Bio-Entities Based on Albumin Nanoparticles and Biomimetic Cell Membranes: Design, Characterization and Biophysical Evaluation
Abstract
1. Introduction
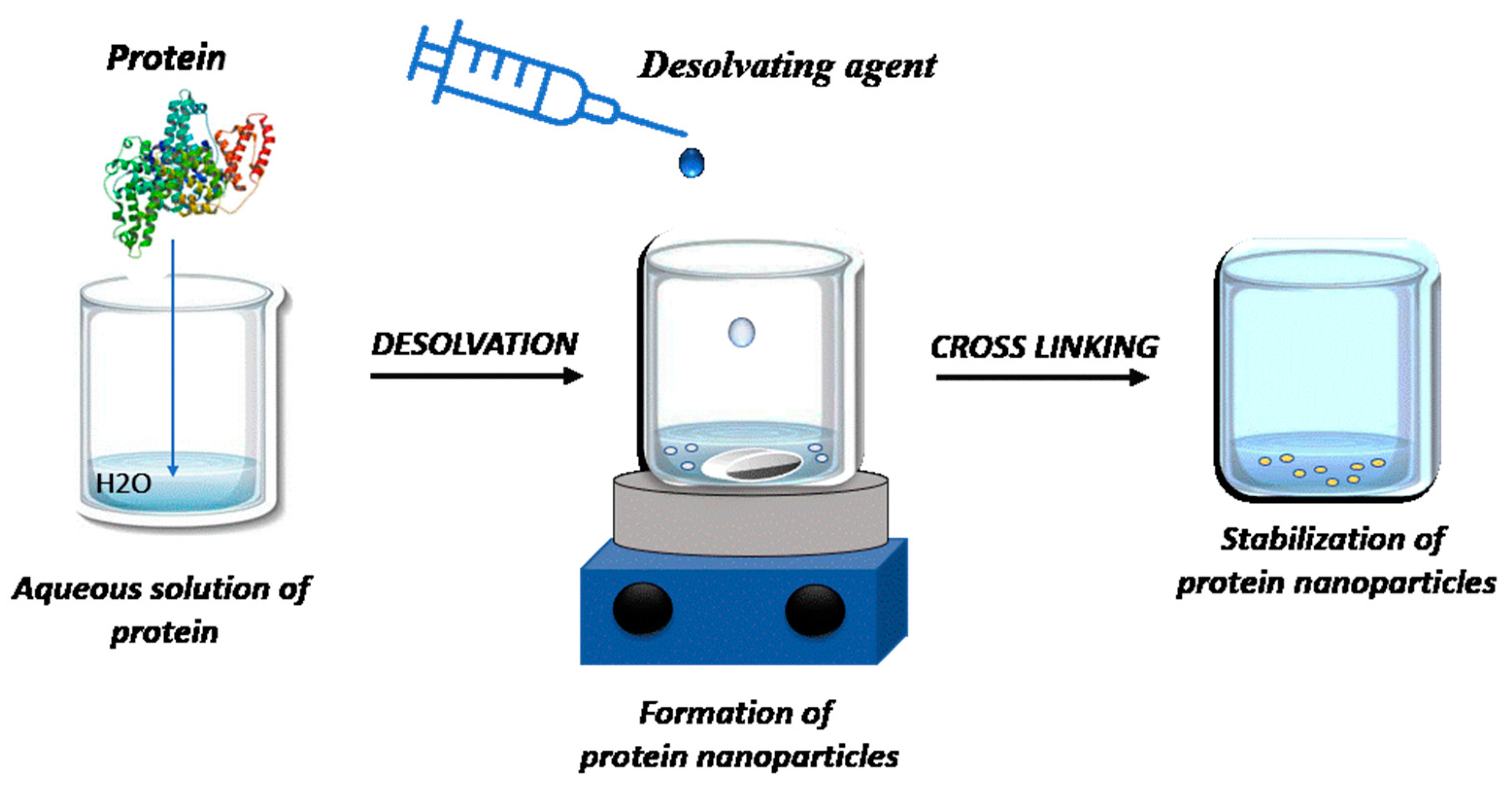
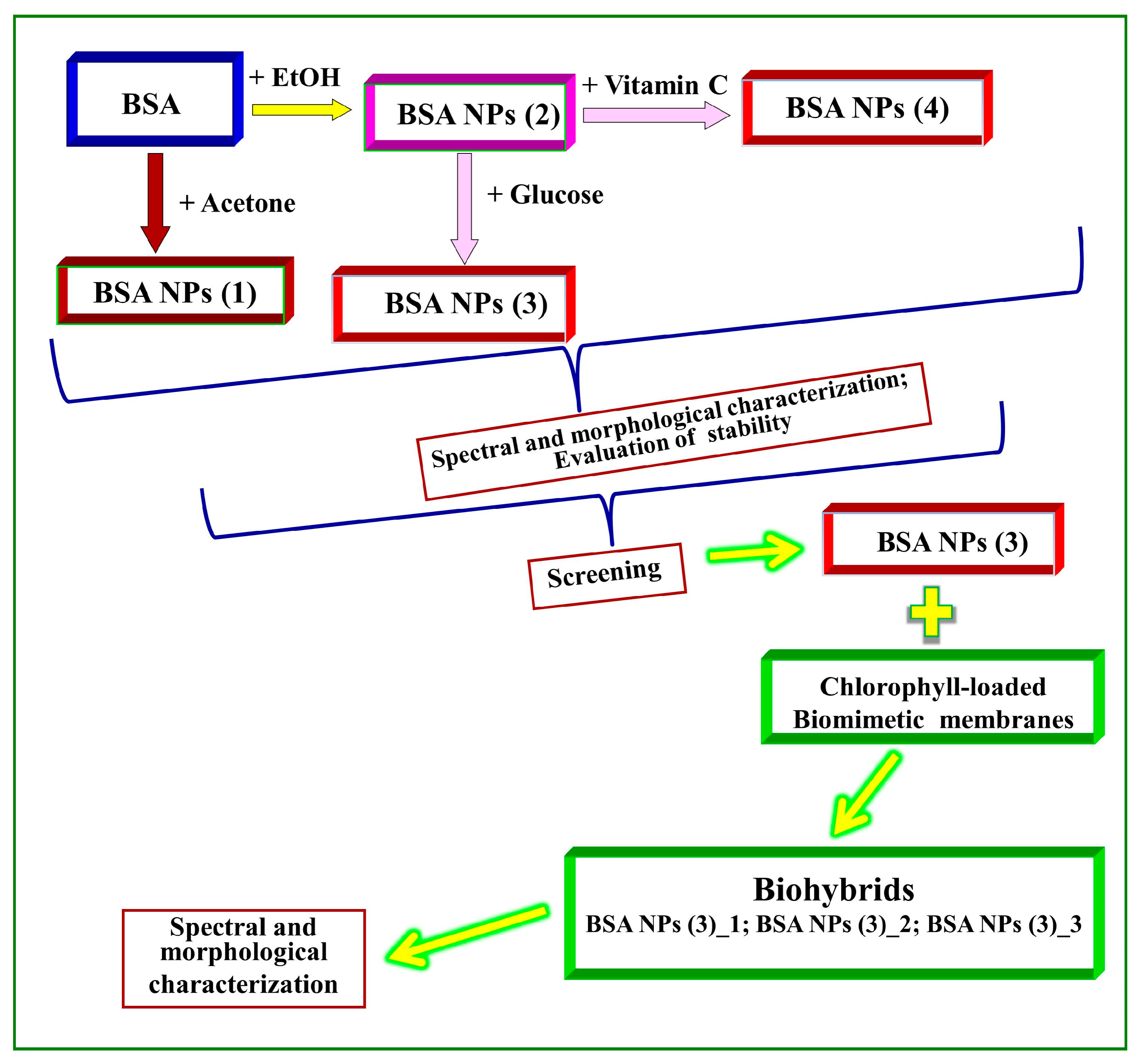
2. Materials and Methods
3. Results and Discussion
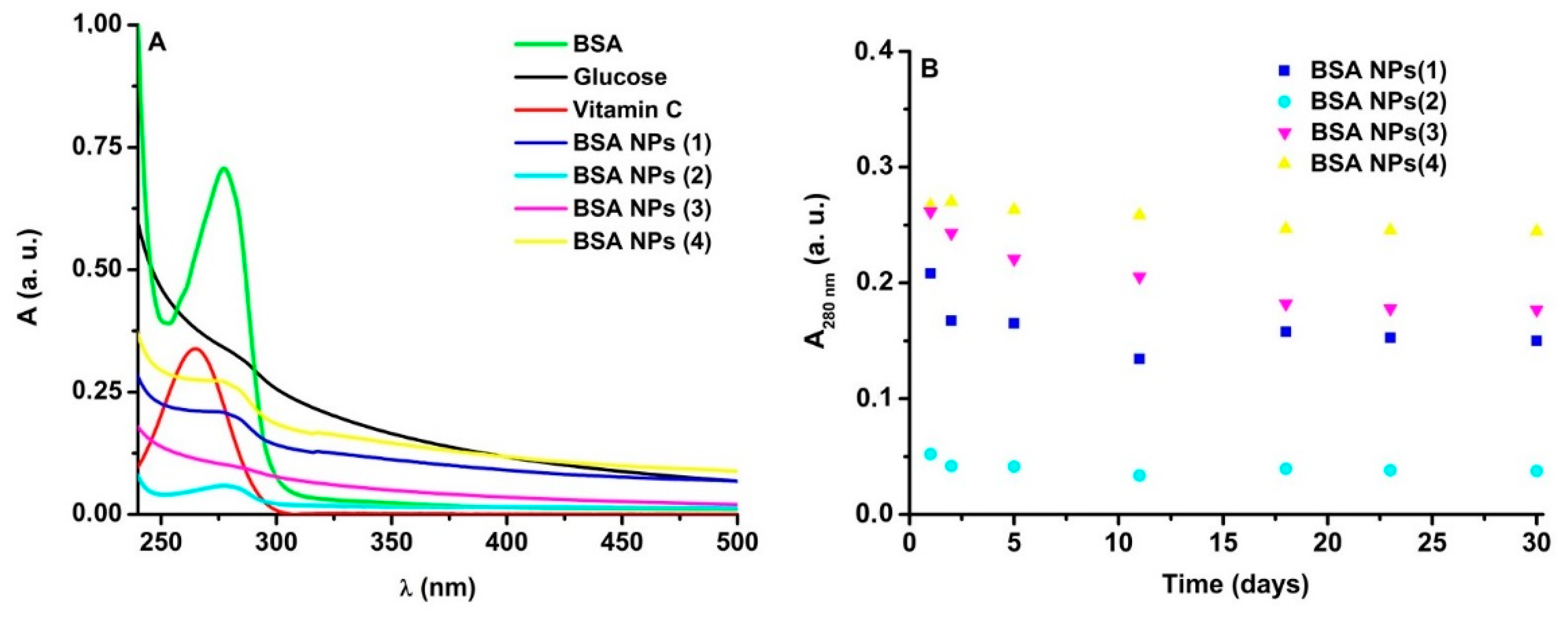
3.1. Characterization of Nanoparticles by UV–Vis Absorption Spectroscopy
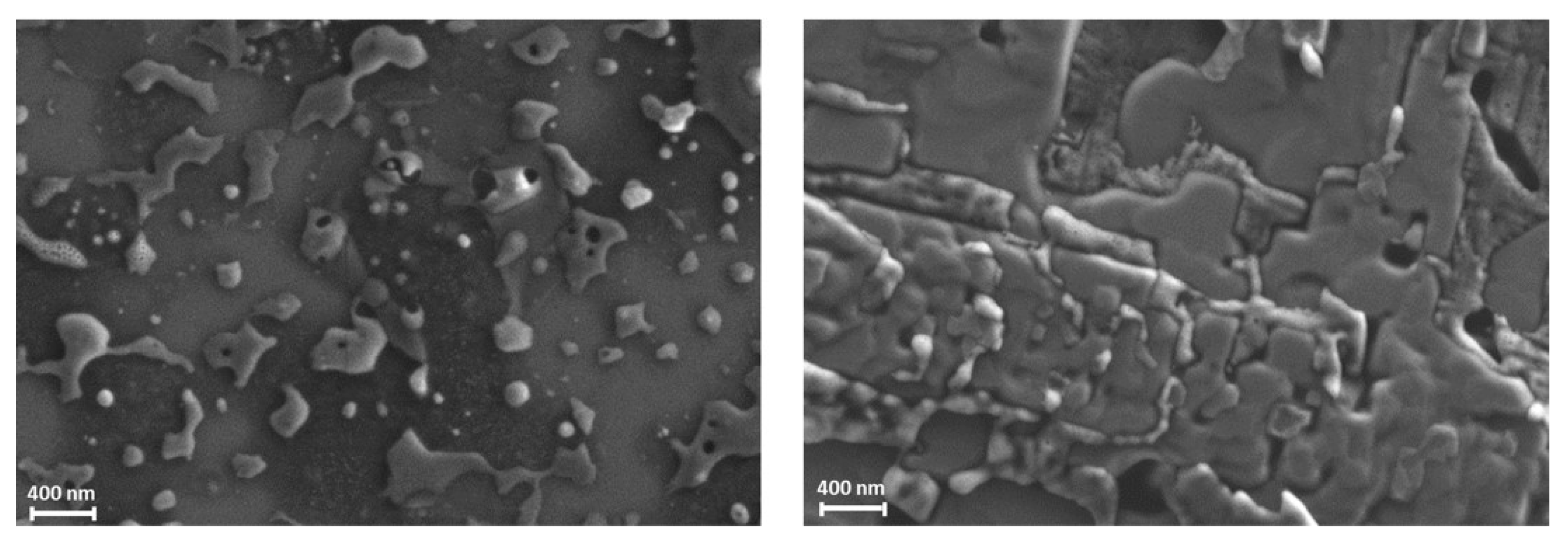
3.2. Characterization of Albumin Nanoparticles by SEM
3.3. Characterization of BSA NPs/Liposomes Bio-Entities
3.3.1. Spectral Characterization of Biohybrids
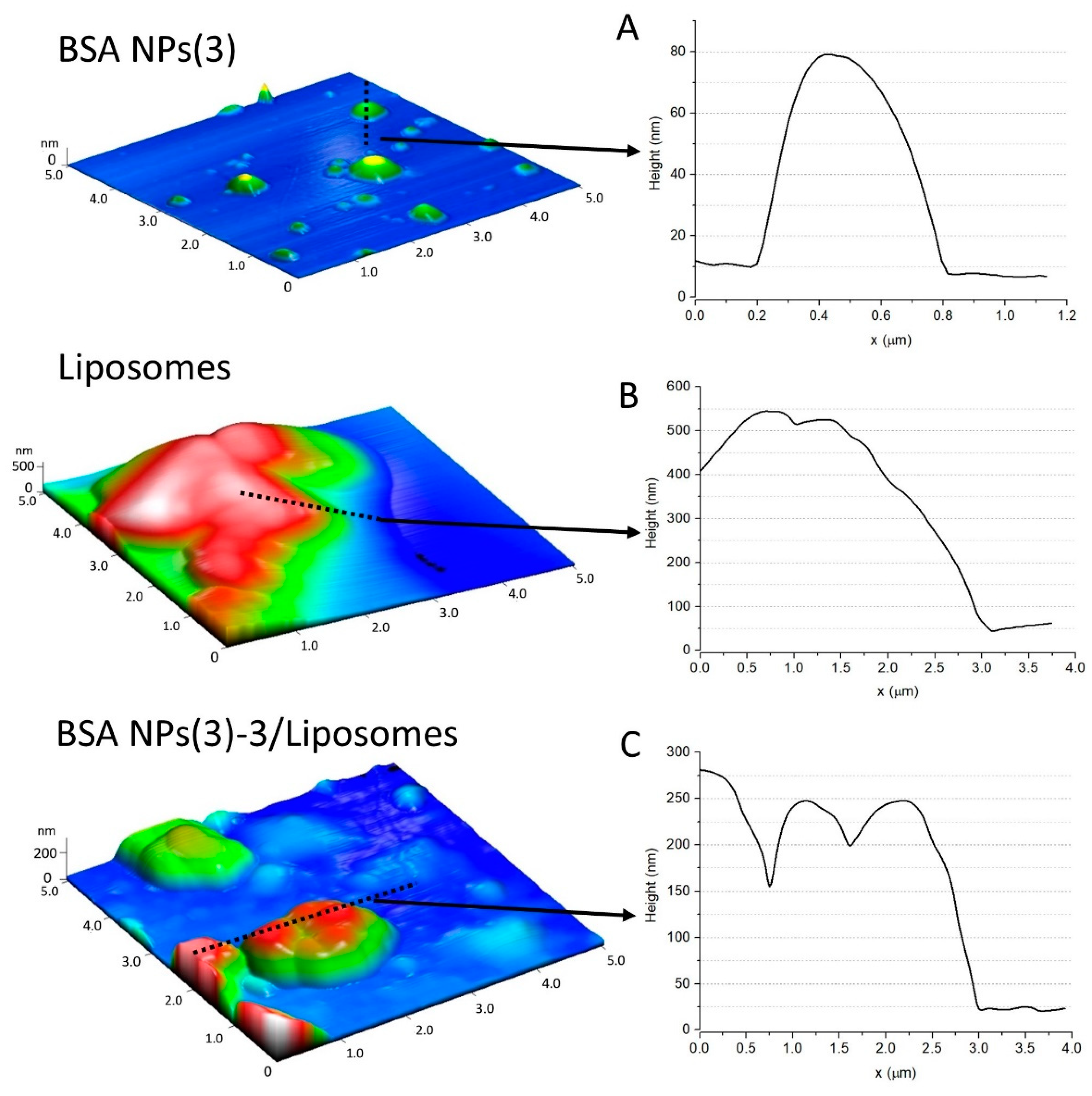
3.3.2. Morphological Characterization of BSA NPs and Their Bio-Entities
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hankins, J. The Role of Albumin in Fluid and Electrolyte Balance. J. Infus. Nurs. 2006, 29, 260–265. [Google Scholar] [CrossRef]
- Francis, G.L. Albumin and mammalian cell culture: Implications for biotechnology applications. Cytotechnology 2010, 62, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Kuntip, N.; Japrung, D.; Pongprayoon, P. How human serum albumin-selective DNA aptamer binds to bovine and canine serum albumins. Biopolymers 2021, 112, e23421. [Google Scholar] [CrossRef] [PubMed]
- Tao, C.; Chuah, Y.J.; Xu, C.; Wang, D.-A. Albumin conjugates and assemblies as versatile bio-functional additives and carriers for biomedical applications. J. Mater. Chem. B 2019, 7, 357–367. [Google Scholar] [CrossRef]
- Chilom, C.; Barangă, G.; Găzdaru, D.; Popescu, A. Characterisation by fluorescence of human and bovine serum albumins in interaction with eosin Y. J. Optoelectron. Adv. Mater. 2013, 15, 311–316. [Google Scholar]
- Chilom, C.G.; Nistorescu, A. A spectroscopic study of the interaction of HSA with tetracaine. IJBB 2016, 53, 206–211. [Google Scholar]
- Sandu, N.; Chilom, C.G.; Popescu, A.I. Spectroscopic Insights on the Binding of Rutin to Bovine Serum Albumin. Rom. J. Phys. 2020, 65, 703. [Google Scholar]
- Neumann, E.; Frei, E.; Funk, D.; Becker, M.D.; Schrenk, H.-H.; Müller-Ladner, U.; Fiehn, C. Native albumin for targeted drug delivery. Expert Opin. Drug Deliv. 2010, 7, 915–925. [Google Scholar] [CrossRef] [PubMed]
- Chilom, C.G.; Bălan, A.; Sandu, N.; Bălăşoiu, M.; Stolyar, S.; Orelovich, O. Exploring the Conformation and Thermal Stability of Human Serum Albumin Corona of Ferrihydrite Nanoparticles. Int. J. Mol. Sci. 2020, 21, 9734. [Google Scholar] [CrossRef] [PubMed]
- Chilom, C.; Barangă, G.; Găzdaru, D.; Popescu, A. A spectroscopic approach of pH effect on thermal denaturation of human and bovine serum albumins. J. Optoelectron. Adv. Mater. 2011, 13, 583–587. [Google Scholar]
- Baler, K.; Martin, O.A.; Carignano, M.A.; Ameer, G.A.; Vila, J.A.; Szleifer, I. Electrostatic Unfolding and Interactions of Albumin Driven by pH Changes: A Molecular Dynamics Study. J. Phys. Chem. B 2014, 118, 921–930. [Google Scholar] [CrossRef]
- Froehlich, E.; Mandeville, J.S.; Jennings, C.J.; Sedaghat-Herati, R.; Tajmir-Riahi, H.A. Dendrimers Bind Human Serum Albumin. J. Phys. Chem. B 2009, 113, 6986–6993. [Google Scholar] [CrossRef]
- Taguchi, K.; Okamoto, Y.; Matsumoto, K.; Otagiri, M.; Chuang, V. When Albumin Meets Liposomes: A Feasible Drug Carrier for Biomedical Applications. Pharmaceuticals 2021, 14, 296. [Google Scholar] [CrossRef]
- Spada, A.; Emami, J.; Tuszynski, J.A.; Lavasanifar, A. The Uniqueness of Albumin as a Carrier in Nanodrug Delivery. Mol. Pharm. 2021, 18, 1862–1894. [Google Scholar] [CrossRef]
- Lu, N.; Sui, Y.; Ding, Y.; Tian, R.; Li, L.; Liu, F. Adsorption of human serum albumin on functionalized single-walled carbon nanotubes reduced cytotoxicity. Chem. Interact. 2018, 295, 64–72. [Google Scholar] [CrossRef] [PubMed]
- Wojnarowska-Nowak, R.; Polit, J.; Zięba, A.; Stolyarchuk, I.; Nowak, S.; Romerowicz-Misielak, M.; Sheregii, E. Colloidal quantum dots conjugated with human serum albumin-interactions and bioimaging properties. Opto-Electron. Rev. 2017, 25, 137–147. [Google Scholar] [CrossRef]
- Karimi, M.; Bahrami, S.; Ravari, S.B.; Zangabad, P.S.; Mirshekari, H.; Bozorgomid, M.; Shahreza, S.; Sori, M.; Hamblin, M.R. Albumin nanostructures as advanced drug delivery systems. Expert Opin. Drug Deliv. 2016, 13, 1609–1623. [Google Scholar] [CrossRef]
- Chilom, C.G.; Zorilă, B.; Bacalum, M.; Bălăşoiu, M.; Yaroslavtsev, R.; Stolyar, S.V.; Tyutyunnicov, S. Ferrihydrite nanopar-ticles interaction with model lipid membranes. Chem. Phys. Lipids 2020, 226, 104851. [Google Scholar] [CrossRef]
- Wu, J. The Enhanced Permeability and Retention (EPR) Effect: The Significance of the Concept and Methods to Enhance Its Application. J. Pers. Med. 2021, 11, 771. [Google Scholar] [CrossRef] [PubMed]
- Chubarov, A.S. Serum Albumin for Magnetic Nanoparticles Coating. Magnetochemistry 2022, 8, 13. [Google Scholar] [CrossRef]
- Niknejad, H.; Mahmoudzadeh, R. Comparison of Different Crosslinking Methods for Preparation of Docetaxel-loaded Albumin Nanoparticles. Iran. J. Pharm. Res. 2015, 14, 385–394. [Google Scholar]
- Hong, S.; Choi, D.W.; Kim, H.N.; Park, C.G.; Lee, W.; Park, H.H. Protein-Based Nanoparticles as Drug Delivery Systems. Pharmaceutics 2020, 12, 604. [Google Scholar] [CrossRef] [PubMed]
- Bloch, K.; Pardesi, K.; Satriano, C.; Ghosh, S. Bacteriogenic Platinum Nanoparticles for Applicationin Nanomedicine. Front. Chem. 2021, 9, 624344. [Google Scholar] [CrossRef]
- Hornok, V. Serum Albumin Nanoparticles: Problems and Prospects. Polymers 2021, 13, 3759. [Google Scholar] [CrossRef] [PubMed]
- Jahanban-Esfahlan, A.; Dastmalchi, S.; Davaran, S. A simple improved desolvation method for the rapid preparation of albumin nanoparticles. Int. J. Biol. Macromol. 2016, 91, 703–709. [Google Scholar] [CrossRef]
- Chu, F.; Thornton, D.T.; Nguyen, H. Chemical cross-linking in the structural analysis of protein assemblies. Methods 2018, 144, 53–63. [Google Scholar] [CrossRef]
- Luo, R.; Lin, M.; Zhang, C.; Shi, J.; Zhang, S.; Chen, Q.; Hu, Y.; Zhang, M.; Zhang, J.; Gao, F. Genipin-crosslinked human serum albumin coating using a tannic acid layer for enhanced oral administration of curcumin in the treatment of ulcerative colitis. Food Chem. 2020, 330, 127241. [Google Scholar] [CrossRef] [PubMed]
- Kistriyani, L.; Sulistyawan, W.; Kesuma, S.H. Characteristic of ascorbic acid in crosslinked chitosan edible film as drug delivery system membrane. MATEC Web Conf. 2018, 154, 01027. [Google Scholar] [CrossRef]
- Salihu, R.; Razak, S.I.A.; Zawawi, N.A.; Kadir, M.R.A.; Ismail, N.I.; Jusoh, N.; Mohamad, M.R.; Nayan, N.H.M. Citric acid: A green cross-linker of biomaterials for biomedical applications. Eur. Polym. J. 2021, 146, 110271. [Google Scholar] [CrossRef]
- Herrera, M.A.; Mathew, A.P.; Oksman, K. Barrier and mechanical properties of plasticized and cross-linked nanocellulose coatings for paper packaging applications. Cellulose 2017, 24, 3969–3980. [Google Scholar] [CrossRef]
- Amighi, F.; Emam-Djomeh, Z.; Labbafi-Mazraeh-Shahi, M. Effect of different cross-linking agents on the preparation of bovine serum albumin nanoparticles. J. Iran. Chem. Soc. 2020, 17, 1223–1235. [Google Scholar] [CrossRef]
- Esim, O.; Gedik, M.E.; Dogan, A.L.; Gunaydin, G.; Hascicek, C. Development of carboplatin loaded bovine serum albumin nanoparticles and evaluation of its effect on an ovarian cancer cell line. J. Drug Deliv. Sci. Technol. 2021, 64, 102655. [Google Scholar] [CrossRef]
- An, F.-F.; Zhang, X.-H. Strategies for Preparing Albumin-based Nanoparticles for Multifunctional Bioimaging and Drug Delivery. Theranostics 2017, 7, 3667–3689. [Google Scholar] [CrossRef]
- Cho, H.; Jeon, S.I.; Ahn, C.-H.; Shim, M.K.; Kim, K. Emerging Albumin-Binding Anticancer Drugs for Tumor-Targeted Drug Delivery: Current Understandings and Clinical Translation. Pharmaceutics 2022, 14, 728. [Google Scholar] [CrossRef] [PubMed]
- Shi, J.-H.; Zhou, K.-L.; Lou, Y.-Y.; Pan, D.-Q. Multi-spectroscopic and molecular modeling approaches to elucidate the binding interaction between bovine serum albumin and darunavir, a HIV protease inhibitor. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2018, 188, 362–371. [Google Scholar] [CrossRef]
- Sandu, N.; Chilom, C.G.; Florescu, M. Molecular insights into binding mechanism of rutin to bovine serum albumin—Levothyroxine complex: Spectroscopic and molecular docking approaches. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2021, 264, 120261. [Google Scholar] [CrossRef] [PubMed]
- Pawar, S.K.; Jaldappagari, S. Interaction of repaglinide with bovine serum albumin: Spectroscopic and molecular docking approaches. J. Pharm. Anal. 2019, 9, 274–283. [Google Scholar] [CrossRef]
- Merlot, A.M.; Kalinowski, D.S.; Richardson, D.R. Unraveling the mysteries of serum albumin-More than just a serum protein. Front. Physiol. 2014, 5, 299. [Google Scholar] [CrossRef]
- Meng, R.; Zhu, H.; Wang, Z.; Hao, S.; Wang, B. Preparation of Drug-Loaded Albumin Nanoparticles and Its Application in Cancer Therapy. J. Nanomater. 2022, 2022, 1–12. [Google Scholar] [CrossRef]
- Desai, N. Nanoparticle Albumin-Bound Anticancer Agents. In Non-Biological Complex Drugs; Crommelin, D.J., de Vlieger, J.S.B., Eds.; Springer International Publishing: Cham, Switzerland, 2015; Volume 20, pp. 335–354. [Google Scholar]
- Tacheva, B.; Paarvanova, B.; Ivanov, I.T.; Tenchov, B.; Georgieva, R.; Karabaliev, M. Drug Exchange between Albumin Na-noparticles and Erythrocyte Membranes. Nanomaterials 2018, 9, 47. [Google Scholar] [CrossRef]
- Teixeira, S.; Carvalho, M.A.; Castanheira, E.M.S. Functionalized Liposome and Albumin-Based Systems as Carriers for Poorly Water-Soluble Anticancer Drugs: An Updated Review. Biomedicines 2022, 10, 486. [Google Scholar] [CrossRef]
- Peng, Q.; Wei, X.-Q.; Yang, Q.; Zhang, S.; Zhang, T.; Shao, X.-R.; Cai, X.-X.; Zhang, Z.-R.; Lin, Y.-F. Enhanced biostability of nanoparticle-based drug delivery systems by albumin corona. Nanomedicine 2015, 10, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.-Q.; Ba, K. Construction a Long-Circulating Delivery System of Liposomal Curcumin by Coating Albumin. ACS Omega 2020, 5, 16502–16509. [Google Scholar] [CrossRef] [PubMed]
- Pasarin, D.; Ghizdareanu, A.-I.; Enascuta, C.E.; Matei, C.B.; Bilbie, C.; Paraschiv-Palada, L.; Veres, P.-A. Coating Materials to Increase the Stability of Liposomes. Polymers 2023, 15, 782. [Google Scholar] [CrossRef] [PubMed]
- Akbarzadeh, A.; Rezaei-Sadabady, R.; Davaran, S.; Joo, S.W.; Zarghami, N.; Hanifehpour, Y.; Samiei, M.; Mohammad Kouhi, M.; Nejati-Koshki, K. Liposome: Classification, preparation, and applications. Nanoscale Res. Lett. 2013, 8, 102. [Google Scholar] [CrossRef]
- Barbinta-Patrascu, M.-E.; Gorshkova, Y.; Ungureanu, C.; Badea, N.; Bokuchava, G.; Lazea-Stoyanova, A.; Bacalum, M.; Zhi-gunov, A.; Petrovič, S.M. Characterization and Antitumoral Activity of Biohybrids Based on Turmeric and Silver/Silver Chloride Nanoparticles. Materials 2021, 14, 4726. [Google Scholar] [CrossRef]
- Gorshkova, Y.; Barbinta-Patrascu, M.-E.; Bokuchava, G.; Badea, N.; Ungureanu, C.; Lazea-Stoyanova, A.; Răileanu, M.; Bacalum, M.; Turchenko, V.; Zhigunov, A.; et al. Biological Performances of Plasmonic Biohybrids Based on Phy-to-Silver/Silver Chloride Nanoparticles. Nanomaterials 2021, 11, 1811. [Google Scholar] [CrossRef]
- Strain, H.H.; Svec, W.A.; Vernon, L.P.; Seely, G.R. The Chlorophylls; Academic Press: New York, NY, USA, 1966; Volume 2, pp. 21–66. [Google Scholar]
- Santos-Rebelo, A.; Kumar, P.; Pillay, V.; Choonara, Y.E.; Eleutério, C.; Figueira, M.; Viana, A.S.; Ascensão, L.; Molpeceres, J.; Rijo, P.; et al. Development and mechanistic insight into the enhanced cytotoxic potential of Parvifloron D albumin nanoparticles in EGFR-overexpressing pancreatic cancer cells. Cancers 2019, 11, 1733. [Google Scholar] [CrossRef]
- Chen, F.; Huang, G.; Huang, H. Sugar ligand-mediated drug delivery. Futur. Med. Chem. 2020, 12, 161–171. [Google Scholar] [CrossRef]
- Barros, J.; Grenha, A.; Carvalho, R. Vitamin C and glucose cross-linked chitosan as a versatile platform for drug delivery. Int. J. Biol. Macromol. 2019, 137, 1117–1125. [Google Scholar]
- Stefanescu, T.; Manole, C.; Parvu, C.; Barbinta-Patrascu, M.E.; Tugulea, L. Supported phospholipid bilayers with chlorophyll for optoelectronic devices. Optoelectron. Adv. Mater. Rapid Commun. 2010, 4, 33–38. [Google Scholar]
- Barbinta-Patrascu, M.E.; Tugulea, L.; Meghea, A. Procaine effects on model membranes with chlorophyll a. Rev. De Chim. 2009, 60, 337–341. [Google Scholar]
- Lichtenthaler, H.K.; Wenzel, O.; Buschmann, C.; Gitelson, A. Plant Stress Detection by Reflectance and Fluorescencea. Ann. N. Y. Acad. Sci. 1998, 851, 271–285. [Google Scholar] [CrossRef]
- Bronze-Uhle, E.S.; Costa, B.C.; Ximenes, V.F.; Lisboa-Filho, P.N. Synthetic nanoparticles of bovine serum albumin with en-trapped salicylic acid. Nanotechnol. Sci. Appl. 2017, 10, 11–21. [Google Scholar] [CrossRef] [PubMed]
- Çağdaş, M.; Sezer, A.D.; Bucak, S. Liposomes as Potential Drug Carrier Systems for Drug Delivery, Application of Nanotechnology in Drug Delivery; Sezer, A.D., Ed.; IntechOpen: London, UK, 2014. [Google Scholar]
- Vavilin, D.V.; Ermakova-Gerdes, S.Y.; Keilty, A.T.; Vermaas, W.F.J. Tryptophan at Position 181 of the D2 Protein of Photo-system II Confers Quenching of Variable Fluorescence of Chlorophyll: Implications for the Mechanism of Energy-Dependent Quenching. Biochemistry 1999, 38, 14690–14696. [Google Scholar] [CrossRef] [PubMed]
- Chaves, O.A.; Amorim, A.P.D.O.; Castro, L.H.E.; Sant’Anna, C.M.R.; De Oliveira, M.C.C.; Cesarin-Sobrinho, D.; Netto-Ferreira, J.C.; Ferreira, A.B.B. Fluorescence and Docking Studies of the Interaction between Human Serum Albumin and Pheophytin. Molecules 2015, 20, 19526–19539. [Google Scholar] [CrossRef]
- Gabizon, A.; Shmeeda, H. Bypassing the RES: Physicochemical barriers to nanocarrier clearance. Adv. Drug Deliv. Rev. 2003, 55, 1171–1190. [Google Scholar]
- Zhou, C.; Zhang, Y.; Li, J.; Liang, Q.; Huang, Z.; Yu, Y. Enhanced therapeutic efficacy of a combination of doxorubicin and bovine serum albumin-conjugated nanoparticles for hepatocellular carcinoma. Drug Deliv. 2019, 26, 259–267. [Google Scholar]
- Zhong, H.; Tarrach, G.; Wu, P.; Drechsler, A.; Wei, D.; Yuan, J. High resolution magnetic force microscopy of patterned L10-FePt dot arrays by nanosphere lithography. Nanotechnology 2008, 19, 095703. [Google Scholar] [CrossRef]
- Rohiwal, S.; Tiwari, A.; Verma, G.; Pawar, S. Preparation and evaluation of bovine serum albumin nanoparticles for ex vivo colloidal stability in biological media. Colloids Surf. A Physicochem. Eng. Asp. 2015, 480, 28–37. [Google Scholar] [CrossRef]
- Leitgeb, M.; Knez, Ž.; Primožič, M. Sustainable technologies for liposome preparation. J. Supercrit. Fluids 2020, 165, 104984. [Google Scholar]
- Zhang, L.; Gu, F.; Chan, J.; Wang, A.; Langer, R.S.; Farokhzad, O.C. Nanoparticles in Medicine: Therapeutic Applications and Developments. Clin. Pharmacol. Ther. 2007, 83, 761–769. [Google Scholar] [CrossRef] [PubMed]
- Peer, D.; Karp, J.M.; Hong, S.; Farokhzad, O.C.; Margalit, R.; Langer, R. Nanocarriers as an emerging platform for cancer therapy. Nat. Nanotechnol. 2007, 2, 751–760. [Google Scholar] [CrossRef] [PubMed]
- Patil, Y.; Sadhukha, T.; Ma, L.; Panyam, J. Nanoparticle-mediated simultaneous and targeted delivery of paclitaxel and tariquidar overcomes tumor drug resistance. J. Control. Release 2009, 136, 21–29. [Google Scholar] [CrossRef] [PubMed]




Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barbinta-Patrascu, M.-E.; Iftimie, S.; Cazacu, N.; Stan, D.L.; Costas, A.; Balan, A.E.; Chilom, C.G. Bio-Entities Based on Albumin Nanoparticles and Biomimetic Cell Membranes: Design, Characterization and Biophysical Evaluation. Coatings 2023, 13, 671. https://doi.org/10.3390/coatings13040671
Barbinta-Patrascu M-E, Iftimie S, Cazacu N, Stan DL, Costas A, Balan AE, Chilom CG. Bio-Entities Based on Albumin Nanoparticles and Biomimetic Cell Membranes: Design, Characterization and Biophysical Evaluation. Coatings. 2023; 13(4):671. https://doi.org/10.3390/coatings13040671
Chicago/Turabian StyleBarbinta-Patrascu, Marcela-Elisabeta, Sorina Iftimie, Nicoleta Cazacu, Diana Lavinia Stan, Andreea Costas, Adriana Elena Balan, and Claudia Gabriela Chilom. 2023. "Bio-Entities Based on Albumin Nanoparticles and Biomimetic Cell Membranes: Design, Characterization and Biophysical Evaluation" Coatings 13, no. 4: 671. https://doi.org/10.3390/coatings13040671
APA StyleBarbinta-Patrascu, M.-E., Iftimie, S., Cazacu, N., Stan, D. L., Costas, A., Balan, A. E., & Chilom, C. G. (2023). Bio-Entities Based on Albumin Nanoparticles and Biomimetic Cell Membranes: Design, Characterization and Biophysical Evaluation. Coatings, 13(4), 671. https://doi.org/10.3390/coatings13040671








