MicroRNAs in Juvenile Idiopathic Arthritis: State of the Art and Future Perspectives
Abstract
Simple Summary
Abstract
1. Introduction
1.1. Juvenile Idiopathic Arthritis: Clinical Features and Treatment
1.2. microRNAs (miRNAs): Role in Disease Pathogenesis and as Potential Biomarkers
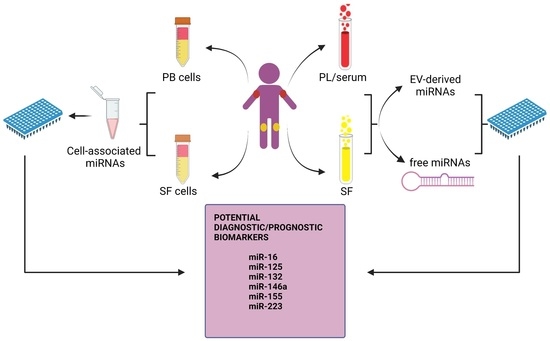
2. miRNAs in JIA: General Considerations
3. Cell-Associated miRNAs in JIA
3.1. Differentially Expressed miRNAs
3.2. Data Comparison among Different Studies
4. miRNAs Released in the Serum and PL of JIA Patients
4.1. Differentially Expressed miRNAs
4.2. Data Comparison among Different Studies
5. miRNAs Released in SF from JIA Joints
5.1. Differentially Expressed miRNAs
5.2. Data Comparison between Studies
6. JIA-Derived EV-miRs
7. Concluding Remarks
8. Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Ravelli, A.; Martini, A. Juvenile idiopathic arthritis. Lancet 2007, 369, 767–778. [Google Scholar] [CrossRef]
- Schiappapietra, B.; Bava, C.; Rosina, S.; Pistorio, A.; Mongelli, F.; Pederzoli, S.; Verazza, S.; Lanni, S.; Muratore, V.; Dalprà, S.; et al. A prediction rule for polyarticular extension in oligoarticular-onset juvenile idiopathic arthritis. Clin. Exp. Rheumatol. 2021, 39, 913–919. [Google Scholar] [CrossRef]
- Martini, A.; Lovell, D.J.; Albani, S.; Brunner, H.I.; Hyrich, K.L.; Thompson, S.D.; Ruperto, N. Juvenile idiopathic arthritis. Nat. Rev. Dis. Primers 2022, 8, 5. [Google Scholar] [CrossRef] [PubMed]
- Thierry, S.; Fautrel, B.; Lemelle, I.; Guillemin, F. Prevalence and incidence of juvenile idiopathic arthritis: A systematic review. Jt. Bone Spine 2014, 81, 112–117. [Google Scholar] [CrossRef]
- Consolaro, A.; Giancane, G.; Alongi, A.; van Dijkhuizen, E.H.P.; Aggarwal, A.; Al-Mayouf, S.M.; Bovis, F.; De, I.J.; Demirkaya, E.; Flato, B.; et al. Phenotypic variability and disparities in treatment and outcomes of childhood arthritis throughout the world: An observational cohort study. Lancet Child Adolesc. Health 2019, 3, 255–263. [Google Scholar] [CrossRef] [PubMed]
- Martini, A.; Lovell, D.J. Juvenile idiopathic arthritis: State of the art and future perspectives. Ann. Rheum. Dis. 2010, 69, 1260–1263. [Google Scholar] [CrossRef]
- Macaubas, C.; Nguyen, K.; Milojevic, D.; Park, J.L.; Mellins, E.D. Oligoarticular and polyarticular JIA: Epidemiology and pathogenesis. Nat. Rev. Rheumatol. 2009, 5, 616–626. [Google Scholar] [CrossRef] [PubMed]
- Bosco, M.C.; Varesio, L. Monocytic Cell Gene Regulation by the Hypoxic Synovial Environment in Juvenile Idiopathic Arthritis: Implications for Disease Pathogenesis. J. Clin. Rheumatol. Musculoskelet. Med. 2010, 1, 47–55. [Google Scholar]
- Bosco, M.C.; Delfino, S.; Ferlito, F.; Battaglia, F.P.M.; Gregorio, A.; Gambini, C.; Gattorno, M.; Martini, A.; Varesio, L. Hypoxic synovial environment and expression of macrophage inflammatory protein MIP-3a/CCL20 in Juvenile Idiopathic Arthritis. Arthritis Rheum. 2008, 58, 1833–1838. [Google Scholar] [CrossRef]
- Bosco, M.C.; Delfino, S.; Ferlito, F.; Puppo, M.; Gregorio, A.; Gambini, C.; Gattorno, M.; Martini, A.; Varesio, L. The hypoxic synovial environment regulates expression of vascular endothelial growth factor and osteopontin in juvenile idiopathic arthritis. J. Rheumatol. 2009, 36, 1318–1329. [Google Scholar] [CrossRef]
- Raggi, F.; Pelassa, S.; Pierobon, D.; Penco, F.; Gattorno, M.; Novelli, F.; Eva, A.; Varesio, L.; Giovarelli, M.; Bosco, M.C. Regulation of Human Macrophage M1-M2 Polarization Balance by Hypoxia and the Triggering Receptor Expressed on Myeloid Cells-1. Front. Immunol. 2017, 8, 1097. [Google Scholar] [CrossRef] [PubMed]
- Giancane, G.; Consolaro, A.; Lanni, S.; Davi, S.; Schiappapietra, B.; Ravelli, A. Juvenile Idiopathic Arthritis: Diagnosis and Treatment. Rheumatol. Ther. 2016, 3, 187–207. [Google Scholar] [CrossRef] [PubMed]
- Viola, S.; Felici, E.; Magni-Manzoni, S.; Pistorio, A.; Buoncompagni, A.; Ruperto, N.; Rossi, F.; Bartoli, M.; Martini, A.; Ravelli, A. Development and validation of a clinical index for assessment of long-term damage in juvenile idiopathic arthritis. Arthritis Rheum. 2005, 52, 2092–2102. [Google Scholar] [CrossRef] [PubMed]
- Petty, R.E.; Southwood, T.R.; Manners, P.; Baum, J.; Glass, D.N.; Goldenberg, J.; He, X.; Maldonado-Cocco, J.; Orozco-Alcala, J.; Prieur, A.M.; et al. International League of Associations for Rheumatology classification of juvenile idiopathic arthritis: Second revision, Edmonton, 2001. J. Rheumatol. 2004, 31, 390–392. [Google Scholar]
- Zaripova, L.N.; Midgley, A.; Christmas, S.E.; Beresford, M.W.; Baildam, E.M.; Oldershaw, R.A. Juvenile idiopathic arthritis: From aetiopathogenesis to therapeutic approaches. Pediatr. Rheumatol. Online J. 2021, 19, 135. [Google Scholar] [CrossRef]
- Rosina, S.; Natoli, V.; Santaniello, S.; Trincianti, C.; Consolaro, A.; Ravelli, A. Novel biomarkers for prediction of outcome and therapeutic response in juvenile idiopathic arthritis. Expert Rev. Clin. Immunol. 2021, 17, 853–870. [Google Scholar] [CrossRef]
- Ahn, J.G. Role of Biomarkers in Juvenile Idiopathic Arthritis. J. Rheum. Dis. 2020, 27, 233–240. [Google Scholar] [CrossRef]
- Trincianti, C.; van Dijkhuizen, E.H.P.; Alongi, A.; Mazzoni, M.; Swart, J.F.; Nikishina, I.; Lahdenne, P.; Rutkowska-Sak, L.; Avcin, T.; Quartier, P.; et al. Definition and validation of the American College of Rheumatology 2021 juvenile arthritis disease activity score cutoffs for disease activity states in juvenile idiopathic arthritis. Arthritis Rheumatol. 2021, 73, 1966–1975. [Google Scholar] [CrossRef]
- Nziza, N.; Duroux-Richard, I.; Apparailly, F. MicroRNAs in juvenile idiopathic arthritis: Can we learn more about pathophysiological mechanisms? Autoimmun. Rev. 2019, 18, 796–804. [Google Scholar] [CrossRef]
- Crayne, C.B.; Beukelman, T. Juvenile Idiopathic Arthritis: Oligoarthritis and Polyarthritis. Pediatr. Clin. N. Am. 2018, 65, 657–674. [Google Scholar] [CrossRef]
- Barut, K.; Adrovic, A.; Şahin, S.; Kasapçopur, Ö. Juvenile Idiopathic Arthritis. Balk. Med. J. 2017, 34, 90–101. [Google Scholar] [CrossRef]
- Ravelli, A.; Felici, E.; Magni-Manzoni, S.; Pistorio, A.; Novarini, C.; Bozzola, E.; Viola, S.; Martini, A. Patients with antinuclear antibody-positive juvenile idiopathic arthritis constitute a homogeneous subgroup irrespective of the course of joint disease. Arthritis Rheum. 2005, 52, 826–832. [Google Scholar] [CrossRef]
- Ravelli, A.; Varnier, G.C.; Oliveira, S.; Castell, E.; Arguedas, O.; Magnani, A.; Pistorio, A.; Ruperto, N.; Magni-Manzoni, S.; Galasso, R.; et al. Antinuclear antibody-positive patients should be grouped as a separate category in the classification of juvenile idiopathic arthritis. Arthritis Rheum. 2011, 63, 267–275. [Google Scholar] [CrossRef] [PubMed]
- Martini, A.; Ravelli, A.; Avcin, T.; Beresford, M.W.; Burgos-Vargas, R.; Cuttica, R.; Ilowite, N.T.; Khubchandani, R.; Laxer, R.M.; Lovell, D.J.; et al. Toward New Classification Criteria for Juvenile Idiopathic Arthritis: First Steps, Pediatric Rheumatology International Trials Organization International Consensus. J. Rheumatol. 2019, 46, 190–197. [Google Scholar] [CrossRef] [PubMed]
- Adrovic, A.; Yildiz, M.; Köker, O.; Şahin, S.; Barut, K.; Kasapçopur, Ö. Biologics in juvenile idiopathic arthritis-main advantages and major challenges: A narrative review. Arch. Rheumatol. 2021, 36, 146–157. [Google Scholar] [CrossRef] [PubMed]
- Wallace, C.A.; Ruperto, N.; Giannini, E. Preliminary criteria for clinical remission for select categories of juvenile idiopathic arthritis. J. Rheumatol. 2004, 31, 2290–2294. [Google Scholar]
- Shenoi, S.; Wallace, C.A. Remission in juvenile idiopathic arthritis: Current facts. Curr. Rheumatol. Rep. 2010, 12, 80–86. [Google Scholar] [CrossRef]
- Shoop-Worrall, S.J.W.; Kearsley-Fleet, L.; Thomson, W.; Verstappen, S.M.M.; Hyrich, K.L. How common is remission in juvenile idiopathic arthritis: A systematic review. Semin. Arthritis Rheum. 2017, 47, 331–337. [Google Scholar] [CrossRef]
- Ravelli, A.; Davi, S.; Bracciolini, G.; Pistorio, A.; Consolaro, A.; van Dijkhuizen, E.H.P.; Lattanzi, B.; Filocamo, G.; Verazza, S.; Gerloni, V.; et al. Intra-articular corticosteroids versus intra-articular corticosteroids plus methotrexate in oligoarticular juvenile idiopathic arthritis: A multicentre, prospective, randomised, open-label trial. Lancet 2017, 389, 909–916. [Google Scholar] [CrossRef]
- Consolaro, A.; Varnier, G.C.; Martini, A.; Ravelli, A. Advances in biomarkers for paediatric rheumatic diseases. Nat. Rev. Rheumatol. 2015, 11, 265–275. [Google Scholar] [CrossRef]
- Duurland, C.L.; Wedderburn, L.R. Current developments in the use of biomarkers for juvenile idiopathic arthritis. Curr. Rheumatol. Rep. 2014, 16, 406. [Google Scholar] [CrossRef] [PubMed]
- Gibson, D.S.; Finnegan, S.; Jordan, G.; Scaife, C.; Brockbank, S.; Curry, J.; McAllister, C.; Pennington, S.; Dunn, M.; Rooney, M.E. Stratification and monitoring of juvenile idiopathic arthritis patients by synovial proteome analysis. J. Proteome Res. 2009, 8, 5601–5609. [Google Scholar] [CrossRef]
- Al-Matar, M.J.; Petty, R.E.; Tucker, L.B.; Malleson, P.N.; Schroeder, M.L.; Cabral, D.A. The early pattern of joint involvement predicts disease progression in children with oligoarticular (pauciarticular) juvenile rheumatoid arthritis. Arthritis Rheum. 2002, 46, 2708–2715. [Google Scholar] [CrossRef] [PubMed]
- Nistala, K.; Moncrieffe, H.; Newton, K.R.; Varsani, H.; Hunter, P.; Wedderburn, L.R. Interleukin-17-producing T cells are enriched in the joints of children with arthritis, but have a reciprocal relationship to regulatory T cell numbers. Arthritis Rheum. 2008, 58, 875–887. [Google Scholar] [CrossRef]
- Hunter, P.J.; Nistala, K.; Jina, N.; Eddaoudi, A.; Thomson, W.; Hubank, M.; Wedderburn, L.R. Biologic predictors of extension of oligoarticular juvenile idiopathic arthritis as determined from synovial fluid cellular composition and gene expression. Arthritis Rheum. 2010, 62, 896–907. [Google Scholar] [CrossRef]
- Consolaro, A.; Ruperto, N.; Bazso, A.; Pistorio, A.; Magni-Manzoni, S.; Filocamo, G.; Malattia, C.; Viola, S.; Martini, A.; Ravelli, A. Development and validation of a composite disease activity score for juvenile idiopathic arthritis. Arthritis Rheum. 2009, 61, 658–666. [Google Scholar] [CrossRef]
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Wightman, B.; Ha, I.; Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 1993, 75, 855–862. [Google Scholar] [CrossRef]
- Leitão, A.L.; Enguita, F.J. A Structural View of miRNA Biogenesis and Function. Noncoding RNA 2022, 8, 10. [Google Scholar] [CrossRef]
- O’Brien, J.; Hayder, H.; Zayed, Y.; Peng, C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front. Endocrinol. 2018, 9, 402. [Google Scholar] [CrossRef]
- Annese, T.; Tamma, R.; De, G.M.; Ribatti, D. microRNAs Biogenesis, Functions and Role in Tumor Angiogenesis. Front. Oncol. 2020, 10, 581007. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, L.A.; Murphy, P.R. MicroRNA: Biogenesis, Function and Role in Cancer. Curr. Genom. 2010, 11, 537–561. [Google Scholar] [CrossRef] [PubMed]
- Jonas, S.; Izaurralde, E. Towards a molecular understanding of microRNA-mediated gene silencing. Nat. Rev. Genet. 2015, 16, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Medley, J.C.; Panzade, G.; Zinovyeva, A.Y. microRNA strand selection: Unwinding the rules. Wiley Interdiscip. Rev. RNA 2021, 12, e1627. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, S. Posttranscriptional upregulation by microRNAs. Wiley Interdiscip. Rev. RNA 2012, 3, 311–330. [Google Scholar] [CrossRef]
- Truesdell, S.S.; Mortensen, R.D.; Seo, M.; Schroeder, J.C.; Lee, J.H.; LeTonqueze, O.; Vasudevan, S. MicroRNA-mediated mRNA translation activation in quiescent cells and oocytes involves recruitment of a nuclear microRNP. Sci. Rep. 2012, 2, 842. [Google Scholar] [CrossRef]
- Bukhari, S.I.A.; Truesdell, S.S.; Vasudevan, S. Analysis of MicroRNA-Mediated Translation Activation of In Vitro Transcribed Reporters in Quiescent Cells. Methods Mol. Biol. 2018, 1686, 251–264. [Google Scholar]
- Luo, B.; Zhou, K.; Liufu, Y.; Huang, X.; Zeng, H.; Zhang, Z. Novel insight into miRNA biology and its role in the pathogenesis of systemic lupus erythematosus. Front. Immunol. 2022, 13, 1059887. [Google Scholar] [CrossRef]
- Cortez, M.A.; Bueso-Ramos, C.; Ferdin, J.; Lopez-Berestein, G.; Sood, A.K.; Calin, G.A. MicroRNAs in body fluids—The mix of hormones and biomarkers. Nat. Rev. Clin. Oncol. 2011, 8, 467–477. [Google Scholar] [CrossRef]
- Chen, X.; Ba, Y.; Ma, L.; Cai, X.; Yin, Y.; Wang, K.; Guo, J.; Zhang, Y.; Chen, J.; Guo, X.; et al. Characterization of microRNAs in serum: A novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res. 2008, 18, 997–1006. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Weber, J.A.; Baxter, D.H.; Zhang, S.; Huang, D.Y.; Huang, K.H.; Lee, M.J.; Galas, D.J.; Wang, K. The microRNA spectrum in 12 body fluids. Clin. Chem. 2010, 56, 1733–1741. [Google Scholar] [CrossRef]
- Montecalvo, A.; Larregina, A.T.; Shufesky, W.J.; Stolz, D.B.; Sullivan, M.L.; Karlsson, J.M.; Baty, C.J.; Gibson, G.A.; Erdos, G.; Wang, Z.; et al. Mechanism of transfer of functional microRNAs between mouse dendritic cells via exosomes. Blood 2012, 119, 756–766. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 2012, 7, e30679. [Google Scholar] [CrossRef]
- Zakeri, Z.; Salmaninejad, A.; Hosseini, N.; Shahbakhsh, Y.; Fadaee, E.; Shahrzad, M.K.; Fadaei, S. MicroRNA and exosome: Key players in rheumatoid arthritis. J. Cell. Biochem. 2019, 120, 10930–10944. [Google Scholar] [CrossRef]
- Van, N.G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar]
- Meldolesi, J. Extracellular vesicles, news about their role in immune cells: Physiology, pathology and diseases. Clin. Exp. Immunol. 2019, 196, 318–327. [Google Scholar] [CrossRef]
- Robbins, P.D.; Dorronsoro, A.; Booker, C.N. Regulation of chronic inflammatory and immune processes by extracellular vesicles. J. Clin. Investig. 2016, 126, 1173–1180. [Google Scholar] [CrossRef]
- Ge, Q.; Zhou, Y.; Lu, J.; Bai, Y.; Xie, X.; Lu, Z. miRNA in plasma exosome is stable under different storage conditions. Molecules 2014, 19, 1568–1575. [Google Scholar] [CrossRef]
- Zhang, Z.Y.; Li, Y.C.; Geng, C.Y.; Wang, H.J.; Chen, W.M. Potential Relationship between Clinical Significance and Serum Exosomal miRNAs in Patients with Multiple Myeloma. BioMed Res. Int. 2019, 2019, 1575468. [Google Scholar] [CrossRef] [PubMed]
- Boukouris, S.; Mathivanan, S. Exosomes in bodily fluids are a highly stable resource of disease biomarkers. Proteom. Clin. Appl. 2015, 9, 358–367. [Google Scholar] [CrossRef] [PubMed]
- Skog, J.; Wurdinger, T.; Van Rijn, S.; Meijer, D.H.; Gainche, L.; Sena-Esteves, M.; Curry, W.T., Jr.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef]
- Cheng, L.; Sharples, R.A.; Scicluna, B.J.; Hill, A.F. Exosomes provide a protective and enriched source of miRNA for biomarker profiling compared to intracellular and cell-free blood. J. Extracell. Vesicles 2014, 3, 23743. [Google Scholar] [CrossRef]
- Baltimore, D.; Boldin, M.P.; O’Connell, R.M.; Rao, D.S.; Taganov, K.D. MicroRNAs: New regulators of immune cell development and function. Nat. Immunol. 2008, 9, 839–845. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.; Rajewsky, K. MicroRNA control in the immune system: Basic principles. Cell 2009, 136, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Ormseth, M.J.; Solus, J.F.; Sheng, Q.; Ye, F.; Wu, Q.; Guo, Y.; Oeser, A.M.; Allen, R.M.; Vickers, K.C.; Stein, C.M. Development and Validation of a MicroRNA Panel to Differentiate Between Patients with Rheumatoid Arthritis or Systemic Lupus Erythematosus and Controls. J. Rheumatol. 2020, 47, 188–196. [Google Scholar] [CrossRef]
- Romo-García, M.F.; Bastian, Y.; Zapata-Zuñiga, M.; Macías-Segura, N.; Castillo-Ortiz, J.D.; Lara-Ramírez, E.E.; Fernández-Ruiz, J.C.; Berlanga-Taylor, A.J.; González-Amaro, R.; Ramos-Remus, C.; et al. Identification of putative miRNA biomarkers in early rheumatoid arthritis by genome-wide microarray profiling: A pilot study. Gene 2019, 720, 144081. [Google Scholar] [CrossRef]
- Hanna, J.; Hossain, G.S.; Kocerha, J. The Potential for microRNA Therapeutics and Clinical Research. Front. Genet. 2019, 10, 478. [Google Scholar] [CrossRef]
- Nana-Sinkam, S.P.; Croce, C.M. MicroRNA regulation of tumorigenesis, cancer progression and interpatient heterogeneity: Towards clinical use. Genome Biol. 2014, 15, 445. [Google Scholar] [CrossRef]
- Brown, D.; Rahman, M.; Nana-Sinkam, S.P. MicroRNAs in respiratory disease. A clinician’s overview. Ann. Am. Thorac. Soc. 2014, 11, 1277–1285. [Google Scholar] [CrossRef]
- Boon, R.A.; Dimmeler, S. MicroRNAs in myocardial infarction. Nat. Rev. Cardiol. 2015, 12, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Liao, Q.; Wang, B.; Li, X.; Jiang, G. miRNAs in acute myeloid leukemia. Oncotarget 2017, 8, 3666–3682. [Google Scholar] [CrossRef] [PubMed]
- Garo, L.P.; Murugaiyan, G. Contribution of MicroRNAs to autoimmune diseases. Cell. Mol. Life Sci. 2016, 73, 2041–2051. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.Q.; Papp, G.; Szodoray, P.; Zeher, M. The role of microRNAs in the pathogenesis of autoimmune diseases. Autoimmun. Rev. 2016, 15, 1171–1180. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wu, H.; Zhao, M.; Lu, Q. Identifying the differentially expressed microRNAs in autoimmunity: A systemic review and meta-analysis. Autoimmunity 2020, 53, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Hashemi, A.; Gorji-Bahri, G. MicroRNA: Promising Roles in Cancer Therapy. Curr. Pharm. Biotechnol. 2020, 21, 1186–1203. [Google Scholar] [CrossRef] [PubMed]
- Condrat, C.E.; Thompson, D.C.; Barbu, M.G.; Bugnar, O.L.; Boboc, A.; Cretoiu, D.; Suciu, N.; Cretoiu, S.M.; Voinea, S.C. miRNAs as Biomarkers in Disease: Latest Findings Regarding Their Role in Diagnosis and Prognosis. Cells 2020, 9, 276. [Google Scholar] [CrossRef]
- Wang, J.; Chen, J.; Sen, S. MicroRNA as Biomarkers and Diagnostics. J. Cell. Physiol. 2016, 231, 25–30. [Google Scholar] [CrossRef]
- Kai, K.; Dittmar, R.L.; Sen, S. Secretory microRNAs as biomarkers of cancer. Semin. Cell Dev. Biol. 2018, 78, 22–36. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, M.; Shan, X.; Zhou, X.; Wang, T.; Zhang, J.; Tao, J.; Cheng, W.; Chen, G.; Li, J.; et al. A panel of seven-miRNA signature in plasma as potential biomarker for colorectal cancer diagnosis. Gene 2019, 687, 246–254. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Cui, J.; Wu, H.; Lu, Q. The emerging role of circulating microRNAs as biomarkers in autoimmune diseases. Autoimmunity 2014, 47, 419–429. [Google Scholar] [CrossRef] [PubMed]
- Churov, A.V.; Oleinik, E.K.; Knip, M. MicroRNAs in rheumatoid arthritis: Altered expression and diagnostic potential. Autoimmun. Rev. 2015, 14, 1029–1037. [Google Scholar] [CrossRef]
- Murata, K.; Yoshitomi, H.; Tanida, S.; Ishikawa, M.; Nishitani, K.; Ito, H.; Nakamura, T. Plasma and synovial fluid microRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis Res. Ther. 2010, 12, R86. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Peng, W.; Ouyang, X.; Li, W.; Dai, Y. Circulating microRNAs as candidate biomarkers in patients with systemic lupus erythematosus. Transl. Res. 2012, 160, 198–206. [Google Scholar] [CrossRef]
- Chen, X.M.; Zhao, Y.; Wu, X.D.; Wang, M.J.; Yu, H.; Lu, J.J.; Hu, Y.J.; Huang, Q.C.; Huang, R.Y.; Lu, C.J. Novel findings from determination of common expressed plasma exosomal microRNAs in patients with psoriatic arthritis, psoriasis vulgaris, rheumatoid arthritis, and gouty arthritis. Discov. Med. 2019, 28, 47–68. [Google Scholar]
- Laurent, L.C.; Abdel-Mageed, A.B.; Adelson, P.D.; Arango, J.; Balaj, L.; Breakefield, X.; Carlson, E.; Carter, B.S.; Majem, B.; Chen, C.C.; et al. Meeting report: Discussions and preliminary findings on extracellular RNA measurement methods from laboratories in the NIH Extracellular RNA Communication Consortium. J. Extracell. Vesicles 2015, 4, 26533. [Google Scholar] [CrossRef]
- Guo, Y.; Bosompem, A.; Zhong, X.; Clark, T.; Shyr, Y.; Kim, A.S. A comparison of microRNA sequencing reproducibility and noise reduction using mirVana and TRIzol isolation methods. Int. J. Comput. Biol. Drug Des. 2014, 7, 102–112. [Google Scholar] [CrossRef]
- Moldovan, L.; Batte, K.E.; Trgovcich, J.; Wisler, J.; Marsh, C.B.; Piper, M. Methodological challenges in utilizing miRNAs as circulating biomarkers. J. Cell Mol. Med. 2014, 18, 371–390. [Google Scholar] [CrossRef]
- Zhang, L.; Wu, H.; Zhao, M.; Chang, C.; Lu, Q. Clinical significance of miRNAs in autoimmunity. J. Autoimmun. 2020, 109, 102438. [Google Scholar] [CrossRef]
- Ye, J.; Xu, M.; Tian, X.; Cai, S.; Zeng, S. Research advances in the detection of miRNA. J. Pharm. Anal. 2019, 9, 217–226. [Google Scholar] [CrossRef] [PubMed]
- Siddika, T.; Heinemann, I.U. Bringing MicroRNAs to Light: Methods for MicroRNA Quantification and Visualization in Live Cells. Front. Bioeng. Biotechnol. 2020, 8, 619583. [Google Scholar] [CrossRef] [PubMed]
- Bryzgunova, O.; Konoshenko, M.; Zaporozhchenko, I.; Yakovlev, A.; Laktionov, P. Isolation of Cell-Free miRNA from Biological Fluids: Influencing Factors and Methods. Diagnostics 2021, 11, 865. [Google Scholar] [CrossRef]
- El-Khoury, V.; Pierson, S.; Kaoma, T.; Bernardin, F.; Berchem, G. Assessing cellular and circulating miRNA recovery: The impact of the RNA isolation method and the quantity of input material. Sci. Rep. 2016, 6, 19529. [Google Scholar] [CrossRef] [PubMed]
- Tiberio, P.; Callari, M.; Angeloni, V.; Daidone, M.G.; Appierto, V. Challenges in using circulating miRNAs as cancer biomarkers. BioMed Res. Int. 2015, 2015, 731479. [Google Scholar] [CrossRef] [PubMed]
- Godoy, P.M.; Barczak, A.J.; DeHoff, P.; Srinivasan, S.; Etheridge, A.; Galas, D.; Das, S.; Erle, D.J.; Laurent, L.C. Comparison of Reproducibility, Accuracy, Sensitivity, and Specificity of miRNA Quantification Platforms. Cell Rep. 2019, 29, 4212–4222. [Google Scholar] [CrossRef]
- Morini, M.; Cangelosi, D.; Segalerba, D.; Marimpietri, D.; Raggi, F.; Castellano, A.; Fruci, D.; de Mora, J.F.; Canete, A.; Yanez, Y.; et al. Exosomal microRNAs from Longitudinal Liquid Biopsies for the Prediction of Response to Induction Chemotherapy in High-Risk Neuroblastoma Patients: A Proof of Concept SIOPEN Study. Cancers 2019, 11, 1476. [Google Scholar] [CrossRef] [PubMed]
- Chugh, P.; Dittmer, D.P. Potential pitfalls in microRNA profiling. Wiley Interdiscip. Rev. RNA 2012, 3, 601–616. [Google Scholar] [CrossRef]
- De Gonzalo-Calvo, D.; Pérez-Boza, J.; Curado, J.; Devaux, Y. Challenges of microRNA-based biomarkers in clinical application for cardiovascular diseases. Clin. Transl. Med. 2022, 12, e585. [Google Scholar] [CrossRef]
- Lashine, Y.A.; Salah, S.; Aboelenein, H.R.; Abdelaziz, A.I. Correcting the expression of miRNA-155 represses PP2Ac and enhances the release of IL-2 in PBMCs of juvenile SLE patients. Lupus 2015, 24, 240–247. [Google Scholar] [CrossRef]
- Kamiya, Y.; Kawada, J.; Kawano, Y.; Torii, Y.; Kawabe, S.; Iwata, N.; Ito, Y. Serum microRNAs as Potential Biomarkers of Juvenile Idiopathic Arthritis. Clin. Rheumatol. 2015, 34, 1705–1712. [Google Scholar] [CrossRef] [PubMed]
- Schulert, G.S.; Fall, N.; Harley, J.B.; Shen, N.; Lovell, D.J.; Thornton, S.; Grom, A.A. Monocyte microRNA expression in active systemic juvenile idiopathic arthritis implicates miR-125a-5p in polarized monocyte phenotypes. Arthritis Rheumatol. 2016, 68, 2300–2313. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Duan, M.; Feng, Y.; Geng, L.; Li, X.; Zhang, W. MiR-146a modulates macrophage polarization in systemic juvenile idiopathic arthritis by targeting INHBA. Mol. Immunol. 2016, 77, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Fan, Z.D.; Cao, Q.; Huang, N.; Ma, L.; Ma, H.H.; Zhang, Y.Y.; Yu, H.G.; Zhou, G.P. MicroRNA-125b regulates Th17/Treg cell differentiation and is associated with juvenile idiopathic arthritis. World J. Pediatr. 2020, 16, 99–110. [Google Scholar] [CrossRef]
- Rajendiran, A.; Klemm, P.; Schippers, A.; Scheufen, A.; Schwarz, T.; Peitz, J.; Brandenburg, L.O.; Wagner, N.; Consolaro, A.; Raggi, F.; et al. miR-23a contributes to T cellular redox metabolism in juvenile idiopathic oligoarthritis. Rheumatology 2021, 61, 2694–2703. [Google Scholar] [CrossRef]
- McAlpine, S.M.; Roberts, S.E.; Hargreaves, B.K.; Bullock, C.; Ramsey, S.; Stringer, E.; Lang, B.; Huber, A.; György, B.; Erdélyi, F.; et al. Differentially Expressed Inflammation-Regulating MicroRNAs in Oligoarticular Juvenile Idiopathic Arthritis. J. Rheumatol. 2023, 50, 227–235. [Google Scholar] [CrossRef]
- Ma, X.; Wu, F.; Xin, L.; Su, G.; He, F.; Yang, Y.; Sun, J.; Liu, Z. Differential plasma microRNAs expression in juvenile idiopathic arthritis. Mod. Rheumatol. 2016, 26, 224–232. [Google Scholar] [CrossRef]
- Demir, F.; Cebi, A.H.; Kalyoncu, M. Evaluation of plasma microRNA expressions in patients with juvenile idiopathic arthritis. Clin. Rheumatol. 2018, 37, 3255–3262. [Google Scholar] [CrossRef]
- Sun, J.; Feng, M.; Wu, F.; Ma, X.; Lu, J.; Kang, M.; Liu, Z. Plasma miR-26a as a Diagnostic Biomarker Regulates Cytokine Expression in Systemic Juvenile Idiopathic Arthritis. J. Rheumatol. 2016, 43, 1607–1614. [Google Scholar] [CrossRef]
- Nziza, N.; Jeziorski, E.; Delpont, M.; Cren, M.; Chevassus, H.; Carbasse, A.; Mahe, P.; Abassi, H.; Joly-Monrigal, P.; Schordan, E.; et al. Synovial-Fluid miRNA Signature for Diagnosis of Juvenile Idiopathic Arthritis. Cells 2019, 8, 1521. [Google Scholar] [CrossRef]
- Raggi, F.; Cangelosi, D.; Consolaro, A.; Rossi, C.; Pelassa, S.; Cortese, K.; Gagliani, M.C.; Morini, M.; Segalerba, D.; Brignole, C.; et al. Extracellular vesicle-derived microRNAs as potential biomarkers in oligoarticular juvenile idiopathic arthritis patients: Methodological challenges and new perspectives. Clin. Transl. Med. 2022, 12, e1067. [Google Scholar] [CrossRef] [PubMed]
- Orczyk, K.; Smolewska, E. The Potential Importance of MicroRNAs as Novel Indicators How to Manage Patients with Juvenile Idiopathic Arthritis More Effectively. J. Immunol. Res. 2021, 2021, 9473508. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, S.; Xie, N.; Cui, H.; Tan, Z.; Yang, S.; Icyuz, M.; Abraham, E.; Liu, G. MicroRNA let-7c regulates macrophage polarization. J. Immunol. 2013, 190, 6542–6549. [Google Scholar] [CrossRef] [PubMed]
- Dufourd, T.; Robil, N.; Mallet, D.; Carcenac, C.; Boulet, S.; Brishoual, S.; Rabois, E.; Houeto, J.L.; de la Grange, P.; Carnicella, S. Plasma or serum? A qualitative study on rodents and humans using high-throughput microRNA sequencing for circulating biomarkers. Biol. Methods Protoc. 2019, 4, bpz006. [Google Scholar] [CrossRef]
- Withrow, J.; Murphy, C.; Liu, Y.; Hunter, M.; Fulzele, S.; Hamrick, M.W. Extracellular vesicles in the pathogenesis of rheumatoid arthritis and osteoarthritis. Arthritis Res. Ther. 2016, 18, 286. [Google Scholar] [CrossRef] [PubMed]
- Raggi, F.; Bartolucci, M.; Cangelosi, D.; Rossi, C.; Pelassa, S.; Trincianti, C.; Petretto, A.; Filocamo, G.; Civino, A.; Eva, A.; et al. Proteomic profiling of extracellular vesicles in synovial fluid and plasma from Oligoarticular Juvenile Idiopathic Arthritis patients reveals novel immunopathogenic biomarkers. Front. Immunol. 2023, 14, 1134747. [Google Scholar] [CrossRef]
- De, T.J.; Herschlik, L.; Waldner, C.; Mongini, C. Emerging roles of exosomes in normal and pathological conditions: New insights for diagnosis and therapeutic applications. Front. Immunol. 2015, 6, 203. [Google Scholar]
- Shah, R.; Patel, T.; Freedman, J.E. Circulating Extracellular Vesicles in Human Disease. N. Engl. J. Med. 2018, 379, 958–966. [Google Scholar] [CrossRef]
- Menck, K.; Sivaloganathan, S.; Bleckmann, A.; Binder, C. Microvesicles in Cancer: Small Size, Large Potential. Int. J. Mol. Sci. 2020, 21, 5373. [Google Scholar] [CrossRef]
- Resaz, R.; Cangelosi, D.; Segalerba, D.; Morini, M.; Uva, P.; Bosco, M.C.; Banderali, G.; Estrella, A.; Wanner, C.; Weinstein, D.A.; et al. Exosomal MicroRNAs as Potential Biomarkers of Hepatic Injury and Kidney Disease in Glycogen Storage Disease Type Ia Patients. Int. J. Mol. Sci. 2021, 23, 328. [Google Scholar] [CrossRef]
- Lipps, C.; Northe, P.; Figueiredo, R.; Rohde, M.; Brahmer, A.; Krämer-Albers, E.M.; Liebetrau, C.; Wiedenroth, C.B.; Mayer, E.; Kriechbaum, S.D.; et al. Non-Invasive Approach for Evaluation of Pulmonary Hypertension Using Extracellular Vesicle-Associated Small Non-Coding RNA. Biomolecules 2019, 9, 666. [Google Scholar] [CrossRef] [PubMed]
- Foers, A.D.; Garnham, A.L.; Chatfield, S.; Smyth, G.K.; Cheng, L.; Hill, A.F.; Wicks, I.P.; Pang, K.C. Extracellular Vesicles in Synovial Fluid from Rheumatoid Arthritis Patients Contain miRNAs with Capacity to Modulate Inflammation. Int. J. Mol. Sci. 2021, 22, 4910. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Patro, P.S.; Aggarwal, A. MicroRNA-132, miR-146a, and miR-155 as potential biomarkers of methotrexate response in patients with rheumatoid arthritis. Clin. Rheumatol. 2019, 38, 877–884. [Google Scholar] [CrossRef]
- Daien, C.; Krogulec, M.; Gineste, P.; Steens, J.M.; Desroys du Roure, L.; Biguenet, S.; Scherrer, D.; Santo, J.; Ehrlich, H.; Durez, P. Safety and efficacy of the miR-124 upregulator ABX464 (obefazimod, 50 and 100 mg per day) in patients with active rheumatoid arthritis and inadequate response to methotrexate and/or anti-TNFα therapy: A placebo-controlled phase II study. Ann. Rheum. Dis. 2022, 81, 1076–1084. [Google Scholar] [CrossRef]
- Castro-Villegas, C.; Pérez-Sánchez, C.; Escudero, A.; Filipescu, I.; Verdu, M.; Ruiz-Limón, P.; Aguirre, M.A.; Jiménez-Gomez, Y.; Font, P.; Rodriguez-Ariza, A.; et al. Circulating miRNAs as potential biomarkers of therapy effectiveness in rheumatoid arthritis patients treated with anti-TNFα. Arthritis Res. Ther. 2015, 17, 49. [Google Scholar] [CrossRef] [PubMed]
- Pei, D.; Cao, J.; Qin, G.; Wang, X. Measurement of circulating miRNA-125a exhibits good value in the management of etanercept-treated psoriatic patients. J. Dermatol. 2020, 47, 140–146. [Google Scholar] [CrossRef] [PubMed]
| Subtype a | Characteristics | Onset Age and Sex | Treatment | Biomarkers b |
|---|---|---|---|---|
| Systemic arthritis | ≤10% JIA patients. Number of affected joints ≥ 5. Systemic inflammatory features (continuous fever for at least 2 weeks, rashes, muscle pain, generalized symmetrical lymphadenopathy, enlargement of liver or spleen, or serositis) [12]. | Throughout childhood, F = M [1] | Thalidomide (rarely), systemic glucocorticoids, NSAIDs csDMARDs: MTX (with predominant joint inflammation and without active systemic symptoms), bDMARDs: anti-IL-1 (Anakinra, Canakinumab, Rilonacept) [15], anti-IL-6 (Tocilizumab) [15]. | S100 proteins IL-18, IL-6, Ferritin [16] RF HLA-B27 ANA Anti-CCP antibodies ESR CRP JADAS [16,17,18] |
| Oligoarthritis | The most common in Western Countries (50–80% cases in Europe and North America) [19]. Number of affected joints within the first six months of disease ≤ 4. High frequency of positivity to ANA. Association with HLA-DRB1*0801 [2]. | 2–4 y, early childhood, F > > > M [1] | Intrarticular corticosteroid injection(s), NSAIDs csDMARDs: MTX [20], bDMARDSs: TNF inhibitor (Etanercept) [15]. | RF HLA-B27 ANA Anti-CCP antibodies ESR CRP JADAS [16,17,18] |
| Persistent oligoarthritis | Number of affected joints ≤ 4 after the first 6 months from onset [14]. | |||
| Extended oligoarthritis | Number of affected joints extends to >4 after the first 6 months of disease [14]. | |||
| Polyarthritis | Number of affected joints ≥ 5 after the first 6 months from onset, in the absence of fever and in the presence or absence of IgM RF [14]. | csDMARDs: MTX, Leflunomide (patients who cannot tolerate MTX), bDMARD: anti-TNF (Etanercept, adalimumab, golimumab), anti-IL-6 (Tocilizumab), anti-CTLA4 (Abatacept-), and Janus kinase (JAK) inhibitors (Tofacitinib) [3]. | RF HLA-B27 ANA Anti-CCP antibodies ESR CRP JADAS [16,17,18] | |
| Rheumatoid-factor-positive polyarthritis | 5% of JIA cases. Number of affected joints ≥ 5 during the first 6 months of disease. Psitivity for RF [14]. | Late childhood or adolescence, F > > M [1] | ||
| Rheumatoid factor negative polyarthritis | 15–20% of JIA cases. Number of affected joints ≥ 5 during the first 6 months of disease. Negativity for RF [14]. | Early peak 2–4 y and later peak at 6–12 y, F > > M [1] | ||
| Enthesitis-related arthritis | Arthritis and enthesitis. Tow or more of the following: presence/history of sacroiliac joint tenderness (with or without lumbosacral pain), HLA-B27 antigen positivity, onset of arthritis in a male >6 years old, acute symptomatic anterior uveitis [14]. | Late childhood or adolescence, M > >F [1] | csDMARDs: Sulfasalazine (patients with moderate activity), bDMARDs: TNF inhibitors (Adalimumab, Etanercept) [15]. | RF HLA-B27 ANA Anti-CCP antibodies ESR CRP JADAS [16,17,18] |
| Psoriatic arthritis | Characterized by arthritis and psoriasis or arthritis and at least two of the following: dactylitis, nail pitting or onycholysis, psoriasis in a first-degree relative. Psoriasis can occur before or after arthritis onset [21]. | Early peak at 2–4 y and later peak at 9–11 y, F > M [1] | csDMARDs: MTX [15], bDMARDs: TNF inhibitors (Adalimumab, Etanercept, Infliximab). | RF HLA-B27 ANA Anti-CCP antibodies ESR CRP JADAS [16,17,18] |
| Undifferentiated arthritis | Not fulfilling any of the listed categories or having criteria formore than one of them. |
| Authors | N° of Patients | Disease Type (N° of Patients) | Types of Samples | Techniques for miRNA Isolation | Techniques for miRNA Analysis | Types of miRNAs |
|---|---|---|---|---|---|---|
| Lashine et al. [100] | 10 | NA | Peripheral blood mononuclear cells | mirVana miRNA extraction kit | RT-qPCR (TaqMan) | Free circulating |
| Kamiya et al. [101] | 6 24 | sJIA (3), pJIA (3) sJIA (8), pJIA (16) | Peripheral blood leukocytes Serum | mirVana PARIS Kit | RT-qPCR (TaqMan) | Free circulating |
| Schulert et al. [102] | 27 | sJIA patients: NOS (7) active established (9) CID (11) | Peripheral blood monocytes | mirVana miRNA isolation kit | Microarray | Free circulating |
| Li et al. [103] | 32 | sJIA patients: | Peripheral blood monocytes | TRIzol® LS reagent | qPCR (TaqMan) | Free circulating |
| NOS (8) | ||||||
| established disease (11) | ||||||
| CID (13) | ||||||
| Fan et al. [104] | 16 | RF-positive pJIA | PBMCs/CD4+ T lymphocytes | qPCR (SYBR Green) | Free circulating | |
| Rajendiran et al. [105] | 9 | oJIA | Synovial Fluid Mononuclear cells | RNeasy Mini Kit | Gene Chip Array | Free circulating |
| McAlpine et al. [106] | 40 | oJIA | Peripheral Blood Leukocytes | miRNeasy Mini Kit | RT-PCR (SYBR Green) | Free circulating |
| 20 | Plasma | miRNeasy Serum/Plasma Kit, ExoRNeasy Serum/Plasma Midi Kit | Digital droplet PCR | Free circulating -Encapsulated | ||
| 20 | Synovial Fluid | miRNeasy Serum/PlasmaKit, ExoRNeasy Serum/Plasma Midi Kit | digital droplet PCR | Free circulating -Encapsulated | ||
| Ma et al. [107] | 109 | oJIA (43) | Plasma | TRIzol® LS reagent | Microarray-RT-qPCR (SYBR® Green) | Free circulating |
| pJIA (37) | ||||||
| JAS (29) | ||||||
| Demir et al. [108] | 31 | oJIA (17) | Plasma | miRNeasy serum/plasma kit | RT-qPCR (SYBR® Green) | Free circulating |
| pJIA (9) | ||||||
| ERA (5) | ||||||
| Sun et al. [109] | 155 | sJIA (20) | Plasma | TRIzol® LS reagent | Microarray | Free circulating |
| JAS (30) | ||||||
| SLE (25) | ||||||
| KD (40) | ||||||
| HSP (40) | ||||||
| Nziza et al. [110] | 34 | oJIA (18) | Serum | miRNeasy Serum/Plasma kit | RT-PCR (TaqMan) | Free circulating |
| SA (16) | Synovial Fluid | |||||
| Raggi et al. [111] | 13 | oJIA | Plasma | exoRNeasy Serum/Plasma Midi kit | Array Card | Encapsulated |
| Synovial Fluid |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pelassa, S.; Raggi, F.; Rossi, C.; Bosco, M.C. MicroRNAs in Juvenile Idiopathic Arthritis: State of the Art and Future Perspectives. Biology 2023, 12, 991. https://doi.org/10.3390/biology12070991
Pelassa S, Raggi F, Rossi C, Bosco MC. MicroRNAs in Juvenile Idiopathic Arthritis: State of the Art and Future Perspectives. Biology. 2023; 12(7):991. https://doi.org/10.3390/biology12070991
Chicago/Turabian StylePelassa, Simone, Federica Raggi, Chiara Rossi, and Maria Carla Bosco. 2023. "MicroRNAs in Juvenile Idiopathic Arthritis: State of the Art and Future Perspectives" Biology 12, no. 7: 991. https://doi.org/10.3390/biology12070991
APA StylePelassa, S., Raggi, F., Rossi, C., & Bosco, M. C. (2023). MicroRNAs in Juvenile Idiopathic Arthritis: State of the Art and Future Perspectives. Biology, 12(7), 991. https://doi.org/10.3390/biology12070991