Genome-Wide Identification of LATERAL ORGAN BOUNDARIES DOMAIN (LBD) Transcription Factors and Screening of Salt Stress Candidates of Rosa rugosa Thunb
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of RrLBD Family
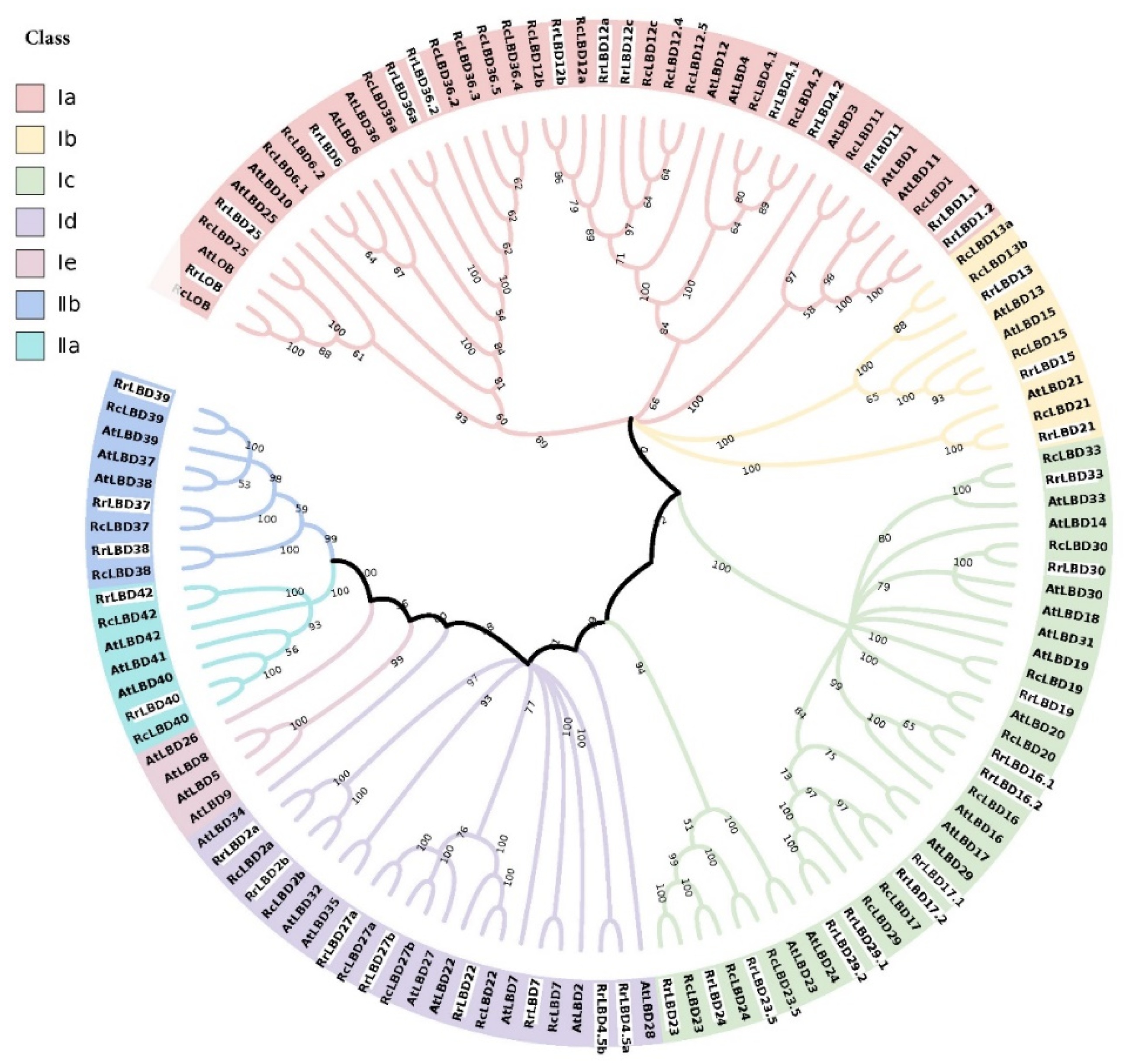
2.2. Phylogenetic Analysis of LBD Family
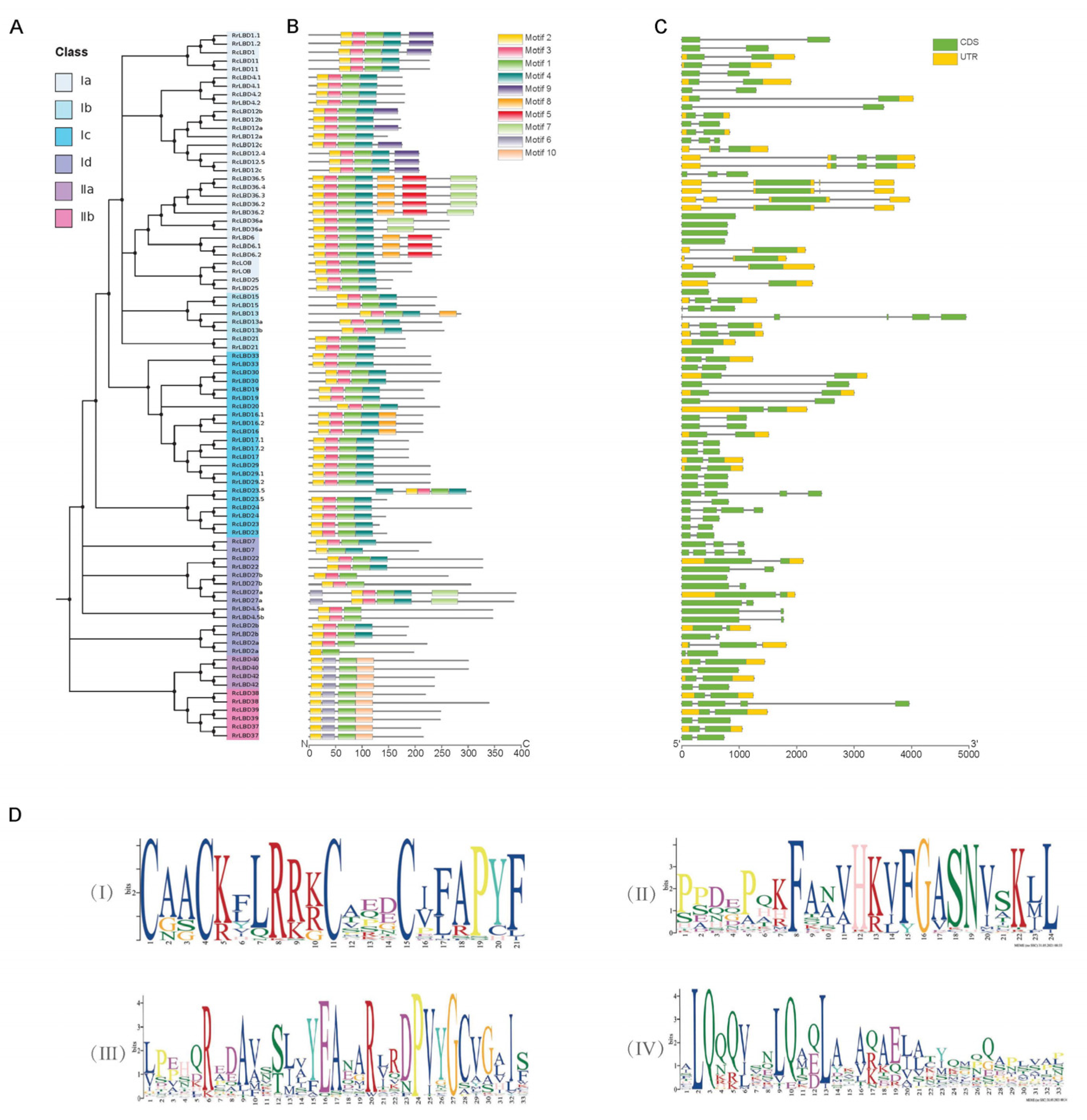
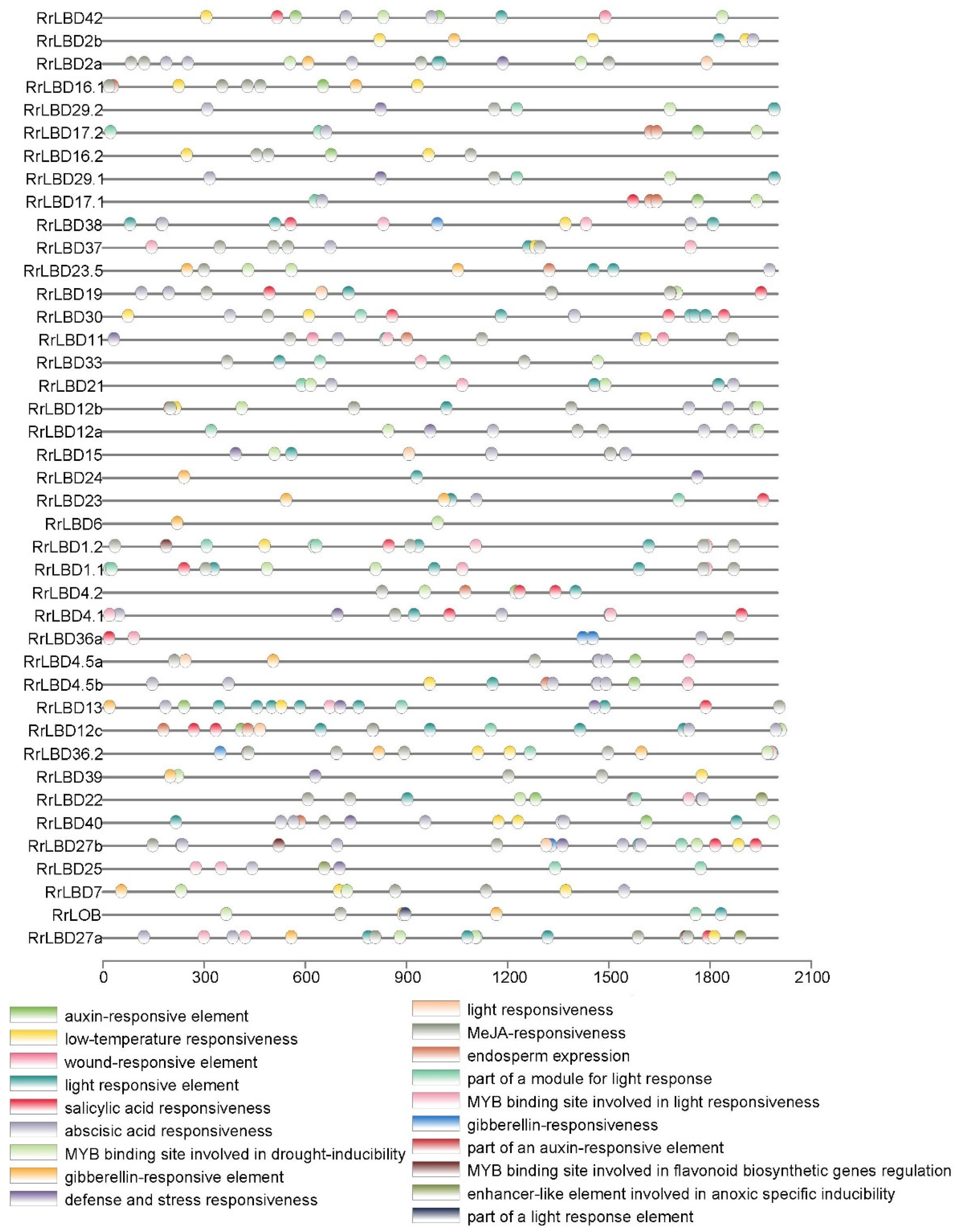
2.3. Genen Structure, Motif Composition, and Cis-Acting Elements Analysis of RrLBD Family
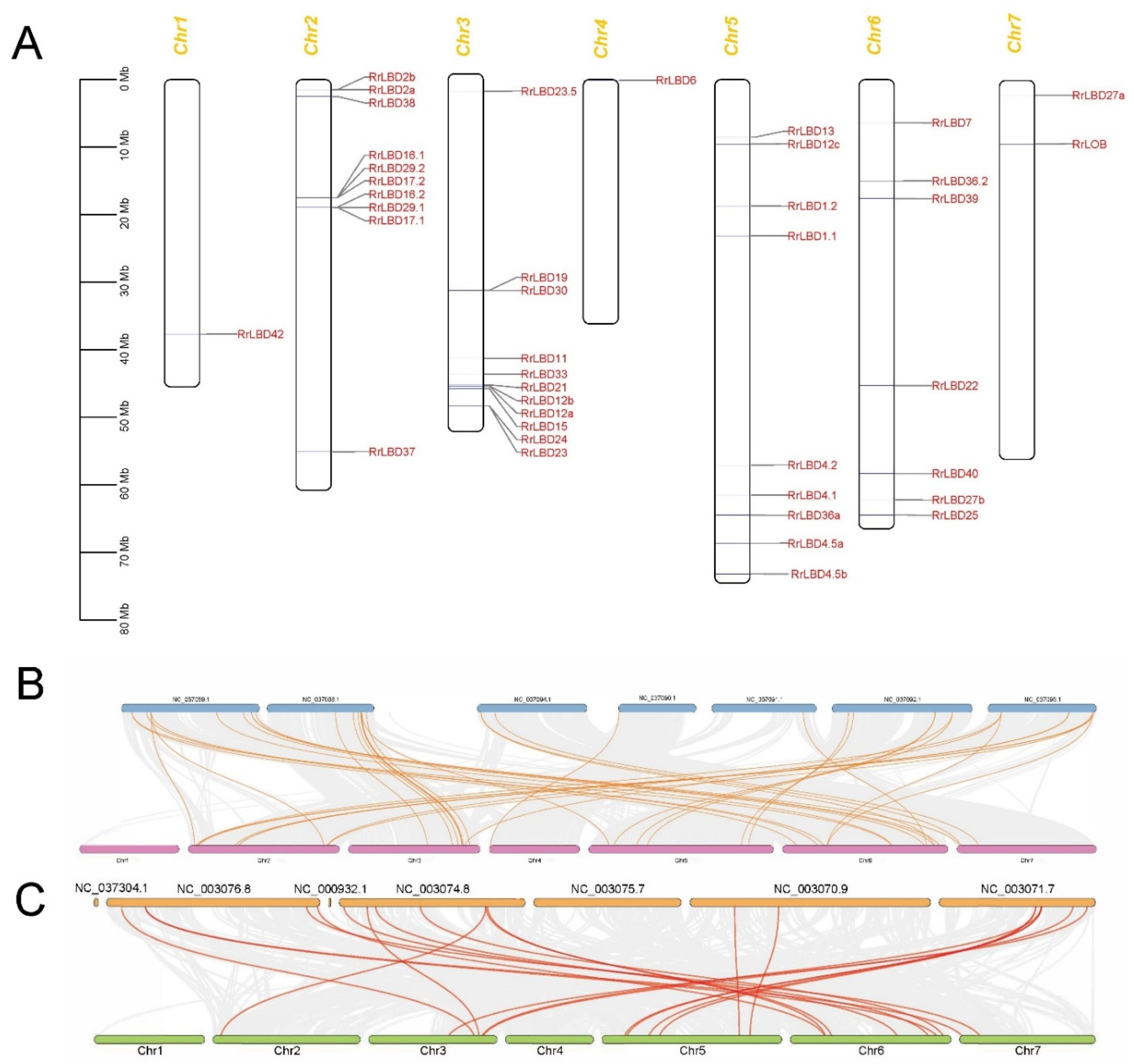
2.4. Synteny Analysis of LBD Genes
2.5. Expression Analysis of RrLBDs in Different Tissues under Salt Stress
3. Results
3.1. Identification and Phylogenetic Analysis of RrLBD Family
3.2. Gene Structure and Motif Composition Analysis of RrLBD Family
3.3. Chromosomal Distribution and Evolutionary Analysis of RrLBD Genes
3.4. Analysis of Cis-Acting Elements of RrLBD Promotors
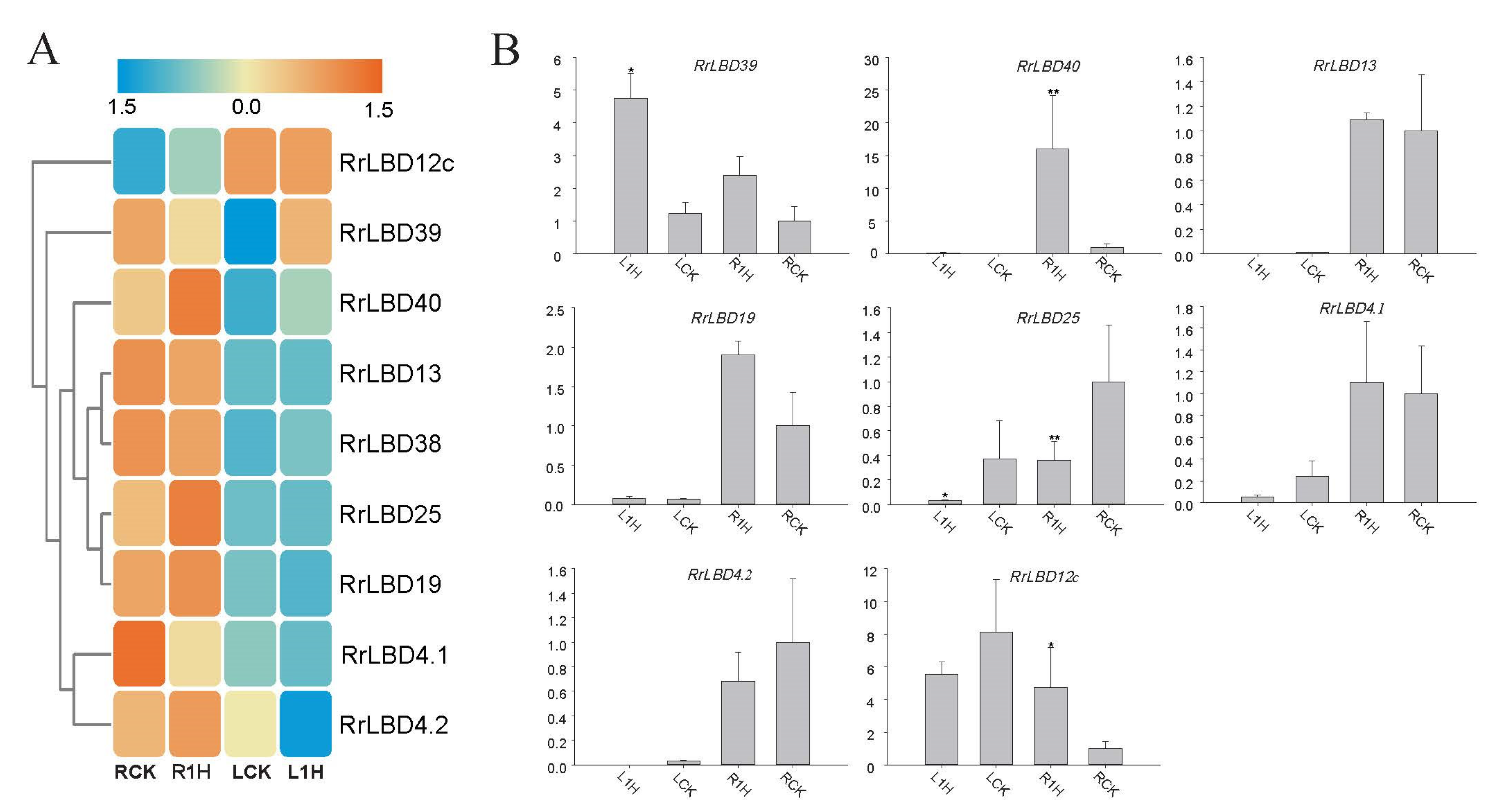
3.5. Salt-Responsive Expression Analysis of RrLBDs
4. Discussion
4.1. High Conservation of RrLBD Family
4.2. Salt Response of RrLBD Candidates
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2019, 48, D1104–D1113. [Google Scholar] [CrossRef] [PubMed]
- Jin, J.; Tian, F.; Yang, D.-C.; Meng, Y.-Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Majer, C.; Hochholdinger, F. Defining the boundaries: Structure and function of LOB domain proteins. Trends Plant Sci. 2011, 16, 47–52. [Google Scholar] [CrossRef]
- Lee, H.W.; Kim, M.-J.; Park, M.Y.; Han, K.-H.; Kim, J. The Conserved Proline Residue in the LOB Domain of LBD18 Is Critical for DNA-Binding and Biological Function. Mol. Plant 2013, 6, 1722–1725. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shuai, B.; Reynaga-Peña, C.G.; Springer, P.S. The Lateral Organ Boundaries Gene Defines a Novel, Plant-Specific Gene Family. Plant Physiol. 2002, 129, 747–761. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.; Wang, S.; Yu, X.; Yu, J.; He, X.; Zhang, S.; Shou, H.; Wu, P. ARL1, a LOB-domain protein required for adventitious root formation in rice. Plant J. 2005, 43, 47–56. [Google Scholar] [CrossRef]
- Yu, J.; Xie, Q.; Li, C.; Dong, Y.; Zhu, S.; Chen, J. Comprehensive characterization and gene expression patterns of LBD gene family in Gossypium. Planta 2020, 251, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Wang, X.; Zhang, S.; Su, L.; Liu, X.; Hao, Y. A Genome-Wide Analysis of the LBD (LATERAL ORGAN BOUNDARIES Domain) Gene Family in Malus domestica with a Functional Characterization of MdLBD11. PLoS ONE 2013, 8, e57044. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, X.; Wang, J.; Zhao, M.; Yuan, H. Identification and expression analysis of LATERAL ORGAN BOUNDARIES DOMAIN (LBD) transcription factor genes in Fragaria vesca. Can. J. Plant Sci. 2017. [Google Scholar] [CrossRef] [Green Version]
- Matsumura, Y.; Iwakawa, H.; Machida, Y.; Machida, C. Characterization of genes in the ASYMMETRIC LEAVES2/LATERAL ORGAN BOUNDARIES(AS2/LOB) family inArabidopsis thaliana, and functional and molecular comparisons betweenAS2and other family members. Plant J. 2009, 58, 525–537. [Google Scholar] [CrossRef] [Green Version]
- Gendron, J.M.; Liu, J.-S.; Fan, M.; Bai, M.-Y.; Wenkel, S.; Springer, P.S.; Barton, M.K.; Wang, Z.-Y. Brassinosteroids regulate organ boundary formation in the shoot apical meristem of Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 21152–21157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, H.; Dai, Z.; Li, L.; Wang, J.; Miao, X.; Shi, Z. OsRAMOSA2 Shapes Panicle Architecture through Regulating Pedicel Length. Front. Plant Sci. 2017, 8, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bortiri, E.; Chuck, G.; Vollbrecht, E.; Rocheford, T.; Martienssen, R.; Hake, S. ramosa2Encodes a LATERAL ORGAN BOUNDARY Domain Protein That Determines the Fate of Stem Cells in Branch Meristems of Maize. Plant Cell 2006, 18, 574–585. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iwakawa, H.; Ueno, Y.; Semiarti, E.; Onouchi, H.; Kojima, S.; Tsukaya, H.; Hasebe, M.; Soma, T.; Ikezaki, M.; Machida, C.; et al. The ASYMMETRIC LEAVES2 Gene of Arabidopsis thaliana, Required for Formation of a Symmetric Flat Leaf Lamina, Encodes a Member of a Novel Family of Proteins Characterized by Cysteine Repeats and a Leucine Zipper. Plant Cell Physiol. 2002, 43, 467–478. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Semiarti, E.; Ueno, Y.; Tsukaya, H.; Iwakawa, H.; Machida, C. The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana regulates formation of a symmetric lamina, establishment of venation and repression of meristem-related homeobox genes in leaves. Development 2001, 128, 1771–1783. [Google Scholar] [CrossRef]
- Lin, W.-C.; Shuai, B.; Springer, P.S. The Arabidopsis LATERAL ORGAN BOUNDARIES-Domain Gene ASYMMETRIC LEAVES2 Functions in the Repression of KNOX Gene Expression and in Adaxial-Abaxial Patterning. Plant Cell 2003, 15, 2241–2252. [Google Scholar] [CrossRef] [Green Version]
- Evans, M.M. Theindeterminate gametophyte1Gene of Maize Encodes a LOB Domain Protein Required for Embryo Sac and Leaf Development. Plant Cell 2007, 19, 46–62. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Wang, F.; Guo, J.; Zhang, X.S. Rice OsAS2 Gene, a Member of LOB Domain Family, Functions in the Regulation of Shoot Differentiation and Leaf Development. J. Plant Biol. 2009, 52, 374–381. [Google Scholar] [CrossRef]
- Zhang, J.; Tang, W.; Huang, Y.; Niu, X.; Zhao, Y.; Han, Y.; Liu, Y. Down-regulation of a LBD-like gene, OsIG1, leads to occurrence of unusual double ovules and developmental abnormalities of various floral organs and megagametophyte in rice. J. Exp. Bot. 2014, 66, 99–112. [Google Scholar] [CrossRef] [Green Version]
- Okushima, Y.; Fukaki, H.; Onoda, M.; Theologis, A.; Tasaka, M. ARF7 and ARF19 Regulate Lateral Root Formation via Direct Activation ofLBD/ASLGenes inArabidopsis. Plant Cell 2007, 19, 118–130. [Google Scholar] [CrossRef] [Green Version]
- Porco, S.; Larrieu, A.; Casimiro, I.; Hill, K.; Benkova, E.; Fukaki, H.; Brady, S.M.; Scheres, B.; Péret, B.; Bennett, M.J.; et al. Lateral root emergence in Arabidopsis is dependent on transcription factor LBD29 regulating auxin influx carrier LAX3. Development 2016, 143, 3340–3349. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, H.W.; Kim, N.Y.; Lee, D.J.; Kim, J. LBD18/ASL20 Regulates Lateral Root Formation in Combination with LBD16/ASL18 Downstream of ARF7 and ARF19 in Arabidopsis. Plant Physiol. 2009, 151, 1377–1389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Inukai, Y.; Sakamoto, T.; Ueguchi-Tanaka, M.; Shibata, Y.; Gomi, K.; Umemura, I.; Hasegawa, Y.; Ashikari, M.; Kitano, H.; Matsuoka, M. Crown rootless1, Which Is Essential for Crown Root Formation in Rice, Is a Target of an AUXIN RESPONSE FACTOR in Auxin Signaling. Plant Cell 2005, 17, 1387–1396. [Google Scholar] [CrossRef] [Green Version]
- Majer, C.; Xu, C.; Berendzen, K.W.; Hochholdinger, F. Molecular interactions of ROOTLESS CONCERNING CROWN AND SEMINAL ROOTS, a LOB domain protein regulating shoot-borne root initiation in maize (Zea mays L.). Philos. Trans. R. Soc. B Biol. Sci. 2012, 367, 1542–1551. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taramino, G.; Sauer, M.; Stauffer, J.L.; Multani, D.; Niu, X.; Sakai, H.; Hochholdinger, F. The maize (Zea mays L.) RTCS gene encodes a LOB domain protein that is a key regulator of embryonic seminal and post-embryonic shoot-borne root initiation. Plant J. 2007, 50, 649–659. [Google Scholar] [CrossRef] [PubMed]
- Rubin, G.; Tohge, T.; Matsuda, F.; Saito, K.; Scheible, W.-R. Members of the LBD Family of Transcription Factors Repress Anthocyanin Synthesis and Affect Additional Nitrogen Responses in Arabidopsis. Plant Cell 2009, 21, 3567–3584. [Google Scholar] [CrossRef] [Green Version]
- Médiène, S.; Pagès, L.; Jordan, M.-O.; Le Bot, J.; Adamowicz, S. Influence of nitrogen availability on shoot development in young peach trees [ Prunus persica (L.) Batsch]. Trees 2002, 16, 547–554. [Google Scholar] [CrossRef]
- Albinsky, D.; Kusano, M.; Higuchi, M.; Hayashi, N.; Kobayashi, M.; Fukushima, A.; Mori, M.; Ichikawa, T.; Matsui, K.; Kuroda, H.; et al. Metabolomic Screening Applied to Rice FOX Arabidopsis Lines Leads to the Identification of a Gene-Changing Nitrogen Metabolism. Mol. Plant 2010, 3, 125–142. [Google Scholar] [CrossRef]
- Huang, X.; Liu, G.; Zhang, W. Genome-wide Analysis of LBD (LATERAL ORGAN BOUNDARIES Domain) Gene Family in Brassica rapa. Braz. Arch. Biol. Technol. 2018, 61, 61. [Google Scholar] [CrossRef]
- Luo, Y.; Ma, B.; Zeng, Q.; Xiang, Z.; He, N. Identification and characterization of Lateral Organ Boundaries Domain genes in mulberry, Morus notabilis. Meta Gene 2016, 8, 44–50. [Google Scholar] [CrossRef]
- Zhang, X.; He, Y.; He, W.; Su, H.; Wang, Y.; Hong, G.; Xu, P. Structural and functional insights into the LBD family involved in abiotic stress and flavonoid synthases in Camellia sinensis. Sci. Rep. 2019, 9, 15651. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yan, H.; Liu, Y.; Yi, Y. Genome-wide analysis of LATERAL ORGAN BOUNDARIES DOMAIN-in Physcomitrella patens and stress responses. Genes Genom. 2020, 42, 651–662. [Google Scholar] [CrossRef] [PubMed]
- Saint-Oyant, L.H.; Ruttink, T.; Hamama, L.; Kirov, I.; Lakhwani, D.; Zhou, N.N.; Bourke, P.M.; Daccord, N.; Leus, L.; Schulz, D.; et al. A high-quality genome sequence of Rosa chinensis to elucidate ornamental traits. Nat. Plants 2018, 4, 473–484. [Google Scholar] [CrossRef] [Green Version]
- Chen, F.; Su, L.; Hu, S.; Xue, J.-Y.; Liu, H.; Liu, G.; Jiang, Y.; Du, J.; Qiao, Y.; Fan, Y.; et al. A chromosome-level genome assembly of rugged rose (Rosa rugosa) provides insights into its evolution, ecology, and floral characteristics. Hortic. Res. 2021, 8, 1–13. [Google Scholar] [CrossRef]
- Finn, R.D.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Mistry, J.; Mitchell, A.L.; Potter, S.C.; Punta, M.; Qureshi, M.; Sangrador-Vegas, A.; et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 2015, 44, D279–D285. [Google Scholar] [CrossRef]
- Bateman, A. The Pfam protein families database. Nucleic Acids Res. 2004, 32, D138–D141. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef] [Green Version]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Russo, C.; Selvatti, A.P. Bootstrap and Rogue Identification Tests for Phylogenetic Analyses. Mol. Biol. Evol. 2018, 35, 2327–2333. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree Of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools-an inte-grative toolkit developed for interactive analyses of big biological data. Molecules 2020.
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2−ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Kong, Y.; Xu, P.; Jing, X.; Chen, L.; Li, L.; Li, X. Decipher the ancestry of the plant-specific LBD gene family. BMC Genom. 2017, 18, 951. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chanderbali, A.S.; He, F.; Soltis, P.S.; Soltis, U.E. Out of the Water: Origin and Diversification of the LBD Gene Family. Mol. Biol. Evol. 2015, 32, 1996–2000. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Y.; Li, Z.; Ma, B.; Hou, Q.; Wan, X. Phylogeny and Functions of LOB Domain Proteins in Plants. Int. J. Mol. Sci. 2020, 21, 2278. [Google Scholar] [CrossRef] [Green Version]
- Zhang, L.; Wu, S.; Chang, X.; Wang, X.; Zhao, Y.; Xia, Y.; Trigiano, R.N.; Jiao, Y.; Chen, F. The ancient wave of polyploidization events in flowering plants and their facilitated adaptation to environmental stress. Plant Cell Environ. 2020, 43, 2847–2856. [Google Scholar] [CrossRef]
- Zhang, Y.-M.; Zhang, S.-Z.; Zheng, C.-C. Genomewide analysis of LATERAL ORGAN BOUNDARIES Domain gene family in Zea mays. J. Genet. 2014, 93, 79–91. [Google Scholar] [CrossRef]
- Jiang, Z.; Zhou, X.; Tao, M.; Yuan, F.; Liu, L.; Wu, F.; Wu, X.; Xiang, Y.; Niu, Y.; Liu, F.; et al. Plant cell-surface GIPC sphingolipids sense salt to trigger Ca2+ influx. Nat. Cell Biol. 2019, 572, 341–346. [Google Scholar] [CrossRef]
- Eulgem, T. Regulation of the Arabidopsis defense transcriptome. Trends Plant Sci. 2005, 10, 71–78. [Google Scholar] [CrossRef]
- Thatcher, L.F.; Anderson, J.P.; Singh, K.B. Plant defence responses: What have we learnt from Arabidopsis? Funct. Plant Biol. 2005, 32, 1–19. [Google Scholar] [CrossRef]
- Mangeon, A.; Bell, E.M.; Lin, W.-C.; Jablonska, B.; Springer, P.S. Misregulation of the LOB domain gene DDA1 suggests possible functions in auxin signalling and photomorphogenesis. J. Exp. Bot. 2010, 62, 221–233. [Google Scholar] [CrossRef] [Green Version]
- Sarker, U.; Oba, S. Drought stress enhances nutritional and bioactive compounds, phenolic acids and antioxidant capacity of Amaranthus leafy vegetable. BMC Plant Biol. 2018, 18, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Petropoulos, S.A.; Levizou, E.; Ntatsi, G.; Fernandes, Â.; Petrotos, K.; Akoumianakis, K.; Barros, L.; Ferreira, I.C.F.R. Salinity effect on nutritional value, chemical composition and bioactive compounds content of Cichorium spinosum L. Food Chem. 2017, 214, 129–136. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Cao, M.; Chen, X.; Ye, M.; Zhao, P.; Nan, Y.; Li, W.; Zhang, C.; Kong, L.; Kong, N.; et al. Genome-Wide Analysis of the Lateral Organ Boundaries Domain (LBD) Gene Family in Solanum tuberosum. Int. J. Mol. Sci. 2019, 20, 5360. [Google Scholar] [CrossRef] [Green Version]
- Lang, D.; Yu, X.; Jia, X.; Li, Z.; Zhang, X. Methyl jasmonate improves metabolism and growth of NaCl-stressed Glycyrrhiza uralensis seedlings. Sci. Hortic. 2020, 266, 109287. [Google Scholar] [CrossRef]
- Gómez, S.; Ferrieri, R.A.; Schueller, M.; Orians, C.M. Methyl jasmonate elicits rapid changes in carbon and nitrogen dynamics in tomato. New Phytol. 2010, 188, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-B.; Song, J.-P.; Wang, Z.-B. ASYMMETRIC LEAVES2-LIKE38, one member of AS2/LOB gene family, involves in regulating ab-adaxial patterning in Arabidopsis lateral organs. Acta Physiol. Plant. 2015, 37. [Google Scholar] [CrossRef]
- Van Ha, C.; Leyva-González, M.A.; Osakabe, Y.; Tran, U.T.; Nishiyama, R.; Watanabe, Y.; Tanaka, M.; Seki, M.; Yamaguchi, S.; Van Dong, N.; et al. Positive regulatory role of strigolactone in plant responses to drought and salt stress. Proc. Natl. Acad. Sci. USA 2014, 111, 851–856. [Google Scholar] [CrossRef] [Green Version]
- Fang, Q.; Wang, Q.; Mao, H.; Xu, J.; Wang, Y.; Hu, H.; He, S.; Tu, J.; Cheng, C.; Tian, G.; et al. AtDIV2, an R-R-type MYB transcription factor of Arabidopsis, negatively regulates salt stress by modulating ABA signaling. Plant Cell Rep. 2018, 37, 1499–1511. [Google Scholar] [CrossRef] [PubMed]
- Zou, C.; Chen, A.; Xiao, L.; Muller, H.M.; Ache, P.; Haberer, G.; Zhang, M.; Jia, W.; Deng, P.; Huang, R.; et al. A high-quality genome assembly of quinoa provides insights into the molecular basis of salt bladder-based salinity tolerance and the exceptional nutritional value. Cell Res. 2017, 27, 1327–1340. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xu, J.-Y.; Jia, W.; Chen, Z.; Xu, Z.-C. Chloride salinity in a chloride-sensitive plant: Focusing on photosynthesis, hormone synthesis and transduction in tobacco. Plant Physiol. Biochem. 2020, 153, 119–130. [Google Scholar] [CrossRef]
- Liu, X.-L.; Zhang, H.; Jin, Y.-Y.; Wang, M.-M.; Yang, H.-Y.; Ma, H.-Y.; Jiang, C.-J.; Liang, Z.-W. Abscisic acid primes rice seedlings for enhanced tolerance to alkaline stress by upregulating antioxidant defense and stress tolerance-related genes. Plant Soil 2019, 438, 39–55. [Google Scholar] [CrossRef]
- Lu, X.; Liang, X.; Li, X.; Shen, P.; Cao, X.; Chen, C.; Song, S.; Wang, D.; Wang, Z.; Zhang, Y. Genome-wide characterisation and expression profiling of the LBD family in Salvia miltiorrhiza reveals the function of LBD50 in jasmonate signaling and phenolic biosyn-thesis. Ind. Crop. Prod. 2020, 144, 112006. [Google Scholar] [CrossRef]
- Yordanov, Y.S.; Regan, S.; Busov, V. Members of the LATERAL ORGAN BOUNDARIES DOMAIN Transcription Factor Family Are Involved in the Regulation of Secondary Growth in Populus. Plant Cell 2010, 22, 3662–3677. [Google Scholar] [CrossRef] [Green Version]
- Liu, S.; Wang, B.; Li, X.; Pan, J.; Qian, X.; Yu, Y.; Xu, P.; Zhu, J.; Xu, X. Lateral Organ Boundaries Domain 19 (LBD19) negative regulate callus formation in Arabidopsis. Plant Cell, Tissue Organ Cult. (PCTOC) 2019, 137, 485–494. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, J.; Zhang, W.; Cheng, Y.; Feng, L. Genome-Wide Identification of LATERAL ORGAN BOUNDARIES DOMAIN (LBD) Transcription Factors and Screening of Salt Stress Candidates of Rosa rugosa Thunb. Biology 2021, 10, 992. https://doi.org/10.3390/biology10100992
Wang J, Zhang W, Cheng Y, Feng L. Genome-Wide Identification of LATERAL ORGAN BOUNDARIES DOMAIN (LBD) Transcription Factors and Screening of Salt Stress Candidates of Rosa rugosa Thunb. Biology. 2021; 10(10):992. https://doi.org/10.3390/biology10100992
Chicago/Turabian StyleWang, Jianwen, Weijie Zhang, Yufei Cheng, and Liguo Feng. 2021. "Genome-Wide Identification of LATERAL ORGAN BOUNDARIES DOMAIN (LBD) Transcription Factors and Screening of Salt Stress Candidates of Rosa rugosa Thunb" Biology 10, no. 10: 992. https://doi.org/10.3390/biology10100992
APA StyleWang, J., Zhang, W., Cheng, Y., & Feng, L. (2021). Genome-Wide Identification of LATERAL ORGAN BOUNDARIES DOMAIN (LBD) Transcription Factors and Screening of Salt Stress Candidates of Rosa rugosa Thunb. Biology, 10(10), 992. https://doi.org/10.3390/biology10100992





