On Origin and Evolution of the Antibody Molecule
Abstract
Simple Summary
Abstract
1. Introduction
2. The Ig Fold in Bacteria
3. The Ig Fold in Viruses
4. The Eukaryotic IgV Domain
5. Somatic Recombination
6. Somatic HyperMutation
7. The Ig Domain in Adaptive Immune Receptors: IgC1
8. Evolution of the Antibody Architecture
8.1. The Modular Structure
8.2. The Dynamic Structure
9. Evolution Sites of the Antibody Molecule Involved in Effector Functions
9.1. Fc Receptors
9.2. Factor C1q of the Complement System
9.3. Superantigens
10. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Janeway, C.A.; Travers, P.; Walport, M.; Shlomchik, M.J. Immunobiology: The Immune System in Health and Disease, 5th ed.; Garland Science: New York, NY, USA, 2001. [Google Scholar]
- Matz, H.; Munir, D.; Logue, J.; Dooley, H. The Immunoglobulins of Cartilaginous Fishes. Dev. Comp. Immunol. 2021, 115, 103873. [Google Scholar] [CrossRef]
- Eason, D.D.; Cannon, J.P.; Haire, R.N.; Rast, J.P.; Ostrov, D.A.; Litman, G.W. Mechanisms of Antigen Receptor Evolution. Semin. Immunol. 2004, 16, 215–226. [Google Scholar] [CrossRef]
- Lesk, A.M.; Chothia, C. Evolution of Proteins Formed by Beta-Sheets. II. The Core of the Immunoglobulin Domains. J. Mol. Biol. 1982, 160, 325–342. [Google Scholar] [CrossRef]
- Feige, M.J.; Hendershot, L.M.; Buchner, J. How Antibodies Fold. Trends Biochem. Sci. 2010, 35, 189–198. [Google Scholar] [CrossRef]
- Hofmann, B.E.; Bender, H.; Schulz, G.E. Three-Dimensional Structure of Cyclodextrin Glycosyltransferase from Bacillus Circulans at 3·4 Å Resolution. J. Mol. Biol. 1989, 209, 793–800. [Google Scholar] [CrossRef]
- Braun, T.; Vos, M.R.; Kalisman, N.; Sherman, N.E.; Rachel, R.; Wirth, R.; Schröder, G.F.; Egelman, E.H. Archaeal Flagellin Combines a Bacterial Type IV Pilin Domain with an Ig-like Domain. Proc. Natl. Acad. Sci. USA 2016, 113, 10352–10357. [Google Scholar] [CrossRef]
- Bateman, A.; Eddy, S.R.; Chothia, C. Members of the Immunoglobulin Superfamily in Bacteria. Protein Sci. 1996, 5, 1939–1941. [Google Scholar] [CrossRef]
- Raman, R.; Rajanikanth, V.; Palaniappan, R.U.M.; Lin, Y.-P.; He, H.; McDonough, S.P.; Sharma, Y.; Chang, Y.-F. Big Domains Are Novel Ca2+-Binding Modules: Evidences from Big Domains of Leptospira Immunoglobulin-Like (Lig) Proteins. PLoS ONE 2010, 5, e14377. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Zhang, J.; Zhang, X.; Xu, C.; Tu, X. Solution Structure of the Big Domain from Streptococcus Pneumoniae Reveals a Novel Ca2+-Binding Module. Sci. Rep. 2013, 3, 1079. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Lu, P.; Pan, Z.; Zhu, Y.; Ma, J.; Zhong, X.; Dong, W.; Lu, C.; Yao, H. SssP1, a Streptococcus Suis Fimbria-Like Protein Transported by the SecY2/A2 System, Contributes to Bacterial Virulence. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Kakuda, S.; Yukitake, H.; Kondo, Y.; Shoji, M.; Takebe, K.; Narita, Y.; Naito, M.; Nakane, D.; Abiko, Y.; et al. Immunoglobulin-like Domains of the Cargo Proteins Are Essential for Protein Stability during Secretion by the Type IX Secretion System. Mol. Microbiol. 2018, 110, 64–81. [Google Scholar] [CrossRef] [PubMed]
- Guttula, D.; Yao, M.; Baker, K.; Yang, L.; Goult, B.T.; Doyle, P.S.; Yan, J. Calcium-Mediated Protein Folding and Stabilization of Salmonella Biofilm-Associated Protein A. J. Mol. Biol. 2019, 431, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Hüttener, M.; Prieto, A.; Aznar, S.; Bernabeu, M.; Glaría, E.; Valledor, A.F.; Paytubi, S.; Merino, S.; Tomás, J.; Juárez, A. Expression of a Novel Class of Bacterial Ig-like Proteins Is Required for IncHI Plasmid Conjugation. PLoS Genet. 2019, 15, e1008399. [Google Scholar] [CrossRef]
- Vance, T.D.R.; Graham, L.A.; Davies, P.L. An Ice-Binding and Tandem Beta-Sandwich Domain-Containing Protein in Shewanella Frigidimarina Is a Potential New Type of Ice Adhesin. FEBS J. 2018, 285, 1511–1527. [Google Scholar] [CrossRef]
- Farré, D.; Martínez-Vicente, P.; Engel, P.; Angulo, A. Immunoglobulin Superfamily Members Encoded by Viruses and Their Multiple Roles in Immune Evasion. Eur. J. Immunol. 2017, 47, 780–796. [Google Scholar] [CrossRef]
- Tan, Y.; Schneider, T.; Leong, M.; Aravind, L.; Zhang, D. Novel Immunoglobulin Domain Proteins Provide Insights into Evolution and Pathogenesis of SARS-CoV-2-Related Viruses. mBio 2020, 11, e00760-20. [Google Scholar] [CrossRef] [PubMed]
- Mijnes, J.D.; Lutters, B.C.; Vlot, A.C.; van Anken, E.; Horzinek, M.C.; Rottier, P.J.; de Groot, R.J. Structure-Function Analysis of the GE-GI Complex of Feline Herpesvirus: Mapping of GI Domains Required for GE-GI Interaction, Intracellular Transport, and Cell-to-Cell Spread. J. Virol. 1997, 71, 8397–8404. [Google Scholar] [CrossRef]
- Hewitt, E.W. The MHC class I antigen presentation pathway: Strategies for viral immune evasion. Immunology 2003, 110, 163–169. [Google Scholar] [CrossRef] [PubMed]
- Vasiliauskaite, I.; Owsianka, A.; England, P.; Khan, A.G.; Cole, S.; Bankwitz, D.; Foung, S.K.H.; Pietschmann, T.; Marcotrigiano, J.; Rey, F.A.; et al. Conformational Flexibility in the Immunoglobulin-Like Domain of the Hepatitis C Virus Glycoprotein E2. mBio 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.S.; Yu, Z.; Maxwell, K.L.; Davidson, A.R. Ig-like Domains on Bacteriophages: A Tale of Promiscuity and Deceit. J. Mol. Biol. 2006, 359, 496–507. [Google Scholar] [CrossRef] [PubMed]
- Williams, A.F.; Barclay, A.N. The Immunoglobulin Superfamily--Domains for Cell Surface Recognition. Annu. Rev. Immunol. 1988, 6, 381–405. [Google Scholar] [CrossRef]
- Jones, E.Y.; Davis, S.J.; Williams, A.F.; Harlos, K.; Stuart, D.I. Crystal Structure at 2.8 A Resolution of a Soluble Form of the Cell Adhesion Molecule CD2. Nature 1992, 360, 232–239. [Google Scholar] [CrossRef]
- Bourgois, A. Evidence for an Ancestral Immunoglobulin Gene Coding for Half a Domain. Immunochemistry 1975, 12, 873–876. [Google Scholar] [CrossRef]
- Cooper, E.L.; Mansour, M.H. Distribution of Thy-1 in Invertebrates and Ectothermic Vertebrates. Immunol. Ser. 1989, 45, 197–219. [Google Scholar]
- Kubrycht, J.; Borecký, J.; Souček, P.; Ježek, P. Sequence Similarities of Protein Kinase Substrates and Inhibitors with Immunoglobulins and Model Immunoglobulin Homologue: Cell Adhesion Molecule from the Living Fossil Sponge Geodia Cydonium. Mapping of Coherent Database Similarities and Implications for Evolution of CDR1 and Hypermutation. Folia Microbiol. 2004, 49, 219–246. [Google Scholar]
- Holland, L.Z.; Albalat, R.; Azumi, K.; Benito-Gutiérrez, È.; Blow, M.J.; Bronner-Fraser, M.; Brunet, F.; Butts, T.; Candiani, S.; Dishaw, L.J.; et al. The Amphioxus Genome Illuminates Vertebrate Origins and Cephalochordate Biology. Genome Res. 2008, 18, 1100–1111. [Google Scholar] [CrossRef]
- Huang, S.; Yuan, S.; Guo, L.; Yu, Y.; Li, J.; Wu, T.; Liu, T.; Yang, M.; Wu, K.; Liu, H.; et al. Genomic Analysis of the Immune Gene Repertoire of Amphioxus Reveals Extraordinary Innate Complexity and Diversity. Genome Res. 2008, 18, 1112–1126. [Google Scholar] [CrossRef]
- Cannon, J.P.; Haire, R.N.; Schnitker, N.; Mueller, M.G.; Litman, G.W. Individual Protochordates Have Unique Immune-Type Receptor Repertoires. Curr. Biol. 2004, 14, R465–R466. [Google Scholar] [CrossRef] [PubMed]
- Dishaw, L.J.; Giacomelli, S.; Melillo, D.; Zucchetti, I.; Haire, R.N.; Natale, L.; Russo, N.A.; Santis, R.D.; Litman, G.W.; Pinto, M.R. A Role for Variable Region-Containing Chitin-Binding Proteins (VCBPs) in Host Gut–Bacteria Interactions. Proc. Natl. Acad. Sci. USA 2011, 108, 16747–16752. [Google Scholar] [CrossRef]
- Hernández Prada, J.A.; Haire, R.N.; Allaire, M.; Jakoncic, J.; Stojanoff, V.; Cannon, J.P.; Litman, G.W.; Ostrov, D.A. Ancient Evolutionary Origin of Diversified Variable Regions Demonstrated by Crystal Structures of an Immune-Type Receptor in Amphioxus. Nat. Immunol. 2006, 7, 875–882. [Google Scholar] [CrossRef]
- Franchi, N.; Ballarin, L. Immunity in Protochordates: The Tunicate Perspective. Front. Immunol. 2017, 8, 674. [Google Scholar] [CrossRef]
- Pancer, Z.; Diehl-Seifert, B.; Rinkevich, B.; Müller, W.E. A Novel Tunicate (Botryllus schlosseri) Putative C-Type Lectin Features an Immunoglobulin Domain. DNA Cell Biol. 1997, 16, 801–806. [Google Scholar] [CrossRef] [PubMed]
- Chen, R.; Zhang, L.; Qi, J.; Zhang, N.; Zhang, L.; Yao, S.; Wu, Y.; Jiang, B.; Wang, Z.; Yuan, H.; et al. Discovery and Analysis of Invertebrate IgVJ-C2 Structure from Amphioxus Provides Insight into the Evolution of the Ig Superfamily. J. Immunol. 2018, 200, 2869–2881. [Google Scholar] [CrossRef]
- Ely, K.R.; Herron, J.N.; Harker, M.; Edmundson, A.B. Three-Dimensional Structure of a Light Chain Dimer Crystallized in Water: Conformational Flexibility of a Molecule in Two Crystal Forms. J. Mol. Biol. 1989, 210, 601–615. [Google Scholar] [CrossRef]
- Strong, S.J.; Mueller, M.G.; Litman, R.T.; Hawke, N.A.; Haire, R.N.; Miracle, A.L.; Rast, J.P.; Amemiya, C.T.; Litman, G.W. A Novel Multigene Family Encodes Diversified Variable Regions. Proc. Natl. Acad. Sci. USA 1999, 96, 15080–15085. [Google Scholar] [CrossRef] [PubMed]
- Litman, G.W.; Hawke, N.A.; Yoder, J.A. Novel Immune-Type Receptor Genes. Immunol. Rev. 2001, 181, 250–259. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, A.; Eastman, Q.M.; Schatz, D.G. Transposition Mediated by RAG1 and RAG2 and Its Implications for the Evolution of the Immune System. Nature 1998, 394, 744–751. [Google Scholar] [CrossRef] [PubMed]
- Hiom, K.; Melek, M.; Gellert, M. DNA Transposition by the RAG1 and RAG2 Proteins: A Possible Source of Oncogenic Translocations. Cell 1998, 94, 463–470. [Google Scholar] [CrossRef]
- Ferraresso, S.; Kuhl, H.; Milan, M.; Ritchie, D.W.; Secombes, C.J.; Reinhardt, R.; Bargelloni, L. Identification and Characterisation of a Novel Immune-Type Receptor (NITR) Gene Cluster in the European Sea Bass, Dicentrarchus Labrax, Reveals Recurrent Gene Expansion and Diversification by Positive Selection. Immunogenetics 2009, 61, 773–788. [Google Scholar] [CrossRef]
- Litman, G.W.; Cannon, J.P.; Dishaw, L.J.; Haire, R.N.; Eason, D.D.; Yoder, J.A.; Prada, J.H.; Ostrov, D.A. Immunoglobulin Variable Regions in Molecules Exhibiting Characteristics of Innate and Adaptive Immune Receptors. Immunol. Res. 2007, 38, 294–304. [Google Scholar] [CrossRef]
- Streltsov, V.A.; Carmichael, J.A.; Nuttall, S.D. Structure of a Shark IgNAR Antibody Variable Domain and Modeling of an Early-Developmental Isotype. Protein Sci. 2005, 14, 2901–2909. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, K.; Deng, A.; Fu, X.; Xu, A.; Liu, X. An Amphioxus RAG1-like DNA Fragment Encodes a Functional Central Domain of Vertebrate Core RAG1. Proc. Natl. Acad. Sci. USA 2014, 111, 397–402. [Google Scholar] [CrossRef] [PubMed]
- Fugmann, S.D.; Messier, C.; Novack, L.A.; Cameron, R.A.; Rast, J.P. An Ancient Evolutionary Origin of the Rag1/2 Gene Locus. Proc. Natl. Acad. Sci. USA 2006, 103, 3728–3733. [Google Scholar] [CrossRef] [PubMed]
- Bonatto, D.; Brendel, M.; Henriques, J.A.P. In Silico Identification and Analysis of New Artemis/Artemis-like Sequences from Fungal and Metazoan Species. Protein J. 2005, 24, 399–411. [Google Scholar] [CrossRef] [PubMed]
- Muramatsu, M.; Sankaranand, V.S.; Anant, S.; Sugai, M.; Kinoshita, K.; Davidson, N.O.; Honjo, T. Specific Expression of Activation-Induced Cytidine Deaminase (AID), a Novel Member of the RNA-Editing Deaminase Family in Germinal Center B Cells. J. Biol. Chem. 1999, 274, 18470–18476. [Google Scholar] [CrossRef]
- Muramatsu, M.; Kinoshita, K.; Fagarasan, S.; Yamada, S.; Shinkai, Y.; Honjo, T. Class Switch Recombination and Hypermutation Require Activation-Induced Cytidine Deaminase (AID), a Potential RNA Editing Enzyme. Cell 2000, 102, 553–563. [Google Scholar] [CrossRef]
- Rada, C.; Ehrenstein, M.R.; Neuberger, M.S.; Milstein, C. Hot Spot Focusing of Somatic Hypermutation in MSH2-Deficient Mice Suggests Two Stages of Mutational Targeting. Immunity 1998, 9, 135–141. [Google Scholar] [CrossRef]
- Krishnan, A.; Iyer, L.M.; Holland, S.J.; Boehm, T.; Aravind, L. Diversification of AID/APOBEC-like Deaminases in Metazoa: Multiplicity of Clades and Widespread Roles in Immunity. Proc. Natl. Acad. Sci. USA 2018, 115, E3201–E3210. [Google Scholar] [CrossRef]
- Newman, E.N.C.; Holmes, R.K.; Craig, H.M.; Klein, K.C.; Lingappa, J.R.; Malim, M.H.; Sheehy, A.M. Antiviral Function of APOBEC3G Can Be Dissociated from Cytidine Deaminase Activity. Curr. Biol. 2005, 15, 166–170. [Google Scholar] [CrossRef]
- Liu, M.-C.; Liao, W.-Y.; Buckley, K.M.; Yang, S.Y.; Rast, J.P.; Fugmann, S.D. AID/APOBEC-like Cytidine Deaminases Are Ancient Innate Immune Mediators in Invertebrates. Nat. Commun. 2018, 9, 1948. [Google Scholar] [CrossRef]
- Conticello, S.G.; Thomas, C.J.F.; Petersen-Mahrt, S.K.; Neuberger, M.S. Evolution of the AID/APOBEC Family of Polynucleotide (Deoxy)Cytidine Deaminases. Mol. Biol. Evol. 2005, 22, 367–377. [Google Scholar] [CrossRef] [PubMed]
- Barreto, V.M.; Pan-Hammarstrom, Q.; Zhao, Y.; Hammarstrom, L.; Misulovin, Z.; Nussenzweig, M.C. AID from Bony Fish Catalyzes Class Switch Recombination. J. Exp. Med. 2005, 202, 733–738. [Google Scholar] [CrossRef] [PubMed]
- Conticello, S.G. The AID/APOBEC family of nucleic acid mutators. Genome Biol. 2008, 9, 229. [Google Scholar] [CrossRef] [PubMed]
- Rogozin, I.B.; Iyer, L.M.; Liang, L.; Glazko, G.V.; Liston, V.G.; Pavlov, Y.I.; Aravind, L.; Pancer, Z. Evolution and Diversification of Lamprey Antigen Receptors: Evidence for Involvement of an AID-APOBEC Family Cytosine Deaminase. Nat. Immunol. 2007, 8, 647–656. [Google Scholar] [CrossRef]
- Dooley, H.; Stanfield, R.L.; Brady, R.A.; Flajnik, M.F. First Molecular and Biochemical Analysis of In Vivo Affinity Maturation in an Ectothermic Vertebrate. Proc. Natl. Acad. Sci. USA 2006, 103, 1846–1851. [Google Scholar] [CrossRef]
- Malecek, K.; Brandman, J.; Brodsky, J.E.; Ohta, Y.; Flajnik, M.F.; Hsu, E. Somatic Hypermutation and Junctional Diversification at Ig Heavy Chain Loci in the Nurse Shark. J. Immunol. 2005, 175, 8105–8115. [Google Scholar] [CrossRef]
- Zhu, C.; Hsu, E. Error-Prone DNA Repair Activity during Somatic Hypermutation in Shark B Lymphocytes. J. Immunol. 2010, 185, 5336–5347. [Google Scholar] [CrossRef]
- Oreste, U.; Coscia, M.R. Specific Features of Immunoglobulin VH Genes of the Antarctic Teleost Trematomus bernacchii. Gene 2002, 295, 199–204. [Google Scholar] [CrossRef]
- Coscia, M.R.; Oreste, U. Limited Diversity of the Immunoglobulin Heavy Chain Variable Domain of the Emerald Rockcod Trematomus bernacchii. Fish Shellfish Immunol. 2003, 14, 71–92. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Springer, T.A. Structural Specializations of Immunoglobulin Superfamily Members for Adhesion to Integrins and Viruses. Immunol. Rev. 1998, 163, 197–215. [Google Scholar] [CrossRef] [PubMed]
- Schluter, S.F.; Bernstein, R.M.; Bernstein, H.; Marchalonis, J.J. “Big Bang” Emergence of the Combinatorial Immune System. Dev. Comp. Immunol. 1999, 23, 107–111. [Google Scholar]
- Potapov, V.; Sobolev, V.; Edelman, M.; Kister, A.; Gelfand, I. Protein–Protein Recognition: Juxtaposition of Domain and Interface Cores in Immunoglobulins and Other Sandwich-like Proteins. J. Mol. Biol. 2004, 342, 665–679. [Google Scholar] [CrossRef]
- Chintalacharuvu, K.R.; Morrison, S.L. Residues critical for H-L disulfide bond formation in human IgA1 and IgA2. J. Immunol. 1996, 157, 3443–3449. [Google Scholar] [PubMed]
- Savan, R.; Aman, A.; Sato, K.; Yamaguchi, R.; Sakai, M. Discovery of a New Class of Immunoglobulin Heavy Chain from Fugu. Eur. J. Immunol. 2005, 35, 3320–3331. [Google Scholar] [CrossRef] [PubMed]
- Zimmerman, A.M.; Moustafa, F.M.; Romanowski, K.E.; Steiner, L.A. Zebrafish Immunoglobulin IgD: Unusual Exon Usage and Quantitative Expression Profiles with IgM and IgZ/T Heavy Chain Isotypes. Mol. Immunol. 2011, 48, 2220–2223. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, A.S.; Avila, D.; Hughes, M.; Hughes, A.; McKinney, E.C.; Flajnik, M.F. A New Antigen Receptor Gene Family That Undergoes Rearrangement and Extensive Somatic Diversification in Sharks. Nature 1995, 374, 168–173. [Google Scholar] [CrossRef] [PubMed]
- Feige, M.J.; Gräwert, M.A.; Marcinowski, M.; Hennig, J.; Behnke, J.; Ausländer, D.; Herold, E.M.; Peschek, J.; Castro, C.D.; Flajnik, M.; et al. The Structural Analysis of Shark IgNAR Antibodies Reveals Evolutionary Principles of Immunoglobulins. Proc. Natl. Acad. Sci. USA 2014, 111, 8155–8160. [Google Scholar] [CrossRef] [PubMed]
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hamers, C.; Songa, E.B.; Bendahman, N.; Hamers, R. Naturally Occurring Antibodies Devoid of Light Chains. Nature 1993, 363, 446–448. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.K.; Brewer, J.W.; Hellman, R.; Hendershot, L.M. BiP and Immunoglobulin Light Chain Cooperate to Control the Folding of Heavy Chain and Ensure the Fidelity of Immunoglobulin Assembly. Mol. Biol. Cell. 1999, 10, 2209–2219. [Google Scholar] [CrossRef] [PubMed]
- Roux, K.H.; Greenberg, A.S.; Greene, L.; Strelets, L.; Avila, D.; McKinney, E.C.; Flajnik, M.F. Structural Analysis of the Nurse Shark (New) Antigen Receptor (NAR): Molecular Convergence of NAR and Unusual Mammalian Immunoglobulins. Proc. Natl. Acad. Sci. USA 1998, 95, 11804–11809. [Google Scholar] [CrossRef]
- Mirete-Bachiller, S.; Olivieri, D.N.; Gambón-Deza, F. Immunoglobulin T Genes in Actinopterygii. Fish Shellfish Immunol. 2021, 108, 86–93. [Google Scholar] [CrossRef]
- Tongsri, P.; Meng, K.; Liu, X.; Wu, Z.; Yin, G.; Wang, Q.; Liu, M.; Xu, Z. The Predominant Role of Mucosal Immunoglobulin IgT in the Gills of Rainbow Trout (Oncorhynchus mykiss) after Infection with Flavobacterium columnare. Fish Shellfish Immunol. 2020, 99, 654–662. [Google Scholar] [CrossRef]
- Zhang, W.; Qin, L.; Cai, X.; Juma, S.N.; Xu, R.; Wei, L.; Wu, Y.; Cui, X.; Chen, G.; Liu, L.; et al. Sequence Structure Character of IgNAR Sec in Whitespotted Bamboo Shark (Chiloscyllium plagiosum). Fish Shellfish Immunol. 2020, 102, 140–144. [Google Scholar] [CrossRef]
- Zhang, Y.-A.; Salinas, I.; Sunyer, J.O. Recent Findings on the Structure and Function of Teleost IgT. Fish Shellfish Immunol. 2011, 31, 627–634. [Google Scholar] [CrossRef]
- Ho, B.K.; Coutsias, E.A.; Seok, C.; Dill, K.A. The Flexibility in the Proline Ring Couples to the Protein Backbone. Protein Sci. 2005, 14, 1011–1018. [Google Scholar] [CrossRef]
- Das, S.; Hirano, M.; Tako, R.; McCallister, C.; Nikolaidis, N. Evolutionary Genomics of Immunoglobulin-Encoding Loci in Vertebrates. Curr. Genom. 2012, 13, 95–102. [Google Scholar]
- Zhao, Y.; Pan-Hammarström, Q.; Yu, S.; Wertz, N.; Zhang, X.; Li, N.; Butler, J.E.; Hammarström, L. Identification of IgF, a Hinge-Region-Containing Ig Class, and IgD in Xenopus Tropicalis. Proc. Natl. Acad. Sci. USA 2006, 103, 12087–12092. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Huang, T.; Hammarström, L.; Zhao, Y. The Immunoglobulins: New Insights, Implications, and Applications. Annu. Rev. Anim. Biosci. 2020, 8, 145–169. [Google Scholar] [CrossRef]
- Adlersberg, J.B. The Immunoglobulin Hinge (Interdomain) Region. Ric. Clin. Lab. 1976, 6, 191. [Google Scholar] [PubMed]
- Sakano, H.; Rogers, J.H.; Hüppi, K.; Brack, C.; Traunecker, A.; Maki, R.; Wall, R.; Tonegawa, S. Domains and the Hinge Region of an Immunoglobulin Heavy Chain Are Encoded in Separate DNA Segments. Nature 1979, 277, 627–633. [Google Scholar] [CrossRef]
- Kawamura, S.; Omoto, K.; Ueda, S. Evolutionary Hypervariability in the Hinge Region of the Immunoglobulin Alpha Gene. J. Mol. Biol. 1990, 215, 201–206. [Google Scholar] [CrossRef]
- Putnam, F.W.; Takahashi, N.; Tetaert, D.; Debuire, B.; Lin, L.C. Amino Acid Sequence of the First Constant Region Domain and the Hinge Region of the Delta Heavy Chain of Human IgD. Proc. Natl. Acad. Sci. USA 1981, 78, 6168–6172. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Calvert, R.A.; Sutton, B.J.; Doré, K.A. IgY: A Key Isotype in Antibody Evolution. Biol. Rev. 2017, 92, 2144–2156. [Google Scholar] [CrossRef]
- Parvari, R.; Avivi, A.; Lentner, F.; Ziv, E.; Tel-Or, S.; Burstein, Y.; Schechter, I. Chicken Immunoglobulin Gamma-Heavy Chains: Limited VH Gene Repertoire, Combinatorial Diversification by D Gene Segments and Evolution of the Heavy Chain Locus. EMBO J. 1988, 7, 739–744. [Google Scholar] [CrossRef]
- Tucker, P.W.; Slightom, J.L.; Blattner, F.R. Mouse IgA Heavy Chain Gene Sequence: Implications for Evolution of Immunoglobulin Hinge Axons. Proc. Natl. Acad. Sci. USA 1981, 78, 7684–7688. [Google Scholar] [CrossRef]
- Coscia, M.R.; Morea, V.; Tramontano, A.; Oreste, U. Analysis of a CDNA Sequence Encoding the Immunoglobulin Heavy Chain of the Antarctic Teleost Trematomus bernacchii. Fish Shellfish Immunol. 2000, 10, 343–357. [Google Scholar] [CrossRef]
- Coscia, M.R.; Varriale, S.; De Santi, C.; Giacomelli, S.; Oreste, U. Evolution of the Antarctic Teleost Immunoglobulin Heavy Chain Gene. Mol. Phylogenetics Evol. 2010, 55, 226–233. [Google Scholar] [CrossRef] [PubMed]
- Giacomelli, S.; Buonocore, F.; Albanese, F.; Scapigliati, G.; Gerdol, M.; Oreste, U.; Coscia, M.R. New Insights into Evolution of IgT Genes Coming from Antarctic Teleosts. Mar. Genom. 2015, 24, 55–68. [Google Scholar] [CrossRef] [PubMed]
- Collins, A.M.; Watson, C.T. Immunoglobulin Light Chain Gene Rearrangements, Receptor Editing and the Development of a Self-Tolerant Antibody Repertoire. Front. Immunol. 2018, 9, 2249. [Google Scholar] [CrossRef] [PubMed]
- Criscitiello, M.F.; Flajnik, M.F. Four Primordial Immunoglobulin Light Chain Isotypes, Including Lambda and Kappa, Identified in the Most Primitive Living Jawed Vertebrates. Eur. J. Immunol. 2007, 37, 2683–2694. [Google Scholar] [CrossRef] [PubMed]
- Mashoof, S.; Criscitiello, M.F. Fish Immunoglobulins. Biology 2016, 5, 45. [Google Scholar] [CrossRef]
- Guselnikov, S.V.; Baranov, K.O.; Najakshin, A.M.; Mechetina, L.V.; Chikaev, N.A.; Makunin, A.I.; Kulemzin, S.V.; Andreyushkova, D.A.; Stöck, M.; Wuertz, S. Diversity of Immunoglobulin Light Chain Genes in Non-Teleost Ray-Finned Fish Uncovers IgL Subdivision into Five Ancient Isotypes. Front. Immunol. 2018, 9, 1079. [Google Scholar] [CrossRef] [PubMed]
- Nikolaidis, N.; Klein, J.; Nei, M. Origin and Evolution of the Ig-like Domains Present in Mammalian Leukocyte Receptors: Insights from Chicken, Frog, and Fish Homologues. Immunogenet 2005, 57, 151–157. [Google Scholar] [CrossRef] [PubMed]
- Akula, S.; Mohammadamin, S.; Hellman, L. Fc Receptors for Immunoglobulins and Their Appearance during Vertebrate Evolution. PLoS ONE 2014, 9, e96903. [Google Scholar] [CrossRef] [PubMed]
- West, A.P.; Herr, A.B.; Bjorkman, P.J. The Chicken Yolk Sac IgY Receptor, a Functional Equivalent of the Mammalian MHC-Related Fc Receptor, Is a Phospholipase A2 Receptor Homolog. Immunity 2004, 20, 601–610. [Google Scholar] [CrossRef]
- Murin, C.D. Considerations of Antibody Geometric Constraints on NK Cell Antibody Dependent Cellular Cytotoxicity. Front. Immunol. 2020, 11. [Google Scholar] [CrossRef]
- Turula, H.; Wobus, C.E. The Role of the Polymeric Immunoglobulin Receptor and Secretory Immunoglobulins during Mucosal Infection and Immunity. Viruses 2018, 10, 237. [Google Scholar] [CrossRef]
- Hamuro, K.; Suetake, H.; Saha, N.R.; Kikuchi, K.; Suzuki, Y. A Teleost Polymeric Ig Receptor Exhibiting Two Ig-Like Domains Transports Tetrameric IgM into the Skin. J. Immunol. 2007, 178, 5682–5689. [Google Scholar] [CrossRef]
- Kaetzel, C.S. Coevolution of Mucosal Immunoglobulins and the Polymeric Immunoglobulin Receptor: Evidence That the Commensal Microbiota Provided the Driving Force. ISRN Immunol. 2014, 2014, 541537. [Google Scholar] [CrossRef]
- Pyzik, M.; Sand, K.M.K.; Hubbard, J.J.; Andersen, J.T.; Sandlie, I.; Blumberg, R.S. The Neonatal Fc Receptor (FcRn): A Misnomer? Front. Immunol. 2019, 10, 1540. [Google Scholar] [CrossRef]
- Matsushita, M.; Matsushita, A.; Endo, Y.; Nakata, M.; Kojima, N.; Mizuochi, T.; Fujita, T. Origin of the Classical Complement Pathway: Lamprey Orthologue of Mammalian C1q Acts as a Lectin. Proc. Natl. Acad. Sci. USA 2004, 101, 10127–10131. [Google Scholar] [CrossRef] [PubMed]
- Kishore, U.; Ghai, R.; Greenhough, T.J.; Shrive, A.K.; Bonifati, D.M.; Gadjeva, M.G.; Waters, P.; Kojouharova, M.S.; Chakraborty, T.; Agrawal, A. Structural and Functional Anatomy of the Globular Domain of Complement Protein C1q. Immunol. Lett. 2004, 95, 113–128. [Google Scholar] [CrossRef] [PubMed]
- Graille, M.; Stura, E.A.; Corper, A.L.; Sutton, B.J.; Taussig, M.J.; Charbonnier, J.-B.; Silverman, G.J. Crystal Structure of a Staphylococcus Aureus Protein A Domain Complexed with the Fab Fragment of a Human IgM Antibody: Structural Basis for Recognition of B-Cell Receptors and Superantigen Activity. Proc. Natl. Acad. Sci. USA 2000, 97, 5399–5404. [Google Scholar] [CrossRef] [PubMed]
- Deisenhofer, J. Crystallographic Refinement and Atomic Models of a Human Fc Fragment and Its Complex with Fragment B of Protein A from Staphylococcus aureus at 2.9- and 2.8-.ANG. Resolution. Biochemistry 1981, 20, 2361–2370. [Google Scholar] [CrossRef]
- Graille, M.; Harrison, S.; Crump, M.P.; Findlow, S.C.; Housden, N.G.; Muller, B.H.; Battail-Poirot, N.; Sibaï, G.; Sutton, B.J.; Taussig, M.J.; et al. Evidence for Plasticity and Structural Mimicry at the Immunoglobulin Light Chain-Protein L Interface. J. Biol. Chem. 2002, 277, 47500–47506. [Google Scholar] [CrossRef]
- Hsu, E.; Flajnik, M.F.; Pasquier, L.D. A Third Immunoglobulin Class in Amphibians. J. Immunol. 1985, 135, 1998–2004. [Google Scholar]
- Vidarsson, G.; Dekkers, G.; Rispens, T. IgG Subclasses and Allotypes: From Structure to Effector Functions. Front. Immunol. 2014, 5, 520. [Google Scholar] [CrossRef]
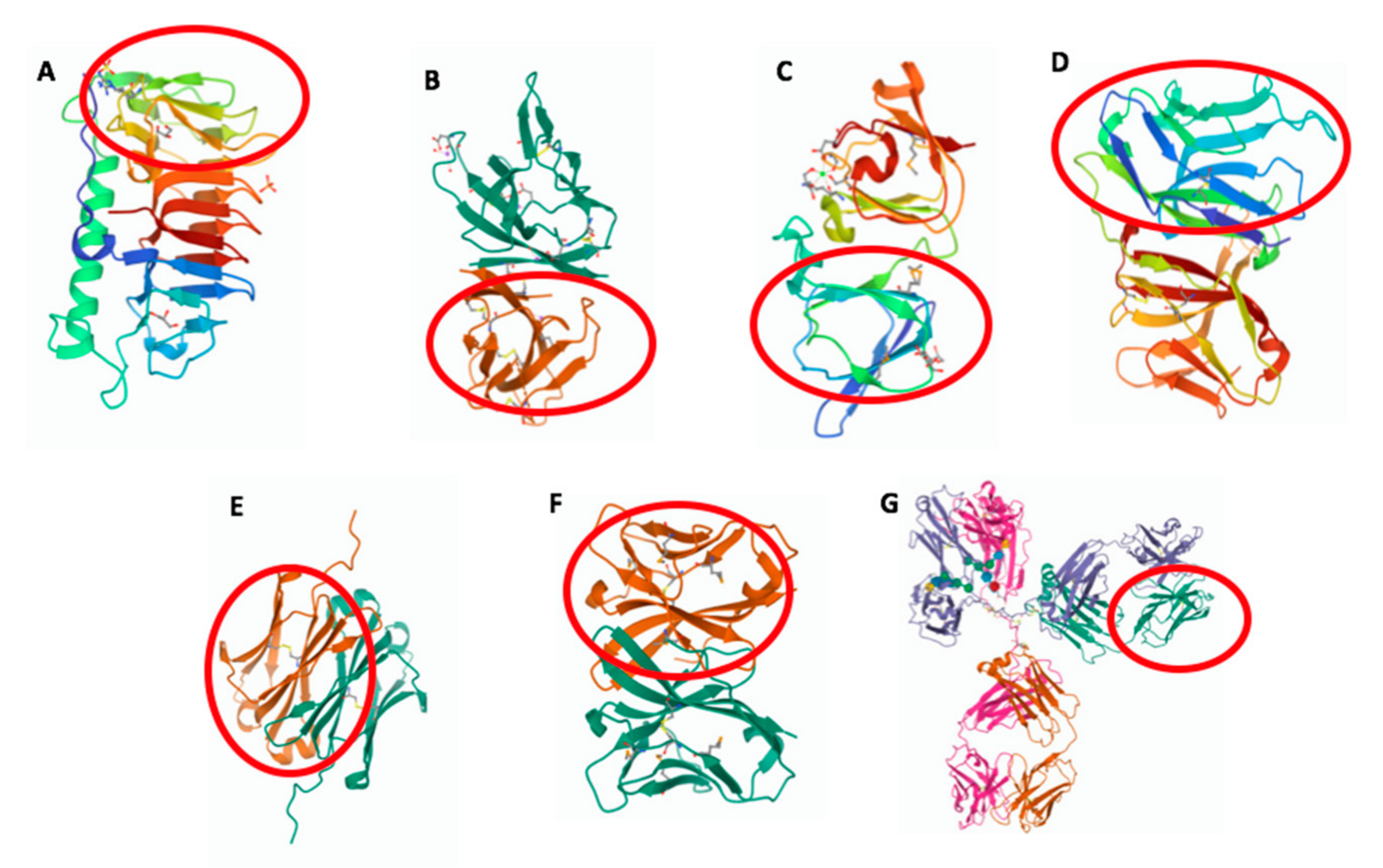
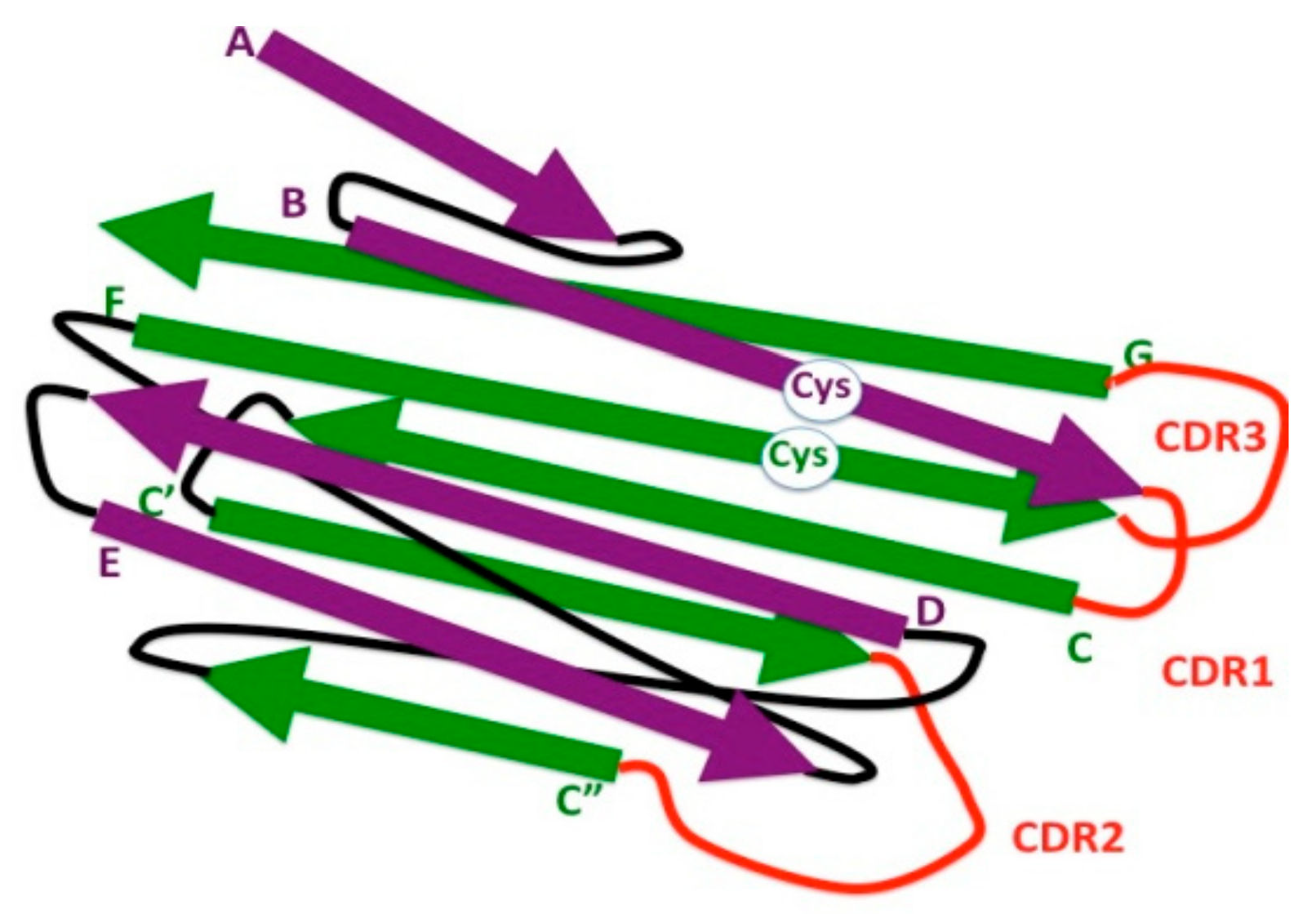
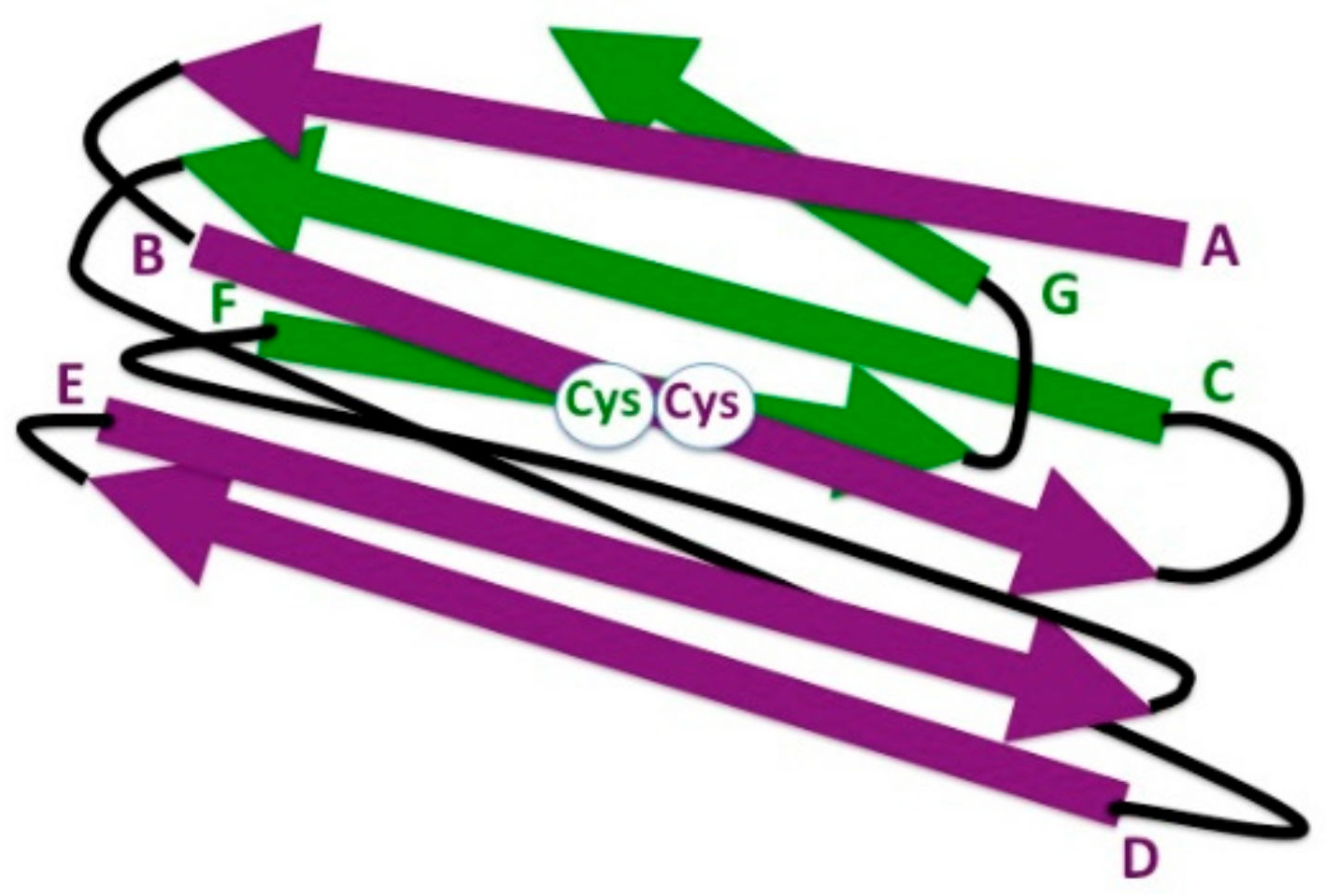
Bacteria
 | Ig domain | [8] |
Porifera
 | IgV-like domain | [26] |
Mollusca
 | RAG homologs | [44] |
Echinodermata
 | AID homologs | [51] |
Protochordates
 | IgV domain | [29] |
Agnatha
 | AID | [49] |
Chondrichtyes
 | RAG, IgC1 domain, IgNAR, IgM, IgD/IgW | [2] |
Osteichtyes
 | IgT, FcR, RAG, pIgR | [72,95,99] |
Amphibia
 | IgX (IgA precursor) | [107] |
Aves
 | IgY (IgG/IgE precursor) | [85] |
Mammals
 | HCAb, IgG, IgE | [69,79] |
Homo sapiens
 | IgG1, IgG2, IgG3, IgG4 subisotype | [108] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oreste, U.; Ametrano, A.; Coscia, M.R. On Origin and Evolution of the Antibody Molecule. Biology 2021, 10, 140. https://doi.org/10.3390/biology10020140
Oreste U, Ametrano A, Coscia MR. On Origin and Evolution of the Antibody Molecule. Biology. 2021; 10(2):140. https://doi.org/10.3390/biology10020140
Chicago/Turabian StyleOreste, Umberto, Alessia Ametrano, and Maria Rosaria Coscia. 2021. "On Origin and Evolution of the Antibody Molecule" Biology 10, no. 2: 140. https://doi.org/10.3390/biology10020140
APA StyleOreste, U., Ametrano, A., & Coscia, M. R. (2021). On Origin and Evolution of the Antibody Molecule. Biology, 10(2), 140. https://doi.org/10.3390/biology10020140






