Diagnostic Underuse and Antimicrobial Resistance Patterns Among Hospitalized Children in a National Referral Hospital in Kenya: A Five-Year Retrospective Study
Abstract
1. Introduction
2. Results
2.1. Patient Demographics
2.2. Infection Types
2.3. Diagnostic Utilization
2.3.1. Culture Requests
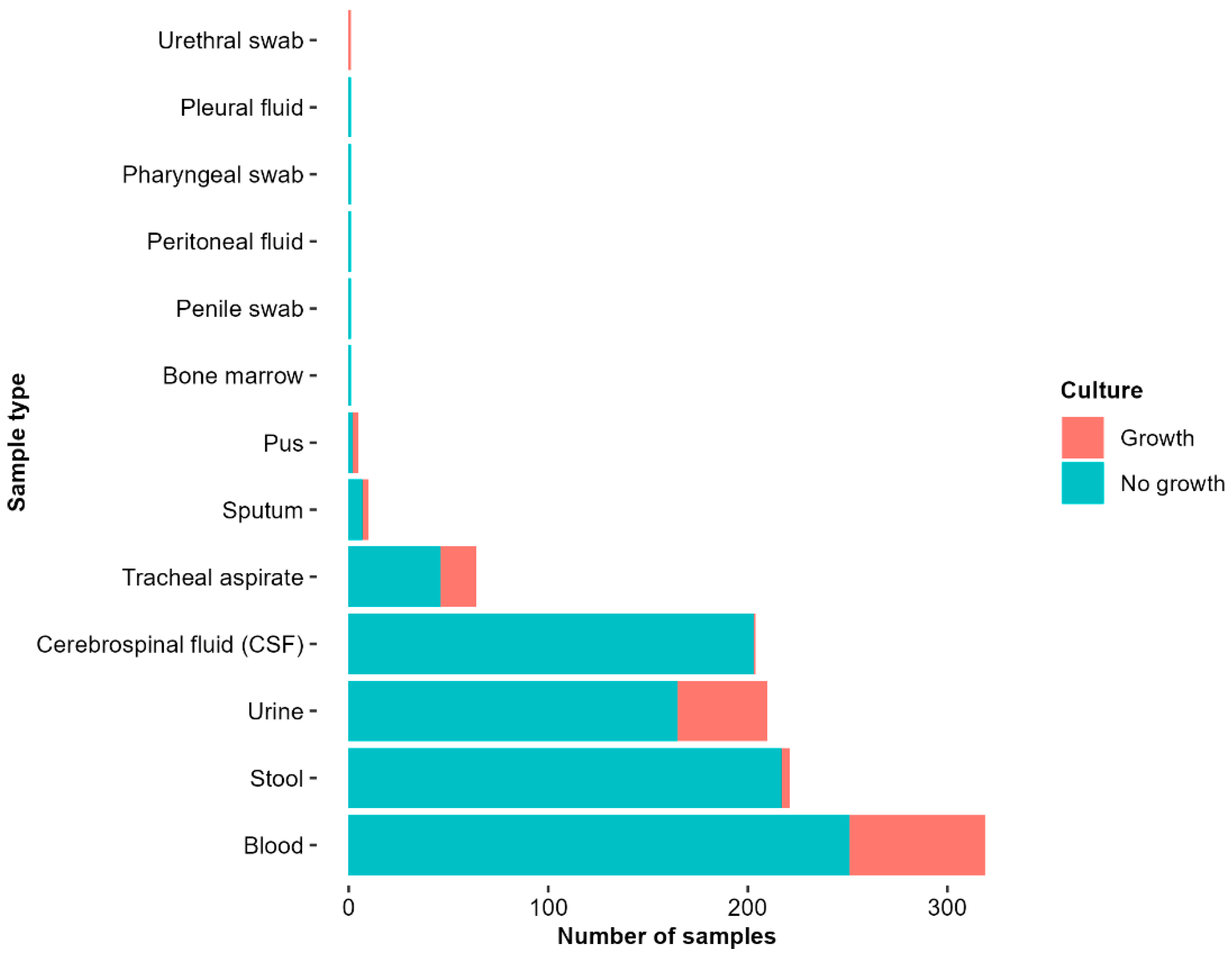
2.3.2. Sample Types and Culture Yield
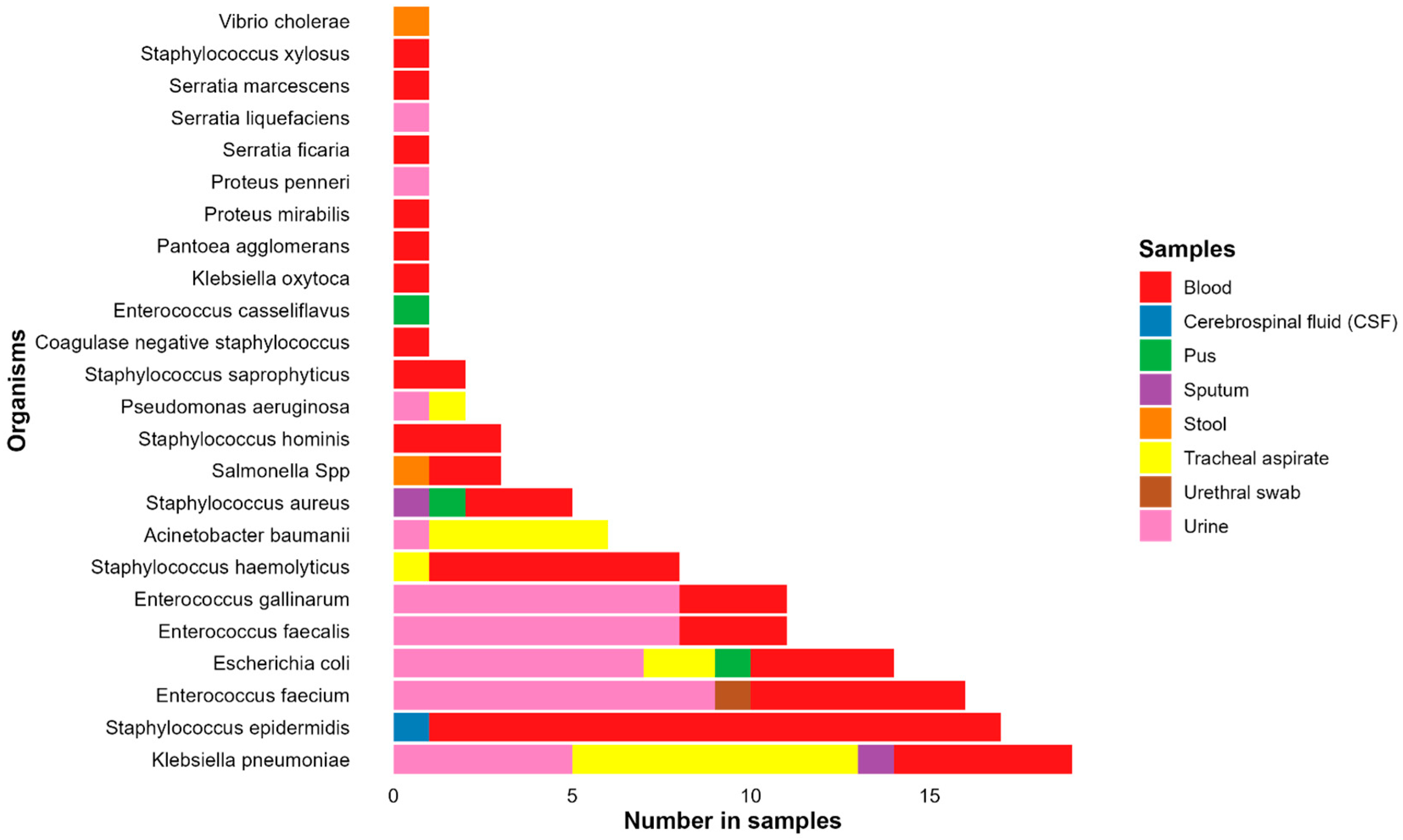
2.3.3. Pathogen Identification and Antimicrobial Susceptibility Testing
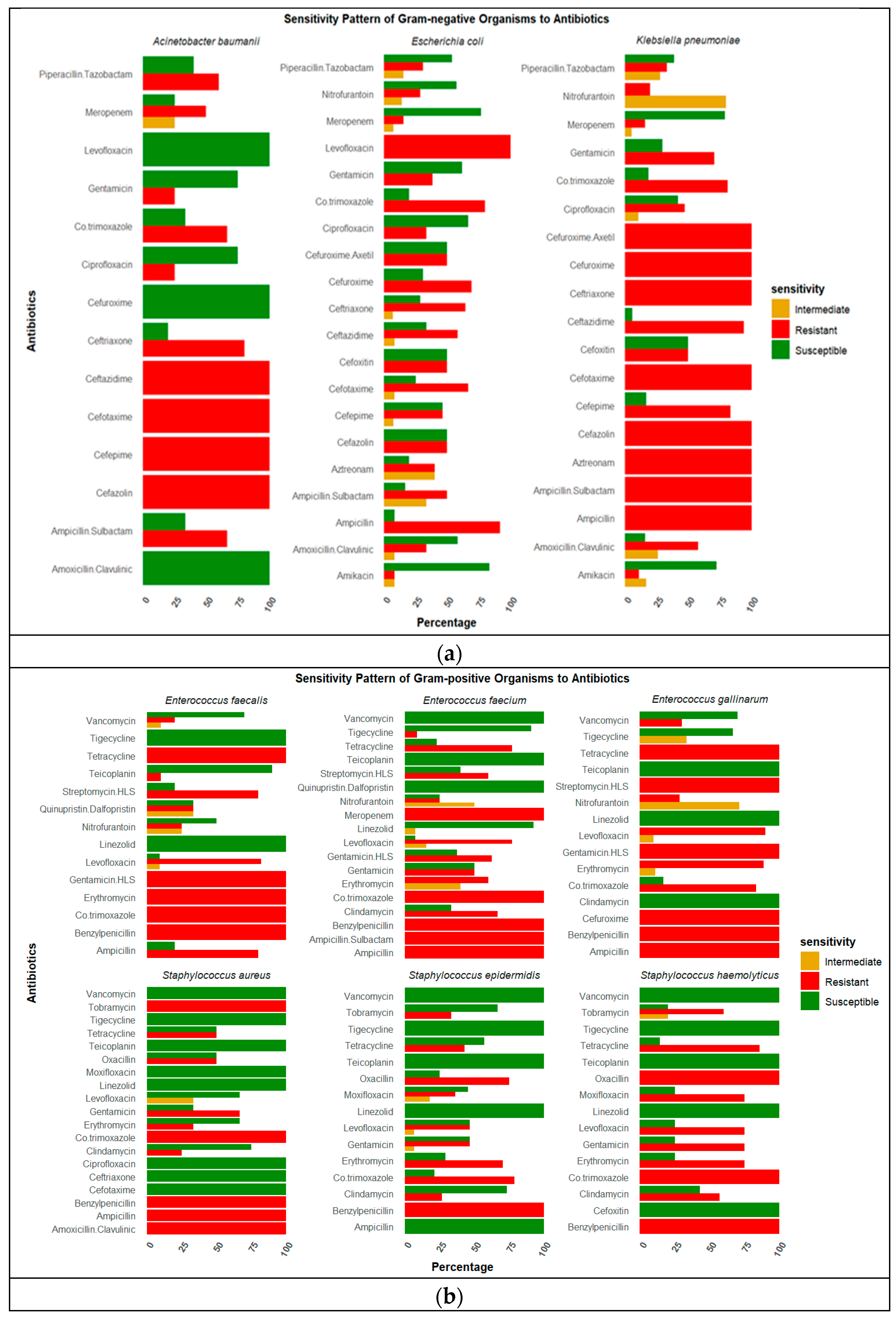
2.4. Antibiotic Resistance
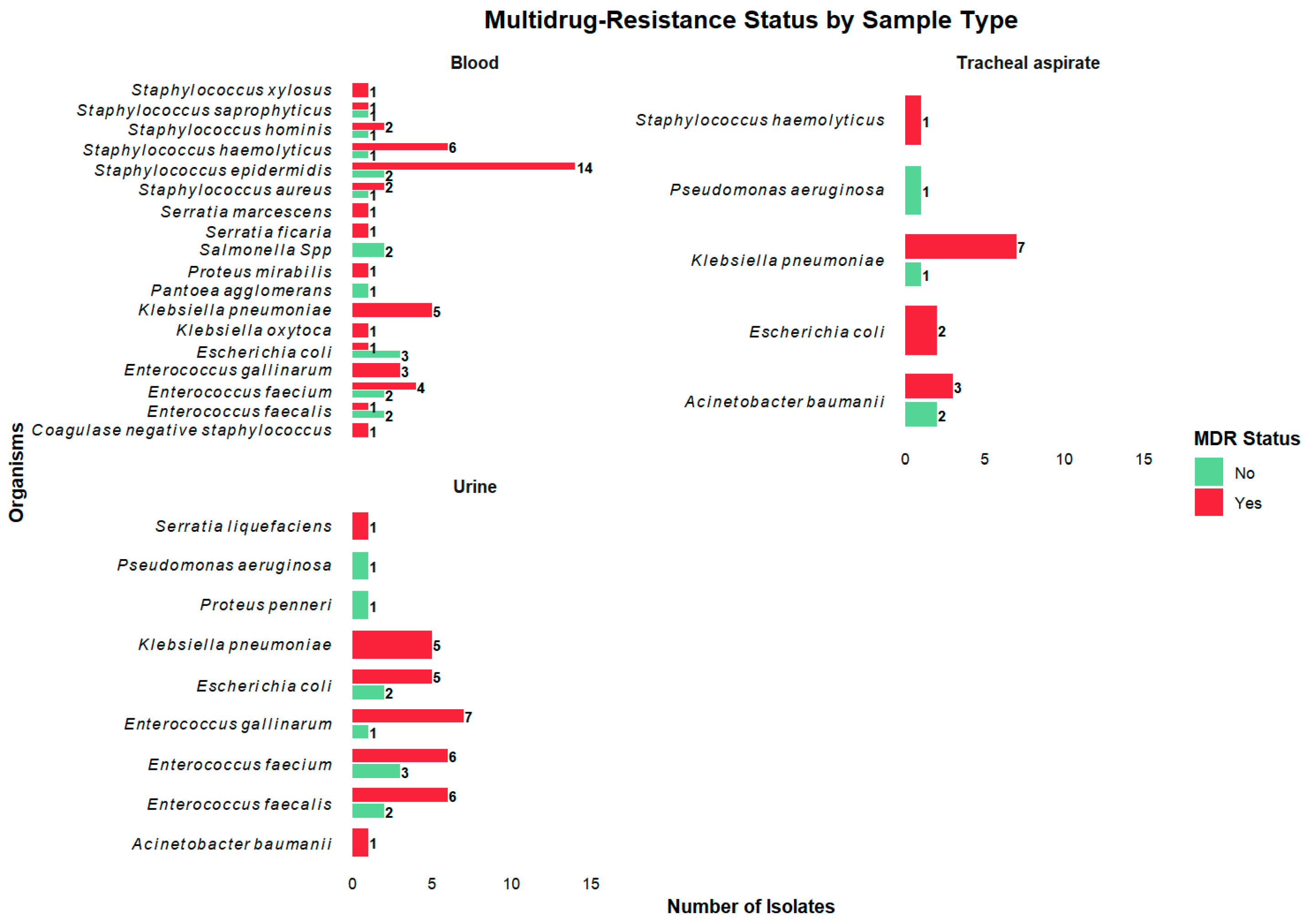
2.5. Multidrug- and Extensively Drug-Resistant Organisms
2.6. Association Between Drug-Resistant Infections and Mortality
2.7. Factors Associated with Acquiring Multidrug-Resistant Organisms
3. Discussion
4. Materials and Methods
4.1. Study Design and Setting
4.2. Study Population and Eligibility Criteria
4.3. Data Collection Procedures
4.4. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- WHO. Antimicrobial Resistance: Global Report on Surveillance; World Health Organization (WHO): Geneva, Switzerland, 2014. [Google Scholar]
- Muteeb, G.; Rehman, M.T.; Shahwan, M.; Aatif, M. Origin of Antibiotics and Antibiotic Resistance, and Their Impacts on Drug Development: A Narrative Review. Pharmaceuticals 2023, 16, 1615. [Google Scholar] [CrossRef]
- Leopold, S.J.; Van, L.F.; Tarekegn, H.; Schultsz, C. Antimicrobial drug resistance among clinically relevant bacterial isolates in sub-Saharan Africa: A systematic review. J. Antimicrob. Chemother. 2014, 69, 2337–2353. [Google Scholar] [CrossRef]
- Williams, P.C.M.; Isaacs, D.; Berkley, J.A. Antimicrobial resistance among children in sub-Saharan Africa. Lancet Infect. Dis. 2018, 18, e33–e44. [Google Scholar] [CrossRef]
- Tadesse, B.T.; Ashley, E.A.; Ongarello, S.; Havumaki, J.; Wijegoonewardena, M.; González, I.J.; Dittrich, S. Antimicrobial resistance in Africa: A systematic review. BMC Infect. Dis. 2017, 17, 616. [Google Scholar] [CrossRef]
- Bebell, L.M.; Muiru, A.N. Antibiotic Use and Emerging Resistance: How Can Resource-Limited Countries Turn the Tide? Glob. Heart 2014, 9, 347–358. [Google Scholar] [CrossRef] [PubMed]
- Sriram, A.; Kalanxhi, E.; Kapoor, G.; Craig, J.; Balasubramanian, R.; Brar, S.; Criscuolo, N.; Hamilton, A.; Klein, E.; Tseng, K.; et al. The State of the World’s Antibiotics 2021:A Global Analysis of Antimicrobial Resistance and Its Drivers; Center for Disease Dynamics, Economics & Policy: Washington, DC, USA, 2021. [Google Scholar]
- Van Boeckel, T.P.; Gandra, S.; Ashok, A.; Caudron, Q.; Grenfell, B.T.; Levin, S.A.; Laxminarayan, R. Global antibiotic consumption 2000 to 2010: An analysis of national pharmaceutical sales data. Lancet Infect. Dis. 2014, 14, 742–750. [Google Scholar] [CrossRef] [PubMed]
- Klein, E.Y.; Van Boeckel, T.P.; Martinez, E.M.; Pant, S.; Gandra, S.; Levin, S.A.; Goossens, H.; Laxminarayan, R. Global increase and geographic convergence in antibiotic consumption between 2000 and 2015. Proc. Natl. Acad. Sci. USA 2018, 115, E3463–E3470. [Google Scholar] [CrossRef]
- Harbarth, S.; Balkhy, H.H.; Goossens, H.; Jarlier, V.; Kluytmans, J.; Laxminarayan, R.; Saam, M.; Van Belkum, A.; Pittet, D. Antimicrobial resistance: One world, one fight! Antimicrob. Resist. Infect. Control 2015, 4, 49. [Google Scholar] [CrossRef]
- Essack, S.Y.; Desta, A.T.; Abotsi, R.E.; Agoba, E.E. Perspectives Antimicrobial resistance in the WHO African region: Current status and roadmap for action. Public Health 2016, 39, 8–13. [Google Scholar]
- Trevas, D.; Caliendo, A.M.; Hanson, K.; Levy, J.; Ginocchio, C.C. Diagnostic Tests Can Stem the Threat of Antimicrobial Resistance: Infectious Disease Professionals Can Help. Clin. Infect. Dis. 2021, 72, e893–e900. [Google Scholar] [CrossRef]
- Burnham, C.-A.D.; Leeds, J.; Nordmann, P.; O’Grady, J.; Patel, J. Diagnosing antimicrobial resistance. Microbiology 2017, 15, 697–703. [Google Scholar] [CrossRef]
- FIND. Accelerating Diagnostics Use to Prevent Antimicrobial Resistance: AMR Strategy 2018; Foundation for Innovative New Diagnostics (FIND): Geneva, Switzerland, 2018. [Google Scholar]
- Ramanathan, K.; Antognini, D.; Combes, A.; Paden, M.; Zakhary, B.; Ogino, M.; Maclaren, G.; Brodie, D. The Lancet Commission on diagnostics: Transforming access to diagnostics Kenneth. Lancet 2021, 398, 1997–2050. [Google Scholar]
- Murray, C.J.L.; Ikuta, K.S.; Sharara, F.; Swetschinski, L.; Aguilar, G.R.; Gray, A.; Han, C.; Bisignano, C.; Rao, P.; Wool, E.; et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar] [CrossRef] [PubMed]
- Peeling, R.W.; Heymann, L.D.; Boeras, D. The Value of Diagnostics in the Fight Against Antimicrobial Resistance; World Health Organization (WHO): Geneva, Switzerland, 2024. [Google Scholar]
- Yu, D.; Zheng, Y.; Shen, A.; Wu, F.; Dmitriev, A.V.; Kilian, M.; Yang, Y. Antimicrobial resistance in pediatric infectious diseases: Antimicrobial resistance, resistance mechanisms and antimicrobial use. Front. Cell. Infect. Microbiol. 2023, 13, 1287051. [Google Scholar] [CrossRef]
- Mustafa, Z.U.; Khan, A.H.; Salman, M.; Syed Sulaiman, S.A.; Godman, B. Antimicrobial Utilization among Neonates and Children: A Multicenter Point Prevalence Study from Leading Children’s Hospitals in Punjab, Pakistan. Antibiotics 2022, 11, 1056. [Google Scholar] [CrossRef]
- Bamford, A.; Masini, T.; Williams, P.; Sharland, M.; Gigante, V.; Dixit, D.; Sati, H.; Huttner, B.; Bin Nisar, Y.; Cappello, B.; et al. Tackling the threat of antimicrobial resistance in neonates and children: Outcomes from the first WHO-convened Paediatric Drug Optimisation exercise for antibiotics. Lancet Child Adolesc. Health 2024, 8, 456–466. [Google Scholar] [CrossRef]
- Sohaili, A.; Asin, J.; Thomas, P.P.M. The Fragmented Picture of Antimicrobial Resistance in Kenya: A Situational Analysis of Antimicrobial Consumption and the Imperative for Antimicrobial Stewardship. Antibiotics 2024, 13, 197. [Google Scholar] [CrossRef]
- Ahmed, S.K.; Hussein, S.; Qurbani, K.; Ibrahim, R.H.; Fareeq, A.; Mahmood, K.A.; Mohamed, M.G. Antimicrobial resistance: Impacts, challenges, and future prospects. J. Med. Surg. Public Health 2024, 2, 100081. [Google Scholar] [CrossRef]
- Iskandar, K.; Molinier, L.; Hallit, S.; Sartelli, M.; Hardcastle, T.C. Surveillance of antimicrobial resistance in low—And middle—Income countries: A scattered picture. Antimicrob. Resist. Infect. Control 2021, 10, 63. [Google Scholar] [CrossRef]
- Walker, C.L.F.; Rudan, I.; Liu, L.; Nair, H.; Theodoratou, E.; Bhutta, Z.A.; O’Brien, K.L.; Campbell, H.; Black, R.E. Global burden of childhood pneumonia and diarrhoea. Lancet 2013, 381, 1405–1416. [Google Scholar] [CrossRef] [PubMed]
- Troeger, C.; Khalil, I.A.; Rao, P.C.; Cao, S.; Blacker, B.F.; Ahmed, T.; Armah, G.; Bines, J.E.; Brewer, T.G.; Colombara, D.V.; et al. Rotavirus Vaccination and the Global Burden of Rotavirus Diarrhea among Children Younger Than 5 Years. JAMA Pediatr. 2018, 172, 958–965. [Google Scholar] [CrossRef]
- WHO. UNICEF Progress on Household Drinking Water, Sanitation and Hygiene 2000–2020: Five Years into the SDGs; Joint Water Supply & Sanitation Monitoring Programme; World Health Organization (WHO) and the United Nations Children’s Fund (UNICEF): Geneva, Switzerland, 2021; ISBN 9789240030848. Available online: http://apps.who.int/bookorders (accessed on 6 July 2025).
- McAllister, D.A.; Liu, L.; Shi, T.; Chu, Y.; Reed, C.; Burrows, J.; Adeloye, D.; Rudan, I.; Black, R.E.; Campbell, H.; et al. Global, regional, and national estimates of pneumonia morbidity and mortality in children younger than 5 years between 2000 and 2015: A systematic analysis. Lancet Glob. Health 2019, 7, e47–e57. [Google Scholar] [CrossRef]
- WHO. Pneumonia in Children; WHO Press: Geneva, Switzerland, 2022; Available online: https://www.who.int/news-room/fact-sheets/detail/pneumonia (accessed on 6 July 2025).
- Nair, H.; Simões, E.A.F.; Rudan, I.; Gessner, B.D.; Azziz-Baumgartner, E.; Zhang, J.S.F.; Feikin, D.R.; MacKenzie, G.A.; Moïsi, J.C.; Roca, A.; et al. Global and regional burden of hospital admissions for severe acute lower respiratory infections in young children in 2010: A systematic analysis. Lancet 2013, 381, 1380–1390. [Google Scholar] [CrossRef]
- Lu, W.; Chen, X.; Ho, D.C.W.; Wang, H. Global burden of acterial antimicrobial resistance in 2019: A systematic analysis. Lancet 2022, 399, 629–655. [Google Scholar]
- Liu, L.; Oza, S.; Hogan, D.; Chu, Y.; Perin, J.; Zhu, J.; Lawn, J.E.; Cousens, S.; Mathers, C.; Black, R.E. Global, regional, and national causes of under-5 mortality in 2000–2015: An updated systematic analysis with implications for the Sustainable Development Goals. Lancet 2016, 388, 3027–3035. [Google Scholar] [CrossRef]
- Rudd, K.E.; Johnson, S.C.; Agesa, K.M.; Shackelford, K.A.; Tsoi, D.; Kievlan, D.R.; Colombara, D.V.; Ikuta, K.S.; Kissoon, N.; Finfer, S.; et al. Global, regional, and national sepsis incidence and mortality, 1990–2017: Analysis for the Global Burden of Disease Study. Lancet 2020, 395, 200–211. [Google Scholar] [CrossRef] [PubMed]
- Obiero, C.W.; Chb, M.B.; Seale, A.C.; Berkley, J.A. Europe PMC Funders Group Empiric Treatment of Neonatal Sepsis in Developing Countries. Pediatr. Infect. Dis. 2019, 34, 659–661. [Google Scholar] [CrossRef]
- Yadav, H.; Shah, D.; Sayed, S.; Horton, S.; Schroeder, L.F. Availability of essential diagnostics in ten low-income and middle-income countries: Results from national health facility surveys. Lancet Glob. Health 2021, 9, e1553–e1560. [Google Scholar] [CrossRef] [PubMed]
- WHO. All for Health, Health for all WHO Investment Case Deep Dive: Antimicrobial Resistance; World Health Organization (WHO): Geneva, Switzerland, 2024. [Google Scholar]
- Leazer, R.; Erickson, N.; Paulson, J.; Zipkin, R.; Stemmle, M.; Schroeder, A.R.; Bendel-Stenzel, M.; Fine, B.R. Epidemiology of cerebrospinal fluid cultures and time to detection in term infants. Pediatrics 2017, 139, 8–10. [Google Scholar] [CrossRef]
- Tam, P.Y.I.; Musicha, P.; Kawaza, K.; Cornick, J.; Denis, B.; Freyne, B.; Everett, D.; Dube, Q.; French, N.; Feasey, N.; et al. Emerging resistance to empiric antimicrobial regimens for pediatric bloodstream infections in Malawi (1998–2017). Clin. Infect. Dis. 2019, 69, 61–68. [Google Scholar]
- Wangai, F.K.; Masika, M.M.; Lule, G.N.; Karari, E.M.; Maritim, M.C.; Jaoko, W.G.; Museve, B.; Kuria, A. Bridging antimicrobial resistance knowledge gaps: The East African perspective on a global problem. PLoS ONE 2019, 14, e0212131. [Google Scholar] [CrossRef]
- Medernach, R.L.; Logan, L.K. The Growing Threat of Antibiotic Resistance in Children. Infect. Dis. Clin. North Am. 2018, 32, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Maina, D.; Omuse, G.; Revathi, G.; Adam, R.D. Spectrum of microbial diseases and resistance patterns at a private teaching hospital in Kenya: Implications for clinical practice. PLoS ONE 2016, 11, e0147659. [Google Scholar] [CrossRef] [PubMed]
- Nazir, A. Neonatal sepsis due to coagulase negative Staphylococci: A study from Kashmir valley, India. Int. J. Contemp. Pediatr. 2019, 6, 650. [Google Scholar] [CrossRef]
- Venkatesh, M.P.; Placencia, F.; Weisman, L.E. Coagulase-Negative Staphylococcal Infections in the Neonate and Child: An Update. Semin. Pediatr. Infect. Dis. 2006, 17, 120–127. [Google Scholar] [CrossRef]
- Chabi, R.; Momtaz, H. Virulence factors and antibiotic resistance properties of the Staphylococcus epidermidis strains isolated from hospital infections in Ahvaz, Iran. Trop. Med. Health 2019, 47, 56. [Google Scholar] [CrossRef]
- Sahal, G.; Bilkay, I.S. Multi drug resistance in strong biofilm forming clinical isolates of Staphylococcus epidermidis. Braz. J. Microbiol. 2014, 45, 539–544. [Google Scholar] [CrossRef]
- Noshak, M.A.; Ahangarzadeh Rezaee, M.; Hasani, A.; Mirzaii, M.; Memar, M.Y.; Azimi, T.; Ebrahimi, M.; Dolatyar Dehkharghani, A. Molecular Detection and Characterization of the Staphylococcus epidermidis and Staphylococcus haemolyticus Isolated from Hospitalized Patients and Healthcare Workers in Iran. BioMed Res. Int. 2023, 2023, 3775142. [Google Scholar] [CrossRef]
- Melese, A.; Genet, C.; Andualem, T. Prevalence of Vancomycin resistant enterococci (VRE) in Ethiopia: A systematic review and meta-analysis. BMC Infect. Dis. 2020, 20, 124. [Google Scholar] [CrossRef] [PubMed]
- Alemayehu, T.; Hailemariam, M. Prevalence of vancomycin-resistant enterococcus in Africa in one health approach: A systematic review and meta-analysis. Sci. Rep. 2020, 10, 20542. [Google Scholar] [CrossRef]
- Ampaire, L.; Muhindo, A.; Orikiriza, P.; Mwanga-Amumpaire, J.; Bebell, L.; Boum, Y. A review of antimicrobial resistance in East Africa. Afr. J. Lab. Med. 2016, 5, a432. [Google Scholar] [CrossRef]
- Mouanga-Ndzime, Y.; Bisseye, C.; Longo-Pendy, N.M.; Bignoumba, M.; Dikoumba, A.C.; Onanga, R. Trends in Escherichia coli and Klebsiella pneumoniae Urinary Tract Infections and Antibiotic Resistance over a 5-Year Period in Southeastern Gabon. Antibiotics 2025, 14, 14. [Google Scholar] [CrossRef]
- Seni, J.; Peirano, G.; Mshana, S.E.; Pitout, J.D.D.; DeVinney, R. The importance of Escherichia coli clonal complex 10 and ST131 among Tanzanian patients on antimicrobial resistance surveillance programs. Eur. J. Clin. Microbiol. Infect. Dis. 2021. ahead of print. [Google Scholar] [CrossRef] [PubMed]
- Mofolorunsho, K.C.; Ocheni, H.O.; Aminu, R.F.; Omatola, C.A.; Olowonibi, O.O. Prevalence and antimicrobial susceptibility of extended-spectrum beta lactamases-producing Escherichia coli and Klebsiella pneumoniae isolated in selected hospitals of anyigba, Nigeria. Afr. Health Sci. 2021, 21, 505–512. [Google Scholar] [CrossRef] [PubMed]
- Iroha, I.R.; Onyia, U.; Moses, I.B.; Ejikeugwu, C.P.; Nwakaeze, A.E.; Ugbo, E.N. Prevalence and antibiotic resistance profiles of extended spectrum β-lactamase-producing Escherichia coli among paediatric patients with urinary tract infection in St. Patricks’ Hospital, Mile Four, Abakaliki, Ebonyi State, Nigeria. Afr. J. Clin. Exp. Microbiol. 2019, 20, 332. [Google Scholar] [CrossRef]
- Tola, M.A.; Abera, N.A.; Gebeyehu, Y.M.; Dinku, S.F.; Tullu, K.D. High prevalence of extended-spectrum betalactamase- producing Escherichia coli and Klebsiella pneumoniae fecal carriage among children under five years in Addis Ababa, Ethiopia. PLoS ONE 2021, 16, e0258117. [Google Scholar] [CrossRef]
- Omulo, S.; Oluka, M.; Achieng, L.; Osoro, E.; Kinuthia, R.; Guantai, A.; Opanga, S.A.; Ongayo, M.; Ndegwa, L.; Verani, J.R.; et al. Point-prevalence survey of antibiotic use at three public referral hospitals in Kenya. PLoS ONE 2022, 17, e0270048. [Google Scholar] [CrossRef] [PubMed]
- Momanyi, L.; Opanga, S.; Nyamu, D.; Oluka, M.; Kurdi, A.; Godman, B. Antibiotic prescribing patterns at a leading referral hospital in Kenya: A point prevalence survey. J. Res. Pharm. Pract. 2019, 8, 149. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]


| Variable | Coefficient | Odds Ratio | 95% CI | p-Value (<0.2) |
|---|---|---|---|---|
| Length of stay | 0.026 | 1.03 | 0.99–1.08 | 0.214 |
| Referral status | 0.310 | 1.36 | 0.64–2.95 | 0.425 |
| Age | −0.299 | 0.74 | 0.58–0.91 | 0.008 |
| Multiple infections | −0.310 | 0.73 | 0.34–1.57 | 0.425 |
| Comorbidity | 0.593 | 1.81 | 0.81–4.01 | 0.144 |
| Surgery | 0.041 | 1.04 | 0.28–4.97 | 0.953 |
| Multiple admissions | 1.540 | 4.47 | 1.27–30.25 | 0.045 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chuchu, V.M.; Ita, T.; Inwani, I.; Oyugi, J.; Thumbi, S.M.; Omulo, S. Diagnostic Underuse and Antimicrobial Resistance Patterns Among Hospitalized Children in a National Referral Hospital in Kenya: A Five-Year Retrospective Study. Antibiotics 2025, 14, 872. https://doi.org/10.3390/antibiotics14090872
Chuchu VM, Ita T, Inwani I, Oyugi J, Thumbi SM, Omulo S. Diagnostic Underuse and Antimicrobial Resistance Patterns Among Hospitalized Children in a National Referral Hospital in Kenya: A Five-Year Retrospective Study. Antibiotics. 2025; 14(9):872. https://doi.org/10.3390/antibiotics14090872
Chicago/Turabian StyleChuchu, Veronicah M., Teresa Ita, Irene Inwani, Julius Oyugi, S. M. Thumbi, and Sylvia Omulo. 2025. "Diagnostic Underuse and Antimicrobial Resistance Patterns Among Hospitalized Children in a National Referral Hospital in Kenya: A Five-Year Retrospective Study" Antibiotics 14, no. 9: 872. https://doi.org/10.3390/antibiotics14090872
APA StyleChuchu, V. M., Ita, T., Inwani, I., Oyugi, J., Thumbi, S. M., & Omulo, S. (2025). Diagnostic Underuse and Antimicrobial Resistance Patterns Among Hospitalized Children in a National Referral Hospital in Kenya: A Five-Year Retrospective Study. Antibiotics, 14(9), 872. https://doi.org/10.3390/antibiotics14090872






