2008, 72, 85–109. [Google Scholar] [CrossRef] [PubMed]
- Leduc, M.; van Heijenoort, J. Autolysis of Escherichia coli. J. Bacteriol. 1980, 142, 52–59. [Google Scholar] [PubMed]
- Leduc, M.; Kasra, R.; van Heijenoort, J. Induction and control of the autolytic system of Escherichia coli. J. Bacteriol. 1982, 152, 26–34. [Google Scholar] [PubMed]
- Wegener, W.S.; Hebeler, B.H.; Morse, S.A. Cell envelope of Neisseria gonorrhoeae: Relationship between autolysis in buffer and the hydrolysis of peptidoglycan. Infect. Immun. 1977, 18, 210–219. [Google Scholar] [PubMed]
- Rayman, M.K.; MacLeod, R.A. Interaction of Mg2+ with peptidoglycan and its relation to the prevention of lysis of a marine pseudomonad. J. Bacteriol. 1975, 122, 650–659. [Google Scholar] [PubMed]
- Holland, I.B.; Jones, H.E.; Campbell, A.K.; Jacq, A. An assessment of the role of intracellular free Ca2+ in E. coli. Biochimie 1999, 81, 901–907. [Google Scholar] [CrossRef]
- Lehtinen, J.; Nuutila, J.; Lilius, E.M. Green fluorescent protein-propidium iodide (GFP-PI) based assay for flow cytometric measurement of bacterial viability. Cytom. A 2004, 60, 165–172. [Google Scholar] [CrossRef]
- Amano, M.; Toyoda, K.; Ichinose, Y.; Yamada, T.; Shiraishi, T. Association between ion fluxes and defense responses in pea and cowpea tissues. Plant Cell Physiol. 1997, 38, 698–706. [Google Scholar] [CrossRef] [PubMed]
- Slonczewski, J.L.; Fujisawa, M.; Dopson, M.; Krulwichta, T.A. Cytoplasmic pH measurement and homeostasis in bacteria and archaea. Adv. Microb. Physiol. 2009, 55, 1–79. [Google Scholar] [PubMed]
- Dominguez, D.C. Calcium signaling in bacteria. Mol. Microbiol. 2004, 54, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Rosch, J.W.; Sublett, J.; Gao, G.; Wang, Y.D.; Tuomanen, E.I. Calcium efflux is essential for bacterial survival in the eukaryotic host. Mol. Microbiol. 2008, 70, 435–444. [Google Scholar] [CrossRef] [PubMed]
- Patrauchan, M.A.; Sarkisova, S.K.; Sauer, K.; Franklin, M.J. Calcium influences cellular and extracellular product formation during biofilm-associated growth of a marine Pseudoalteromonas sp. Micobiology 2005, 151, 2885–2897. [Google Scholar]
- Furutani, M.; Suzuki, T.; Ishihara, H.; Hashimoto, H.; Kunoh, H.; Takada, J. Initial assemblage of bacterial saccharic fibrils and element deposition to form an immature sheath in cultured Leptothrix sp. strain OUMS1. Minerals 2011, 1, 157–166. [Google Scholar]
- Jensen, R.B.; Shapiro, L. Chromosome segregation during the prokaryotic cell division cycle. Curr. Opin. Cell Biol. 1999, 11, 726–731. [Google Scholar] [CrossRef] [PubMed]
- Birnboim, H.C.; Doly, J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucl. Acids Res. 1979, 7, 1513–1523. [Google Scholar] [CrossRef] [PubMed]
- Balke, V.L.; Gralla, J.D. Changes in the linking number of supercoiled DNA accompany growth transitions in Escherichia coli. J. Bacteriol. 1987, 169, 4499–4506. [Google Scholar] [PubMed]
- Vázquez-Laslop, N.; Lee, H.; Hu, R.; Neyfakh, A.A. Molecular sieve mechanism of selective release of cytoplasmic proteins by osmotically shocked Escherichia coli. J. Bacteriol. 2001, 183, 2399–2404. [Google Scholar] [CrossRef] [PubMed]
- Vollmer, W. Structural variation in the glycan strands of bacterial peptidoglycan. FEMS Microbiol. Rev. 2008, 32, 287–306. [Google Scholar] [CrossRef] [PubMed]
- Kianoush, N.; Adler, C.J.; Nguyen, K.A.; Browne, G.V.; Simonian, M.; Hunter, N. Bacterial profile of dentine caries and the impact of pH on bacterial population diversity. PLOS ONE 2014, 9e, 92940. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kunoh, T.; Suzuki, T.; Shiraishi, T.; Kunoh, H.; Takada, J. Treatment of Leptothrix Cells with Ultrapure Water Poses a Threat to Their Viability. Biology 2015, 4, 50-66. https://doi.org/10.3390/biology4010050
Kunoh T, Suzuki T, Shiraishi T, Kunoh H, Takada J. Treatment of Leptothrix Cells with Ultrapure Water Poses a Threat to Their Viability. Biology. 2015; 4(1):50-66. https://doi.org/10.3390/biology4010050
Chicago/Turabian StyleKunoh, Tatsuki, Tomoko Suzuki, Tomonori Shiraishi, Hitoshi Kunoh, and Jun Takada. 2015. "Treatment of Leptothrix Cells with Ultrapure Water Poses a Threat to Their Viability" Biology 4, no. 1: 50-66. https://doi.org/10.3390/biology4010050
APA StyleKunoh, T., Suzuki, T., Shiraishi, T., Kunoh, H., & Takada, J. (2015). Treatment of Leptothrix Cells with Ultrapure Water Poses a Threat to Their Viability. Biology, 4(1), 50-66. https://doi.org/10.3390/biology4010050





