Investigation of In Vitro Susceptibility and Resistance Mechanisms in Skin Pathogens: Perspectives for Fluoroquinolone Therapy in Canine Pyoderma
Abstract
1. Introduction
2. Results
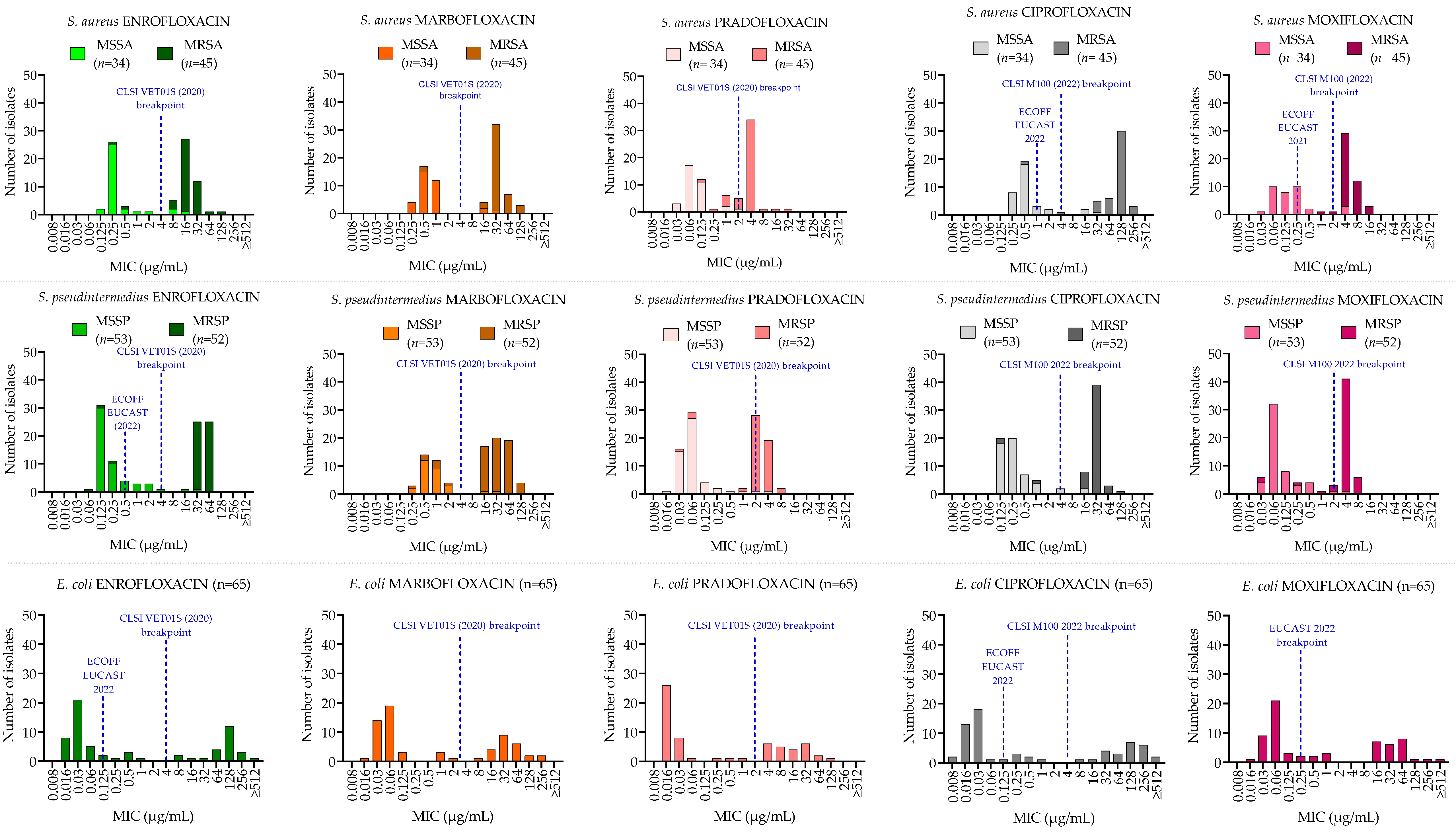
2.1. Minimum Inhibitory Concentration
2.2. Minimum Bactericidal Concentration
2.3. Resistance Mechanism Detection
3. Discussion
4. Materials and Methods
4.1. Bacterial Pathogens
4.2. Antimicrobial Agents Tested
4.3. Minimum Inhibitory and Bactericidal Concentrations
4.4. Efflux Pump Overexpression
4.5. Chromosomal Mutations and PMQR Genes
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hillier, A.; Alcorn, J.R.; Cole, L.K.; Kowalski, J.J. Pyoderma caused by Pseudomonas aeruginosa infection in dogs: 20 cases. Vet. Dermatol. 2006, 17, 432–439. [Google Scholar] [CrossRef] [PubMed]
- Gortel, K. Recognizing pyoderma: More difficult than it may seem. Vet. Clin. Small Anim. 2013, 43, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Petersen, A.D.; Walker, R.D.; Bowman, M.M.; Schott, H.C.; Rosser, E.J., Jr. Frequency of isolation and antimicrobial susceptibility patterns of Staphylococcus intermedius and Pseudomonas aeruginosa isolates from canine skin and ear samples over a 6-year period (1992–1997). J. Am. Anim. Hosp. Assoc. 2002, 38, 407–413. [Google Scholar] [CrossRef] [PubMed]
- Summers, J.F.; Brodbelt, D.C.; Forsythe, P.J.; Loeffler, A.; Hendricks, A. The effectiveness of systemic antimicrobial treatment in canine superficial and deep pyoderma: A systematic review. Vet. Dermatol. 2012, 23, 305-e61. [Google Scholar] [CrossRef] [PubMed]
- Frank, L.A.; Loeffler, A. Meticillin-resistant Staphylococcus pseudintermedius: Clinical challenge and treatment options. Vet. Dermatol. 2012, 23, 256–283. [Google Scholar] [CrossRef]
- Loncaric, I.; Lepuschitz, S.; Ruppitsch, W.; Trstan, A.; Andreadis, T.; Bouchlis, N.; Marbach, H.; Schauer, B.; Szostak, M.P.; Feßler, A.T. Increased genetic diversity of methicillin-resistant Staphylococcus aureus (MRSA) isolated from companion animals. Vet. Microbiol. 2019, 235, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Morris, D.O.; Loeffler, A.; Davis, M.F.; Guardabassi, L.; Weese, J.S. Recommendations for approaches to methicillin-resistant staphylococcal infections of small animals: Diagnosis, therapeutic considerations and preventative measures. Clinical Consensus Guidelines of the World Association for Veterinary Dermatology. Vet. Dermatol. 2017, 28, 304-e69. [Google Scholar] [CrossRef]
- Lord, J.; Millis, N.; Jones, R.D.; Johnson, B.; Kania, S.A. Patterns of antimicrobial, multidrug and methicillin resistance among Staphylococcus spp. isolated from canine specimens submitted to a diagnostic laboratory in Tennessee, USA: A descriptive study. BMC Vet. Res. 2022, 18, 1–16. [Google Scholar] [CrossRef]
- Ishii, S.; Meyer, K.P.; Sadowsky, M.J. Relationship between phylogenetic groups, genotypic clusters, and virulence gene profiles of Escherichia coli strains from diverse human and animal sources. Appl. Environ. Microbiol. 2007, 73, 5703–5710. [Google Scholar] [CrossRef]
- Johnson, J.R.; Clabots, C.; Kuskowski, M.A. Multiple-host sharing, long-term persistence, and virulence of Escherichia coli clones from human and animal household members. J. Clin. Microbiol. 2008, 46, 4078–4082. [Google Scholar] [CrossRef] [PubMed]
- Platell, J.L.; Cobbold, R.N.; Johnson, J.R.; Heisig, A.; Heisig, P.; Clabots, C.; Kuskowski, M.A.; Trott, D.J. Commonality among fluoroquinolone-resistant sequence type ST131 extraintestinal Escherichia coli isolates from humans and companion animals in Australia. Antimicrob. Agents Chemother. 2011, 55, 3782–3787. [Google Scholar] [CrossRef]
- Carlotti, D.N.; Guaguere, E.; Pin, D.; Jasmin, P.; Thomas, E.; Guiral, V. Therapy of difficult cases of canine pyoderma with marbofloxacin: A report of 39 dogs. J. Small Anim. Pract. 1999, 40, 265–270. [Google Scholar] [CrossRef]
- Paradis, M.; Abbey, L.; Baker, B.; Coyne, M.; Hannigan, M.; Joffe, D.; Pukay, B.; Trettien, A.; Waisglass, S.; Wellington, J. Evaluation of the clinical efficacy of marbofloxacin (Zeniquin) tablets for the treatment of canine pyoderma: An open clinical trial. Vet. Dermatol. 2001, 12, 163–169. [Google Scholar] [CrossRef]
- Horspool, L.J.I.; Van Laar, P.; Van Den Bos, R.; Mawhinney, I. Treatment of canine pyoderma with ibafloxacin and marbofloxacin–fluoroquinolones with different pharmacokinetic profiles. J. Vet. Pharmacol. Ther. 2004, 27, 147–153. [Google Scholar] [CrossRef]
- Mueller, R.S.; Stephan, B. Pradofloxacin in the treatment of canine deep pyoderma: A multicentred, blinded, randomized parallel trial. Vet. Dermatol. 2007, 18, 144–151. [Google Scholar] [CrossRef]
- Restrepo, C.; Ihrke, P.J.; White, S.D.; Spiegel, I.B.; Affolter, V.K. Evaluation of the clinical efficacy of pradofloxacin tablets for the treatment of canine pyoderma. J. Am. Anim. Hosp. Assoc. 2010, 46, 301–311. [Google Scholar] [CrossRef]
- Beco, L.; Guaguere, E.; Méndez, C.L.; Noli, C.; Nuttall, T.; Vroom, M. Suggested guidelines for using systemic antimicrobials in bacterial skin infections: Part 2—Antimicrobial choice, treatment regimens and compliance. Vet. Rec. 2013, 172, 156. [Google Scholar] [CrossRef] [PubMed]
- Hillier, A.; Lloyd, D.H.; Weese, J.S.; Blondeau, J.M.; Boothe, D.; Breitschwerdt, E.; Guardabassi, L.; Papich, M.G.; Rankin, S.; Turnidge, J.D. Guidelines for the diagnosis and antimicrobial therapy of canine superficial bacterial folliculitis (A ntimicrobial G uidelines W orking G roup of the I nternational S ociety for C ompanion A nimal I nfectious D iseases). Vet. Dermatol. 2014, 25, 163-e43. [Google Scholar] [CrossRef]
- European Medicine Agency. Categorisation of Antibiotics in the European Union. Available online: https://www.ema.europa.eu/en/documents/report/categorisation-antibiotics-european-union-answer-request-european-commission-updating-scientific_en.pdf (accessed on 26 August 2022).
- World Health Organization (WHO). Critically Important Antimicrobials for Human Medicine—6th Revision 2018—Ranking of Medically Important Antimicrobials for Risk Management of Antimicrobial Resistance Due to Non-Human Use. Available online: https://www.who.int/publications/i/item/9789241515528 (accessed on 26 August 2022).
- Ihrke, P.J.; Papich, M.G.; Demanuelle, T.C. The use of fluoroquinolones in veterinary dermatology. Vet. Dermatol. 1999, 10, 193–204. [Google Scholar] [CrossRef]
- Authier, S.; Paquette, D.; Labrecque, O.; Messier, S. Comparison of susceptibility to antimicrobials of bacterial isolates from companion animals in a veterinary diagnostic laboratory in Canada between 2 time points 10 years apart. Can. Vet. J. 2006, 47, 774. [Google Scholar]
- Guardabassi, L.; Schwarz, S.; Lloyd, D.H. Pet animals as reservoirs of antimicrobial-resistant bacteria. J. Antimicrob. Chemother. 2004, 54, 321–332. [Google Scholar] [CrossRef] [PubMed]
- Piddock, L.J.V. Mechanisms of fluoroquinolone resistance: An update 1994–1998. Drugs 1999, 58, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.C. Mechanisms of action and resistance of older and newer fluoroquinolones. Clin. Infect. Dis. 2000, 31, S24–S28. [Google Scholar] [CrossRef] [PubMed]
- Ng, E.Y.; Trucksis, M.; Hooper, D.C. Quinolone resistance mediated by norA: Physiologic characterization and relationship to flqB, a quinolone resistance locus on the Staphylococcus aureus chromosome. Antimicrob. Agents Chemother. 1994, 38, 1345–1355. [Google Scholar] [CrossRef]
- Everett, M.J.; Jin, Y.F.; Ricci, V.; Piddock, L.J. Contributions of individual mechanisms to fluoroquinolone resistance in 36 Escherichia coli strains isolated from humans and animals. Antimicrob. Agents Chemother. 1996, 40, 2380–2386. [Google Scholar] [CrossRef] [PubMed]
- Périchon, B.; Courvalin, P.; Galimand, M. Transferable resistance to aminoglycosides by methylation of G1405 in 16S rRNA and to hydrophilic fluoroquinolones by QepA-mediated efflux in Escherichia coli. Antimicrob. Agents Chemother. 2007, 51, 2464–2469. [Google Scholar] [CrossRef] [PubMed]
- Yamane, K.; Wachino, J.; Suzuki, S.; Kimura, K.; Shibata, N.; Kato, H.; Shibayama, K.; Konda, T.; Arakawa, Y. New plasmid-mediated fluoroquinolone efflux pump, QepA, found in an Escherichia coli clinical isolate. Antimicrob. Agents Chemother. 2007, 51, 3354–3360. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.B.; Wang, M.; Park, C.H.; Kim, E.C.; Jacoby, G.A.; Hooper, D.C. oqxAB encoding a multidrug efflux pump in human clinical isolates of Enterobacteriaceae. Antimicrob. Agents Chemother. 2009, 53, 3582–3584. [Google Scholar] [CrossRef]
- Gebru Awji, E.; Tassew, D.D.; Lee, J.S.; Lee, S.J.; Choi, M.J.; Reza, M.A.; Rhee, M.H.; Kim, T.H.; Park, S.C. Comparative mutant prevention concentration and mechanism of resistance to veterinary fluoroquinolones in Staphylococcus pseudintermedius. Vet. Dermatol. 2012, 23, 376-e69. [Google Scholar] [CrossRef]
- Shimizu, T.; Harada, K.; Kataoka, Y. Mutant prevention concentration of orbifloxacin: Comparison between Escherichia coli, Pseudomonas aeruginosa, and Staphylococcus pseudintermedius of canine origin. Acta Vet. Scand. 2013, 55, 37. [Google Scholar] [CrossRef] [PubMed]
- Robicsek, A.; Strahilevitz, J.; Jacoby, G.A.; Macielag, M.; Abbanat, D.; Park, C.H.; Bush, K.; Hooper, D.C. Fluoroquinolone-modifying enzyme: A new adaptation of a common aminoglycoside acetyltransferase. Nat. Med. 2006, 12, 83–88. [Google Scholar] [CrossRef] [PubMed]
- Xiong, X.; Bromley, E.H.C.; Oelschlaeger, P.; Woolfson, D.N.; Spencer, J. Structural insights into quinolone antibiotic resistance mediated by pentapeptide repeat proteins: Conserved surface loops direct the activity of a Qnr protein from a gram-negative bacterium. Nucleic Acids Res. 2011, 39, 3917–3927. [Google Scholar] [CrossRef] [PubMed]
- Redgrave, L.S.; Sutton, S.B.; Webber, M.A.; Piddock, L.J.V. Fluoroquinolone resistance: Mechanisms, impact on bacteria, and role in evolutionary success. Trends Microbiol. 2014, 22, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Shaheen, B.W.; Boothe, D.M.; Oyarzabal, O.A.; Wang, C.; Johnson, C.M. Evaluation of the contribution of gyrA mutation and efflux pumps to fluoroquinolone and multidrug resistance in pathogenic Escherichia coli isolates from dogs and cats. Am. J. Vet. Res. 2011, 72, 25–32. [Google Scholar] [CrossRef]
- Shaheen, B.W.; Nayak, R.; Foley, S.L.; Boothe, D.M. Chromosomal and plasmid-mediated fluoroquinolone resistance mechanisms among broad-spectrum-cephalosporin-resistant Escherichia coli isolates recovered from companion animals in the USA. J. Antimicrob Chemother 2013, 68, 1019–1024. [Google Scholar] [CrossRef]
- Vingopoulou, E.I.; Delis, G.A.; Batzias, G.C.; Kaltsogianni, F.; Koutinas, A.; Kristo, I.; Pournaras, S.; Saridomichelakis, M.N.; Siarkou, V.I. Prevalence and mechanisms of resistance to fluoroquinolones in Pseudomonas aeruginosa and Escherichia coli isolates recovered from dogs suffering from otitis in Greece. Vet. Microbiol. 2018, 213, 102–107. [Google Scholar] [CrossRef]
- Ruiz-Ripa, L.; Simón, C.; Ceballos, S.; Ortega, C.; Zarazaga, M.; Torres, C.; Gómez-Sanz, E. S. pseudintermedius and S. aureus lineages with transmission ability circulate as causative agents of infections in pets for years. BMC Vet. Res. 2021, 17, 1–10. [Google Scholar] [CrossRef]
- Saputra, S.; Jordan, D.; Worthing, K.A.; Norris, J.M.; Wong, H.S.; Abraham, R.; Trott, D.J.; Abraham, S. Antimicrobial resistance in coagulase-positive staphylococci isolated from companion animals in Australia: A one year study. PLoS ONE 2017, 12, e0176379. [Google Scholar] [CrossRef]
- Priyantha, R.; Gaunt, M.C.; Rubin, J.E. Antimicrobial susceptibility of Staphylococcus pseudintermedius colonizing healthy dogs in Saskatoon, Canada. Can. Vet. J. 2016, 57, 65. [Google Scholar]
- Schink, A.K.; Kadlec, K.; Hauschild, T.; Brenner Michael, G.; Dorner, J.C.; Ludwig, C.; Werckenthin, C.; Hehnen, H.R.; Stephan, B.; Schwarz, S. Susceptibility of canine and feline bacterial pathogens to pradofloxacin and comparison with other fluoroquinolones approved for companion animals. Vet. Microbiol. 2013, 162, 119–126. [Google Scholar] [CrossRef]
- Meunier, D.d.; Acar, J.F.; Martel, J.L.; Kroemer, S.; Valle, M. A seven-year survey of susceptibility to marbofloxacin of pathogenic strains isolated from pets. Int. J. Antimicrob. Agents 2004, 24, 592–598. [Google Scholar] [CrossRef] [PubMed]
- Ganière, J.P.; Médaille, C.; Limet, A.; Ruvoen, N.; André-Fontaine, G. Antimicrobial activity of enrofloxacin against Staphylococcus intermedius strains isolated from canine pyodermas. Vet. Dermatol. 2001, 12, 171–175. [Google Scholar] [CrossRef] [PubMed]
- Descloux, S.; Rossano, A.; Perreten, V. Characterization of new staphylococcal cassette chromosome mec (SCC mec) and topoisomerase genes in fluoroquinolone-and methicillin-resistant Staphylococcus pseudintermedius. J. Clin. Microbiol. 2008, 46, 1818–1823. [Google Scholar] [CrossRef] [PubMed]
- Ganiere, J.P.; Medaille, C.; Mangion, C. Antimicrobial drug susceptibility of Staphylococcus intermedius clinical isolates from canine pyoderma. J. Vet. Med. Ser. B 2005, 52, 25–31. [Google Scholar] [CrossRef]
- Peterson, L.R. Quinolone molecular structure-activity relationships: What we have learned about improving antimicrobial activity. Clin. Infect. Dis. 2001, 33, S180–S186. [Google Scholar] [CrossRef]
- Drlica, K.; Malik, M. Fluoroquinolones: Action and resistance. Curr. Top. Med. Chem. 2003, 3, 249–282. [Google Scholar] [CrossRef]
- Drlica, K.; Malik, M.; Kerns, R.J.; Zhao, X. Quinolone-mediated bacterial death. Antimicrob. Agents Chemother. 2008, 52, 385–392. [Google Scholar] [CrossRef]
- Wetzstein, H.G. Comparative mutant prevention concentrations of pradofloxacin and other veterinary fluoroquinolones indicate differing potentials in preventing selection of resistance. Antimicrob. Agents Chemother. 2005, 49, 4166–4173. [Google Scholar] [CrossRef]
- Plowgian, C.; Blondeau, J.M.; Levinson, M.; Rosenkrantz, W. A pilot study on the comparative minimum inhibitory and mutant prevention concentration values for moxifloxacin and pradofloxacin against canine and human isolates of Staphylococcus pseudintermedius and S. schleiferi. Vet. Dermatol. 2019, 30, 481-e142. [Google Scholar] [CrossRef]
- Lees, P. Pharmacokinetics, pharmacodynamics and therapeutics of pradofloxacin in the dog and cat. J. Vet. Pharmacol. Ther. 2013, 36, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Messias, A.; Gekeler, F.; Wegener, A.; Dietz, K.; Kohler, K.; Zrenner, E. Retinal safety of a new fluoroquinolone, pradofloxacin, in cats: Assessment with electroretinography. Doc. Ophthalmol. 2008, 116, 177–191. [Google Scholar] [CrossRef] [PubMed]
- Silley, P.; Stephan, B.; Greife, H.A.; Pridmore, A. Bactericidal properties of pradofloxacin against veterinary pathogens. Vet. Microbiol. 2012, 157, 106–111. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.-T.; Wang, X.-M.; Liao, X.-P.; Sun, J.; Zhu, H.-Q.; Chen, X.-Y.; Liu, Y.-H. Plasmid-mediated quinolone resistance determinants oqxAB and aac (6′)-Ib-cr and extended-spectrum β-lactamase gene bla CTX-M-24 co-located on the same plasmid in one Escherichia coli strain from China. J. Antimicrob. Chemother. 2011, 66, 1638–1639. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Boothe, D.M.; Jin, Y.; Thungrat, K. In vitro potency and efficacy favor later generation fluoroquinolones for treatment of canine and feline Escherichia coli uropathogens in the United States. World J. Microbiol. Biotechnol. 2013, 29, 347–354. [Google Scholar] [CrossRef]
- Ludwig, C.; de Jong, A.; Moyaert, H.; El Garch, F.; Janes, R.; Klein, U.; Morrissey, I.; Thiry, J.; Youala, M. Antimicrobial susceptibility monitoring of dermatological bacterial pathogens isolated from diseased dogs and cats across Europe (ComPath results). J. Appl. Microbiol. 2016, 121, 1254–1267. [Google Scholar] [CrossRef]
- Dancer, S.J. The effect of antibiotics on methicillin-resistant Staphylococcus aureus. J. Antimicrob. Chemother. 2007, 61, 246–253. [Google Scholar] [CrossRef]
- Harbarth, S.; Fankhauser, C.; Schrenzel, J.; Christenson, J.; Gervaz, P.; Bandiera-Clerc, C.; Renzi, G.; Vernaz, N.; Sax, H.; Pittet, D. Universal screening for methicillin-resistant Staphylococcus aureus at hospital admission and nosocomial infection in surgical patients. JAMA 2008, 299, 1149–1157. [Google Scholar] [CrossRef]
- Bemis, D.A.; Jones, R.D.; Frank, L.A.; Kania, S.A. Evaluation of susceptibility test breakpoints used to predict mecA-mediated resistance in Staphylococcus pseudintermedius isolated from dogs. J. Vet. Diagn. Investig. 2009, 21, 53–58. [Google Scholar] [CrossRef]
- Berrington, A.W.; Perry, J.D.; Gould, F.K. Bactericidal activity of moxifloxacin against Staphylococcus aureus. Clin. Microbiol. Infect. 2001, 7, 161–163. [Google Scholar] [CrossRef][Green Version]
- Hansen, G.T.; Blondeau, J.M. Comparison of the minimum inhibitory, mutant prevention and minimum bactericidal concentrations of ciprofloxacin, levofloxacin and garenoxacin against enteric Gram-negative urinary tract infection pathogens. J. Chemother. 2005, 17, 484–492. [Google Scholar] [CrossRef]
- Smith, S.M.; Eng, R.H. Activity of ciprofloxacin against methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 1985, 27, 688–691. [Google Scholar] [CrossRef] [PubMed]
- Intorre, L.; Vanni, M.; Di Bello, D.; Pretti, C.; Meucci, V.; Tognetti, R.; Soldani, G.; Cardini, G.; Jousson, O. Antimicrobial susceptibility and mechanism of resistance to fluoroquinolones in Staphylococcus intermedius and Staphylococcus schleiferi. J. Vet. Pharmacol. Ther. 2007, 30, 464–469. [Google Scholar] [CrossRef] [PubMed]
- Onuma, K.; Tanabe, T.; Sato, H. Antimicrobial resistance of Staphylococcus pseudintermedius isolates from healthy dogs and dogs affected with pyoderma in Japan. Vet. Dermatol. 2012, 23, 17-e15. [Google Scholar] [CrossRef] [PubMed]
- Hiasa, H. The Glu-84 of the ParC subunit plays critical roles in both topoisomerase IV—Quinolone and Topoisomerase IV—DNA interactions. Biochemistry 2002, 41, 11779–11785. [Google Scholar] [CrossRef]
- Wohlkonig, A.; Chan, P.F.; Fosberry, A.P.; Homes, P.; Huang, J.; Kranz, M.; Leydon, V.R.; Miles, T.J.; Pearson, N.D.; Perera, R.L. Structural basis of quinolone inhibition of type IIA topoisomerases and target-mediated resistance. Nat. Struct. Mol. Biol. 2010, 17, 1152–1153. [Google Scholar] [CrossRef]
- Sáenz, Y.; Zarazaga, M.; Briñas, L.; Ruiz-Larrea, F.; Torres, C. Mutations in gyrA and parC genes in nalidixic acid-resistant Escherichia coli strains from food products, humans and animals. J. Antimicrob. Chemother. 2003, 51, 1001–1005. [Google Scholar] [CrossRef]
- Neyfakh, A.A.; Borsch, C.M.; Kaatz, G.W. Fluoroquinolone resistance protein NorA of Staphylococcus aureus is a multidrug efflux transporter. Antimicrob. Agents Chemother. 1993, 37, 128–129. [Google Scholar] [CrossRef]
- Kaatz, G.W.; Seo, S.M. Inducible NorA-mediated multidrug resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 1995, 39, 2650–2655. [Google Scholar] [CrossRef]
- Markham, P.N.; Neyfakh, A.A. Inhibition of the multidrug transporter NorA prevents emergence of norfloxacin resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 1996, 40, 2673–2674. [Google Scholar] [CrossRef]
- Tejedor, M.a.T.; Martín, J.L.; Navia, M.; Freixes, J.; Vila, J. Mechanisms of fluoroquinolone resistance in Pseudomonas aeruginosa isolates from canine infections. Vet. Microbiol. 2003, 94, 295–301. [Google Scholar] [CrossRef]
- Harada, K.; Arima, S.; Niina, A.; Kataoka, Y.; Takahashi, T. Characterization of Pseudomonas aeruginosa isolates from dogs and cats in Japan: Current status of antimicrobial resistance and prevailing resistance mechanisms. Microbiol. Immunol. 2012, 56, 123–127. [Google Scholar] [CrossRef]
- Schmitz, F.-J.; Fluit, A.C.; Lückefahr, M.; Engler, B.; Hofmann, B.; Verhoef, J.; Heinz, H.P.; Hadding, U.; Jones, M.E. The effect of reserpine, an inhibitor of multidrug efflux pumps, on the in-vitro activities of ciprofloxacin, sparfloxacin and moxifloxacin against clinical isolates of Staphylococcus aureus. J. Antimicrob. Chemother. 1998, 42, 807–810. [Google Scholar] [CrossRef]
- Guillard, T.; de Jong, A.; Limelette, A.; Lebreil, A.L.; Madoux, J.; De Champs, C.; ComPath Study, G. Characterization of quinolone resistance mechanisms in Enterobacteriaceae recovered from diseased companion animals in Europe. Vet. Microbiol. 2016, 194, 23–29. [Google Scholar] [CrossRef]
- Cusack, T.P.; Ashley, E.A.; Ling, C.L.; Rattanavong, S.; Roberts, T.; Turner, P.; Wangrangsimakul, T.; Dance, D.A.B. Impact of CLSI and EUCAST breakpoint discrepancies on reporting of antimicrobial susceptibility and AMR surveillance. Clin. Microbiol. Infect. 2019, 25, 910–911. [Google Scholar] [CrossRef]
- Brakstad, O.G.; Aasbakk, K.; Maeland, J.A. Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J. Clin. Microbiol. 1992, 30, 1654–1660. [Google Scholar] [CrossRef]
- Becker, K.; von Eiff, C.; Keller, B.; Brück, M.; Etienne, J.; Peters, G. Thermonuclease gene as a target for specific identification of Staphylococcus intermedius isolates: Use of a PCR-DNA enzyme immunoassay. Diagn. Microbiol. Infect. Dis. 2005, 51, 237–244. [Google Scholar] [CrossRef]
- McDaniels, A.E.; Rice, E.W.; Reyes, A.L.; Johnson, C.H.; Haugland, R.A.; Stelma Jr, G.N. Confirmational identification of Escherichia coli, a comparison of genotypic and phenotypic assays for glutamate decarboxylase and beta-D-glucuronidase. Appl. Environ. Microbiol. 1996, 62, 3350–3354. [Google Scholar] [CrossRef]
- Anastasi, E.M.; Matthews, B.; Gundogdu, A.; Vollmerhausen, T.L.; Ramos, N.L.; Stratton, H.; Ahmed, W.; Katouli, M. Prevalence and persistence of Escherichia coli strains with uropathogenic virulence characteristics in sewage treatment plants. Appl. Environ. Microbiol. 2010, 76, 5882–5886. [Google Scholar] [CrossRef]
- Brakstad, O.G.; MÆLand, J.A.; Tveten, Y. Multiplex polymerase chain reaction for detection of genes for Staphylococcus aureus thermonuclease and methicillin resistance and correlation with oxacillin resistance. Apmis 1993, 101, 681–688. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standard Institue (CLSI). Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals; Approved Standard—Fourth Edition—CLSI VET01-A4. 2013. Available online: https://clsi.org/media/1531/vet01a4_sample.pdf (accessed on 1 April 2022).
- European Committee on Antimicrobial Susceptibility Testing (EUCAST). Available online: https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Disk_test_documents/2022_manuals/Reading_guide_BMD_v_4.0_2022.pdf (accessed on 1 April 2022).
- Clinical and Laboratory Standard Institue (CLSI). Performance Standards for Antimicrobial Disk and Dilution Susceptibility Tests for Bacteria Isolated From Animals, 5th Edition—CLSI VET01S. 2020. Available online: https://clsi.org/media/m1gppkx2/vet01sed5_sample.pdf (accessed on 1 April 2022).
- Clinical and laboratory standard institue (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 32nd Edition—CLSI M100. 2022. Available online: https://clsi.org/media/2663/m100ed29_sample.pdf (accessed on 1 April 2022).
- European Committee on Antimicrobial Susceptibility Testing (EUCAST). Available online: https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_12.0_Breakpoint_Tables.pdf (accessed on 1 April 2022).
- Clinical and Laboratory Standard institue (CLSI). Methods for Determining Bactericidal Activity of Antimicrobial Agents; Approved Guideline—M26-A. 1999. Available online: https://clsi.org/media/1462/m26a_sample.pdf (accessed on 1 April 2022).
- Kim, H.B.; Park, C.H.; Kim, C.J.; Kim, E.-C.; Jacoby, G.A.; Hooper, D.C. Prevalence of plasmid-mediated quinolone resistance determinants over a 9-year period. Antimicrob. Agents Chemother. 2009, 53, 639–645. [Google Scholar] [CrossRef]
- Schmitz, F.-J.; Jones, M.E.; Hofmann, B.; Hansen, B.; Scheuring, S.; Lückefahr, M.; Fluit, A.; Verhoef, J.; Hadding, U.; Heinz, H.-P. Characterization of grlA, grlB, gyrA, and gyrB mutations in 116 unrelated isolates of Staphylococcus aureus and effects of mutations on ciprofloxacin MIC. Antimicrob. Agents Chemother. 1998, 42, 1249–1252. [Google Scholar] [CrossRef] [PubMed]
- Robicsek, A.; Strahilevitz, J.; Sahm, D.F.; Jacoby, G.A.; Hooper, D.C. qnr prevalence in ceftazidime-resistant Enterobacteriaceae isolates from the United States. Antimicrob. Agents Chemother. 2006, 50, 2872–2874. [Google Scholar] [CrossRef] [PubMed]
- Ciesielczuk, H.; Hornsey, M.; Choi, V.; Woodford, N.; Wareham, D.W. Development and evaluation of a multiplex PCR for eight plasmid-mediated quinolone-resistance determinants. J. Med. Microbiol. 2013, 62, 1823–1827. [Google Scholar] [CrossRef] [PubMed]
| Bacterial Type | Fluoroquinolone | MIC (μg/mL) | MBC (μg/mL) | MBC/MIC Ratio * | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| MIC50 | MIC90 | Range | MBC50 | MBC90 | Range | MBC50/MIC50 | MBC90/MIC90 | Range | ||
| MSSA (n = 34) | ENR | 0.25 | 2 | 0.125–16 | 0.25 | 2 | 0.125–16 | 1 | 2 | 1–8 |
| MAR | 0.5 | 1 | 0.25–32 | 1 | 2 | 0.25–32 | 1 | 2 | 1–4 | |
| PRA | 0.06 | 0.125 | 0.03–2 | 0.125 | 0.25 | 0.03–4 | 1 | 2 | 1–2 | |
| CIP | 0.5 | 2 | 0.25–32 | 0.5 | 8 | 0.25–64 | 1 | 2 | 1–4 | |
| MOX | 0.125 | 0.5 | 0.03–4 | 0.25 | 0.5 | 0.06–4 | 1 | 2 | 1–4 | |
| MRSA (n = 45) | ENR | 16 | 32 | 0.25–128 | 32 | 64 | 0.5– ≥512 | 2 | 4 | 1–8 |
| MAR | 32 | 64 | 0.5–128 | 64 | 64 | 0.5–128 | 1 | 2 | 1–2 | |
| PRA | 4 | 4 | 0.125–32 | 4 | 8 | 0.25–32 | 1 | 2 | 1–4 | |
| CIP | 128 | 128 | 0.5–256 | 256 | ≥512 | 0.5 – ≥512 | 4 | 16 | 1–32 | |
| MOX | 4 | 8 | 0.125–16 | 8 | 16 | 0.125–32 | 1 | 2 | 1–8 | |
| MSSP (n = 53) | ENR | 0.125 | 2 | 0.125–32 | 0.25 | 2 | 0.125–64 | 1 | 2 | 1–4 |
| MAR | 0.25 | 2 | 0.25–32 | 0.5 | 2 | 0.25–64 | 1 | 2 | 1–4 | |
| PRA | 0.06 | 0.25 | 0.016–4 | 0.06 | 0.25 | 0.03–4 | 1 | 2 | 1–8 | |
| CIP | 0.25 | 1 | 0.125–16 | 0.25 | 2 | 0.125– ≥512 | 1 | 2 | 1–64 | |
| MOX | 0.06 | 0.5 | 0.03–4 | 0.06 | 0.5 | 0.03–8 | 1 | 2 | 1–2 | |
| MRSP (n = 52) | ENR | 32 | 64 | 0.06–64 | 64 | ≥512 | 0.125– ≥512 | 2 | 32 | 1–64 |
| MAR | 32 | 64 | 0.25–64 | 64 | 64 | 0.5–64 | 1 | 2 | 1–2 | |
| PRA | 2 | 4 | 0.03–8 | 4 | 4 | 0.06–16 | 1 | 2 | 1–2 | |
| CIP | 32 | 32 | 0.125–128 | 256 | 512 | 0.125– ≥512 | 8 | 16 | 1–64 | |
| MOX | 4 | 8 | 0.06–8 | 0.06 | 8 | 0.125–32 | 2 | 2 | 1–8 | |
| E. coli (n = 65) | ENR | 0.06 | 128 | 0.016–512 | 0.06 | 256 | 0.016–512 | 1 | 2 | 1–4 |
| MAR | 0.06 | 64 | 0.016–256 | 0.125 | 64 | 0.016–512 | 1 | 2 | 1–8 | |
| PRA | 0.03 | 32 | 0.016–128 | 0.03 | 32 | 0.016–128 | 1 | 2 | 1–4 | |
| CIP | 0.03 | 256 | 0.008– ≥512 | 0.03 | 256 | 0.008– ≥512 | 1 | 2 | 1–8 | |
| MOX | 0.125 | 64 | 0.016–512 | 0.125 | 64 | 0.016–512 | 1 | 2 | 1–4 | |
| ISOLATE ID | MIC Enrofloxacin (µg/mL) | MIC Marbofloxacin (µg/mL) | MIC Pradofloxacin (µg/mL) | MIC Ciprofloxacin (µg/mL) | MIC Moxifloxacin (µg/mL) | gyrA | grlA (parC) | PMQR | MIC/MICepi |
|---|---|---|---|---|---|---|---|---|---|
| MSSA B053 | 16 | 32 | 1 | 16 | 4 | Ser84Leu | Ser80Phe | NA | 1 |
| MSSA B071 | 8 | 16 | 1 | 16 | 4 | No mutation | No mutation | NA | 16 * |
| MSSA B074 | 8 | 16 | 2 | 32 | 4 | Ser84Leu | Ser80Phe | NA | 1 |
| MRSA A019 | 16 | 32 | 1 | 128 | 4 | Ser84Leu | Ser80Phe | NA | 1 |
| MRSA A069 | 64 | 128 | 4 | 256 | 1 | Ser84Leu, Gly90Cys | Ser80Phe | NA | 1 |
| MRSA A132 | 16 | 64 | 32 | 256 | 16 | Ser84Leu, Gly90Cys | Ser80Phe, Glu84Lys | NA | 1 |
| MSSP 098 | 4 | 4 | 0.5 | 4 | 0.5 | Glu88Gly | Ser80Arg | NA | 1 |
| MSSP 115 | 16 | 16 | 2 | 16 | 2 | Ser84Leu | Asp84Asn | NA | 1 |
| MSSP 099 | 32 | 32 | 4 | 16 | 4 | Ser84Leu | Ser80Ile | NA | 2 |
| MRSP 045 | 32 | 32 | 1 | 32 | 4 | Ser84Leu | Ser80Ile | NA | 1 |
| MRSP 038 | 64 | 32 | 2 | 32 | 2 | Ser84Leu | Ser80Ile | NA | 1 |
| MRSP 067 | 64 | 64 | 4 | 32 | 4 | Ser84Leu | Ser80Ile | NA | 2 |
| E. coli 10L-2253 | 8 | 32 | 4 | 32 | 16 | Ser83Leu, Asp87Asn | Ser80Ile, Ala108Val | - | 4 * |
| E. coli 16L-4063 | 64 | 32 | 4 | 128 | 16 | Ser83Leu, Asp87Asn | Ser80Ile, Glu84Val | aac-(6’)-lb-cr | 2 |
| E. coli 15L-3275 | 128 | 64 | 32 | 64 | 64 | Ser83Leu, Asp87Asn | Ser80Ile | QnrB | 4 * |
| E. coli 282305 | 128 | 64 | 32 | 64 | 64 | Ser83Leu, Asp87Asn | Ser80Ile | QnrB | 8 * |
| E. coli 13L-4865 | 256 | 128 | 64 | 256 | 128 | Ser83Leu, Asp87Asn | Ser80Ile, Ala108Val | QnrS | 8 * |
| E. coli 13L-5283 | 128 | 256 | 64 | 128 | 256 | Ser83Leu, Asp87Asn | Ser80Ile | QnrS, aac-(6’)-lb-cr | 8 * |
| E. coli 13L-6009 | 512 | 256 | 128 | 256 | 512 | Ser83Leu, Asp87Asn | Ser80Ile | QnrB, QnrS, aac-(6’)-lb-cr | 8 * |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Azzariti, S.; Bond, R.; Loeffler, A.; Zendri, F.; Timofte, D.; Chang, Y.-M.; Pelligand, L. Investigation of In Vitro Susceptibility and Resistance Mechanisms in Skin Pathogens: Perspectives for Fluoroquinolone Therapy in Canine Pyoderma. Antibiotics 2022, 11, 1204. https://doi.org/10.3390/antibiotics11091204
Azzariti S, Bond R, Loeffler A, Zendri F, Timofte D, Chang Y-M, Pelligand L. Investigation of In Vitro Susceptibility and Resistance Mechanisms in Skin Pathogens: Perspectives for Fluoroquinolone Therapy in Canine Pyoderma. Antibiotics. 2022; 11(9):1204. https://doi.org/10.3390/antibiotics11091204
Chicago/Turabian StyleAzzariti, Stefano, Ross Bond, Anette Loeffler, Flavia Zendri, Dorina Timofte, Yu-Mei Chang, and Ludovic Pelligand. 2022. "Investigation of In Vitro Susceptibility and Resistance Mechanisms in Skin Pathogens: Perspectives for Fluoroquinolone Therapy in Canine Pyoderma" Antibiotics 11, no. 9: 1204. https://doi.org/10.3390/antibiotics11091204
APA StyleAzzariti, S., Bond, R., Loeffler, A., Zendri, F., Timofte, D., Chang, Y.-M., & Pelligand, L. (2022). Investigation of In Vitro Susceptibility and Resistance Mechanisms in Skin Pathogens: Perspectives for Fluoroquinolone Therapy in Canine Pyoderma. Antibiotics, 11(9), 1204. https://doi.org/10.3390/antibiotics11091204






