Developing Diagnostic and Therapeutic Approaches to Bacterial Infections for a New Era: Implications of Globalization
Abstract
1. Introduction
2. State-of-the-Art of Diagnostic Tests for Bacterial Pathogens
2.1. Pathogen Culture
2.2. Antibody-Based Tests
2.3. Molecular Techniques
2.4. High-Throughput Sequencing (HTS)
2.5. Biosensors
2.6. Lateral Flow Chromatography
2.7. Microfluidic Chips
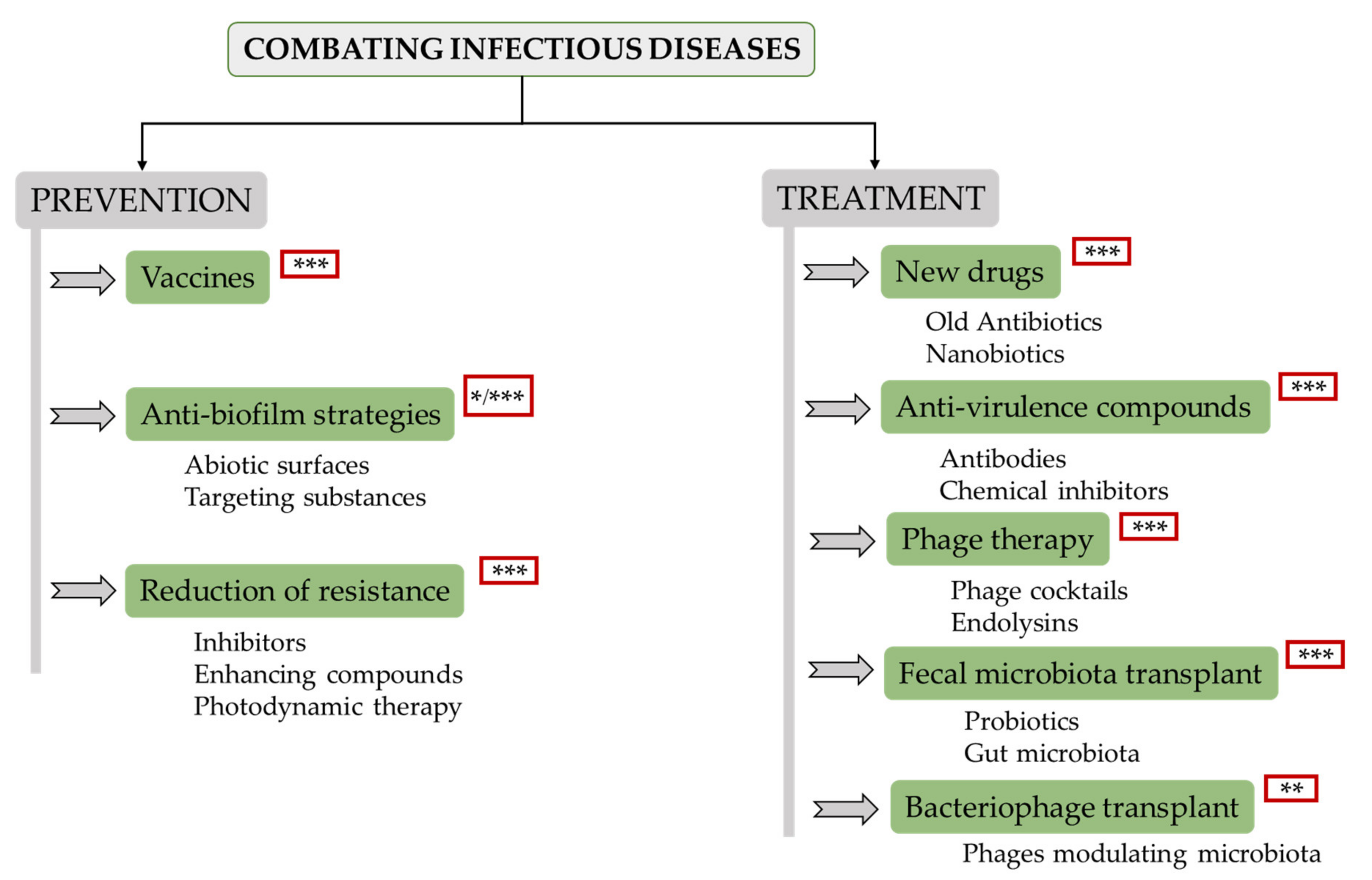
3. State-of-the-Art of New Therapies for Fighting Infectious Bacterial Diseases
3.1. Vaccines
3.2. Antibiofilm Strategies
3.3. Reduction of Antibiotic Resistance
3.4. New Drugs
3.5. Antivirulence Treatments
3.6. Phage Therapy
3.7. Faecal Microbiota Transplant
3.8. Bacteriophage Transplant
4. Future Challenges
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Saker, L.; Lee, K.; Cannito, B.; Gilmore, A.; Campbell-Lendrum, D.; Phil, D. Globalization and infectious diseases: A review of the linkages. World Health Organization 2004. Available online: https://apps.who.int/iris/handle/10665/68726 (accessed on 15 January 2020).
- WHO. List of Blueprint Priority Diseases. 2018 Annual Review. 2019. Available online: https://www.who.int/blueprint/priority-diseases/en/ (accessed on 16 August 2019).
- Centers for Disease Control and Prevention. Antibiotic Resistant Threats in the United States 2019. Available online: https://www.cdc.gov/drugresistance/pdf/threats-report/2019-ar-threats-report-508.pdf (accessed on 15 January 2020).
- Cassini, A.; Högberg, L.D.; Plachouras, D.; Quattrocchi, A.; Hoxha, A.; Simonsen, G.S.; Colomb-Cotinat, M.; Kretzschmar, M.E.; Devleesschauwer, B.; Cecchini, M.; et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Lancet Infect. Dis. 2019, 19, 56–66. [Google Scholar] [CrossRef]
- Ayukekbong, J.A.; Ntemgwa, M.; Atabe, A.N. The threat of antimicrobial resistance in developing countries: Causes and control strategies. Antimicrob. Resist. Infect. Cont. 2017, 6, 47. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.S.; Liu, C.Y.; Qiu, X.Y.; Xie, S.S.; Li, W.Y.; Zhu, L.; Zhu, L.Y. Novel nucleic acid detection strategies based on CRISPR-Cas systems: From construction to application. Biotechn. Bioeng. 2020, 117, 2279–2294. [Google Scholar] [CrossRef] [PubMed]
- Lai, C.C.; Shih, T.P.; Ko, W.C.; Tang, H.J.; Hsueh, P.R. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and coronavirus disease-2019 (COVID-19): The epidemic and the challenges. Int. J. Antimicrob. Agents 2020, 55, 105924. [Google Scholar] [CrossRef] [PubMed]
- CenterWatch. FDA Approved Drugs. 2020. Available online: https://www.centerwatch.com/directories/1067-fda-approved-drugs/topic/546-bacterial-infections (accessed on 10 January 2020).
- Funke, G.; Funke-Kissling, P. Evaluation of the new VITEK 2 card for identification of clinically relevant gram-negative rods. J. Clin. Microbiol. 2004, 42, 4067–4071. [Google Scholar] [CrossRef]
- Martinez, R.M.; Wolk, D.M. Bloodstream infections. Microbiol. Spectr. 2016, 4, 4. [Google Scholar] [CrossRef]
- Chowdhury, I.H.; Sen, A.; Bahar, B.; Hazra, A.; Chakraborty, U.; Choudhuri, S.; Goswami, A.; Pal, N.K.; Bhattacharya, B. A molecular approach to identification and profiling of first-line-drug-resistant mycobacteria from sputum of pulmonary tuberculosis patients. J. Clin. Microbiol. 2012, 50, 2082–2084. [Google Scholar] [CrossRef][Green Version]
- Reja, A.H.; Biswas, N.; Biswas, S.; Dasgupta, S.; Chowdhury, I.H.; Banerjee, S.; Chakraborty, T.; Dutta, P.K.; Bhattacharya, B. Fite-Faraco staining in combination with multiplex polymerase chain reaction: A new approach to leprosy diagnosis. Indian J. Derm. Venereol. Leprol. 2013, 79, 693–700. [Google Scholar] [CrossRef]
- Iha, K.; Inada, M.; Kawada, N.; Nakaishi, K.; Watabe, S.; Tan, Y.H.; Shen, C.; Ke, L.-Y.; Yoshimura, T.; Ito, E. Ultrasensitive ELISA developed for diagnosis. Diagnostics 2019, 9, 78. [Google Scholar] [CrossRef]
- Sakashita, K.; Takeuchi, R.; Takeda, K.; Takamori, M.; Ito, K.; Igarashi, Y.; Hayashi, E.; Iguchi, M.; Ono, M.; Kashiyama, T.; et al. Ultrasensitive enzyme-linked immunosorbent assay for the detection of MPT64 secretory antigen to evaluate Mycobacterium tuberculosis viability in sputum. Int. J. Infect. Dis. 2020, 96, 244–253. [Google Scholar] [CrossRef]
- Salimnia, H.; Fairfax, M.R.; Lephart, P.R.; Schreckenberger, P.; DesJarlais, S.M.; Johnson, J.K.; Robinson, G.; Carroll, K.C.; Greer, A.; Morgan, M.; et al. Evaluation of the FilmArray blood culture identification panel: Results of a multicenter controlled trial. J. Clin. Microbiol. 2016, 54, 687–698. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Bai, R.; Zhao, Z.; Tao, L.; Ma, M.; Ji, Z.; Jian, M.; Ding, Z.; Dai, X.; Bao, F.; et al. Application of droplet digital PCR to detect the pathogens of infectious diseases. Biosci Rep. 2018, 38, BSR20181170. [Google Scholar] [CrossRef] [PubMed]
- Ecker, D.J.; Massire, C.; Blyn, L.B.; Hofstadler, S.A.; Hannis, J.C.; Eshoo, M.W.; Hall, T.A.; Sampath, R. Molecular genotyping of microbes by multilocus PCR and mass spectrometry: A new tool for hospital infection control and public health surveillance. Methods Mol. Biol. 2009, 13, e0197436. [Google Scholar] [CrossRef]
- Wilson, M.R.; O’Donovan, B.D.; Gelfand, J.M.; Sample, H.A.; Chow, F.C.; Betjemann, J.P.; Shah, M.P.; Richie, M.B.; Gorman, M.P.; Hajj-Ali, R.A.; et al. Chronic meningitis investigated via metagenomic next-generation sequencing. JAMA Neurol. 2018, 75, 947–955. [Google Scholar] [CrossRef]
- Zhang, X.; Wu, Z.; Wang, K. Diagnosis of Streptococcus suis meningoencephalitis with metagenomic next-generation sequencing of the cerebrospinal fluid: A case report with literature review. BMC Infect. Dis. 2020, 20, 884. [Google Scholar] [CrossRef]
- Petersen, L.M.; Martin, I.W.; Moschetti, W.E.; Kershaw, C.M.; Tsongalis, G.J. Third generation sequencing in the clinical laboratory: Exploring the advantages and challenges of nanopore sequencing. J. Clin. Microbiol. 2020, 58, e01315–e01319. [Google Scholar] [CrossRef]
- Vincent, M.; Xu, Y.; Kong, H. Helicase-dependent isothermal DNA amplification. EMBO Rep. 2004, 5, 795–800. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic. Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef]
- Guatelli, J.C.; Whitfield, K.M.; Kwoh, D.Y.; Barringer, K.J.; Richman, D.D.; Gingeras, T.R. Isothermal, in vitro amplification of nucleic acids by a multienzyme reaction modeled after retroviral replication. Proc. Natl. Acad. Sci. USA 1990, 87, 1874–1878. [Google Scholar] [CrossRef]
- Lizardi, P.M.; Huang, X.; Zhu, Z.; Bray-Ward, P.; Thomas, D.C.; Ward, D.C. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 1998, 19, 225–232. [Google Scholar] [CrossRef]
- Kozel, T.R.; Burnham-Marusich, A.R. Point-of-care testing for infectious diseases: Past, present, and future. J. Clin. Microbiol. 2017, 55, 2313–2320. [Google Scholar] [CrossRef] [PubMed]
- Blair, E.O.; Corrigan, D.K. A review of microfabricated electrochemical biosensors for DNA detection. Biosens. Bioelectron. 2019, 1, 57–67. [Google Scholar] [CrossRef] [PubMed]
- Usachev, E.V.; Usacheva, O.V.; Aganovski, I.E. Surface plasmon resonance-based bacterial aerosol detection. J. App. Microbiol. 2014, 17, 1655–1662. [Google Scholar] [CrossRef] [PubMed]
- Yeh, Y.; Zhang, Y.; Sabestien, A.; Chou, T.; Zhou, B.; Zhong, L.; Albert, I.; Lu, H.; Swaminathan, V.; Ghedin, E.; et al. A rapid and label-free platform for virus capture and identification from clinical samples. Proc. Natl. Acd. Sci. USA 2020, 117, 895–901. [Google Scholar] [CrossRef]
- Rafael, M.E.; Taylor, T.; Magill, A.; Lim, Y.W.; Girosi, F.; Allan, R. Reducing the burden of childhood malaria in Africa: The role of improved. Nature 2006, 1, 39–48. [Google Scholar] [CrossRef]
- Nasseri, B.; Soleimani, N.; Rabiee, N.; Kalbasi, A.; Karimi, M.; Hambling, M.R. Point-of-care microfluidic devices for pathogen detection. Biosen. Bioelectron. 2018, 117, 112–128. [Google Scholar] [CrossRef]
- Rodriguez-Manzano, J.; Moser, N.; Malpartida-Cardenas, K.; Moniri, A.; Fisarova, L.; Pennisi, I.; Boonyasiri, A.; Jauneikaite, E.; Abdolrasouli, A.; Otter, J.A.; et al. Rapid detection of mobilized colistin resistance using a nucleic acid based Lab-on-a-Chip diagnostic system. Sci. Rep. 2020, 10, 8448. [Google Scholar] [CrossRef]
- WHO. 2017. Available online: https://www.who.int/news-room/detail/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed (accessed on 15 January 2020).
- Creech, C.B.; Frenck, R.W.; Fiquet, A.; Feldman, R.; Kankam, M.K.; Pathirana, S.; Baber, J.; Radley, D.; Cooper, D.; Eiden, J.; et al. Persistence of immune responses through 36 months in healthy adults after vaccination with a novel Staphylococcus aureus 4-Antigen Vaccine (SA4Ag). Open Forum. Infect. Dis. 2020, 7, ofz532. [Google Scholar] [CrossRef]
- Kay, E.; Cuccui, J.; Wren, B.W. Recent advances in the production of recombinant glycoconjugate vaccines. NPJ Vaccines 2019, 4, 16. [Google Scholar] [CrossRef]
- Ingolotti, M.; Kawalekar, O.; Shedlock, D.J.; Muthumani, K.; Weiner, D.B. DNA vaccines for targeting bacterial infections. Expert Rev. Vaccines 2010, 9, 747–763. [Google Scholar] [CrossRef]
- Caskey, M.; Lefebvre, F.; Filali-Mouhim, A.; Cameron, M.J.; Goulet, J.P.; Haddad, E.K.; Breton, G.; Trumpfheller, C.; Pollak, S.; Shimeliovich, I.; et al. Synthetic double-stranded RNA induces innate immune responses similar to a live viral vaccine in humans. J. Exp. Med. 2011, 208, 2357–2366. [Google Scholar] [CrossRef] [PubMed]
- Diaz Blanco, C.; Ortner, A.; Dimitrov, R.; Navarro, A.; Mendoza, E.; Tzanov, T. Building an antifouling zwitterionic coating on urinary catheters using an enzymatically triggered bottom-up approach. ACS Appl. Mater Interfaces 2014, 6, 11385–11393. [Google Scholar] [CrossRef] [PubMed]
- Murugayah, S.A.; Gerth, M.L. Engineering quorum quenching enzymes: Progress and perspectives. Biochem. Soc. Trans. 2019, 47, 793–800. [Google Scholar] [CrossRef] [PubMed]
- Defraine, V.; Fauvart, M.; Michiels, J. Fighting bacterial persistence: Current and emerging anti-persister strategies and therapeutics. Drug Resist Updat. 2018, 38, 12–26. [Google Scholar] [CrossRef]
- Parlet, C.P.; Kavanaugh, J.S.; Crosby, H.A.; Raja, H.A.; El-Elimat, T.; Todd, D.A.; Pearce, C.J.; Cech, N.B.; Oberlies, N.H.; Horswill, A.R. Apicidin attenuates MRSA virulence through quorum-sensing inhibition and enhanced host defense. Cell Rep. 2019, 27, 187–198.e6. [Google Scholar] [CrossRef]
- Hentzer, M.; Eberl, L.; Nielsen, J.; Givskov, M. Quorum sensing: A novel target for the treatment of biofilm infections. BioDrugs 2003, 17, 241–250. [Google Scholar] [CrossRef]
- Fleming, D.; Chahin, L.; Rumbaugh, K. Glycoside hydrolases degrade polymicrobial bacterial biofilms in wounds. Antimicrob. Agents Chemother. 2017, 61, e01998-16. [Google Scholar] [CrossRef]
- Bush, K.; Bradford, P.A. Interplay between β-lactamases and new β-lactamase inhibitors. Nat. Rev. Microbiol. 2019, 17, 295–306. [Google Scholar] [CrossRef]
- Grimsey, E.M.; Fais, C.; Marshall, R.L.; Ricci, V.; Ciusa, M.L.; Stone, J.W.; Ivens, A.; Malloci, G.; Ruggerone, P.; Vargiu, A.V.; et al. Chlorpromazine and amitriptyline are substrates and inhibitors of the acrb multidrug efflux pump. MBio 2020, 11, e00465-20. [Google Scholar] [CrossRef]
- Fukumoto, A.; Kim, Y.P.; Hanaki, H.; Shiomi, K.; Tomoda, H.; Omura, S. Cyslabdan, a new potentiator of imipenem activity against methicillin-resistant Staphylococcus aureus, produced by Streptomyces sp. K04-0144: II. Biological activities. J. Antibiot (Tokyo) 2008, 61, 7–10. [Google Scholar] [CrossRef]
- Nieves, I.; Hally, C.; Viappiani, C.; Agut, M.; Nonell, S. A porphycene-gentamicin conjugate for enhanced photodynamic inactivation of bacteria. Bioorg. Chem. 2020, 97, 103661. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Palanisami, A.; Ashraf, S.; Bhayana, B.; Hasan, T. Photodynamic inactivation of bacterial carbapenemases restores bacterial carbapenem susceptibility and enhances carbapenem antibiotic effectiveness. Photodiagnosis Photodyn. 2020, 30, 101693. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.S.; Sun, W.; Xu, M.; Shen, M.; Khraiwesh, M.; Sciotti, R.J.; Zheng, W. Repurposing screen identifies unconventional drugs with activity against multidrug resistant Acinetobacter baumannii. Front. Cell Infect. Microbiol. 2019, 8, 438. [Google Scholar] [CrossRef]
- Talele, T.T.; Khedkar, S.A.; Rigby, A.C. Successful applications of computer aided drug discovery: Moving drugs from concept to the clinic. Curr. Top. Med. Chem. 2010, 10, 127–141. [Google Scholar] [CrossRef] [PubMed]
- Batalha, I.L.; Bernut, A.; Schiebler, M.; Ouberai, M.M.; Passemar, C.; Klapholz, C.; Kinna, S.; Michel, S.; Sader, K.; Castro-Hartmann, P.; et al. Polymeric nanobiotics as a novel treatment for mycobacterial infections. J. Control. Release 2019, 314, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-López, E.; Gomes, D.; Esteruelas, G.; Bonilla, L.; Lopez-Machado, A.L.; Galindo, R.; Cano, A.; Espina, M.; Ettcheto, M.; Camins, A.; et al. Metal-based nanoparticles as antimicrobial agents: An overview. Nanomaterials 2020, 10, 292. [Google Scholar] [CrossRef] [PubMed]
- Skoura, N.; Wang-Jairaj, J.; Della Pasqua, O.; Chandrasekaran, V.; Billiard, J.; Yeakey, A.; Smith, W.; Steel, H.; Tan, L.K. Effect of raxibacumab on immunogenicity of Anthrax Vaccine Adsorbed: A phase 4, open-label, parallel-group, randomised non-inferiority study. Lancet Infect. Dis. 2020, 20, 983–991. [Google Scholar] [CrossRef]
- Wilcox, M.H.; Gerding, D.N.; Poxton, I.R.; Kelly, C.; Nathan, R.; Birch, T.; Cornely, O.A.; Rahav, G.; Bouza, E.; Lee, C.; et al. Bezlotoxumab for prevention of recurrent Clostridium difficile infection. N. Engl. J. Med. 2017, 376, 305–317. [Google Scholar] [CrossRef]
- Marshall, N.C.; Brett Finlay, B. Targeting the type III secretion system to treat bacterial infections. Expert Opin. Targets 2014, 18, 137–152. [Google Scholar] [CrossRef]
- Saleh, M.M.; Sadeq, R.A.; Abdel Latif, H.K.; Abbas, H.A.; Askoura, M. Zinc oxide nanoparticles inhibits quorum sensing and virulence in Pseudomonas aeruginosa. Afr. Health Sci. 2019, 19, 2043–2055. [Google Scholar] [CrossRef]
- DiGiandomenico, A.; Keller, A.E.; Gao, C.; Rainey, G.J.; Warrener, P.; Camara, M.M.; Bonnell, J.; Fleming, R.; Bezabeh, B.; Dimasi, N.; et al. A multifunctional bispecific antibody protects against Pseudomonas aeruginosa. Sci. Transl. Med. 2014, 6, 262ra155. [Google Scholar] [CrossRef] [PubMed]
- Varshney, A.K.; Kuzmicheva, G.A.; Lin, J.; Sunley, K.M.; Bowling, R.A., Jr.; Kwan, T.Y.; Mays, H.R.; Rambhadran, A.; Zhang, Y.; Martin, R.L.; et al. A natural human monoclonal antibody targeting Staphylococcus protein A protects against Staphylococcus aureus bacteremia. PLoS ONE 2018, 13, e0190537. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Zhou, L.; Li, Y.; Bai, W.; Liu, N.; Li, W.; Gao, Y.; Liu, Z.; Han, R. High-density lipoprotein acts as an opsonin to enhance phagocytosis of group A Streptococcus by U937 cells. Microbiol. Immunol. 2015, 59, 419–425. [Google Scholar] [CrossRef] [PubMed]
- Pirnay, J.P.; Verbeken, G.; Ceyssens, P.J.; Huys, I.; De Vos, D.; Ameloot, C.; Fauconnier, A. The Magistral Phage. Viruses 2018, 10, 64. [Google Scholar] [CrossRef]
- Ferry, T.; Boucher, F.; Fevre, C.; Perpoint, T.; Chateau, J.; Petitjean, C.; Josse, J.; Chidiac, C.; L’hostis, G.; Leboucher, G.; et al. Innovations for the treatment of a complex bone and joint infection due to XDR Pseudomonas aeruginosa including local application of a selected cocktail of bacteriophages. J. Antimicrob. Chemother. 2018, 73, 2901–2903. [Google Scholar] [CrossRef]
- Dedrick, R.M.; Guerrero-Bustamante, C.A.; Garlena, R.A.; Russell, D.A.; Ford, K.; Harris, K.; Gilmour, K.C.; Soothill, J.; Jacobs-Sera, D.; Schooley, R.T.; et al. Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat. Med. 2019, 25, 730–733. [Google Scholar] [CrossRef]
- Hatoum-Aslan, A. Phage genetic engineering using CRISPR–Cas systems. Viruses 2018, 10, 335. [Google Scholar] [CrossRef]
- Briers, Y.; Walmagh, M.; Van Puyenbroeck, V.; Cornelissen, A.; Cenens, W.; Aertsen, A.; Oliveira, H.; Azeredo, J.; Verween, G.; Pirnay, J.P.; et al. Engineered endolysin-based ‘Artilysins’ to combat multidrug-resistant gram-negative pathogens. MBio 2014, 5, e01379–14. [Google Scholar] [CrossRef]
- Hvas, C.L.; Dahl Jørgensen, S.M.; Jørgensen, S.P.; Storgaard, M.; Lemming, L.; Hansen, M.M.; Erikstrup, C.; Dahlerup, J.F. Fecal microbiota transplantation is superior to fidaxomicin for treatment of recurrent Clostridium difficile infection. Gastroenterology 2019, 156, 1324–1332.e3. [Google Scholar] [CrossRef]
- Ozdemir, T.; Fedorec, A.J.H.; Danino, T.; Barnes, C.P. Synthetic biology and engineered live biotherapeutics: Toward increasing system complexity. Cell Syst. 2018, 7, 5–16. [Google Scholar] [CrossRef]
- Bar-Yoseph, H.; Carasso, S.; Shklar, S.; Korytny, A.; Even Dar, R.; Daoud, H.; Nassar, R.; Maharshak, N.; Hussein, K.; Geffen, Y.; et al. Oral capsulized fecal microbiota transplantation for eradication of carbapenemase-producing Enterobacteriaceae colonization with a metagenomic perspective. Clin. Infect. Dis. 2020, cia737. [Google Scholar] [CrossRef] [PubMed]
- Hsu, B.B.; Gibson, T.E.; Yeliseyev, V.; Liu, Q.; Lyon, L.; Bry, L.; Silver, P.A.; Gerber, G.K. Dynamic modulation of the gut microbiota and metabolome by bacteriophages in a mouse model. Cell Host. Microbe. 2019, 25, 803–814.e5. [Google Scholar] [CrossRef] [PubMed]
- Draper, L.A.; Ryan, F.J.; Smith, M.K.; Jalanka, J.; Mattila, E.; Arkkila, P.A.; Ross, R.P.; Satokari, R.; Hill, C. Long-term colonisation with donor bacteriophages following successful faecal microbial transplantation. Microbiome 2018, 6, 220. [Google Scholar] [CrossRef] [PubMed]
- Kosack, C.S.; Page, A.-L.; Klatse, P.R. A guide to aid the selection of diagnostic tests. Bull. World Health Organ. 2017, 95, 639–645. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernández, L.; Cima-Cabal, M.D.; Duarte, A.C.; Rodriguez, A.; García, P.; García-Suárez, M.d.M. Developing Diagnostic and Therapeutic Approaches to Bacterial Infections for a New Era: Implications of Globalization. Antibiotics 2020, 9, 916. https://doi.org/10.3390/antibiotics9120916
Fernández L, Cima-Cabal MD, Duarte AC, Rodriguez A, García P, García-Suárez MdM. Developing Diagnostic and Therapeutic Approaches to Bacterial Infections for a New Era: Implications of Globalization. Antibiotics. 2020; 9(12):916. https://doi.org/10.3390/antibiotics9120916
Chicago/Turabian StyleFernández, Lucía, María Dolores Cima-Cabal, Ana Catarina Duarte, Ana Rodriguez, Pilar García, and María del Mar García-Suárez. 2020. "Developing Diagnostic and Therapeutic Approaches to Bacterial Infections for a New Era: Implications of Globalization" Antibiotics 9, no. 12: 916. https://doi.org/10.3390/antibiotics9120916
APA StyleFernández, L., Cima-Cabal, M. D., Duarte, A. C., Rodriguez, A., García, P., & García-Suárez, M. d. M. (2020). Developing Diagnostic and Therapeutic Approaches to Bacterial Infections for a New Era: Implications of Globalization. Antibiotics, 9(12), 916. https://doi.org/10.3390/antibiotics9120916







