Antimicrobial Drug Interactions: Systematic Evaluation of Protein and Nucleic Acid Synthesis Inhibitors
Abstract
1. Introduction
2. Materials and Methods
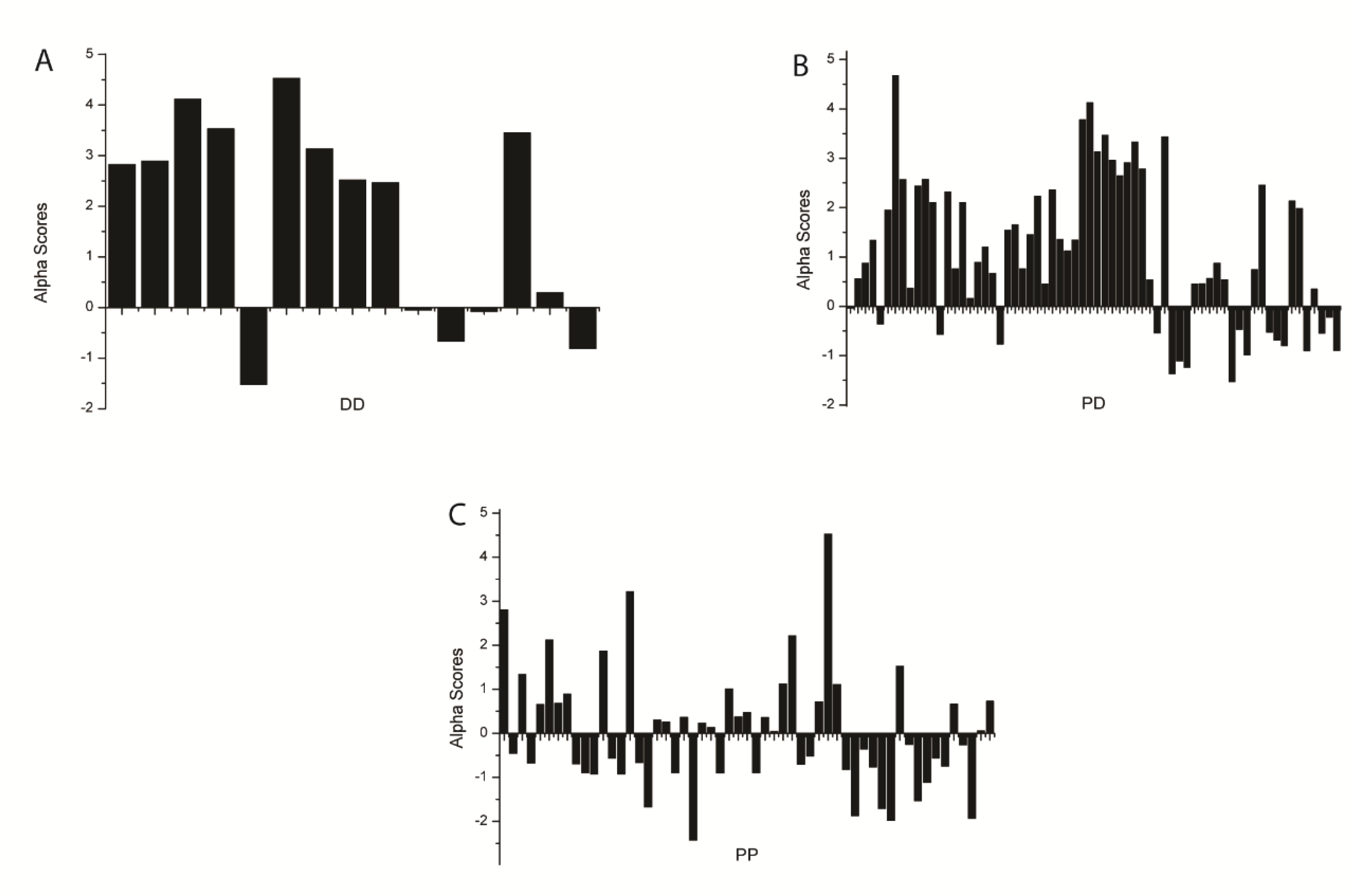
3. Results
4. Discussion
5. Conclusions
Funding
Conflicts of Interest
References
- Foxman, B.; Brown, P. Epidemiology of urinary tract infections: Transmission and risk factors, incidence, and costs. Infect. Dis. Clin. North Am. 2003, 17, 227–241. [Google Scholar] [CrossRef]
- Cokol, M.; Chua, H.N.; Tasan, M.; Mutlu, B.; Weinstein, Z.B.; Suzuki, Y.; Nergiz, M.E.; Costanzo, M.; Baryshnikova, A.; Giaever, G.; et al. Systematic exploration of synergistic drug pairs. Mol. Syst. Biol. 2011, 7, 544. [Google Scholar] [CrossRef] [PubMed]
- Chait, R.; Craney, A.; Kishony, R. Antibiotic interactions that select against resistance. Nature 2007, 446, 668–671. [Google Scholar] [CrossRef] [PubMed]
- Yeh, P.; Tschumi, A.I.; Kishony, R. Functional classification of drugs by properties of their pairwise interactions. Nat. Genet. 2006, 38, 489–494. [Google Scholar] [CrossRef] [PubMed]
- Lehar, J.; Krueger, A.S.; Avery, W.; Heilbut, A.M.; Johansen, L.M.; Price, E.R.; Rickles, R.J.; Short, G.F.; Staunton, J.E.; Jin, X.; et al. Synergistic drug combinations tend to improve therapeutically relevant selectivity. Nat. Biotechnol. 2009, 27, 659–666. [Google Scholar] [CrossRef] [PubMed]
- Torella, J.P.; Chait, R.; Kishony, R. Optimal drug synergy in antimicrobial treatments. PLoS Comput. Biol. 2010, 6, e1000796. [Google Scholar] [CrossRef]
- Messina, G.; Rosadini, D.; Burgassi, S.; Messina, D.; Nante, N.; Tani, M.; Cevenini, G. Tanning the bugs—A pilot study of an innovative approach to stethoscope disinfection. J. Hosp. Infect. 2017, 95, 228–230. [Google Scholar] [CrossRef] [PubMed]
- Fattorini, M.; Ceriale, E.; Nante, N.; Lenzi, D.; Manzi, P.; Basagni, C.; Messina, G. Use of a fluorescent marker for assessing hospital bathroom cleanliness. Am. J. Infect. Control 2016, 44, 1066–1068. [Google Scholar] [CrossRef] [PubMed]
- Messina, G.; Fattorini, M.; Nante, N.; Rosadini, D.; Serafini, A.; Tani, M.; Cevenini, G. Time Effectiveness of Ultraviolet C Light (UVC) Emitted by Light Emitting Diodes (LEDs) in Reducing Stethoscope Contamination. Int. J. Environ. Res. Public Health 2016, 13, 940. [Google Scholar] [CrossRef] [PubMed]
- Orhan, G.; Bayram, A.; Zer, Y.; Balci, I. Synergy tests by E test and checkerboard methods of antimicrobial combinations against Brucella melitensis. J. Clin. Microbiol. 2005, 43, 140–143. [Google Scholar] [CrossRef] [PubMed]
- Loewe, S. The problem of synergism and antagonism of combined drugs. Arzneim. Forsch. 1953, 3, 285–290. [Google Scholar]
- Worthington, R.J.; Melander, C. Combination approaches to combat multidrug-resistant bacteria. Trends Biotechnol. 2013, 31, 177–184. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandri, R.M.; McNeely, D.J.; Kluge, R.M. Antibiotic synergy and antagonism against clinical isolates of Klebsiella species. Antimicrob. Agents Chemother. 1976, 10, 889–892. [Google Scholar] [CrossRef] [PubMed]
- Ocampo, P.S.; Lazar, V.; Papp, B.; Arnoldini, M.; zur Wiesch, P.A.; Busa-Fekete, R.; Fekete, G.; Pal, C.; Ackermann, M.; Bonhoeffer, S. Antagonism between Bacteriostatic and Bactericidal Antibiotics Is Prevalent. Antimicrob. Agents Chemother. 2014, 58, 4573–4582. [Google Scholar] [CrossRef] [PubMed]
- Yilancioglu, K.; Unlu, O. Multidrug resistance stimulated antagonistic antibiotic interactions. Rom. J. Leg. Med. 2017, 25, 331–336. [Google Scholar]
- Yilancioglu, K.; Weinstein, Z.B.; Meydan, C.; Akhmetov, A.; Toprak, I.; Durmaz, A.; Iossifov, I.; Kazan, H.; Roth, F.P.; Cokol, M. Target-Independent Prediction of Drug Synergies Using Only Drug Lipophilicity. J. Chem. Inf. Model. 2014, 54, 2286–2293. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, S.; Cokol-Cakmak, M.; Sahin, N.; Yilancioglu, K.; Kazan, H.; Collins, J.J.; Cokol, M. Chemogenomics and orthology-based design of antibiotic combination therapies. Mol. Syst. Biol. 2016, 12. [Google Scholar] [CrossRef] [PubMed]
- Jawetz, E.; Gunnison, J.B. Antibiotic synergism and antagonism; an assessment of the problem. Pharmacol. Rev. 1953, 5, 175–192. [Google Scholar] [PubMed]
- Oda, Y. Induction of SOS responses in Escherichia coli by 5-fluorouracil. Mutat. Res. DNA Repair Rep. 1987, 183, 103–108. [Google Scholar] [CrossRef]
- Phillips, I.; Culebras, E.; Moreno, F.; Baquero, F. Induction of the SOS response by new 4-quinolones. J. Antimicrob. Chemother. 1987, 20, 631–638. [Google Scholar] [CrossRef] [PubMed]
| Compounds | Abbreviation | Mechanism of Action | MIC/LB (mg/mL) |
|---|---|---|---|
| Amikacin | AMK | Protein synthesis, 30S inhibition | 13 |
| Gentamicin | GEN | Protein synthesis, 30S inhibition | 7 |
| Tobramycin | TOB | Protein synthesis, 30S inhibition | 0.7 |
| Tetracycline | TET | Protein synthesis, 30S inhibition | 5 |
| Spectinomycin | SPE | Protein synthesis, 30S inhibition | 2 |
| Clarithromycin | CLA | Protein synthesis, 50S inhibition | 9 |
| Erythromycin | ERY | Protein synthesis, 50S inhibition | 15 |
| Chloramphenicol | CHL | Protein synthesis, 50S inhibition | 3.5 |
| Fusidic acid | FUS | Elongation factor, protein synthesis inhibition | 80 |
| Mupirocin | MUP | Isoleucyl transfer, RNA (tRNA) synthetase inhibition | 0.4 |
| Roxithromycin | ROX | Protein synthesis, 50S inhibition | 0.3 |
| Ciprofloxacin | CIP | DNA gyrase inhibition | 0.05 |
| Levofloxacin | LEV | DNA gyrase inhibition | 0.01 |
| Nalidixic acid | NAL | DNA gyrase inhibition | 8 |
| Trimethoprim | TRI | Folic acid biosynthesis inhibition | 1.5 |
| Rifampicin | RIF | RNA polymerase inhibition | 0.5 |
| 5-Fluorouracil | 5FU | Inhibition of the formation of thymidylate from uracil | 2 |
| Combination Type | N | Min | Q1 | Median | Q3 | Max |
|---|---|---|---|---|---|---|
| PP | 55 | −2.4 | −0.89 | −0.25 | 0.72 | 4.5 |
| NN | 15 | −1.5 | −0.07 | 2.51 | 3.45 | 4.5 |
| U | Z | Prob > |U| | ||||
| 214 | −2.8 | 0.004 | ||||
| N | Min | Q1 | Median | Q3 | Max | |
| PN | 66 | −1.5 | −0.25 | 0.87 | 2.33 | 4.67 |
| NN | 15 | −1.5 | −0.07 | 2.51 | 3.45 | 4.52 |
| U | Z | Prob > |U| | ||||
| 385 | −1.33 | 0.18 | ||||
| N | Min | Q1 | Median | Q3 | Max | |
| PN | 66 | −1.52 | −0.25 | 0.87 | 2.33 | 4.67 |
| PP | 55 | −2.42 | −0.89 | −0.25 | 0.72 | 4.52 |
| U | Z | Prob > |U| | ||||
| 2552 | 3.83 | 1.26× 10−4 |
| DRUG PAIR | ALPHA | 1. DRUG | 2. DRUG | DRUG PAIR | ALPHA | 1. DRUG | 2. DRUG |
|---|---|---|---|---|---|---|---|
| PN | −0.03 | AMK | CIP | PP | 1.52 | AMK | ROX |
| PN | 0.55 | AMK | LEV | PP | 2.8 | AMK | CHL |
| PN | 0.87 | AMK | NAL | PP | −0.45 | AMK | CLA |
| PN | 1.34 | AMK | RIF | PP | 1.34 | AMK | ERY |
| PN | −0.35 | AMK | TRI | PP | −0.67 | AMK | FUS |
| PN | −0.68 | AMK | 5FU | PP | 0.65 | AMK | GEN |
| PN | 1.94 | CHL | CIP | PP | 2.12 | AMK | SPE |
| PN | 4.67 | CHL | LEV | PP | 0.68 | AMK | TET |
| PN | 2.56 | CHL | NAL | PP | 0.89 | AMK | TOB |
| PN | 0.36 | CHL | RIF | PP | 0.05 | AMK | MUP |
| PN | 2.43 | CHL | TRI | PP | −0.68 | CHL | CLA |
| PN | −1.52 | CHL | 5FU | PP | −0.89 | CHL | ERY |
| PN | 2.57 | CLA | LEV | PP | −0.92 | CHL | FUS |
| PN | 2.1 | CLA | NAL | PP | 1.87 | CHL | GEN |
| PN | −0.56 | CLA | RIF | PP | −0.56 | CHL | SPE |
| PN | 2.32 | CLA | TRI | PP | −0.92 | CHL | TET |
| PN | −0.89 | CLA | 5FU | PP | 3.21 | CHL | TOB |
| PN | 2.36 | CIP | CLA | PP | −0.26 | CHL | ROX |
| PN | 1.35 | CIP | ERY | PP | −0.56 | CHL | MUP |
| PN | 1.12 | CIP | FUS | PP | −0.66 | CLA | ERY |
| PN | 1.34 | CIP | GEN | PP | −1.67 | CLA | FUS |
| PN | 3.78 | CIP | SPE | PP | 0.30 | CLA | GEN |
| PN | 4.12 | CIP | TET | PP | 0.25 | CLA | SPE |
| PN | 2.13 | CIP | MUP | PP | −0.89 | CLA | TET |
| PN | 3.12 | CIP | TOB | PP | 0.36 | CLA | TOB |
| PN | 3.46 | LEV | SPE | PP | −1.11 | CLA | MUP |
| PN | 2.95 | LEV | TET | PP | −0.25 | CLA | ROX |
| PN | 2.64 | LEV | TOB | PP | −2.42 | ERY | FUS |
| PN | 0.76 | LEV | ERY | PP | 0.23 | ERY | GEN |
| PN | 1.2 | LEV | FUS | PP | 0.13 | ERY | SPE |
| PN | −0.21 | LEV | ROX | PP | −0.90 | ERY | TET |
| PN | 0.35 | LEV | GEN | PP | 1.0 | ERY | TOB |
| PN | 2.45 | LEV | MUP | PP | −0.81 | ERY | MUP |
| PN | 2.1 | ERY | NAL | PP | −0.35 | ERY | ROX |
| PN | 0.16 | ERY | RIF | PP | 0.37 | FUS | GEN |
| PN | 0.89 | ERY | TRI | PP | 0.47 | FUS | SPE |
| PN | −0.52 | ERY | 5FU | PP | −0.89 | FUS | TET |
| PN | 0.67 | FUS | NAL | PP | 0.35 | FUS | TOB |
| PN | −0.76 | FUS | RIF | PP | −0.76 | FUS | MUP |
| PN | 1.54 | FUS | TRI | PP | −1.53 | FUS | ROX |
| PN | 0.54 | FUS | 5FU | PP | 2.21 | GEN | SPE |
| PN | 1.65 | GEN | RIF | PP | −0.70 | GEN | TET |
| PN | 0.76 | GEN | TRI | PP | −0.51 | GEN | TOB |
| PN | 1.98 | GEN | 5FU | PP | 0.04 | GEN | ROX |
| PN | 0.45 | GEN | NAL | PP | −0.74 | GEN | MUP |
| PN | 2.90 | NAL | SPE | PP | 0.72 | SPE | TET |
| PN | 3.32 | NAL | TET | PP | 4.52 | SPE | TOB |
| PN | 2.78 | NAL | TOB | PP | −1.87 | SPE | ROX |
| PN | −0.89 | NAL | ROX | PP | −1.70 | SPE | MUP |
| PN | 0.87 | NAL | MUP | PP | 1.10 | TET | TOB |
| PN | −1.36 | ROX | 5FU | PP | −1.98 | TET | ROX |
| PN | −1.10 | ROX | RIF | PP | −1.93 | TET | MUP |
| PN | −1.23 | ROX | TRI | PP | 1.12 | ROX | MUP |
| PN | 0.56 | ROX | CIP | PP | 0.66 | ROX | TOB |
| PN | 0.54 | RIF | SPE | PP | 0.72 | TOB | MUP |
| PN | −0.54 | RIF | TET | NN | 4.12 | 5FU | TRI |
| PN | 3.43 | RIF | TOB | NN | 3.53 | 5FU | NAL |
| PN | 0.45 | RIF | MUP | NN | 4.52 | 5FU | LEV |
| PN | 1.45 | TRI | SPE | NN | 3.13 | 5FU | RIF |
| PN | 2.23 | TRI | TET | NN | 3.45 | 5FU | CIP |
| PN | 0.45 | TRI | TOB | NN | 2.46 | NAL | CIP |
| PN | −0.46 | TRI | MUP | NN | −0.05 | NAL | RIF |
| PN | −0.98 | 5FU | SPE | NN | 2.82 | NAL | TRI |
| PN | 0.74 | 5FU | MUP | NN | −1.52 | NAL | LEV |
| PN | −0.79 | 5FU | TOB | NN | −0.07 | LEV | CIP |
| PN | −0.54 | 5FU | TET | NN | 2.89 | LEV | RIF |
| NN | −0.81 | LEV | TRI | ||||
| NN | −0.67 | TRI | CIP | ||||
| NN | 0.29 | TRI | RIF | ||||
| NN | 2.51 | CIP | RIF |
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yilancioglu, K. Antimicrobial Drug Interactions: Systematic Evaluation of Protein and Nucleic Acid Synthesis Inhibitors. Antibiotics 2019, 8, 114. https://doi.org/10.3390/antibiotics8030114
Yilancioglu K. Antimicrobial Drug Interactions: Systematic Evaluation of Protein and Nucleic Acid Synthesis Inhibitors. Antibiotics. 2019; 8(3):114. https://doi.org/10.3390/antibiotics8030114
Chicago/Turabian StyleYilancioglu, Kaan. 2019. "Antimicrobial Drug Interactions: Systematic Evaluation of Protein and Nucleic Acid Synthesis Inhibitors" Antibiotics 8, no. 3: 114. https://doi.org/10.3390/antibiotics8030114
APA StyleYilancioglu, K. (2019). Antimicrobial Drug Interactions: Systematic Evaluation of Protein and Nucleic Acid Synthesis Inhibitors. Antibiotics, 8(3), 114. https://doi.org/10.3390/antibiotics8030114




