Trends in Antibiotic Resistance of Nosocomial and Community-Acquired Infections in Italy
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Collecting Data
4.2. Collection, Identification, and Drug Susceptibility Test of Microorganisms
4.3. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- World Health Organization (WHO). Prioritization of Pathogens to Guide Discovery, Research and Development of New Antibiotics for Drug-Resistant Bacterial Infections, Including Tuberculosis; World Health Organization: Geneva, Switzerland, 2017.
- Ferri, M.; Ranucci, E.; Romagnoli, P.; Giaccone, V. Antimicrobial Resistance: A Global Emerging Threat to Public Health Systems. Crit. Rev. Food Sci. Nutr. 2017, 57, 2857–2876. [Google Scholar] [CrossRef] [PubMed]
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial Resistance: A Global Multifaceted Phenomenon. Pathog. Glob. Health 2015, 109, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Hollenbeck, B.L.; Rice, L.B. Intrinsic and Acquired Resistance Mechanisms in Enterococcus. Virulence 2012, 3, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Levy, S.B.; Bonnie, M. Antibacterial Resistance Worldwide: Causes, Challenges and Responses. Nat. Med. 2004, 10, S122–S129. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.L. Epidemiology of Drug Resistance: Implications for a Post-Antimicrobial Era. Science 1992, 257, 1050–1055. [Google Scholar] [CrossRef]
- De Oliveira, D.M.P.; Forde, B.M.; Kidd, T.J.; Harris, P.N.A.; Schembri, M.A.; Beatson, S.A.; Paterson, D.L.; Walker, M.J. Antimicrobial Resistance in ESKAPE Pathogens. Clin. Microbiol. Rev. 2020, 33, e00181-19. [Google Scholar] [CrossRef]
- Ecdc. Antimicrobial Resistance Surveillance in Europe, 2022–2020 Data; Ecdc: Solna, Sweden, 2022. [CrossRef]
- Suay-García, B.; Pérez-Gracia, M.T. Present and Future of Carbapenem-Resistant Enterobacteriaceae (CRE) Infections. Antibiotics 2019, 8, 122. [Google Scholar] [CrossRef]
- Otto, M. Community-Associated MRSA: What Makes Them Special? Int. J. Med. Microbiol. 2013, 303, 324–330. [Google Scholar] [CrossRef]
- Mediavilla, J.R.; Chen, L.; Mathema, B.; Kreiswirth, B.N. Global Epidemiology of Community-Associated Methicillin Resistant Staphylococcus Aureus (CA-MRSA). Curr. Opin. Microbiol. 2012, 15, 588–595. [Google Scholar] [CrossRef]
- Wu, C.-J.; Lee, H.-C.; Lee, N.-Y.; Shih, H.-I.; Ko, N.-Y.; Wang, L.-R.; Ko, W.-C. Predominance of Gram-Negative Bacilli and Increasing Antimicrobial Resistance in Nosocomial Bloodstream Infections at a University Hospital in Southern Taiwan, 1996–2003. J. Microbiol. Immunol. Infect. 2006, 39, 135–143. [Google Scholar]
- Llaca-Díaz, J.M.; Mendoza-Olazarán, S.; Camacho-Ortiz, A.; Flores, S.; Garza-González, E. One-Year Surveillance of ESKAPE Pathogens in an Intensive Care Unit of Monterrey, Mexico. Chemotherapy 2012, 58, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Katchanov, J.; Asar, L.; Klupp, E.-M.; Both, A.; Rothe, C.; König, C.; Rohde, H.; Kluge, S.; Maurer, F.P. Carbapenem-Resistant Gram-Negative Pathogens in a German University Medical Center: Prevalence, Clinical Implications and the Role of Novel β-Lactam/β-Lactamase Inhibitor Combinations. PLoS ONE 2018, 13, e0195757. [Google Scholar] [CrossRef] [PubMed]
- Mao, T.; Zhai, H.; Duan, G.; Yang, H. Patterns of Drug-Resistant Bacteria in a General Hospital, China, 2011–2016. Pol. J. Microbiol. 2019, 68, 225–232. [Google Scholar] [CrossRef] [PubMed]
- Spellberg, B.; Guidos, R.; Gilbert, D.; Bradley, J.; Boucher, H.W.; Scheld, W.M.; Bartlett, J.G.; Edwards, J.; Infectious Diseases Society of America. The Epidemic of Antibiotic-Resistant Infections: A Call to Action for the Medical Community from the Infectious Diseases Society of America. Clin. Infect. Dis. 2008, 46, 155–164. [Google Scholar] [CrossRef]
- Church, D.L. Major Factors Affecting the Emergence and Re-Emergence of Infectious Diseases. Clin. Lab. Med. 2004, 24, 559–586. [Google Scholar] [CrossRef]
- Mustafai, M.M.; Hafeez, M.; Munawar, S.; Basha, S.; Rabaan, A.A.; Halwani, M.A.; Alawfi, A.; Alshengeti, A.; Najim, M.A.; Alwarthan, S.; et al. Prevalence of Carbapenemase and Extended-Spectrum β-Lactamase Producing Enterobacteriaceae: A Cross-Sectional Study. Antibiotics 2023, 12, 148. [Google Scholar] [CrossRef]
- Nordmann, P.; Gniadkowski, M.; Giske, C.G.; Poirel, L.; Woodford, N.; Miriagou, V.; European Network on Carbapenemases. Identification and Screening of Carbapenemase-Producing Enterobacteriaceae. Clin. Microbiol. Infect. 2012, 18, 432–438. [Google Scholar] [CrossRef]
- Walsh, T.R. Emerging Carbapenemases: A Global Perspective. Int. J. Antimicrob. Agents 2010, 36 (Suppl. S3), S8–S14. [Google Scholar] [CrossRef]
- Joshi, S.; Shallal, A.; Zervos, M. Vancomycin-Resistant Enterococci: Epidemiology, Infection Prevention, and Control. Infect. Dis. Clin. N. Am 2021, 35, 953–968. [Google Scholar] [CrossRef]
- Fukushige, M.; Syue, L.-S.; Morikawa, K.; Lin, W.-L.; Lee, N.-Y.; Chen, P.-L.; Ko, W.-C. Trend in Healthcare-Associated Infections Due to Vancomycin-Resistant Enterococcus at a Hospital in the Era of COVID-19: More than Hand Hygiene Is Needed. J. Microbiol. Immunol. Infect. 2022, 55 Pt 2, 1211–1218. [Google Scholar] [CrossRef]
- Levitus, M.; Rewane, A.; Perera, T.B. Vancomycin-Resistant Enterococci; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Mestrovic, T.; Robles Aguilar, G.; Swetschinski, L.R.; Ikuta, K.S.; Gray, A.P.; Davis Weaver, N.; Han, C.; Wool, E.E.; Gershberg Hayoon, A.; Hay, S.I.; et al. The Burden of Bacterial Antimicrobial Resistance in the WHO European Region in 2019: A Cross-Country Systematic Analysis. Lancet Public Health 2022, 7, e897–e913. [Google Scholar] [CrossRef] [PubMed]
- Scaglione, V.; Reale, M.; Davoli, C.; Mazzitelli, M.; Serapide, F.; Lionello, R.; la Gamba, V.; Fusco, P.; Bruni, A.; Procopio, D.; et al. Prevalence of Antibiotic Resistance Over Time in a Third-Level University Hospital. Microb. Drug Resist. 2022, 28, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Suetens, C.; Latour, K.; Kärki, T.; Ricchizzi, E.; Kinross, P.; Moro, M.L.; Jans, B.; Hopkins, S.; Hansen, S.; Lyytikäinen, O.; et al. Prevalence of Healthcare-Associated Infections, Estimated Incidence and Composite Antimicrobial Resistance Index in Acute Care Hospitals and Long-Term Care Facilities: Results from Two European Point Prevalence Surveys, 2016 to 2017. Euro Surveill. 2018, 23, 1800516. [Google Scholar] [CrossRef] [PubMed]
- Ecdc. Antimicrobial Resistance in the EU/EEA (EARS-Net); Ecdc: Solna, Sweden, 2020.
- Rangel, K.; Chagas, T.P.G.; De-Simone, S.G. Acinetobacter Baumannii Infections in Times of COVID-19 Pandemic. Pathogens 2021, 10, 1006. [Google Scholar] [CrossRef] [PubMed]
- Stefanini, I.; de Renzi, G.; Foddai, E.; Cordani, E.; Mognetti, B. Profile of Bacterial Infections in COVID-19 Patients: Antimicrobial Resistance in the Time of SARS-CoV-2. Biology 2021, 10, 822. [Google Scholar] [CrossRef]
- Jiang, Y.; Ding, Y.; Wei, Y.; Jian, C.; Liu, J.; Zeng, Z. Carbapenem-Resistant Acinetobacter Baumannii: A Challenge in the Intensive Care Unit. Front. Microbiol. 2022, 13, 1045206. [Google Scholar] [CrossRef]
- Al Sulayyim, H.J.; Ismail, R.; Al Hamid, A.; Ghafar, N.A. Antibiotic Resistance during COVID-19: A Systematic Review. Int. J. Environ. Res. Public Health 2022, 19, 11931. [Google Scholar] [CrossRef]
- Langford, B.J.; Soucy, J.-P.R.; Leung, V.; So, M.; Kwan, A.T.H.; Portnoff, J.S.; Bertagnolio, S.; Raybardhan, S.; MacFadden, D.R.; Daneman, N. Antibiotic Resistance Associated with the COVID-19 Pandemic: A Systematic Review and Meta-Analysis. Clin. Microbiol. Infect. 2023, 29, 302–309. [Google Scholar] [CrossRef]
- Magnasco, L.; Mikulska, M.; Giacobbe, D.R.; Taramasso, L.; Vena, A.; Dentone, C.; Dettori, S.; Tutino, S.; Labate, L.; di Pilato, V.; et al. Spread of Carbapenem-Resistant Gram-Negatives and Candida Auris during the COVID-19 Pandemic in Critically Ill Patients: One Step Back in Antimicrobial Stewardship? Microorganisms 2021, 9, 95. [Google Scholar] [CrossRef]
- Serwecińska, L. Antimicrobials and Antibiotic-Resistant Bacteria: A Risk to the Environment and to Public Health. Water 2020, 12, 3313. [Google Scholar] [CrossRef]
- Polidori, I.; Marullo, L.; Ialongo, C.; Tomassetti, F.; Colombo, R.; di Gaudio, F.; Calugi, G.; Marrone, G.; Noce, A.; Bernardini, S.; et al. Characterization of Gut Microbiota Composition in Type 2 Diabetes Patients: A Population-Based Study. Int. J. Environ. Res. Public Health 2022, 19, 15913. [Google Scholar] [CrossRef] [PubMed]
- Langdon, A.; Schwartz, D.J.; Bulow, C.; Sun, X.; Hink, T.; Reske, K.A.; Jones, C.; Burnham, C.-A.D.; Dubberke, E.R.; Dantas, G. Microbiota Restoration Reduces Antibiotic-Resistant Bacteria Gut Colonization in Patients with Recurrent Clostridioides Difficile Infection from the Open-Label PUNCH CD Study. Genome Med. 2021, 13, 28. [Google Scholar] [CrossRef]
- Wang, X.; Ryu, D.; Houtkooper, R.H.; Auwerx, J. Antibiotic Use and Abuse: A Threat to Mitochondria and Chloroplasts with Impact on Research, Health, and Environment. Bioessays 2015, 37, 1045–1053. [Google Scholar] [CrossRef]
- Rhedin, S.; Elfving, K.; Berggren, A. Novel Biomarkers Differentiating Viral from Bacterial Infection in Febrile Children: Future Perspectives for Management in Clinical Praxis. Children 2021, 8, 1070. [Google Scholar] [CrossRef] [PubMed]
- Nicolai, E.; Pieri, M.; Gratton, E.; Motolese, G.; Bernardini, S. Bacterial Infection Diagnosis and Antibiotic Prescription in 3 h as an Answer to Antibiotic Resistance: The Case of Urinary Tract Infections. Antibiotics 2021, 10, 1168. [Google Scholar] [CrossRef] [PubMed]
- Nicolai, E.; Garau, S.; Favalli, C.; D’Agostini, C.; Gratton, E.; Motolese, G.; Rosato, N. Evaluation of BiesseBioscreen as a new methodology for bacteriuria screening. New Microbiol. 2014, 37, 495–501. [Google Scholar]
- Toosky, M.N.; Grunwald, J.T.; Pala, D.; Shen, B.; Zhao, W.; D’Agostini, C.; Coghe, F.; Angioni, G.; Motolese, G.; Abram, T.; et al. A rapid, point-of-care antibiotic susceptibility test for urinary tract infections. J. Med. Microbiol. 2020, 69, 52–62. [Google Scholar] [CrossRef]
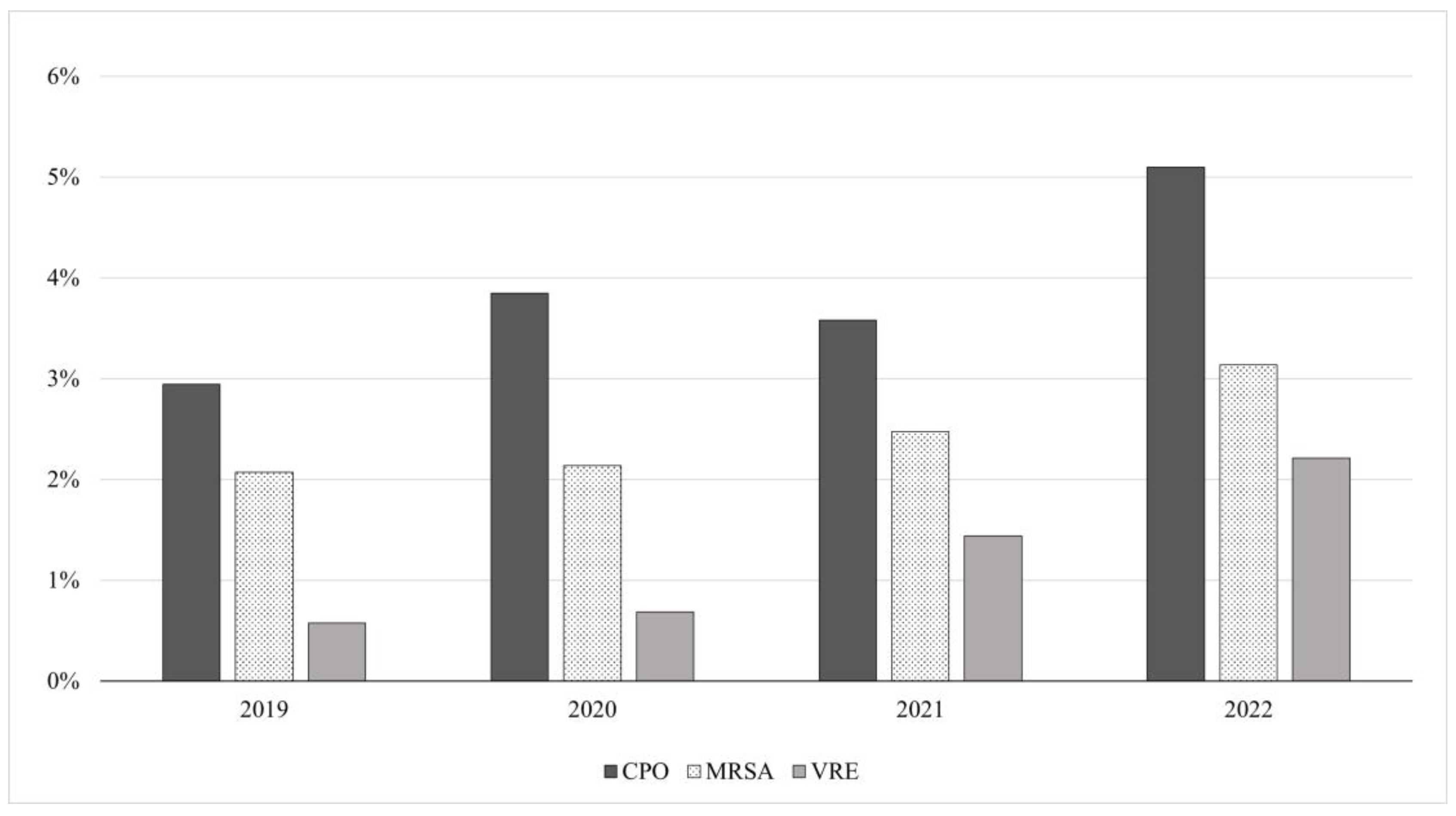

| 2019 | 2020 | 2021 | 2022 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Total | Positive | % | Total | Positive | % | Total | Positive | % | Total | Positive | % | |
| CPO | 17,092 | 564 | 2.62% | 20,504 | 895 | 3.43% | 24,494 | 1009 | 320% | 26,734 | 1570 | 4.56% |
| MRSA | 2069 | 397 | 1.84% | 2745 | 497 | 1.90% | 3686 | 697 | 2.21% | 4062 | 966 | 2.81% |
| VRE | 2385 | 124 | 0.58% | 2850 | 179 | 0.69% | 3400 | 455 | 1.44% | 3614 | 761 | 2.21% |
| TOTAL | 21.546 | 26.099 | 31.580 | 34.410 | ||||||||
| 2019 | 2020 | 2021 | 2022 | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Nosocomial | Community | χ2 Test | p-Value | Nosocomial | Community | χ2 Test | p-Value | Nosocomial | Community | χ2 Test | p-Value | Nosocomial | Community | χ2 Test | p-Value | |
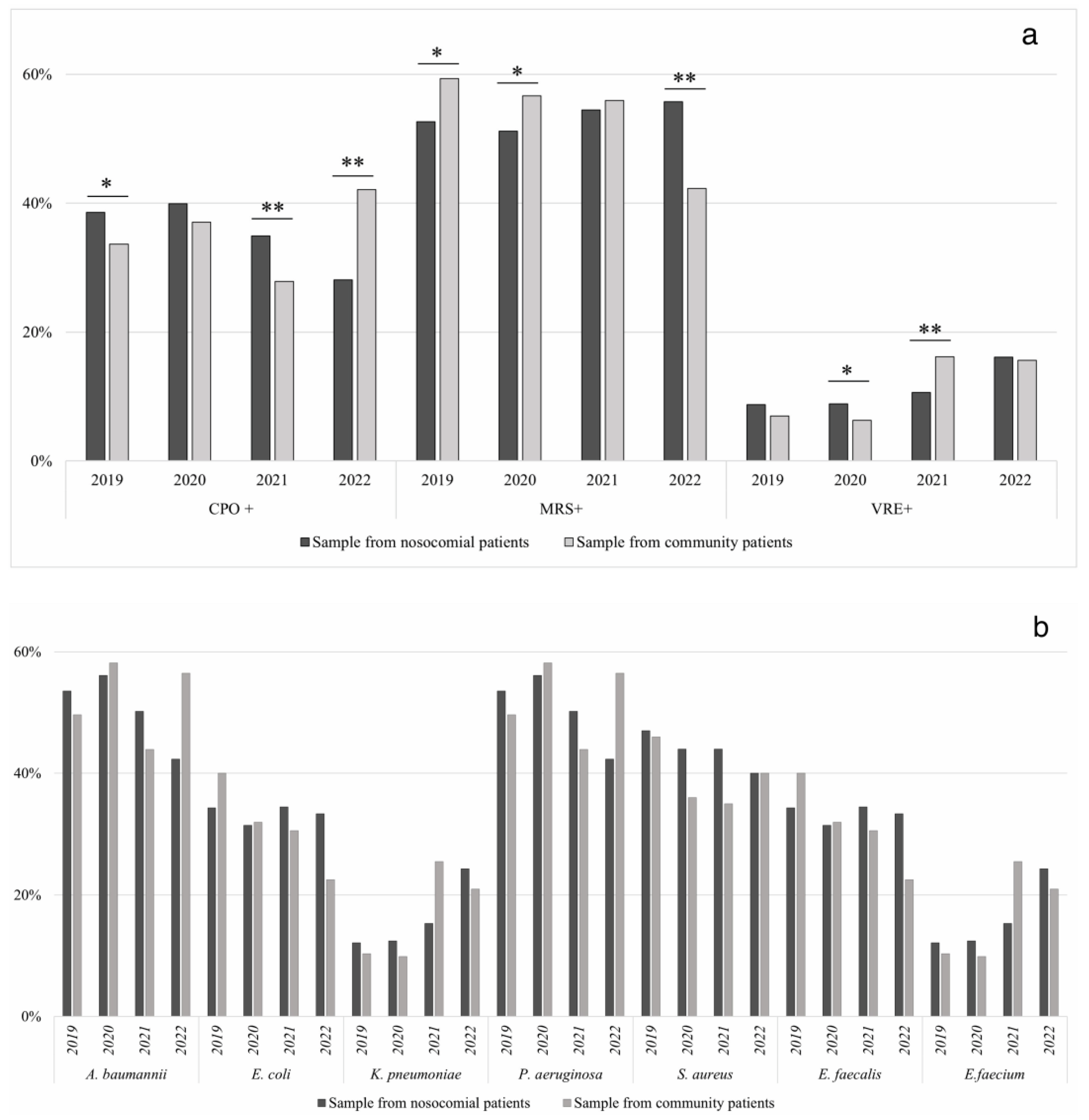
| CPO+ | 39% | 34% | 3.94 | p < 0.05 | 40% | 37% | 1.98 | p: ns | 35% | 28% | 18.46 | p < 0.001 | 28% | 42% | 95.44 | p < 0.001 |
| Acinetobacter baumannii | 37% | 36% | 0.02 | p: ns | 30% | 29% | 0.20 | p: ns | 20% | 24% | 2.30 | p: ns | 24% | 4% | 121.60 | p < 0.001 |
| Escherichia coli | 9% | 7% | 0.54 | p: ns | 7% | 0% | 28.09 | p < 0.001 | 4% | 4% | 0.05 | p: ns | 6% | 1% | 19.18 | p < 0.001 |
| Klebsiella pneumoniae | 41% | 39% | 0.34 | p: ns | 43% | 52% | 7.11 | p < 0.05 | 56% | 51% | 2.56 | p: ns | 51% | 76% | 102.01 | p < 0.001 |
| Pseudomonas aeruginosa | 13% | 18% | 2.62 | p: ns | 19% | 19% | 0.03 | p: ns | 20% | 21% | 0.28 | p: ns | 19% | 19% | 0.05 | p: ns |
| MRSA+ | 53% | 59% | 6.79 | p < 0.05 | 51% | 57% | 10.06 | p < 0.05 | 54% | 56% | 0.73 | p: ns | 56% | 42% | 78.55 | p < 0.001 |
| Staphilococcus aureus | 47% | 46% | 0.14 | p: ns | 44% | 36% | 7.70 | p < 0.05 | 44% | 35% | 4.31 | p < 0.05 | 40% | 40% | 0.00 | p: ns |
| VRE+ | 9% | 7% | 1.55 | p: ns | 9% | 6% | 2.11 | p: ns | 11% | 16% | 20.30 | p < 0.001 | 16% | 16% | 0.19 | p: ns |
| Enterococcus faecalis | 51% | 64% | 0.61 | p: ns | 53% | 59% | 0.19 | p: ns | 55% | 34% | 16.31 | p < 0.001 | 17% | 29% | 10.98 | p < 0.001 |
| Enterococcus faecium | 49% | 36% | 0.87 | p: ns | 47% | 41% | 0.25 | p: ns | 45% | 66% | 4.87 | p < 0.05 | 83% | 71% | 2.07 | p: ns |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cerini, P.; Meduri, F.R.; Tomassetti, F.; Polidori, I.; Brugneti, M.; Nicolai, E.; Bernardini, S.; Pieri, M.; Broccolo, F. Trends in Antibiotic Resistance of Nosocomial and Community-Acquired Infections in Italy. Antibiotics 2023, 12, 651. https://doi.org/10.3390/antibiotics12040651
Cerini P, Meduri FR, Tomassetti F, Polidori I, Brugneti M, Nicolai E, Bernardini S, Pieri M, Broccolo F. Trends in Antibiotic Resistance of Nosocomial and Community-Acquired Infections in Italy. Antibiotics. 2023; 12(4):651. https://doi.org/10.3390/antibiotics12040651
Chicago/Turabian StyleCerini, Paola, Francesca Rita Meduri, Flaminia Tomassetti, Isabella Polidori, Marta Brugneti, Eleonora Nicolai, Sergio Bernardini, Massimo Pieri, and Francesco Broccolo. 2023. "Trends in Antibiotic Resistance of Nosocomial and Community-Acquired Infections in Italy" Antibiotics 12, no. 4: 651. https://doi.org/10.3390/antibiotics12040651
APA StyleCerini, P., Meduri, F. R., Tomassetti, F., Polidori, I., Brugneti, M., Nicolai, E., Bernardini, S., Pieri, M., & Broccolo, F. (2023). Trends in Antibiotic Resistance of Nosocomial and Community-Acquired Infections in Italy. Antibiotics, 12(4), 651. https://doi.org/10.3390/antibiotics12040651









