Employing Genome Mining to Unveil a Potential Contribution of Endophytic Bacteria to Antimicrobial Compounds in the Origanum vulgare L. Essential Oil
Abstract
1. Introduction
2. Results and Discussion
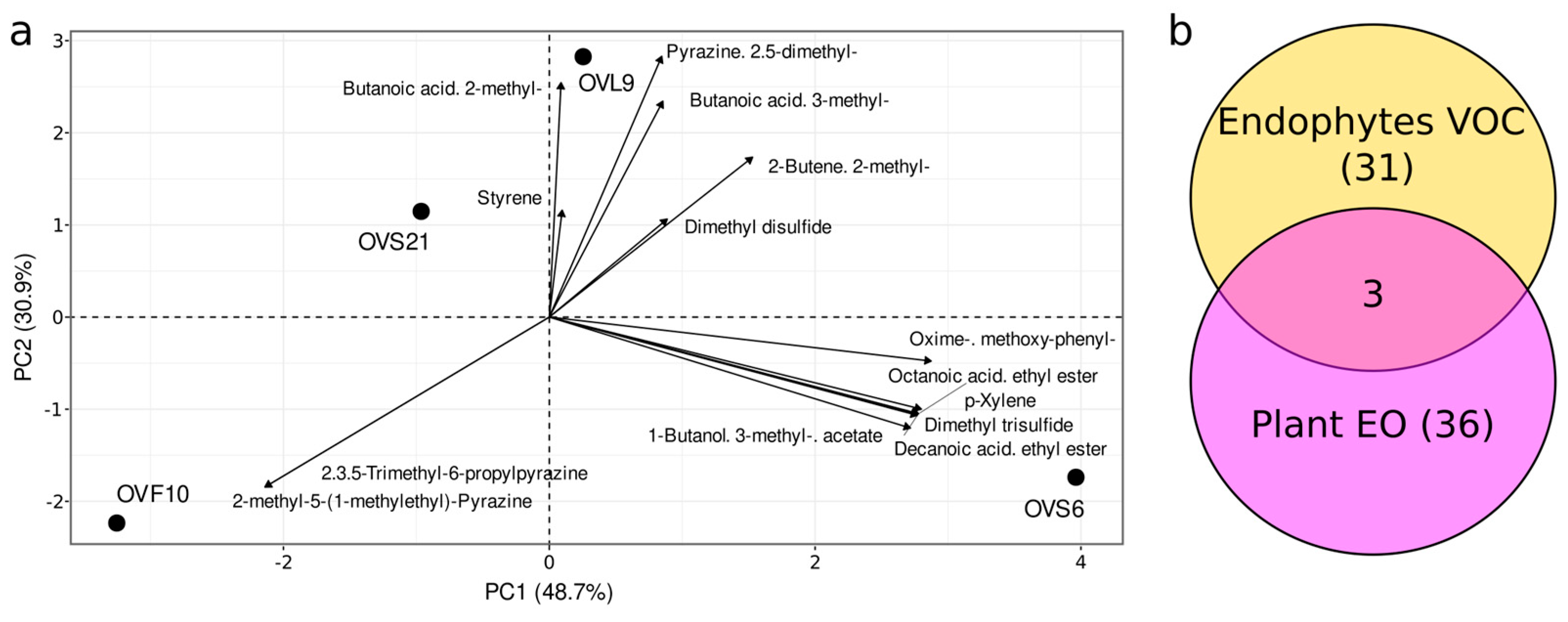
2.1. Comparison of VOCs Produced by Endophytic Strains and EO Composition
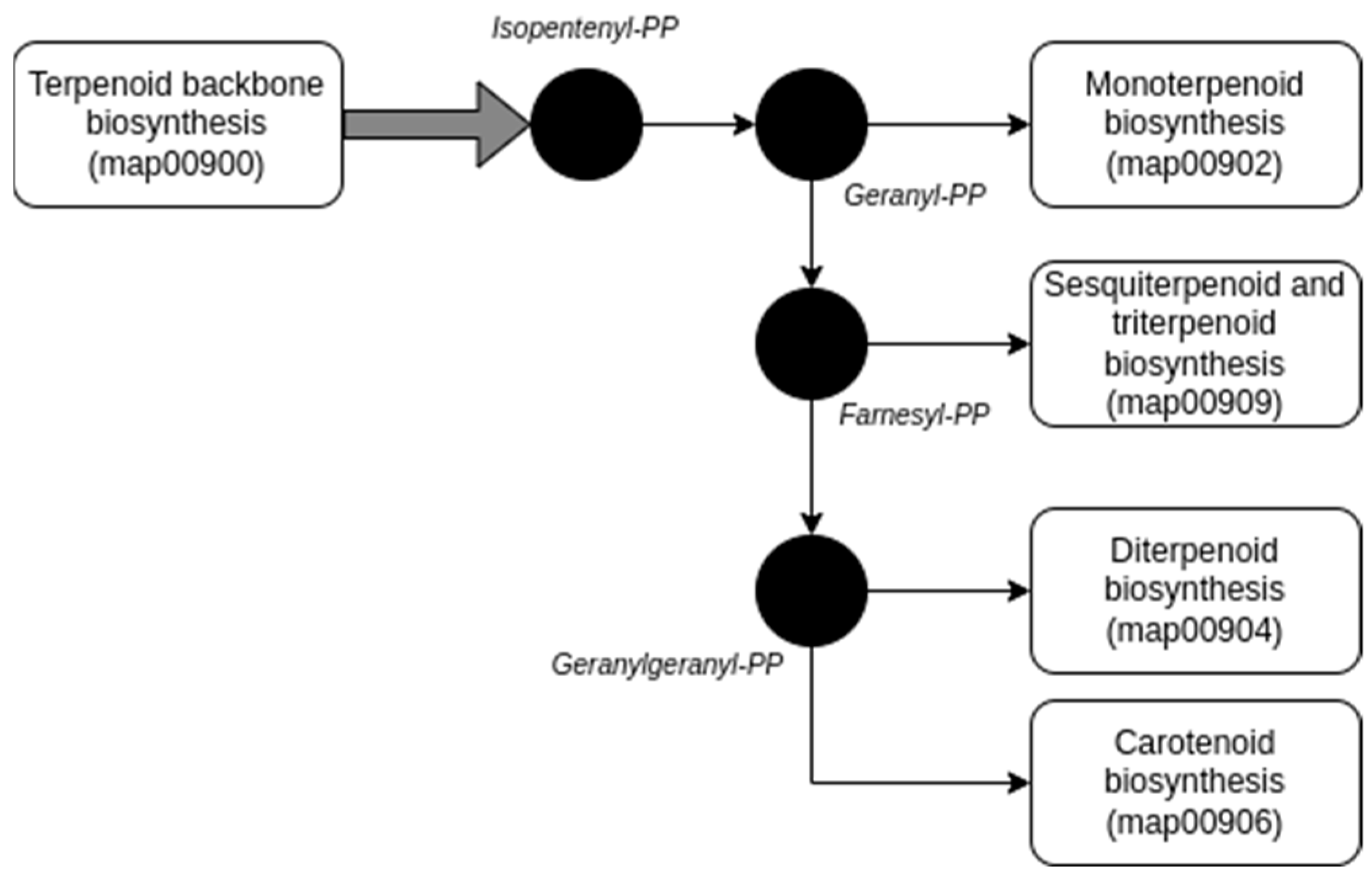
2.2. Exploring the Potential Genomic Contribution of Bacterial Endophytes to VOCs in the EO
2.2.1. Endophytes’ Genomic Basis for Plant Growth Promoting Activity
2.2.2. Strain-Level Differential Genomic Features in VOC Production and PGP Activities
3. Materials and Methods
3.1. Genome Sequencing, Assembly, and Annotation
3.2. VOC Measurements
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Langeveld, W.T.; Veldhuizen, E.J.; Burt, S.A. Synergy between essential oil components and antibiotics: A review. Crit. Rev. Microbiol. 2014, 40, 76–94. [Google Scholar] [CrossRef]
- Yap, P.S.X.; Yiap, B.C.; Ping, H.C.; Lim, S.H.E. Essential oils, a new horizon in combating bacterial antibiotic resistance. Open Microbiol. J. 2014, 8, 6. [Google Scholar] [CrossRef]
- Ayaz, M.; Sadiq, A.; Junaid, M.; Ullah, F.; Subhan, F.; Ahmed, J. Neuroprotective and anti-aging potentials of essential oils from aromatic and medicinal plants. Front. Aging Neurosci. 2017, 9, 168. [Google Scholar] [CrossRef]
- Reddy, D.N. Essential oils extracted from medicinal plants and their applications. In Natural Bio-Active Compounds: Volume 1: Production and Applications; Springer: Singapore, 2019; pp. 237–283. [Google Scholar]
- Abd El-Hack, M.E.; El-Saadony, M.T.; Saad, A.M.; Salem, H.M.; Ashry, N.M.; Ghanima, M.M.A.; Shukry, M.; Swelum, A.A.; Taha, A.E.; El-Tahan, A.M.; et al. Essential oils and their nanoemulsions as green alternatives to antibiotics in poultry nutrition: A comprehensive review. Poult. Sci. 2022, 101, 101584. [Google Scholar] [CrossRef]
- Kant, R.; Kumar, A. Review on essential oil extraction from aromatic and medicinal plants: Techniques, performance and economic analysis. Sustain. Chem. Pharm. 2022, 30, 100829. [Google Scholar] [CrossRef]
- Sharma, P.; Jambhulkar, P.P.; Raja, M.; Sain, S.K.; Javeria, S. Trichoderma spp. in consortium and their rhizospheric interactions. In Trichoderma: Host Pathogen Interactions and Applications; Springer: Singapore, 2020; pp. 267–292. [Google Scholar]
- Wińska, K.; Mączka, W.; Łyczko, J.; Grabarczyk, M.; Czubaszek, A.; Szumny, A. Essential oils as antimicrobial agents—Myth or real alternative? Molecules 2019, 24, 2130. [Google Scholar] [CrossRef]
- Ramos da Silva, L.R.; Ferreira, O.O.; Cruz, J.N.; de Jesus Pereira Franco, C.; Oliveira dos Anjos, T.; Cascaes, M.M.; Almeida da Costa, W.; Helena de Aguiar Andrade, E.; Santana de Oliveira, M. Lamiaceae essential oils, phytochemical profile, antioxidant, and biological activities. Evid. Based Complement. Alternat. Med. 2021, 2021, 6748052. [Google Scholar] [CrossRef]
- Bekut, M.; Brkić, S.; Kladar, N.; Dragović, G.; Gavarić, N.; Božin, B. Potential of selected Lamiaceae plants in anti(retro)viral therapy. Pharmacol. Res. 2018, 133, 301–314. [Google Scholar] [CrossRef]
- Soltani, S.; Shakeri, A.; Iranshahi, M.; Boozari, M. A Review of the Phytochemistry and Antimicrobial Properties of Origanum vulgare L. and Subspecies. Iran. J. Pharm. Res. 2021, 20, 268. [Google Scholar]
- Castronovo, L.M.; Calonico, C.; Ascrizzi, R.; Del Duca, S.; Delfino, V.; Chioccioli, S.; Vassallo, A.; Strozza, I.; De Leo, M.; Biffi, S.; et al. The cultivable bacterial microbiota associated to the medicinal plant Origanum vulgare L.: From antibiotic resistance to growth-inhibitory properties. Front. Microbiol. 2020, 11, 862. [Google Scholar] [CrossRef]
- Leach, J.E.; Triplett, L.R.; Argueso, C.T.; Trivedi, P. Communication in the phytobiome. Cell 2017, 169, 587–596. [Google Scholar] [CrossRef]
- Raza, W.; Shen, Q. Volatile organic compounds mediated plant-microbe interactions in soil. In Molecular Aspects of Plant Beneficial Microbes in Agriculture; Elsevier: Amsterdam, The Netherlands, 2020; pp. 209–219. [Google Scholar]
- Maggini, V.; Miceli, E.; Fagorzi, C.; Maida, I.; Fondi, M.; Perrin, E.; Mengoni, A.; Bogani, P.; Chiellini, C.; Mocali, S.; et al. Antagonism and antibiotic resistance drive a species-specific plant microbiota differentiation in Echinacea spp. FEMS Microbiol. Ecol. 2018, 94, fiy118. [Google Scholar] [CrossRef]
- Chiellini, C.; Maida, I.; Emiliani, G.; Mengoni, A.; Mocali, S.; Fabiani, A.; Biffi, S.; Maggini, V.; Gori, L.; Vannacci, A.; et al. Endophytic and rhizospheric bacterial communities isolated from the medicinal plants Echinacea purpurea and Echinacea angustifolia. Int. Microbiol. 2014, 17, 165–174. [Google Scholar]
- Emiliani, G.; Mengoni, A.; Maida, I.; Perrin, E.; Chiellini, C.; Fondi, M.; Gallo, E.; Gori, L.; Maggini, V.; Vannacci, A.; et al. Linking bacterial endophytic communities to essential oils: Clues from Lavandula angustifolia Mill. Evid.-Based Complement. Altern. Med. 2014, 2014, 650905. [Google Scholar] [CrossRef]
- Mengoni, A.; Maida, I.; Chiellini, C.; Emiliani, G.; Mocali, S.; Fabiani, A.; Fondi, M.; Firenzuoli, F.; Fani, R. Antibiotic resistance differentiates Echinacea purpurea endophytic bacterial communities with respect to plant organs. Res. Microbiol. 2014, 165, 686–694. [Google Scholar] [CrossRef]
- Maida, I.; Chiellini, C.; Mengoni, A.; Bosi, E.; Firenzuoli, F.; Fondi, M.; Fani, R. Antagonistic interactions between endophytic cultivable bacterial communities isolated from the medicinal plant E chinacea purpurea. Environ. Microbiol. 2016, 18, 2357–2365. [Google Scholar] [CrossRef]
- Polito, G.; Semenzato, G.; Del Duca, S.; Castronovo, L.M.; Vassallo, A.; Chioccioli, S.; Borsetti, D.; Calabretta, V.; Puglia, A.M.; Fani, R. Endophytic Bacteria and Essential Oil from Origanum vulgare ssp. vulgare Share Some VOCs with an Antibacterial Activity. Microorganisms 2022, 10, 1424. [Google Scholar] [CrossRef]
- Semenzato, G.; Alonso-Vásquez, T.; Del Duca, S.; Vassallo, A.; Riccardi, C.; Zaccaroni, M.; Mucci, N.; Padula, A.; Emiliani, G.; Palumbo Piccionello, A.; et al. Genomic Analysis of Endophytic Bacillus-Related Strains Isolated from the Medicinal Plant Origanum vulgare L. Revealed the Presence of Metabolic Pathways Involved in the Biosynthesis of Bioactive Compounds. Microorganisms 2022, 10, 919. [Google Scholar]
- Rehman, R.; Asif Hanif, M.; Mushtaq, Z.; Al-Sadi, A.M. Biosynthesis of essential oils in aromatic plants: A review. Food Rev. Int. 2016, 32, 117–160. [Google Scholar] [CrossRef]
- Bornowski, N.; Hamilton, J.P.; Liao, P.; Wood, J.C.; Dudareva, N.; Buell, C.R. Genome sequencing of four culinary herbs reveals terpenoid genes underlying chemodiversity in the Nepetoideae. DNA Res. 2020, 27, dsaa016. [Google Scholar] [CrossRef]
- Dagenhardt, J.; Köllner, T.G.; Gershenzon, J. Monoterpene and sesquiterpene synthases and the origin of terpene skeletal diversity in plants. Phytochemistry 2009, 70, 1621–1637. [Google Scholar] [CrossRef]
- Yamada, Y.; Kuzuyama, T.; Komatsu, M.; Shin-Ya, K.; Omura, S.; Cane, D.E.; Ikeda, H. Terpene synthases are widely distributed in bacteria. Proc. Natl. Acad. Sci. USA 2015, 112, 857–862. [Google Scholar] [CrossRef]
- Reddy, G.K.; Leferink, N.G.; Umemura, M.; Ahmed, S.T.; Breitling, R.; Scrutton, N.S.; Takano, E. Exploring novel bacterial terpene synthases. PLoS ONE 2020, 15, e0232220. [Google Scholar] [CrossRef]
- Schulz-Bohm, K.; Martín-Sánchez, L.; Garbeva, P. Microbial volatiles: Small molecules with an important role in intra-and inter-kingdom interactions. Front. Microbiol. 2017, 8, 2484. [Google Scholar] [CrossRef]
- Çakmakçı, R.; Mosber, G.; Milton, A.H.; Alatürk, F.; Ali, B. The effect of auxin and auxin-producing bacteria on the growth, essential oil yield, and composition in medicinal and aromatic plants. Curr. Microbiol. 2020, 77, 564–577. [Google Scholar] [CrossRef]
- Kutlu, M.; Cakmakci, R.; Hosseinpour, A.; Karagöz, H. The use of plant growth promoting rhizobacteria (PGPR)’s effect on essential oil rate, essential oil content, some morphological parameters and nutrient uptake of Turkish oregano. Appl. Ecol. Environ. Res. 2019, 17, 1641–1653. [Google Scholar] [CrossRef]
- Banchio, E.; Bogino, P.C.; Santoro, M.; Torres, L.; Zygadlo, J.; Giordano, W. Systemic induction of monoterpene biosynthesis in Origanum × majoricum by soil bacteria. J. Agric. Food Chem. 2010, 58, 650–654. [Google Scholar] [CrossRef]
- Mengoni, A.; Mocali, S.; Surico, G.; Tegli, S.; Fani, R. Fluctuation of endophytic bacteria and phytoplasmosis in elm trees. Microbiol. Res. 2003, 158, 363–369. [Google Scholar] [CrossRef]
- Manning, S.R. Microalgal lipids: Biochemistry and biotechnology. Curr. Opin. Biotechnol. 2022, 74, 1–7. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Kawashima, M.; Furumichi, M.; Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016, 44, D457–D462. [Google Scholar] [CrossRef]
- Luo, W.; Brouwer, C. Pathview: An R/Biocondutor package for pathway-based data integration and visualization. Bioinformatics 2013, 29, 1830–1831. [Google Scholar] [CrossRef]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.; Yu, B.; et al. PubChem 2023 update. Nucleic Acids Res. 2023, 51, D1373–D1380. [Google Scholar] [CrossRef]
- Wohlgemuth, G.; Haldiya, P.K.; Willighagen, E.; Kind, T.; Fiehn, O. The Chemical Translation Service—A web-based tool to improve standardization of metabolomic reports. Bioinformatics 2010, 26, 2647–2648. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vitali, F.; Frascella, A.; Semenzato, G.; Del Duca, S.; Palumbo Piccionello, A.; Mocali, S.; Fani, R.; Emiliani, G. Employing Genome Mining to Unveil a Potential Contribution of Endophytic Bacteria to Antimicrobial Compounds in the Origanum vulgare L. Essential Oil. Antibiotics 2023, 12, 1179. https://doi.org/10.3390/antibiotics12071179
Vitali F, Frascella A, Semenzato G, Del Duca S, Palumbo Piccionello A, Mocali S, Fani R, Emiliani G. Employing Genome Mining to Unveil a Potential Contribution of Endophytic Bacteria to Antimicrobial Compounds in the Origanum vulgare L. Essential Oil. Antibiotics. 2023; 12(7):1179. https://doi.org/10.3390/antibiotics12071179
Chicago/Turabian StyleVitali, Francesco, Arcangela Frascella, Giulia Semenzato, Sara Del Duca, Antonio Palumbo Piccionello, Stefano Mocali, Renato Fani, and Giovanni Emiliani. 2023. "Employing Genome Mining to Unveil a Potential Contribution of Endophytic Bacteria to Antimicrobial Compounds in the Origanum vulgare L. Essential Oil" Antibiotics 12, no. 7: 1179. https://doi.org/10.3390/antibiotics12071179
APA StyleVitali, F., Frascella, A., Semenzato, G., Del Duca, S., Palumbo Piccionello, A., Mocali, S., Fani, R., & Emiliani, G. (2023). Employing Genome Mining to Unveil a Potential Contribution of Endophytic Bacteria to Antimicrobial Compounds in the Origanum vulgare L. Essential Oil. Antibiotics, 12(7), 1179. https://doi.org/10.3390/antibiotics12071179











