Metabolomics in Retinal Diseases: An Update
Abstract
:Simple Summary
Abstract
1. Introduction
2. Metabolomics: Brief Overview
3. Metabolomics: Technological Advances
3.1. Mass Spectrometry
3.2. Nuclear Magnetic Resonance (NMR) Spectroscopy
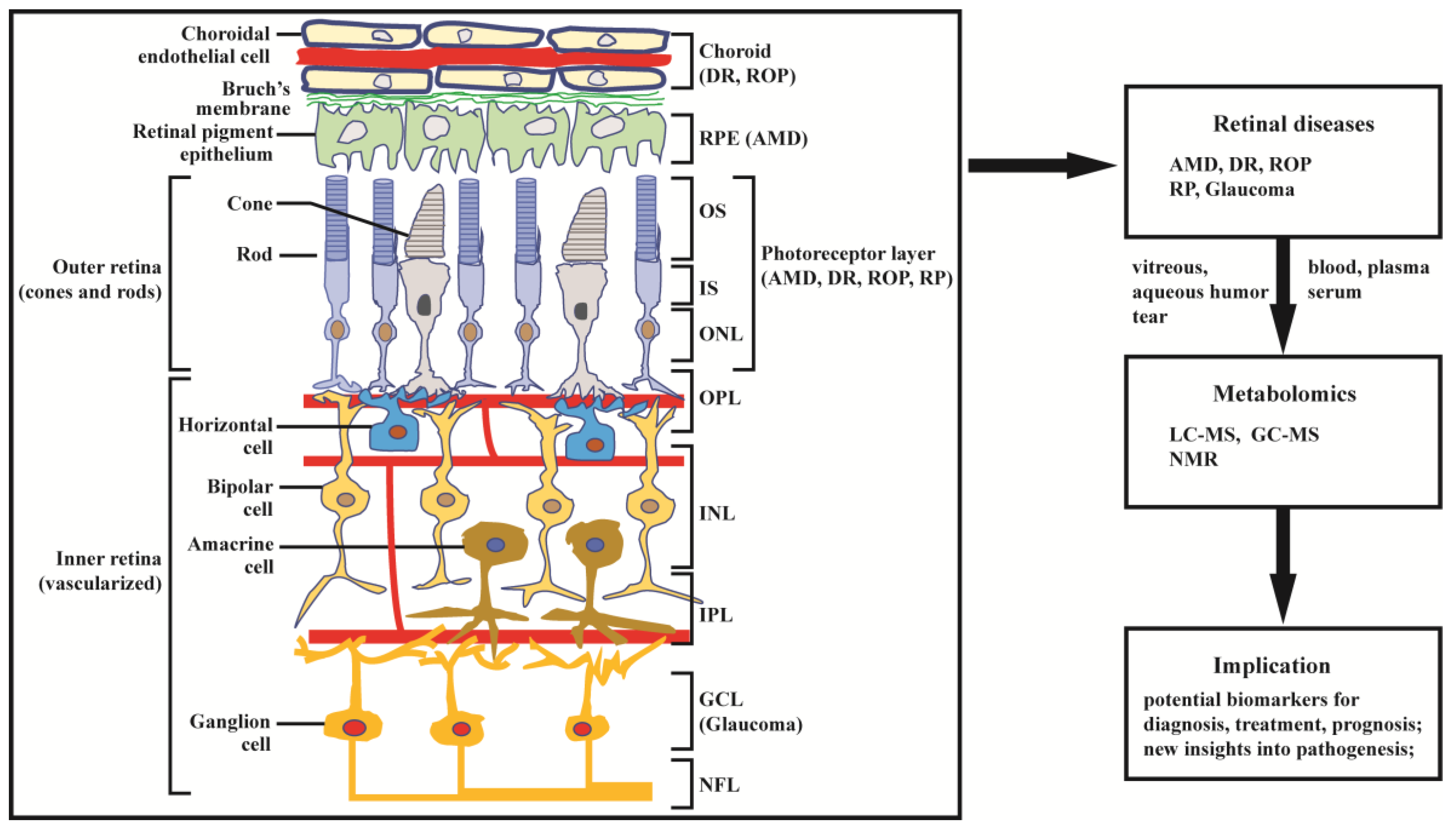
4. Metabolomics in Retinal Diseases
4.1. Age-Related Macular Degeneration (AMD)
| AMD Stage | Samples | Metabolic Biomarkers/Pathway | Analytical Platform | Untargeted/TARGETED | Study Design | References |
|---|---|---|---|---|---|---|
| late AMD (wet) | plasma (−) | Phe, Tyr, Gln, Asp, His-Arg, Trp-Phe, GCA, GDCA, GUDCA; tyrosine metabolism, sulfur amino acid metabolism, and amino acids related to urea metabolism pathway | LC-MS | Untargeted | case-control | [52] |
| early, intermediate, and late AMD | plasma (fasting) | acetate, acetoacetate, creatine, dimethyl sulfone, β-hydroxybutyrate, pyruvate, Ala, Gln, His | NMR | Untargeted | cross-sectional | [53] |
| late AMD (wet) | plasma (fasting) | N-acetyl-L-alanine, N1-methyl-2-pyridone-5-carboxamide, Tyr, Phe, Arg, Met, palmitoylcarnitine, isomaltose, hydrocortisone, biliverdin | GC-MS, LC-MS | Untargeted | case-control | [54] |
| early, intermediate, and late AMD | plasma (fasting) | linoleoyl-arachidonoyl-glycerol (18:2/20:4), stearoyl-arachidonoyl-glycerol (18:0/20:4), oleoyl-arachidonoyl-glycerol (18:1/20:4), 1-palmitoyl-2-arachidonoyl-GPC (16:0/20:4n6), 1-stearoyl-2-arachidonoyl-GPC (18:0/20:4), adenosine; diacylglycerol, glycerophospholipids pathway, purine metabolism | LC-MS | Untargeted | cross-sectional | [55] |
| late AMD (wet) | plasma (−) | L-oxalylalbizziine, isopentyl β-D-glucoside, LysoPC(P-18:0), LysoPC(P-18:1(9Z)), LysoPC(16:1(9Z)), 1-Lyso-2-arachidonoyl-phosphatidate, 9-hexadecenoylcarnitine, heptadecanoylcarnitine, 11Z-octadecenylcarnitine, L-palmitoylcarnitine, stearoylcarnitine, N-ornithyl-L-taurine; carnitine shuttle pathway, bile acid biosynthesis pathway | LC-MS | Untargeted | − | [56] |
| early, intermediate, and late AMD | plasma (fasting) | taurine, β-citrylglutamate, serotonin, N-acetylmethionine, Asp, hypotaurine, N-acetylasparagine, S-adenosylhomocysteine, maltotriose, maltose, nicotinamide, adenosine, cytidine, guanine, inosine, hypoxanthine, adenine, isoleucylglycine, 1-stearoyl-2-oleoyl-GPS(18:0/18:1), PE, PC, sphingosine, 1-(1-enyl-palmitoyl)-GPE (P-16:0), 14-HDoHE/17-HDoHE, 12-HETE, sphinganine, 1-(1-enyl-oleoyl)-GPE (P-18:1), 1-(1-enyl-stearoyl)-GPE (P-18:0); glycerophospholipid, purine, taurine and hypotaurine, and nitrogen metabolism | LC-MS | Untargeted | cross-sectional | [57] |
| late AMD (wet) | plasma (fasting) | Val, Lys, Pro, carnitine, valerylcarnitine, carnosine (Ala-His) | LC-MS | Targeted (IDQ p180 kit) | case-control | [58] |
| early, intermediate, and late AMD | plasma, serum (+) | HDL and VLDL lipoprotein particles, fatty acids, citrate, Ala, Ile, Leu, Phe, Tyr | NMR | Untargeted | − | [38] |
| early, intermediate AMD) | serum (non-fasting) | Gln, Glu:Gln ratio, glutaminolysis, phosphatidylcholine diacyl C28:1; glutamine pathway | LC-MS | Targeted (IDQ p180 kit) | case-control | [59] |
| late AMD (wet) | serum (fasting) | lactate, lipoproteins | NMR | Untargeted | − | [48] |
| late AMD (wet) | serum (−) | GPC, LysoPC (18:2), PS (18:0/20:4) | LC-MS | Untargeted | case-control | [50] |
| AMD subtype (PCV) | serum (fasting) | LPA (18:2), LysoPC (20:4), PC (20:1p/19:1), SM (d16:0/22:2), PAF (35:4), PC (16:0/22:5), PC (18:1/20:4); glycerophospholipid metabolism, ether lipid metabolism, glycerolipid metabolism pathway | LC-MS | Untargeted(lipidomic) | − | [60] |
| early, intermediate, and late AMD | urine (fasting) | 4-hydroxyphenylacetate, formate, s-inositol, sucrose, citrate, Val | NMR | Untargeted | cross-sectional | [37] |
| late AMD (wet) | aqueous humor | carnitine, deoxycarnitine, N6-trimethyl-L-lysine, cis-aconitic acid, itaconatic acid, mesaconic acid, Gly, betaine, creatine; carnitine-associated mitochondrial oxidation pathway, carbohydrate metabolism pathway, osmoprotection pathway | LC-MS/MS | Untargeted | case-control | [61] |
4.2. Diabetic Retinopathy (DR)
| DR Stage | Samples | Metabolic Biomarkers/Pathway | Analytical Platform | Untargeted/Targeted | Study Design | References |
|---|---|---|---|---|---|---|
| pre-DR, NPDR, PDR | plasma (−) | pyruvate, Asp, glycerol, cholesterol | GC-MS | Untargeted | − | [84] |
| NPDR | plasma (−) | 2-deoxyribonic acid, 3,4-dihydroxybutyric acid, erythritol, gluconic acid, ribose; pentose phosphate pathway | GC-MS | Untargeted | case-control | [81] |
| NPDR | plasma (−) | 15-oxo-ETE, 4-HDoHE, 11-HEPE, LTB4, PGD2, RvD2, PGD3, PGF2α, 5,6-DiHETE, 8-HDoHE, 5-oxo-ETE, RvD1, 7-HDoHE, 6R-LXA4, 15d-PGJ2, PGJ2, 10-HDoHE, PGE3 | LC-MS/MS | Targeted (eicosanoids) | − | [85] |
| NPDR, PDR | plasma (−) | Glu, Gln, Gln/Glu | GC-MS, LC-MS | Untargeted | − | [71] |
| PDR | plasma (fasting) | fumaric acid, uridine, acetate, cytidine | LC-MS | Untargeted | case-control | [86] |
| NPDR, PDR | plasma (−) | Arg, citrulline, glutamic γ-semialdehyde, dehydroxycarnitine, carnitine | LC-MS | Untargeted | case-control | [87] |
| NPDR, PDR | plasma, serum (−) | 2,4-DHBA, 3,4-DHBA, ribonic acid, ribitol, the triglycerides 50:1 and 50:2 | GC-MS, LC-MS | Untargeted | cross-sectional | [88] |
| NPDR | serum (−) | ribitol, GPC, UDP-Glc-NAc, fructose-6-phosphate | NMR | Untargeted | − | [89] |
| NPDR, PDR | serum (−) | dimethylarginine, Trp, Pro, PC, kynurenine, propionylcarnitine, butyrylcarnitine, hexose | LC-MS | Targeted (IDQ p180 kit) | cross-sectional | [90] |
| NPDR, PDR | serum (fasting) | 12-HETE, 2-piperidone | GC-MS, LC-MS | Untargeted | − | [80] |
| mild DR | Serum (−) | Cer(d18:1/24:0), ChE 20:3, ChE 20:4, ChE 22:6, DG(16:0_18:2), DG(16:1_18:2), DG(18:2_20:4), DG(18:2_22:6), FA(14:0), FA(16:0) | LC-MS | Untargeted (lipidomic) | − | [91] * |
| NPDR, PDR | serum (fasting) | linoleic acid, nicotinamide, ornithine, phenylacetylglutamine | LC-MS | Targeted | case-control | [72] |
| NPDR, PDR | vitreous humor | 5-HETE, CYP-derived epoxyeicosatrienoic acids | LC-MS/MS | Targeted (lipidomic) | − | [92] |
| PDR | vitreous humor | galactitol, ascorbic acid, lactate | NMR | Untargeted | − | [93] |
| PDR | vitreous humor | Arg, Pro, Met, allantoin, citrulline, ornithine, octanoylcarnitine, decanoylcarnitine; arginine, proline, acylcarnitine metabolism pathway | LC-MS/MS | Untargeted | − | [82] |
| PDR | vitreous humor | pyruvate, inosine, hypoxanthine, urate, allantoate, pentose phosphates, xanthine; glucose metabolism, purine metabolism, pentose phosphate pathway | LC-MS | Untargeted | − | [83] |
| PDR | vitreous humor | 5-HETE, 12-HETE, 20-HETE, 20-COOH-AA | LC-MS/MS | Targeted (eicosanoid) | − | [94] |
| PDR | vitreous humour | Pro, pyruvate, lactate, allantoin, creatine, dimethyl glycine, N-acetyl serine, succinate, α-ketoglutarate | LC-MS/MS | Untargeted | cross-sectional | [95] |
| PDR | vitreous, aqueous humor | CysSSH, Cys, GSSSG, cystine | LC-MS/MS | Targeted (polysulfides) | − | [96] |
| PDR | vitreous, aqueous humor | d-2,3-dihydroxypropanoic acid, isocitric acid, fructose 6-phosphate, lactate; pyroglutamic acid, pyruvate; gluconeogenesis, ascorbate-aldarate metabolism, valine–leucine–isoleucine biosynthesis, and arginine–proline metabolism pathway | GC-MS | Untargeted | − | [97] |
| DR | aqueous humor | His, Thr, Gln, Asn, dimethylamine, lactate, succinate, 2-hydroxybutyrate; alanine, aspartate, and glutamate metabolic pathway | NMR | Untargeted | − | [98] |
4.3. Retinopathy of Prematurity (ROP)
| Disease | Samples | Metabolic Biomarkers/Pathway | Analytical Platform | Untargeted/Targeted | Study Design | References |
|---|---|---|---|---|---|---|
| ROP | plasma (−) | N1-methyl-2-pyridone-5-carboxamide, biliverdin, linoleic acid, 4-guanidinobutyric acid, adenosine, thioetheramide-PC, citrulline, GCDC, cis-9-palmitoleic acid, sunitinib, vanillin, trehalose, 1-aminocyclopropanecarboxylic acid | LC-MS | Untargeted | − | [108] |
| ROP | Blood (−) | Gly, Glu, Leu, Ser, Val, Trp, piperidine, citrulline, malonyl carnitine, homocysteine | LC-MS | Targeted | − | [109] |
4.4. Glaucoma
| Disease | Samples | Metabolic Biomarkers/Pathway | Analytical Platform | Untargeted/ Targeted | Study Design | References |
|---|---|---|---|---|---|---|
| POAG | plasma (−) | palmitoylcarnitine, hydroxyergocalciferol, C17 sphinganine, ergostanol | LC-MS/MS | Untargeted | case-control | [125] * |
| POAG | plasma (fasting) | octadecadienyl-carnitine (C18:1), methionine sulfoxide, propionyl-carnitine, PC (34:2), PC (34:4), PC (36:4) | LC-MS | Targeted (IDQ p180 kit) | − | [126] |
| POAG | plasma (fasting) | nicotinamide, hypoxanthine, xanthine, 1-methyl-6,7-dihydroxy-1,2,3,4-tetrahydroisoquinoline, cystathionine, N-acetyl-L-leucine, Arg, rac-glycerol 1-myristate, 1-oleoyl-rac-glycerol | LC-MS | Untargeted | − | [127] |
| PACG | plasma (fasting) | myristic acid, stearic acid, oleic acid, arachidic acid, eicosenoic acid, eicosadienoic acid, eicosatrienoic acid, AA, eicosapentaenoic acid, docosapentaenoic acid, erucic acid, docosatetraenoioc acid, docosahexaenoic acid, tetracosanoic acid | LC-MS/MS | Targeted (FFAs, lipids) | cross-sectional | [128] |
| PACG | serum (fasting) | palmitoleic acid, linoleic acid, γ-linolenic acid, AA | GC-MS | Targeted (FFAs) | − | [129] |
| POAG | serum (fasting) | Gly, Gly-Pro, Asp-Pro, citric acid, Lys, Glu-Ala, MHPG, hypoxanthine, 17-hydroxypregnenolone sulfate, 3α,7α-dihydroxycholanoic acid | GC-MS | Untargeted | − | [116] |
| OAG | aqueous humor | diacylglycerophosphocholines and 1-ether, 2-acylglycerophosphocholines, SM (d18:2/16:0), SM (d18:1/18:0) | LC-MS | Untargeted (lipidomic) | − | [130] * |
| POAG | aqueous humor | taurine, spermine, creatinine, carnitine, propionylcarnitine, acetylcarnitine, Gln, Gly, Ala, Leu, Ile, hydroxyl-proline, acetyl-ornithine, SM (C18:1), LysoPC (C28:1), PC (34:1), PC (36:2), PC (36:4), PC (38:4), PC (32:1) | FIA-MS/MS | Untargeted (lipidomic) | case-control | [131] * |
| PCG | aqueous humor | Gly, Phe, urea | GC-MS | Untargeted | − | [132] |
| POAG, PEXG | aqueous humor | Arg, Lys, Gln, Tyr, His, creatine, 2,4-diacetamido-2,4,6-trideoxy-beta-l-altrose, 5-hydroxypentanoate, N(6)-acetonyllysine, propylene glycol, 1-aminocyclopropane-1-carboxylate | NMR, LC-MS/MS | Untargeted | − | [117] |
| POAG | aqueous humor | biotin, glucose-1-phosphate, methylmalonic acid, N-cyclohexylformamide, sorbitol, spermidine, 2-mercaptoethanesulfonic acid, galactose, mannose, talose, erythronolactone, dehydroascorbic acid | GC-MS | Untargeted | case-control | [133] |
| POAG | aqueous humor | Lys, Arg, Cys, Gly, Gln, Phe, anthranilate, ascorbate, 4-hydroxybenzoate, myo-inositol, acetate, propylene glycol, 2-hydroxy-butyrate, creatine, choline, 4-aminobutanoate, isopropanol | NMR, LC-MS/MS | Untargeted | − | [118] |
| POAG | aqueous humor | betaine, taurine, Glu | NMR | Untargeted | cross-sectional | [65] |
| POAG | aqueous humor | adenine, N-acetyl alanine, hypoxanthine, Lys, Phe-Glu, nicotinamide, 2-aminobutyraldehyde, acetate | LC-MS | Targeted | − | [134] |
| POAG | aqueous humor, plasma (fasting) | cyclic AMP, methylbenzoic acid, 3′-sialyllactose, N-lactoyl-phenylalanine | LC-MS | Targeted | − | [115] |
| POAG | tear | Ala, Arg, Gln/Lys, Leu/Ile/Pro-OH, Met, Phe, Pro, Val, acetylcarnitine, LysoPC (C22:0), LysoPC (C24:0) | DI-MS | Targeted | − | [135] |
4.5. Retinitis Pigmentosa (RP)
5. Discussion
5.1. Samples
5.2. Study Design
5.3. Differential Metabolites/Biomarkers and Pathway
6. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Steinmetz, J.D.; Bourne, R.R.A.; Briant, P.S.; Flaxman, S.R.; Taylor, H.R.B.; Jonas, J.B.; Abdoli, A.A.; Abrha, W.A.; Abualhasan, A.; Abu-Gharbieh, E.G.; et al. Causes of blindness and vision impairment in 2020 and trends over 30 years, and prevalence of avoidable blindness in relation to VISION 2020: The Right to Sight: An analysis for the Global Burden of Disease Study. Lancet Glob. Health 2021, 9, e144–e160. [Google Scholar] [CrossRef]
- Bourne, R.; Steinmetz, J.D.; Flaxman, S.; Briant, P.S.; Taylor, H.R.; Resnikoff, S.; Casson, R.J.; Abdoli, A.; Abu-Gharbieh, E.; Afshin, A.; et al. Trends in prevalence of blindness and distance and near vision impairment over 30 years: An analysis for the Global Burden of Disease Study. Lancet Glob. Health 2021, 9, e130–e143. [Google Scholar] [CrossRef]
- Wishart, D.S. Metabolomics for investigating physiological and pathophysiological processes. Physiol. Rev. 2019, 99, 1819–1875. [Google Scholar] [CrossRef]
- Nazifova-Tasinova, N.; Radeva, M.; Galunska, B.; Grupcheva, C. Metabolomic analysis in ophthalmology. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc Czech Repub. 2020, 164, 236–246. [Google Scholar] [CrossRef]
- Laíns, I.; Gantner, M.; Murinello, S.; Lasky-Su, J.A.; Miller, J.W.; Friedlander, M.; Husain, D. Metabolomics in the study of retinal health and disease. Prog. Retin. Eye Res. 2019, 69, 57–79. [Google Scholar] [CrossRef] [PubMed]
- Alseekh, S.; Aharoni, A.; Brotman, Y.; Contrepois, K.; D’Auria, J.; Ewald, J.; Ewald, J.C.; Fraser, P.D.; Giavalisco, P.; Hall, R.D.; et al. Mass spectrometry-based metabolomics: A guide for annotation, quantification and best reporting practices. Nat. Methods 2021, 18, 747–756. [Google Scholar] [CrossRef]
- Karu, N.; Deng, L.; Slae, M.; Guo, A.C.; Sajed, T.; Huynh, H.; Wine, E.; Wishart, D.S. A review on human fecal metabolomics: Methods, applications and the human fecal metabolome database. Anal. Chim. Acta 2018, 1030, 1–24. [Google Scholar] [CrossRef]
- Chaleckis, R.; Meister, I.; Zhang, P.; Wheelock, C.E. Challenges, progress and promises of metabolite annotation for LC-MS-based metabolomics. Curr. Opin. Biotech. 2019, 55, 44–50. [Google Scholar] [CrossRef]
- Karnovsky, A.; Li, S. Pathway analysis for targeted and untargeted metabolomics. In Computational Methods and Data Analysis for Metabolomics; Li, S., Ed.; Humana: New York, NY, USA, 2020; pp. 387–400. [Google Scholar]
- Deng, P.; Li, X.; Petriello, M.C.; Wang, C.; Morris, A.J.; Hennig, B. Application of metabolomics to characterize environmental pollutant toxicity and disease risks. Rev. Environ. Health 2019, 34, 251–259. [Google Scholar] [CrossRef]
- Olesti, E.; González-Ruiz, V.; Wilks, M.F.; Boccard, J.; Rudaz, S. Approaches in metabolomics for regulatory toxicology applications. Analyst 2021, 146, 1820–1834. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Tian, Y.; Jiang, P.; Lin, Y.; Liu, X.; Yang, H. Recent advances in the application of metabolomics for food safety control and food quality analyses. Crit. Rev. Food Sci. Nutr. 2021, 61, 1448–1469. [Google Scholar] [CrossRef] [PubMed]
- Pinu, F.R.; Goldansaz, S.A.; Jaine, J. Translational metabolomics: Current challenges and future opportunities. Metabolites 2019, 9, 108. [Google Scholar] [CrossRef] [Green Version]
- Li, D.; Gaquerel, E. Next-generation mass spectrometry metabolomics revives the functional analysis of plant metabolic diversity. Annu. Rev. Plant Biol. 2021, 72, 867–891. [Google Scholar] [CrossRef]
- Zhang, X.; Li, Q.; Xu, Z.; Dou, J. Mass spectrometry-based metabolomics in health and medical science: A systematic review. RSC Adv. 2020, 10, 3092–3104. [Google Scholar] [CrossRef] [Green Version]
- Delvaux, A.; Rathahao-Paris, E.; Alves, S. Different ion mobility-mass spectrometry coupling techniques to promote metabolomics. Mass Spectrom. Rev. 2021, 1–27. [Google Scholar] [CrossRef]
- Wang, R.; Yin, Y.; Zhu, Z. Advancing untargeted metabolomics using data-independent acquisition mass spectrometry technology. Anal. Bioanal. Chem. 2019, 411, 4349–4357. [Google Scholar] [CrossRef] [PubMed]
- Heiles, S. Advanced tandem mass spectrometry in metabolomics and lipidomics—Methods and applications. Anal. Bioanal. Chem. 2021, 413, 5927–5948. [Google Scholar] [CrossRef]
- Petras, D.; Jarmusch, A.K.; Dorrestein, P.C. From single cells to our planet—recent advances in using mass spectrometry for spatially resolved metabolomics. Curr. Opin. Chem. Biol. 2017, 36, 24–31. [Google Scholar] [CrossRef] [PubMed]
- Pereiro, X.; Fernández, R.; Barreda-Gómez, G.; Ruzafa, N.; Acera, A.; Araiz, J.; Astigarraga, E.; Vecino, E. Comparative lipidomic analysis of mammalian retinal ganglion cells and Müller glia in situ and in vitro using high-resolution imaging mass spectrometry. Sci. Rep. 2020, 10, 20053. [Google Scholar] [CrossRef]
- Geier, B.; Sogin, E.M.; Michellod, D.; Janda, M.; Kompauer, M.; Spengler, B.; Dubilier, N.; Liebeke, M. Spatial metabolomics of in situ host-microbe interactions at the micrometre scale. Nat. Microbiol. 2020, 5, 498–510. [Google Scholar] [CrossRef] [PubMed]
- Edison, A.S.; Le Guennec, A.; Delaglio, F.; Kupče, Ē. Practical guidelines for 13C-based NMR metabolomics. In NMR-Based Metabolomics; Gowda, G.A.N., Raftery, D., Eds.; Humana: New York, NY, USA, 2019; pp. 69–95. [Google Scholar]
- Gargallo-Garriga, A.; Sardans, J.; Llusià, J.; Peguero, G.; Asensio, D.; Ogaya, R.; Urbina, I.; Langenhove, L.V.; Verryckt, L.T.; Courtois, E.A.; et al. 31P-NMR metabolomics revealed species-specific use of phosphorous in trees of a French Guiana Rainforest. Molecules 2020, 25, 3960. [Google Scholar] [CrossRef]
- Martineau, E.; Dumez, J.N.; Giraudeau, P. Fast quantitative 2D NMR for metabolomics and lipidomics: A tutorial. Magn. Reson. Chem. 2020, 58, 390–403. [Google Scholar] [CrossRef] [PubMed]
- Marchand, J.; Martineau, E.; Guitton, Y.; Le Bizec, B.; Dervilly-Pinel, G.; Giraudeau, P. A multidimensional 1H NMR lipidomics workflow to address chemical food safety issues. Metabolomics 2018, 14, 60. [Google Scholar] [CrossRef]
- Hu, K.; Westler, W.M.; Markley, J.L. Simultaneous quantification and identification of individual chemicals in metabolite mixtures by two-dimensional extrapolated time-zero 1H−13C HSQC (HSQC0). J. Am. Chem. Soc. 2011, 133, 1662–1665. [Google Scholar] [CrossRef] [PubMed]
- Peterson, D.J.; Loening, N.M. QQ-HSQC: A quick, quantitative heteronuclear correlation experiment for NMR spectroscopy. Magn. Reson. Chem. 2007, 45, 937–941. [Google Scholar] [CrossRef] [PubMed]
- Mäkelä, V.; Helminen, J.; Kilpeläinen, I.; Heikkinen, S. Quantitative, equal carbon response HSQC experiment, QEC-HSQC. J. Magn. Reson. 2016, 271, 34–39. [Google Scholar] [CrossRef]
- Wishart, D.S. NMR metabolomics: A look ahead. J. Magn. Reson. 2019, 306, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Wong, W.L.; Su, X.Y.; Li, X.; Cheung, C.M.G.; Klein, R.; Cheng, C.Y.; Wong, T.Y. Global prevalence of age-related macular degeneration and disease burden projection for 2020 and 2040: A systematic review and meta-analysis. Lancet Glob. Health 2014, 2, e106–e116. [Google Scholar] [CrossRef] [Green Version]
- Datta, S.; Cano, M.; Ebrahimi, K.; Wang, L.; Handa, J.T. The impact of oxidative stress and inflammation on RPE degeneration in non-neovascular AMD. Prog. Retin. Eye Res. 2017, 60, 201–218. [Google Scholar] [CrossRef] [PubMed]
- Semeraro, F.; Morescalchi, F.; Duse, S.; Parmeggiani, F.; Gambicorti, E.; Costagliola, C. Aflibercept in wet AMD: Specific role and optimal use. Drug Des. Devel. Ther. 2013, 7, 711–722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hussain, R.M.; Shaukat, B.A.; Ciulla, L.M.; Berrocal, A.M.; Sridhar, J. Vascular endothelial growth factor antagonists: Promising players in the treatment of neovascular age-related macular degeneration. Drug Des. Devel. Ther. 2021, 15, 2653–2665. [Google Scholar] [CrossRef]
- Assel, M.J.; Li, F.; Wang, Y.; Allen, A.S.; Baggerly, K.A.; Vickers, A.J. Genetic polymorphisms of CFH and ARMS2 do not predict response to antioxidants and zinc in patients with age-related macular degeneration: Independent statistical evaluations of data from the age-related eye disease study. Ophthalmology 2018, 125, 391–397. [Google Scholar] [CrossRef]
- Kersten, E.; Paun, C.C.; Schellevis, R.L.; Hoyng, C.B.; Delcourt, C.; Lengyel, I.; Peto, T.; Ueffing, M.; Klaver, C.C.W.; Dammeier, S.; et al. Systemic and ocular fluid compounds as potential biomarkers in age-related macular degeneration. Surv. Ophthalmol. 2018, 63, 9–39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hou, X.W.; Wang, Y.; Pan, C.W. Metabolomics in age-related macular degeneration: A systematic review. Invest. Ophthalmol. Vis. Sci. 2020, 61, 13. [Google Scholar] [CrossRef] [PubMed]
- Laíns, I.; Duarte, D.; Barros, A.S.; Martins, A.S.; Carneiro, T.J.; Gil, J.Q.; Miller, J.B.; Marques, M.; Mesquita, T.S.; Barreto, P.; et al. Urine nuclear magnetic resonance (NMR) metabolomics in age-related macular degeneration. J. Proteome Res. 2019, 18, 1278–1288. [Google Scholar] [CrossRef]
- Acar, İ.E.; Lores-Motta, L.; Colijn, J.M.; Meester-Smoor, M.A.; Verzijden, T.; Cougnard-Gregoire, A.; Ajana, S.; Merle, B.M.J.; De Breuk, A.; Heesterbeek, T.J.; et al. Integrating metabolomics, genomics, and disease pathways in age-related macular degeneration: The EYE-RISK Consortium. Ophthalmology 2020, 127, 1693–1709. [Google Scholar] [CrossRef]
- Chao, J.R.; Knight, K.; Engel, A.L.; Jankowski, C.; Wang, Y.; Manson, M.A.; Gu, H.; Djukovic, D.; Raftery, D.; Hurley, J.B.; et al. Human retinal pigment epithelial cells prefer proline as a nutrient and transport metabolic intermediates to the retinal side. J. Biol. Chem. 2017, 292, 12895–12905. [Google Scholar] [CrossRef] [Green Version]
- Zhang, R.; Engel, A.L.; Wang, Y.; Li, B.; Shen, W.; Gillies, M.C.; Chao, J.R.; Du, J. Inhibition of mitochondrial respiration impairs nutrient consumption and metabolite transport in human retinal pigment epithelium. J. Proteome Res. 2021, 20, 909–922. [Google Scholar] [CrossRef]
- Alamri, A.; Biswas, L.; Watson, D.G.; Shu, X. Deletion of TSPO resulted in change of metabolomic profile in retinal pigment epithelial cells. Int. J. Mol. Sci. 2019, 20, 1387. [Google Scholar] [CrossRef] [Green Version]
- Zhang, M.; Jiang, N.; Chu, Y.; Postnikova, O.; Varghese, R.; Horvath, A.; Cheema, A.K.; Golestaneh, N. Dysregulated metabolic pathways in age-related macular degeneration. Sci. Rep. 2020, 10, 2464. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Go, Y.M.; Zhang, J.; Fernandes, J.; Litwin, C.; Chen, R.; Wensel, T.G.; Jones, D.P.; Cai, J.; Chen, Y. MTOR-initiated metabolic switch and degeneration in the retinal pigment epithelium. FASEB J. 2020, 34, 12502–12520. [Google Scholar] [CrossRef]
- Rosell, M.; Giera, M.; Brabet, P.; Shchepinov, M.S.; Guichardant, M.; Durand, T.; Vercauteren, J.; Galano, J.M.; Crauste, C. Bis-allylic deuterated DHA alleviates oxidative stress in retinal epithelial cells. Antioxidants 2019, 8, 447. [Google Scholar] [CrossRef] [Green Version]
- Wang, J.; Westenskow, P.D.; Fang, M.; Friedlander, M.; Siuzdak, G. Quantitative metabolomics of photoreceptor degeneration and the effects of stem cell-derived retinal pigment epithelium transplantation. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2016, 374, 20150376. [Google Scholar] [CrossRef] [Green Version]
- Zhang, R.; Shen, W.; Du, J.; Gillies, M.C. Selective knockdown of hexokinase 2 in rods leads to age-related photoreceptor degeneration and retinal metabolic remodeling. Cell Death Dis. 2020, 11, 885. [Google Scholar] [CrossRef] [PubMed]
- Mendez, K.M.; Kim, J.; Laíns, I.; Nigalye, A.; Katz, R.; Pundik, S.; Kim, I.K.; Liang, L.; Vavvas, D.G.; Miller, J.B.; et al. Association of human plasma metabolomics with delayed dark adaptation in age-related macular degeneration. Metabolites 2021, 11, 183. [Google Scholar] [CrossRef] [PubMed]
- Lambert, V.; Hansen, S.; Schoumacher, M.; Lecomte, J.; Leenders, J.; Hubert, P.; Herfs, M.; Blacher, S.; Carnet, O.; Yip, C.; et al. Pyruvate dehydrogenase kinase/lactate axis: A therapeutic target for neovascular age-related macular degeneration identified by metabolomics. J. Mol. Med. 2020, 98, 1737–1751. [Google Scholar] [CrossRef]
- Homma, K.; Toda, E.; Osada, H.; Nagai, N.; Era, T.; Tsubota, K.; Okano, H.; Ozawa, Y. Taurine rescues mitochondria-related metabolic impairments in the patient-derived induced pluripotent stem cells and epithelial-mesenchymal transition in the retinal pigment epithelium. Redox Biol. 2021, 41, 101921. [Google Scholar] [CrossRef]
- Gao, Y.; Teo, Y.C.K.; Beuerman, R.W.; Wong, T.Y.; Zhou, L.; Cheung, C.M.G. A serum metabolomics study of patients with nAMD in response to anti-VEGF therapy. Sci. Rep. 2020, 10, 1341. [Google Scholar] [CrossRef]
- Liu, K.; Fang, J.; Jin, J.; Zhu, S.; Xu, X.; Xu, Y.; Ye, B.; Lin, S.H.; Xu, X. Serum metabolomics reveals personalized metabolic patterns for macular neovascular disease patient stratification. J. Proteome Res. 2020, 19, 699–707. [Google Scholar] [CrossRef] [PubMed]
- Osborn, M.P.; Park, Y.; Parks, M.B.; Burgess, L.G.; Uppal, K.; Lee, K.; Jones, D.P.; Brantley, M.A., Jr. Metabolome-wide association study of neovascular age-related macular degeneration. PLoS ONE 2013, 8, e72737. [Google Scholar] [CrossRef]
- Laíns, I.; Duarte, D.; Barros, A.S.; Martins, A.S.; Gil, J.; Miller, J.B.; Marques, M.; Mesquita, T.; Kim, I.K.; Cachulo, M.D.L.; et al. Human plasma metabolomics in age-related macular degeneration (AMD) using nuclear magnetic resonance spectroscopy. PLoS ONE 2017, 12, e0177749. [Google Scholar] [CrossRef] [Green Version]
- Luo, D.; Deng, T.; Yuan, W.; Deng, H.; Jin, M. Plasma metabolomic study in Chinese patients with wet age-related macular degeneration. BMC Ophthalmol. 2017, 17, 165. [Google Scholar] [CrossRef] [Green Version]
- Laíns, I.; Kelly, R.S.; Miller, J.B.; Silva, R.; Vavvas, D.G.; Kim, I.K.; Murta, J.N.; Lasky-Su, J.; Miller, J.W.; Husain, D. Human plasma metabolomics study across all stages of age-related macular degeneration identifies potential lipid biomarkers. Ophthalmology 2018, 125, 245–254. [Google Scholar] [CrossRef]
- Mitchell, S.L.; Uppal, K.; Williamson, S.M.; Liu, K.; Burgess, L.G.; Tran, V.; Umfress, A.C.; Jarrell, K.L.; Cooke Bailey, J.N.; Agarwal, A.; et al. The carnitine shuttle pathway is altered in patients with neovascular age-related macular degeneration. Invest. Ophthalmol. Vis. Sci. 2018, 59, 4978–4985. [Google Scholar] [CrossRef] [Green Version]
- Laíns, I.; Chung, W.; Kelly, R.S.; Gil, J.; Marques, M.; Barreto, P.; Murta, J.N.; Kim, I.K.; Vavvas, D.G.; Miller, J.B.; et al. Human plasma metabolomics in age-related macular degeneration: Meta-analysis of two cohorts. Metabolites 2019, 9, 127. [Google Scholar] [CrossRef] [Green Version]
- Chao de la Barca, J.M.; Rondet-Courbis, B.; Ferré, M.; Muller, J.; Buisset, A.; Leruez, S.; Plubeau, G.; Macé, T.; Moureauzeau, L.; Chupin, S.; et al. A plasma metabolomic profiling of exudative age-related macular degeneration showing carnosine and mitochondrial deficiencies. J. Clin. Med. 2020, 9, 631. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kersten, E.; Dammeier, S.; Ajana, S.; Groenewoud, J.M.M.; Codrea, M.; Klose, F.; Lechanteur, Y.T.; Fauser, S.; Ueffing, M.; Delcourt, C.; et al. Metabolomics in serum of patients with non-advanced age-related macular degeneration reveals aberrations in the glutamine pathway. PLoS ONE 2019, 14, e0218457. [Google Scholar] [CrossRef] [Green Version]
- Li, M.; Zhang, X.; Liao, N.; Ye, B.; Peng, Y.; Ji, Y.; Wen, F. Analysis of the serum lipid profile in polypoidal choroidal vasculopathy. Sci. Rep. 2016, 6, 38342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Han, G.; Wei, P.; He, M.; Teng, H.; Chu, Y. Metabolomic profiling of the aqueous humor in patients with wet age-related macular degeneration using UHPLC-MS/MS. J. Proteome Res. 2020, 19, 2358–2366. [Google Scholar] [CrossRef] [PubMed]
- Sabanayagam, C.; Banu, R.; Chee, M.L.; Lee, R.; Wang, Y.X.; Tan, G.; Jonas, J.B.; Lamoureux, E.L.; Cheng, C.Y.; Klein, B.E.K.; et al. Incidence and progression of diabetic retinopathy: A systematic review. Lancet Diabetes Endocrinol. 2019, 7, 140–149. [Google Scholar] [CrossRef]
- Eid, S.; Sas, K.M.; Abcouwer, S.F.; Feldman, E.L.; Gardner, T.W.; Pennathur, S.; Fort, P.E. New insights into the mechanisms of diabetic complications: Role of lipids and lipid metabolism. Diabetologia 2019, 62, 1539–1549. [Google Scholar] [CrossRef] [Green Version]
- Antonetti, D.A.; Silva, P.S.; Stitt, A.W. Current understanding of the molecular and cellular pathology of diabetic retinopathy. Nat. Rev. Endocrinol. 2021, 17, 195–206. [Google Scholar] [CrossRef]
- Early Treatment Diabetic Retinopathy Study Research Group. Grading diabetic retinopathy from stereoscopic color fundus photographs—An extension of the modified Airlie House Classification: ETDRS Report Number 10. Ophthalmology 2020, 127, S99–S119. [Google Scholar] [CrossRef] [PubMed]
- Early Treatment Diabetic Retinopathy Study Research Group. Fundus photographic risk factors for progression of diabetic retinopathy: ETDRS Report Number 12. Ophthalmology 1991, 98, 823–833. [Google Scholar] [CrossRef]
- Midena, E.; Frizziero, L.; Midena, G.; Pilotto, E. Intraocular fluid biomarkers (liquid biopsy) in human diabetic retinopathy. Graefe’s Arch. Clin. Exp. Ophthalmol. 2021. [Google Scholar] [CrossRef]
- Aderemi, A.V.; Ayeleso, A.O.; Oyedapo, O.O.; Mukwevho, E. Metabolomics: A scoping review of its role as a tool for disease biomarker discovery in selected non-communicable diseases. Metabolites 2021, 11, 418. [Google Scholar] [CrossRef]
- Marcovecchio, M.L. Importance of identifying novel biomarkers of microvascular damage in type 1 diabetes. Mol. Diagn. Ther. 2020, 24, 507–515. [Google Scholar] [CrossRef]
- Ting, D.S.; Tan, K.A.; Phua, V.; Tan, G.S.; Wong, C.W.; Wong, T.Y. Biomarkers of diabetic retinopathy. Curr. Diab. Rep. 2016, 16, 125. [Google Scholar] [CrossRef] [PubMed]
- Rhee, S.Y.; Jung, E.S.; Park, H.M.; Jeong, S.J.; Kim, K.; Chon, S.; Yu, S.Y.; Woo, J.T.; Lee, C.H. Plasma glutamine and glutamic acid are potential biomarkers for predicting diabetic retinopathy. Metabolomics 2018, 14, 89. [Google Scholar] [CrossRef] [Green Version]
- Zuo, J.; Lan, Y.; Hu, H.; Hou, X.; Li, J.; Wang, T.; Zhang, H.; Zhang, N.; Guo, C.; Peng, F.; et al. Metabolomics-based multidimensional network biomarkers for diabetic retinopathy identification in patients with type 2 diabetes mellitus. BMJ Open Diabetes Res. Care 2021, 9, e001443. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, V.; Yanes, O.; Aguilar, E.; Wang, M.; Friedlander, D.; Moreno, S.; Storm, K.; Zhan, M.; Naccache, S.; Nemerow, G.; et al. Differential macrophage polarization promotes tissue remodeling and repair in a model of ischemic retinopathy. Sci. Rep. 2011, 1, 76. [Google Scholar] [CrossRef]
- Sas, K.M.; Lin, J.; Rajendiran, T.M.; Soni, T.; Nair, V.; Hinder, L.M.; Jagadish, H.V.; Gardner, T.W.; Abcouwer, S.F.; Brosius, F.C.; et al. Shared and distinct lipid-lipid interactions in plasma and affected tissues in a diabetic mouse model. J. Lipid Res. 2018, 59, 173–183. [Google Scholar] [CrossRef] [Green Version]
- Patrick, A.T.; He, W.; Madu, J.; Sripathi, S.R.; Choi, S.; Lee, K.; Samson, F.P.; Powell, F.L.; Bartoli, M.; Jee, D.; et al. Mechanistic dissection of diabetic retinopathy using the protein-metabolite interactome. J. Diabetes Metab. Disord. 2020, 19, 829–848. [Google Scholar] [CrossRef] [PubMed]
- Wiggenhauser, L.M.; Qi, H.; Stoll, S.J.; Metzger, L.; Bennewitz, K.; Poschet, G.; Krenning, G.; Hillebrands, J.L.; Hammes, H.P.; Kroll, J. Activation of retinal angiogenesis in hyperglycemic pdx1−/− Zebrafish mutants. Diabetes 2020, 69, 1020–1031. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, A.; Cervia, D.; Catalani, E.; Gevi, F.; Zolla, L.; Casini, G. Protective effects of the neuropeptides PACAP, substance P and the somatostatin analogue octreotide in retinal ischemia: A metabolomic analysis. Mol. Biosyst. 2014, 10, 1290–1304. [Google Scholar] [CrossRef] [PubMed]
- Ge, N.; Kong, L.; Zhang, A.H.; Sun, Y.; Zhao, M.Q.; Zhang, B.; Xu, L.; Ke, X.; Sun, H.; Wang, X.J. Identification of key lipid metabolites during metabolic dysregulation in the diabetic retinopathy disease mouse model and efficacy of Keluoxin capsule using an UHPLC-MS-based non-targeted lipidomics approach. RSC Adv. 2021, 11, 5491–5505. [Google Scholar] [CrossRef]
- Wang, X.; Li, Y.; Xie, M.; Deng, L.; Zhang, M.; Xie, X. Urine metabolomics study of Bushen Huoxue prescription on diabetic retinopathy rats by UPLC-Q-exactive Orbitrap-MS. Biomed. Chromatogr. 2020, 34, e4792. [Google Scholar] [CrossRef] [PubMed]
- Xuan, Q.; Ouyang, Y.; Wang, Y.; Wu, L.; Li, H.; Luo, Y.; Zhao, X.; Feng, D.; Qin, W.; Hu, C.; et al. Multiplatform metabolomics reveals novel serum metabolite biomarkers in diabetic retinopathy subjects. Adv. Sci. 2020, 7, 2001714. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Cheng, C.Y.; Choi, H.; Ikram, M.K.; Sabanayagam, C.; Tan, G.S.; Tian, D.; Zhang, L.; Venkatesan, G.; Tai, E.S.; et al. Plasma metabonomic profiling of diabetic retinopathy. Diabetes 2016, 65, 1099–1108. [Google Scholar] [CrossRef] [Green Version]
- Paris, L.P.; Johnson, C.H.; Aguilar, E.; Usui, Y.; Cho, K.; Hoang, L.T.; Feitelberg, D.; Benton, H.P.; Westenskow, P.D.; Kurihara, T.; et al. Global metabolomics reveals metabolic dysregulation in ischemic retinopathy. Metabolomics 2016, 12, 15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Haines, N.R.; Manoharan, N.; Olson, J.L.; D’Alessandro, A.; Reisz, J.A. Metabolomics analysis of human vitreous in diabetic retinopathy and rhegmatogenous retinal detachment. J. Proteome Res. 2018, 17, 2421–2427. [Google Scholar] [CrossRef]
- Li, X.; Luo, X.; Lu, X.; Duan, J.; Xu, G. Metabolomics study of diabetic retinopathy using gas chromatography-mass spectrometry: A comparison of stages and subtypes diagnosed by Western and Chinese medicine. Mol. Biosyst. 2011, 7, 2228–2237. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Sun, B.; Liu, M.; Huang, J.; Liu, Y.; Xie, Z.; He, J.; Chen, L.; Wang, D.; Zhu, Y.; et al. Plasma metabolic profile reveals PGF2α protecting against non-proliferative diabetic retinopathy in patients with type 2 diabetes. Biochem. Biophys. Res. Commun. 2018, 496, 1276–1283. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.R.; Yang, F.Y.; Lu, J.; Zhang, H.R.; Sun, R.; Zhou, J.B.; Yang, J.K. Plasma metabolomic profiling of proliferative diabetic retinopathy. Nutr. Metab. 2019, 16, 37. [Google Scholar] [CrossRef] [Green Version]
- Sumarriva, K.; Uppal, K.; Ma, C.; Herren, D.J.; Wang, Y.; Chocron, I.M.; Warden, C.; Mitchell, S.L.; Burgess, L.G.; Goodale, M.P.; et al. Arginine and carnitine metabolites are altered in diabetic retinopathy. Invest. Ophthalmol. Vis. Sci. 2019, 60, 3119–3126. [Google Scholar] [CrossRef] [Green Version]
- Curovic, V.R.; Suvitaival, T.; Mattila, I.; Ahonen, L.; Trošt, K.; Theilade, S.; Hansen, T.W.; Legido-Quigley, C.; Rossing, P. Circulating metabolites and lipids are associated to diabetic retinopathy in individuals with type 1 diabetes. Diabetes 2020, 69, 2217–2226. [Google Scholar] [CrossRef]
- Guha Mazumder, A.; Chatterjee, S.; Chatterjee, S.; Gonzalez, J.J.; Bag, S.; Ghosh, S.; Mukherjee, A.; Chatterjee, J. Spectropathology-corroborated multimodal quantitative imaging biomarkers for neuroretinal degeneration in diabetic retinopathy. Clin. Ophthalmol. 2017, 11, 2073–2089. [Google Scholar] [CrossRef] [Green Version]
- Yun, J.H.; Kim, J.M.; Jeon, H.J.; Oh, T.; Choi, H.J.; Kim, B.J. Metabolomics profiles associated with diabetic retinopathy in type 2 diabetes patients. PLoS ONE 2020, 15, e0241365. [Google Scholar] [CrossRef] [PubMed]
- Xuan, Q.; Zheng, F.; Yu, D.; Ouyang, Y.; Zhao, X.; Hu, C.; Xu, G. Rapid lipidomic profiling based on ultra-high performance liquid chromatography-mass spectrometry and its application in diabetic retinopathy. Anal. Bioanal. Chem. 2020, 412, 3585–3594. [Google Scholar] [CrossRef]
- Schwartzman, M.L.; Iserovich, P.; Gotlinger, K.; Bellner, L.; Dunn, M.W.; Sartore, M.; Grazia Pertile, M.; Leonardi, A.; Sathe, S.; Beaton, A.; et al. Profile of lipid and protein autacoids in diabetic vitreous correlates with the progression of diabetic retinopathy. Diabetes 2010, 59, 1780–1788. [Google Scholar] [CrossRef] [Green Version]
- Barba, I.; Garcia-Ramírez, M.; Hernández, C.; Alonso, M.A.; Masmiquel, L.; García-Dorado, D.; Simó, R. Metabolic fingerprints of proliferative diabetic retinopathy: An 1H-NMR-based metabonomic approach using vitreous humor. Invest. Ophthalmol. Vis. Sci. 2010, 51, 4416–4421. [Google Scholar] [CrossRef]
- Lin, A.L.; Roman, R.J.; Regan, K.A.; Bolch, C.A.; Chen, C.J.; Iyer, S.S.R. Eicosanoid profiles in the vitreous humor of patients with proliferative diabetic retinopathy. Int. J. Mol. Sci. 2020, 21, 7451. [Google Scholar] [CrossRef]
- Tomita, Y.; Cagnone, G.; Fu, Z.; Cakir, B.; Kotoda, Y.; Asakage, M.; Wakabayashi, Y.; Hellström, A.; Joyal, J.S.; Talukdar, S.; et al. Vitreous metabolomics profiling of proliferative diabetic retinopathy. Diabetologia 2021, 64, 70–82. [Google Scholar] [CrossRef]
- Kunikata, H.; Ida, T.; Sato, K.; Aizawa, N.; Sawa, T.; Tawarayama, H.; Murayama, N.; Fujii, S.; Akaike, T.; Nakazawa, T. Metabolomic profiling of reactive persulfides and polysulfides in the aqueous and vitreous humors. Sci. Rep. 2017, 7, 41984. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Fang, J.; Chen, F.; Sun, Q.; Xu, X.; Lin, S.H.; Liu, K. Metabolomic profile of diabetic retinopathy: A GC-TOFMS-based approach using vitreous and aqueous humor. Acta Diabetol. 2020, 57, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Jin, H.; Zhu, B.; Liu, X.; Jin, J.; Zou, H. Metabolic characterization of diabetic retinopathy: An 1H-NMR-based metabolomic approach using human aqueous humor. J. Pharm. Biomed. Anal. 2019, 174, 414–421. [Google Scholar] [CrossRef] [PubMed]
- Smith, L.E.H. Pathogenesis of retinopathy of prematurity. Growth Horm. IGF Res. 2004, 14, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.J.; Port, A.D.; Swan, R.; Campbell, J.P.; Chan, R.V.P.; Chiang, M.F. Retinopathy of prematurity: A review of risk factors and their clinical significance. Surv. Ophthalmol. 2018, 63, 618–637. [Google Scholar] [CrossRef] [PubMed]
- Adams, G.G.W.; Bunce, C.; Xing, W.; Butler, L.; Long, V.; Reddy, A.; Dahlmann-Noor, A.H. Treatment trends for retinopathy of prematurity in the UK: Active surveillance study of infants at risk. BMJ Open 2017, 7, e013366. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sen, P.; Wu, W.C.; Chandra, P.; Vinekar, A.; Manchegowda, P.T.; Bhende, P. Retinopathy of prematurity treatment: Asian perspectives. Eye 2020, 34, 632–642. [Google Scholar] [CrossRef] [PubMed]
- Georgakopoulou, I.; Chasapi, S.A.; Bariamis, S.E.; Varvarigou, A.; Spraul, M.; Spyroulias, G.A. Metabolic changes in early neonatal life: NMR analysis of the neonatal metabolic profile to monitor postnatal metabolic adaptations. Metabolomics 2020, 16, 58. [Google Scholar] [CrossRef] [PubMed]
- Mussap, M.; Antonucci, R.; Noto, A.; Fanos, V. The role of metabolomics in neonatal and pediatric laboratory medicine. Clin. Chim. Acta 2013, 426, 127–138. [Google Scholar] [CrossRef] [PubMed]
- Bjerkhaug, A.U.; Granslo, H.N.; Klingenberg, C. Metabolic responses in neonatal sepsis—A systematic review of human metabolomic studies. Acta Paediatr. 2021, 110, 2316–2325. [Google Scholar] [CrossRef] [PubMed]
- Lu, F.; Liu, Y.; Guo, Y.; Gao, Y.; Piao, Y.; Tan, S.; Tang, Y. Metabolomic changes of blood plasma associated with two phases of rat OIR. Exp. Eye Res. 2020, 190, 107855. [Google Scholar] [CrossRef] [PubMed]
- Singh, C.; Hoppe, G.; Tran, V.; McCollum, L.; Bolok, Y.; Song, W.; Sharma, A.; Brunengraber, H.; Sears, J.E. Serine and 1-carbon metabolism are required for HIF-mediated protection against retinopathy of prematurity. JCI Insight 2019, 4, e129398. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Xu, Y.; Zhang, X.; Zhao, P.; Gong, X.; He, M.; Cao, J.; Jiang, B.; Yoshida, S.; Li, Y. Plasma metabolites in treatment-requiring retinopathy of prematurity: Potential biomarkers identified by metabolomics. Exp. Eye Res. 2020, 199, 108198. [Google Scholar] [CrossRef]
- Yang, Y.; Wu, Z.; Li, S.; Yang, M.; Xiao, X.; Lian, C.; Wen, W.; He, H.; Zeng, J.; Wang, J.; et al. Targeted blood metabolomic study on retinopathy of prematurity. Invest. Ophthalmol. Vis. Sci. 2020, 61, 12. [Google Scholar] [CrossRef] [Green Version]
- Tham, Y.C.; Li, X.; Wong, T.Y.; Quigley, H.A.; Aung, T.; Cheng, C.Y. Global prevalence of glaucoma and projections of glaucoma burden through 2040: A systematic review and meta-analysis. Ophthalmology 2014, 121, 2081–2090. [Google Scholar] [CrossRef]
- Kumarasamy, N.A.; Lam, F.S.; Wang, A.L.; Theoharides, T.C. Glaucoma: Current and developing concepts for inflammation, pathogenesis and treatment. Eur. J. Inflamm. 2006, 4, 129–137. [Google Scholar] [CrossRef] [Green Version]
- Stein, J.D.; Khawaja, A.P.; Weizer, J.S. Glaucoma in adults—Screening, diagnosis, and management: A review. J. Am. Med. Assoc. 2021, 325, 164–174. [Google Scholar] [CrossRef]
- Barbosa-Breda, J.; Himmelreich, U.; Ghesquière, B.; Rocha-Sousa, A.; Stalmans, I. Clinical metabolomics and glaucoma. Ophthalmic. Res. 2018, 59, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hou, X.W.; Liang, G.; Pan, C.W. Metabolomics in glaucoma: A systematic review. Invest. Ophthalmol. Vis. Sci. 2021, 62, 9. [Google Scholar] [CrossRef]
- Tang, Y.; Pan, Y.; Chen, Y.; Kong, X.; Chen, J.; Zhang, H.; Tang, G.; Wu, J.; Sun, X. Metabolomic profiling of aqueous humor and plasma in primary open angle glaucoma patients points towards novel diagnostic and therapeutic strategy. Front. Pharmacol. 2021, 12, 621146. [Google Scholar] [CrossRef] [PubMed]
- Gong, H.; Zhang, S.; Li, Q.; Zuo, C.; Gao, X.; Zheng, B.; Lin, M. Gut microbiota compositional profile and serum metabolic phenotype in patients with primary open-angle glaucoma. Exp. Eye Res. 2020, 191, 107921. [Google Scholar] [CrossRef]
- Myer, C.; Abdelrahman, L.; Banerjee, S.; Khattri, R.B.; Merritt, M.E.; Junk, A.K.; Lee, R.K.; Bhattacharya, S.K. Aqueous humor metabolite profile of pseudoexfoliation glaucoma is distinctive. Mol. Omics 2020, 16, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Myer, C.; Perez, J.; Abdelrahman, L.; Mendez, R.; Khattri, R.B.; Junk, A.K.; Bhattacharya, S.K. Differentiation of soluble aqueous humor metabolites in primary open angle glaucoma and controls. Exp. Eye Res. 2020, 194, 108024. [Google Scholar] [CrossRef] [PubMed]
- Barbosa Breda, J.; Croitor Sava, A.; Himmelreich, U.; Somers, A.; Matthys, C.; Rocha Sousa, A.; Vandewalle, E.; Stalmans, I. Metabolomic profiling of aqueous humor from glaucoma patients-The metabolomics in surgical ophthalmological patients (MISO) study. Exp. Eye Res. 2020, 201, 108268. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, X.; Wen, G.; Wu, G.; Zhang, X. Proton magnetic resonance spectroscopy (1H-MRS) reveals geniculocalcarine and striate area degeneration in primary glaucoma. PLos ONE 2013, 8, e73197. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Takayanagi, Y.; Takai, Y.; Kaidzu, S.; Tanito, M. Evaluation of redox profiles of the serum and aqueous humor in patients with primary open-angle glaucoma and exfoliation glaucoma. Antioxidants 2020, 9, 1305. [Google Scholar] [CrossRef]
- Hysi, P.G.; Khawaja, A.P.; Menni, C.; Tamraz, B.; Wareham, N.; Khaw, K.T.; Foster, P.J.; Benet, L.Z.; Spector, T.D.; Hammond, C.J. Ascorbic acid metabolites are involved in intraocular pressure control in the general population. Redox Biol. 2019, 20, 349–353. [Google Scholar] [CrossRef]
- Harder, J.M.; Guymer, C.; Wood, J.P.M.; Daskalaki, E.; Chidlow, G.; Zhang, C.; Balasubramanian, R.; Cardozo, B.H.; Foxworth, N.E.; Deering, K.E.; et al. Disturbed glucose and pyruvate metabolism in glaucoma with neuroprotection by pyruvate or rapamycin. Proc. Natl. Acad. Sci. USA 2020, 117, 33619–33627. [Google Scholar] [CrossRef]
- Tribble, J.R.; Otmani, A.; Sun, S.; Ellis, S.A.; Cimaglia, G.; Vohra, R.; Jöe, M.; Lardner, E.; Venkataraman, A.P.; Domínguez-Vicent, A.; et al. Nicotinamide provides neuroprotection in glaucoma by protecting against mitochondrial and metabolic dysfunction. Redox Biol. 2021, 43, 101988. [Google Scholar] [CrossRef] [PubMed]
- Burgess, L.G.; Uppal, K.; Walker, D.I.; Roberson, R.M.; Tran, V.; Parks, M.B.; Wade, E.A.; May, A.T.; Umfress, A.C.; Jarrell, K.L.; et al. Metabolome-wide association study of primary open angle glaucoma. Invest. Ophthalmol. Vis. Sci. 2015, 56, 5020–5028. [Google Scholar] [CrossRef]
- Leruez, S.; Marill, A.; Bresson, T.; De Saint Martin, G.; Buisset, A.; Muller, J.; Tessier, L.; Gadras, C.; Verny, C.; Gohier, P.; et al. A metabolomics profiling of glaucoma points to mitochondrial dysfunction, senescence, and polyamines deficiency. Invest. Ophthalmol. Vis. Sci. 2018, 59, 4355–4361. [Google Scholar] [CrossRef]
- Kouassi Nzoughet, J.; Guehlouz, K.; Leruez, S.; Gohier, P.; Bocca, C.; Muller, J.; Blanchet, O.; Bonneau, D.; Simard, G.; Milea, D.; et al. A data mining metabolomics exploration of glaucoma. Metabolites 2020, 10, 49. [Google Scholar] [CrossRef] [Green Version]
- Qin, Y.; Feng, X.; Luo, H.; Liu, S.; Wang, X.; Wang, X.; Yang, Q.; Zhao, Q.; Cao, Y.; Li, J.; et al. Association between plasma free fatty acid levels and primary angle-closure glaucoma based on a mass spectrometry metabolomics analysis. Acta Ophthalmol. 2021. [Google Scholar] [CrossRef]
- Rong, S.; Li, Y.; Guan, Y.; Zhu, L.; Zhou, Q.; Gao, M.; Pan, H.; Zou, L.; Chang, D. Long-chain unsaturated fatty acids as possible important metabolites for primary angle-closure glaucoma based on targeted metabolomic analysis. Biomed. Chromatogr. 2017, 31, e3963. [Google Scholar] [CrossRef] [PubMed]
- Cabrerizo, J.; Urcola, J.A.; Vecino, E. Changes in the lipidomic profile of aqueous humor in open-angle glaucoma. J. Glaucoma 2017, 26, 349–355. [Google Scholar] [CrossRef] [PubMed]
- Buisset, A.; Gohier, P.; Leruez, S.; Muller, J.; Amati-Bonneau, P.; Lenaers, G.; Bonneau, D.; Simard, G.; Procaccio, V.; Annweiler, C.; et al. Metabolomic profiling of aqueous humor in glaucoma points to taurine and spermine deficiency: Findings from the Eye-D study. J. Proteome Res. 2019, 18, 1307–1315. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Chen, Y.; Wang, L.; Sun, X. Metabolomics of the aqueous humor in patients with primary congenital glaucoma. Mol. Vis. 2019, 25, 489–501. [Google Scholar] [PubMed]
- Pan, C.W.; Ke, C.; Chen, Q.; Tao, Y.J.; Zha, X.; Zhang, Y.P.; Zhong, H. Differential metabolic markers associated with primary open-angle glaucoma and cataract in human aqueous humor. BMC Ophthalmol. 2020, 20, 183. [Google Scholar] [CrossRef]
- Pulukool, S.K.; Bhagavatham, S.K.S.; Kannan, V.; Sukumar, P.; Dandamudi, R.B.; Ghaisas, S.; Kunchala, H.; Saieesh, D.; Naik, A.A.; Pargaonkar, A.; et al. Elevated dimethylarginine, ATP, cytokines, metabolic remodeling involving tryptophan metabolism and potential microglial inflammation characterize primary open angle glaucoma. Sci. Rep. 2021, 11, 9766. [Google Scholar] [CrossRef]
- Rossi, C.; Cicalini, I.; Cufaro, M.C.; Agnifili, L.; Mastropasqua, L.; Lanuti, P.; Marchisio, M.; De Laurenzi, V.; Del Boccio, P.; Pieragostino, D. Multi-omics approach for studying tears in treatment-naïve glaucoma patients. Int. J. Mol. Sci. 2019, 20, 4029. [Google Scholar] [CrossRef] [Green Version]
- DiCarlo, J.E.; Mahajan, V.B.; Tsang, S.H. Gene therapy and genome surgery in the retina. J. Clin. Invest. 2018, 128, 2177–2188. [Google Scholar] [CrossRef] [Green Version]
- Hamel, C. Retinitis pigmentosa. Orphanet. J. Rare Dis. 2006, 1, 40. [Google Scholar] [CrossRef]
- Micera, A.; Balzamino, B.O.; Di Zazzo, A.; Dinice, L.; Bonini, S.; Coassin, M. Biomarkers of neurodegeneration and precision therapy in retinal disease. Front. Pharmacol. 2021, 11, 2028. [Google Scholar] [CrossRef] [PubMed]
- Donato, L.; Scimone, C.; Alibrandi, S.; Abdalla, E.M.; Nabil, K.M.; D’Angelo, R.; Sidoti, A. New omics—derived perspectives on retinal dystrophies: Could ion channels-encoding or related genes act as modifier of pathological phenotype? Int. J. Mol. Sci. 2021, 22, 70. [Google Scholar] [CrossRef] [PubMed]
- Donato, L.; Abdalla, E.M.; Scimone, C.; Alibrandi, S.; Rinaldi, C.; Nabil, K.M.; D’Angelo, R.; Sidoti, A. Impairments of photoreceptor outer segments renewal and phototransduction due to a peripherin rare haplotype variant: Insights from molecular modeling. Int. J. Mol. Sci. 2021, 22, 3484. [Google Scholar] [CrossRef]
- Arroba, A.I.; Campos-Caro, A.; Aguilar-Diosdado, M.; Valverde, Á.M. IGF-1, Inflammation and retinal degeneration: A close network. Front. Aging Neurosci. 2018, 10, 203. [Google Scholar] [CrossRef]
- Scimone, C.; Donato, L.; Alibrandi, S.; Vadala, M.; Giglia, G.; Sidoti, A.; D’Angelo, R. N-retinylidene-N-retinylethanolamine adduct induces expression of chronic inflammation cytokines in retinal pigment epithelium cells. Exp. Eye Res. 2021, 209, 108641. [Google Scholar] [CrossRef] [PubMed]
- Gallenga, C.E.; Lonardi, M.; Pacetti, S.; Violanti, S.S.; Tassinari, P.; Di Virgilio, F.; Tognon, M.; Perri, P. Molecular mechanisms related to oxidative stress in retinitis pigmentosa. Antioxidants 2021, 10, 848. [Google Scholar] [CrossRef]
- Kutsyr, O.; Maestre-Carballa, L.; Lluesma-Gomez, M.; Martinez-Garcia, M.; Cuenca, N.; Lax, P. Retinitis pigmentosa is associated with shifts in the gut microbiome. Sci. Rep. 2021, 11, 6692. [Google Scholar] [CrossRef]
- Zhang, L.; Justus, S.; Xu, Y.; Pluchenik, T.; Hsu, C.W.; Yang, J.; Duong, J.K.; Lin, C.S.; Jia, Y.; Bassuk, A.G.; et al. Reprogramming towards anabolism impedes degeneration in a preclinical model of retinitis pigmentosa. Hum. Mol. Genet. 2016, 25, 4244–4255. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhang, E.; Ryu, J.; Levi, S.R.; Oh, J.K.; Hsu, C.W.; Cui, X.; Lee, T.T.; Wang, N.K.; Lima de Carvalho, J.R.; Tsang, S.H. PKM2 ablation enhanced retinal function and survival in a preclinical model of retinitis pigmentosa. Mamm. Genome 2020, 31, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Weiss, E.R.; Osawa, S.; Xiong, Y.; Dhungana, S.; Carlson, J.; McRitchie, S.; Fennell, T.R. Broad spectrum metabolomics for detection of abnormal metabolic pathways in a mouse model for retinitis pigmentosa. Exp. Eye Res. 2019, 184, 135–145. [Google Scholar] [CrossRef]
- Park, K.S.; Xu, C.L.; Cui, X.; Tsang, S.H. Reprogramming the metabolome rescues retinal degeneration. Cell Mol. Life Sci. 2018, 75, 1559–1566. [Google Scholar] [CrossRef]
- Wert, K.J.; Velez, G.; Kanchustambham, V.L.; Shankar, V.; Evans, L.P.; Sengillo, J.D.; Zare, R.N.; Bassuk, A.G.; Tsang, S.H.; Mahajan, V.B. Metabolite therapy guided by liquid biopsy proteomics delays retinal neurodegeneration. EBioMedicine 2020, 52, 102636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Osada, H.; Toda, E.; Homma, K.; Guzman, N.A.; Nagai, N.; Ogawa, M.; Negishi, K.; Arita, M.; Tsubota, K.; Ozawa, Y. ADIPOR1 deficiency-induced suppression of retinal ELOVL2 and docosahexaenoic acid levels during photoreceptor degeneration and visual loss. Cell Death Dis. 2021, 12, 458. [Google Scholar] [CrossRef]
- Civan, M.M.; Macknight, A.D.C. The ins and outs of aqueous humour secretion. Exp. Eye Res. 2004, 78, 625–631. [Google Scholar] [CrossRef] [PubMed]
- Agnifili, L.; Carpineto, P.; Fasanella, V.; Mastropasqua, R.; Zappacosta, A.; Di Staso, S.; Costagliola, C.; Mastropasqua, L. Conjunctival findings in hyperbaric and low-tension glaucoma: An in vivo confocal microscopy study. Acta Ophthalmol. 2012, 90, e132–e137. [Google Scholar] [CrossRef]
- Heberle, H.; Meirelles, G.V.; Da Silva, F.R.; Telles, G.P.; Minghim, R. InteractiVenn: A web-based tool for the analysis of sets through Venn diagrams. BMC Bioinform. 2015, 16, 169. [Google Scholar] [CrossRef] [PubMed]
- Lefevere, E.; Toft-Kehler, A.K.; Vohra, R.; Kolko, M.; Moons, L.; Van Hove, I. Mitochondrial dysfunction underlying outer retinal diseases. Mitochondrion 2017, 36, 66–76. [Google Scholar] [CrossRef] [PubMed]
- Yam, M.; Engel, A.L.; Wang, Y.; Zhu, S.; Hauer, A.; Zhang, R.; Lohner, D.; Huang, J.; Dinterman, M.; Zhao, C.; et al. Proline mediates metabolic communication between retinal pigment epithelial cells and the retina. J. Biol. Chem. 2019, 294, 10278–10289. [Google Scholar] [CrossRef]
- Szabados, L.; Savouré, A. Proline: A multifunctional amino acid. Trends Plant Sci. 2010, 15, 89–97. [Google Scholar] [CrossRef]
- Pang, Z.; Chong, J.; Zhou, G.; De Lima Morais, D.A.; Chang, L.; Barrette, M.; Gauthier, C.; Jacques, P.É.; Li, S.; Xia, J. MetaboAnalyst 5.0: Narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 2021, 49, W388–W396. [Google Scholar] [CrossRef] [PubMed]
- Sissler, M.; González-Serrano, L.E.; Westhof, E. Recent advances in mitochondrial aminoacyl-tRNA synthetases and disease. Trends Mol. Med. 2017, 23, 693–708. [Google Scholar] [CrossRef] [Green Version]
- Pareek, V.; Tian, H.; Winograd, N.; Benkovic, S.J. Metabolomics and mass spectrometry imaging reveal channeled de novo purine synthesis in cells. Science 2020, 368, 283. [Google Scholar] [CrossRef] [PubMed]


| Disease | Common Differential Metabolites/Biomarkers |
|---|---|
| AMD-DR | 12-HETE, acetate, Arg, Asp, carnitine, creatine, cytidine, Gln, GPC, His, hypoxanthine, inosine, lactate, Met, nicotinamide, PC, Pro, pyruvate |
| AMD-Glaucoma | acetate, adenine, Ala, Arg, betaine, carnitine, creatine, Gln, Gly, His, hypoxanthine, Ile, Leu, Lys, Met, nicotinamide, palmitoylcarnitine, Phe, Pro, taurine, Tyr, Val |
| DR-Glaucoma | acetate, Arg, carnitine, creatine, Cys, Gln, Glu, His, hypoxanthine, linoleic acid, Met, nicotinamide, Pro, propionylcarnitine, xanthine |
| AMD-DR-Glaucoma | acetate, Arg, carnitine, creatine, Gln, His, hypoxanthine, Met, nicotinamide, Pro |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, X.; Cai, S.; He, Z.; Reilly, J.; Zeng, Z.; Strang, N.; Shu, X. Metabolomics in Retinal Diseases: An Update. Biology 2021, 10, 944. https://doi.org/10.3390/biology10100944
Li X, Cai S, He Z, Reilly J, Zeng Z, Strang N, Shu X. Metabolomics in Retinal Diseases: An Update. Biology. 2021; 10(10):944. https://doi.org/10.3390/biology10100944
Chicago/Turabian StyleLi, Xing, Shichang Cai, Zhiming He, James Reilly, Zhihong Zeng, Niall Strang, and Xinhua Shu. 2021. "Metabolomics in Retinal Diseases: An Update" Biology 10, no. 10: 944. https://doi.org/10.3390/biology10100944
APA StyleLi, X., Cai, S., He, Z., Reilly, J., Zeng, Z., Strang, N., & Shu, X. (2021). Metabolomics in Retinal Diseases: An Update. Biology, 10(10), 944. https://doi.org/10.3390/biology10100944







