Comprehensive Analysis of Imipenemase (IMP)-Type Metallo-β-Lactamase: A Global Distribution Threatening Asia
Abstract
1. Introduction
2. Materials and Methods
2.1. Review of Literature
2.2. blaIMP Gene Sequence Retrieval and Analysis
2.3. Phylogenetic Tree Estimation
3. Results
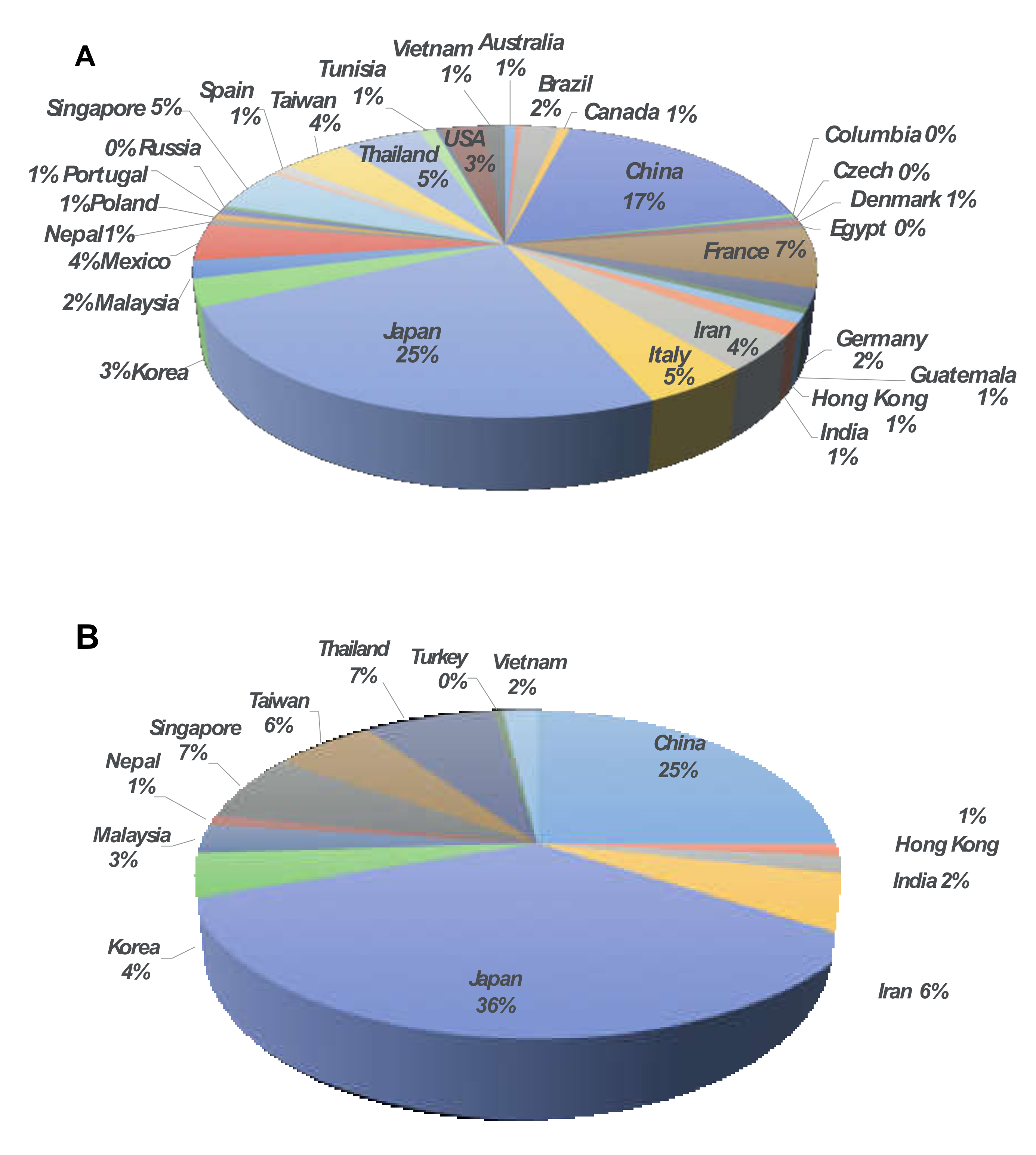
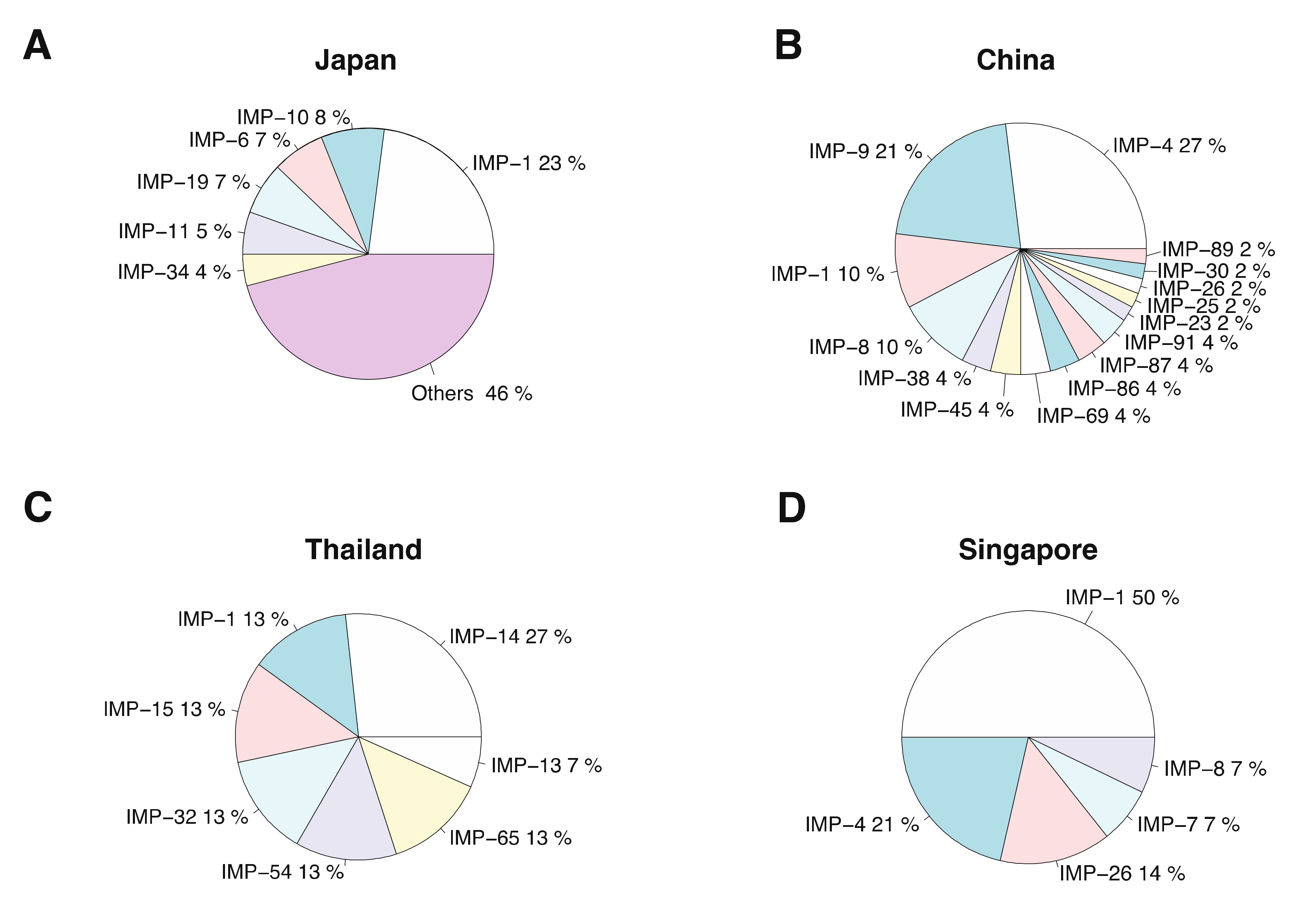
3.1. Distribution of IMP-Type MBLs
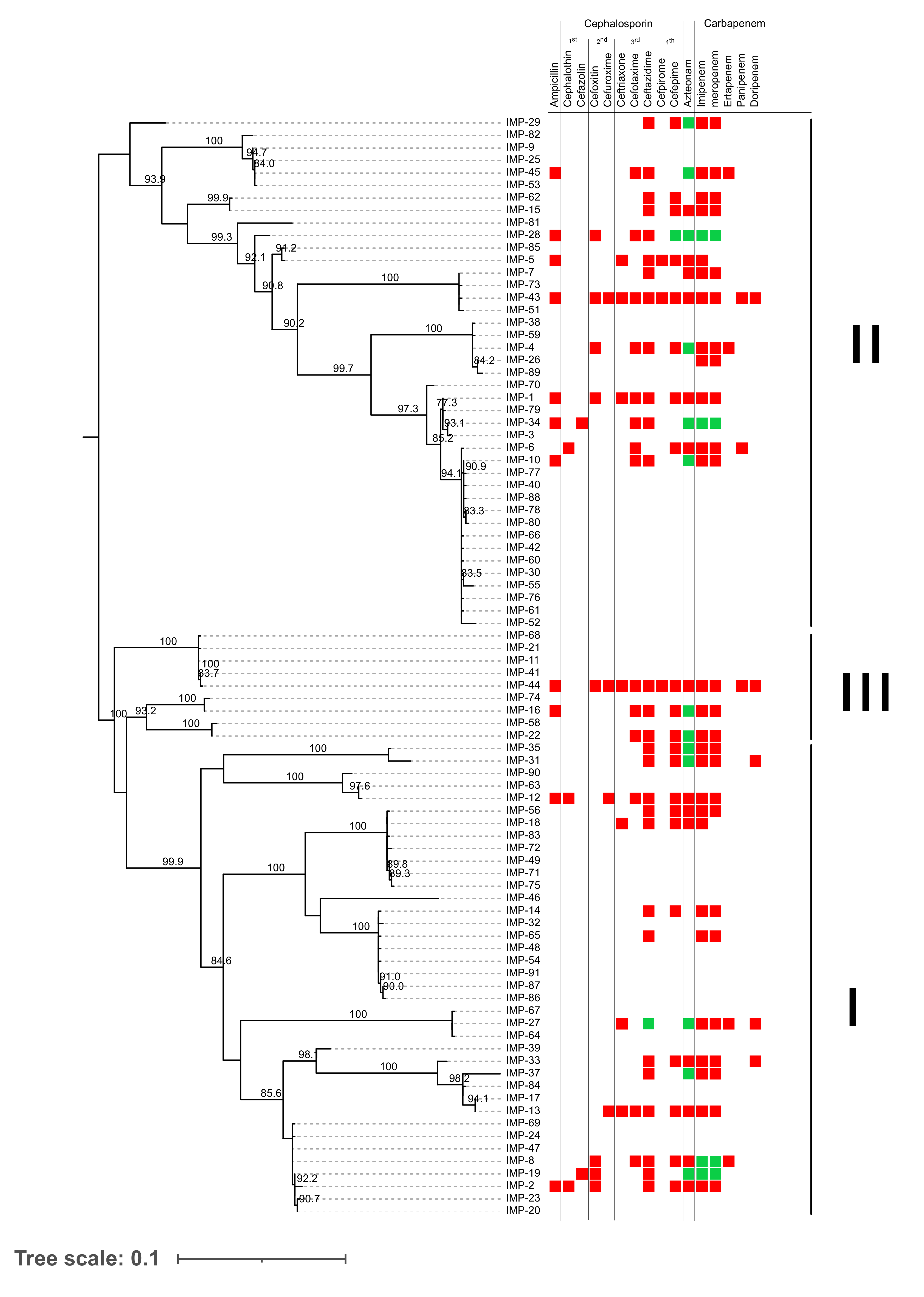
3.2. In Silico Analysis of IMP-Type MBLs
3.3. Resistance of IMP MBL Variants-Carrying Strains
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Serra-Burriel, M.; Keys, M.; Campillo-Artero, C.; Agodi, A.; Barchitta, M.; Gikas, A.; Palos, C.; López-Casasnovas, G. Impact of multi-drug resistant bacteria on economic and clinical outcomes of healthcare-associated infections in adults: Systematic review and meta-analysis. PLoS ONE 2020, 15, e0227139. [Google Scholar] [CrossRef] [PubMed]
- Majiduddin, F.K.; Materon, I.C.; Palzkill, T.G. Molecular analysis of beta-lactamase structure and function. Int. J. Med. Microbiol. 2002, 292, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Hall, B.G.; Barlow, M. Revised ambler classification of β-lactamases. J. Antimicrob. Chemother. 2005, 55, 1050–1051. [Google Scholar] [CrossRef] [PubMed]
- Queenan, A.M.; Bush, K. Carbapenemases: The versatile β-lactamases. Clinical Microbiol. Rev. 2007, 20, 440. [Google Scholar] [CrossRef]
- Deng, Y.; Bao, X.; Ji, L.; Chen, L.; Liu, J.; Miao, J.; Chen, D.; Bian, H.; Li, Y.; Yu, G. Resistance integrons: Class 1, 2 and 3 integrons. Ann. Clin. Microbiol. Antimicrob. 2015, 14, 45. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Iyobe, S.; Inoue, M.; Mitsuhashi, S. Transferable imipenem resistance in Pseudomonas Aeruginosa. Antimicrob. Agents Chemother. 1991, 35, 147. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Oueslati, S.; Bonnin, R.A.; Dabos, M.L.; Zavala, A.; Dortet, L.; Retailleau, P.; Iorga, B.I. Beta-lactamase database (BLDB)–Structure and function. J. Enzyme Inhib. Med. Chem. 2017, 32, 917. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree 2-Approximately maximum-likelihood trees for large alignments. PLoS ONE 2010, 5, e9490. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive tree of life (ITOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef] [PubMed]
- Boonkerd, N.; Pibalpakdi, P.; Tiloklurs, M.; Niumsup, P.R. Class 1 integron containing metallo β-lactamase gene BlaIMP-1 in carbapenem-resistant Pseudomonas Aeruginosa in Thailand. J. Infect. Chemother. 2009, 15, 257–261. [Google Scholar] [CrossRef] [PubMed]
- Senda, K.; Arakawa, Y.; Ichiyama, S.; Nakashima, K.; Ito, H.; Ohsuka, S.; Shimokata, K.; Kato, N.; Ohta, M. PCR Detection of metallo-beta-lactamase gene (BlaIMP) in gram-negative rods resistant to broad-spectrum beta-lactams. J. Clin. Microbiol. 1996, 34, 2909–2913. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Lee, W.G.; Uh, Y.; Ha, G.Y.; Cho, J.; Chong, Y.; Kang, J.O.; Kim, M.Y.; Lee, N.Y.; Kim, M.N.; et al. VIM-and IMP-type metallo-β-lactamase–producing Pseudomonas spp. and Acinetobacter spp. in Korean hospitals-Volume 9, Number 7—July 2003. Emerg. Infect. Dis. J. CDC 2003, 9, 868–871. [Google Scholar] [CrossRef]
- Yan, J.J.; Ko, W.C.; Chuang, C.L.; Wu, J.J. Metallo-β-lactamase-producing enterobacteriaceae isolates in a university hospital in Taiwan: Prevalence of IMP-8 in Enterobacter Cloacae and first identification of VIM-2 in Citrobacter Freundii. J. Antimicrob. Chemother. 2002, 50, 503–511. [Google Scholar] [CrossRef]
- Salahuddin, P.; Khan, A.U. Studies on structure-based sequence alignment and phylogenies of beta-lactamases. Bioinformation 2014, 10, 308. [Google Scholar] [CrossRef][Green Version]
- Khuntayaporn, P.; Yamprayoonswat, W.; Yasawong, M.; Chomnawang, M.T. Dissemination of carbapenem-resistance among multidrug resistant Pseudomonas Aeruginosa carrying metallo-beta-lactamase genes, including the novel BlaIMP-65 gene in Thailand. Infect. Chemother. 2019, 51, 107–118. [Google Scholar] [CrossRef]
- Bush, K.; Bradford, P.A. Epidemiology of β-lactamase-producing pathogens. Clin. Microbiol. Rev. 2020, 33, e00047-19. [Google Scholar] [CrossRef]
- Abdul Momin, M.H.F.; Liakopoulos, A.; Phee, L.M.; Wareham, D.W. Emergence and nosocomial spread of carbapenem-resistant OXA-232-producing Klebsiella Pneumoniae in Brunei Darussalam. J. Glob. Antimicrob. Resist. 2017, 9, 96–99. [Google Scholar] [CrossRef]
- Hsu, L.Y.; Apisarnthanarak, A.; Khan, E.; Suwantarat, N.; Ghafur, A.; Tambyah, P. Carbapenem-resistant acinetobacter Baumannii and Enterobacteriaceae in South and Southeast Asia. Clin. Microbiol. Rev. 2017, 30, 1. [Google Scholar] [CrossRef] [PubMed]
- Stokes, H.W.; Gillings, M.R. Gene flow, mobile genetic elements and the recruitment of antibiotic resistance genes into gram-negative pathogens. FEMS Microbiol. Rev. 2011, 35, 790–819. [Google Scholar] [CrossRef] [PubMed]
- Rowe-Magnus, D.A.; Mazel, D. The role of integrons in antibiotic resistance gene capture. Int. J. Med. Microbiol. 2002, 292, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.S.; Apisarnthanarak, A.; Hsu, L.Y. Mechanisms of β-Lactam antimicrobial resistance and epidemiology of major community- and healthcare-associated multidrug-resistant bacteria. Adv. Drug Deliv. Rev. 2014, 78, 3–13. [Google Scholar] [CrossRef] [PubMed]
| IMP Type | Host | Country of Isolation | Reference or Accession | Source of Isolates |
|---|---|---|---|---|
| IMP−1 | Achromobacter xylosoxidans | Japan | EF027105.1, KF032823.1, KF032821.1, KF032820.1 | Hospital |
| Comamonas thiooxydans | Japan | AP025194.1 | Hospital | |
| Pseudomonas aeruginosa | Japan | AB983593.1 | Hospital | |
| Thailand | [13] | Hospital | ||
| Malaysia | KX987869.1 | Hospital | ||
| China | AY386702.1, AY912485.1 | Hospital | ||
| Iran | KR703251.1, JX648311.1, JX644173.1, JQ766530.1 | Hospital | ||
| Nepal | LC636409.1 | Hospital | ||
| Singapore | AY168635.1, AY625689.1, AY625688.1, AY625687.1, AY625686.1 | Hospital | ||
| Egypt | KX452681.1 | Hospital | ||
| (Direct submission from Brazil) | GU831553.1, GU831552.1, GU831551.1, GU831550.1, GU831549.1, GU831548.1, GU831547.1, GU831546.1 | N/A | ||
| (Submitted from the UK, unpublished) | MH594579.1 | Hospital | ||
| Turkey | DQ842025.1 | Hospital | ||
| India | KF570107.1 | Hospital | ||
| USA | MK388919.1, MF479262.1 | N/A | ||
| Pseudomonas putida | Singapore | AY251052.1 | Hospital | |
| Pseudomonas fluorescens | Singapore | AY250709.1 | Hospital | |
| Serratia marcescens | Japan | AB162950.1, AB162949.1, AB162948.1, AB162947.1, NG_049172.1 | Hospital | |
| Klebsiella pneumoniae | Iran | LC512050.1, LC512051.1 | Hospital | |
| Klebsiella pneumoniae | Japan | [14] | Hospital | |
| Acinetobacter spp. | Korea | [15] | Hospital | |
| Acinetobacter bereziniae | Korea | EU014166.1, EU686386.1 | Hospital | |
| Acinetobacter calcoaceticus | Thailand | HM185482.1 | Hospital | |
| Acinetobacter baumannii | Japan | [15] | Hospital | |
| (Submitted from Korea, unpublished) | EF375699.1 | Hospital | ||
| Iran | KR080548.1, KF723585.1 | Hospital | ||
| (Submitted from Brazil, unpublished) | KF381490.1, KF381489.1, KF381488.1, KF381487.1 | Hospital | ||
| Thailand | HM036079.1 | Hospital | ||
| Acinetobacter pittii | Korea | GQ288398.1, GQ288393.1 | Hospital | |
| Taiwan | GU064942.1, GU064941.1 | N/A | ||
| GQ864268.1 | Hospital | |||
| Japan | AB753459.1 | N/A | ||
| Acinetobacter nosocomialis | Korea | GQ288394.1 | Hospital | |
| Taiwan | GU064940.1, GU064939.1, GU064938.1 | N/A | ||
| Citrobacter freundii | Japan | AB754498.1 | N/A | |
| Citrobacter youngae | (Direct submission from Ireland) | MW847603.1 | Hospital | |
| Enterobacter aerogenes | Japan | [15] | Hospital | |
| Enterobacter cloacae | (Direct submission from Japan) | LC508022.1 | Hospital | |
| Japan | [15] | Hospital | ||
| China | MK088089.1 | Hospital | ||
| Enterobacter hormaechei | China | MG287118.1 | N/A | |
| Escherichia coli | Japan | [16] | Hospital | |
| Iran | LC512049.1 | Hospital | ||
| Proteus mirabilis | Brazil | KY057362.1 | Hospital | |
| Proteus vulgaris | Japan | [16] | Hospital | |
| Providencia rettgeri | Japan | AB754496.1 | N/A | |
| Leclercia adecarboxylata | China | KJ531212.1 | Hospital | |
| IMP−2 | Acinetobacter baumannii | Italy | AJ243491.1, NG_049183.1 | Hospital |
| India | KC588963.1 | Hospital | ||
| Serratia marcescens | Japan | AB182996.1 | N/A | |
| Pseudomonas aeruginosa | India | KC588963.1 | Hospital | |
| IMP−3 | Shigella flexneri | (Published in USA) | NG_049194.1 | N/A |
| IMP−4 | Acinetobacter baumannii | Hong Kong | NG_049203.1, AF445082.1, AF244145.1 | Hospital |
| Singapore | DQ532122.1, AY795963.1, AY590475.1 | Hospital | ||
| Acinetobacter calcoaceticus | (Direct submission from Malaysia, unpublished) | DQ307573.1 | N/A | |
| Citrobacter freundii | China | EU368857.1 | Hospital | |
| JQ818252.1 | N/A | |||
| Escherichia coli | China | AB636651.1 | N/A | |
| (Direct submission from India) | MF169878.1 | N/A | ||
| Enterobacter cloacae | China | KF699334.1 | Hospital | |
| Korea | KY884003.1 | N/A | ||
| Japan | LC198842.1 | Hospital | ||
| Enterobacter aerogenes | China | KF184385.1 | Hospital | |
| Klebsiella pneumoniae | China | EU368858.1, KF184388.1, FJ384365.1 | Hospital | |
| JQ808503.1, JN106667.1, KF680003.1 | N/A | |||
| Klebsiella oxytoca | China | JQ820404.1 | N/A | |
| KY913900.1 | Animal | |||
| Pseudomonas aeruginosa | China | DQ297664.1 | N/A | |
| Malaysia | GQ221782.1 | Hospital | ||
| IMP−5 | Acinetobacter baumannii | Portugal | NG_049212.1, JF810083.1 | Hospital |
| IMP−6 | Escherichia coli | Japan | AB753460.1 | N/A |
| Serratia marcescens | Japan | NG_049220.1, AB040994.1 | Hospital | |
| Providencia rettgeri | Japan | AB754497.1 | N/A | |
| Pseudomonas aeruginosa | Japan | AB188812.1 | Hospital | |
| Korea | EU117233.1 | Hospital | ||
| IMP−7 | Pseudomonas aeruginosa | Canada | NG_049221.1, AF318077.1 | Hospital |
| Czech | JX982232.1 | Hospital | ||
| Japan | LC091209.2, LC091210.2 | Hospital | ||
| Malaysia | GQ221781.1, AF416736.2, GU213192.1 | Hospital | ||
| India | HM641894.1 | Hospital | ||
| Singapore | AY625685.1 | Hospital | ||
| Slovakia | EF601914.1 | Hospital | ||
| IMP−8 | Acinetobacter baumannii | Taiwan | EF127959.1 | Hospital |
| China | DQ845788.1 | Hospital | ||
| Escherichia coli | Singapore | KF534724.1 | Hospital | |
| Enterobacter cloacae | Taiwan | [16] | Hospital | |
| China | JQ820405.1 | N/A | ||
| Klebsiella pneumoniae | China | JQ820406.1, EU368856.1 | Hospital | |
| Taiwan | NG_049222.1, AF322577.2 | Hospital | ||
| Tunisia | HE605039.1 | Non-hospital | ||
| Klebsiella oxytoca | China | HQ651093.1 | Hospital | |
| Serratia marcescens | Taiwan | EU042136.1 | N/A | |
| IMP−9 | Pseudomonas aeruginosa | China | AY033653, EU176818.1 | Hospital |
| KF184386.1, KF255597.1, KF255596.1, KF255595.1 | N/A | |||
| (Direct submission from China) | HM106459.1 | N/A | ||
| IMP−10 | Achromobacter xylosoxidans | Japan | AB074435.1, AB195638.1 | Hospital |
| Pseudomonas aeruginosa | Japan | AB074434.1, AB074433.1, NG_049173.1, AB195637.1 | Hospital | |
| (Direct submission from Japan, Unpublished) | DQ288156.1 | Hospital | ||
| Pseudomonas putida | Italy | AJ420864.1 | Hospital | |
| Klebsiella pneumoniae | Tunisia | HE605040.1 | Non-hospital | |
| IMP−11 | Pseudomonas aeruginosa | Japan | AB074437.1 | Hospital |
| Acinetobacter baumannii | Japan | AB074436, NG_049174.1 | Hospital | |
| Enterobacter cloacae | Japan | LC628821.1 | N/A | |
| IMP−12 | Pseudomonas putida | Italy | NG_049175.1 | Hospital |
| IMP−13 | Pseudomonas aeruginosa | Italy | FJ172676.1, FJ172674.1, AJ512502.1, NG_049176.1 | Hospital |
| France | JX131371.1 | Hospital | ||
| Thailand | GU207399.1 | Hospital | ||
| Pseudomonas monteilii | Italy | JN091097.1 | Hospital | |
| Klebsiella pneumoniae | Tunisia | HE605041.1 | Non-hospital | |
| IMP−14 | Achromobacter xylosoxidans | Thailand | KJ406506.2, KJ406505.2 | Hospital |
| Pseudomonas aeruginosa | Thailand | AY553332.1, NG_049177.1 | Hospital | |
| IMP−15 | Pseudomonas aeruginosa | Thailand | NG_049178.1, AY553333.1 | Hospital |
| Vietnam | LC075716.1 | N/A | ||
| Spain | KC310496.1 | Hospital | ||
| IMP−16 | Pseudomonas aeruginosa | Brazil | AJ584652.2, NG_049179.1 | Hospital |
| IMP−17 | Pseudomonas aeruginosa | Italy | NG_049180.1 | Hospital |
| IMP−18 | Pseudomonas aeruginosa | USA | AY780674.2, NG_049181.1 | Hospital |
| Mexico | HM138673.1 | N/A | ||
| (Direct submission from Costa Rica, unpublished) | KC907377.2 | Hospital | ||
| (Direct submission from Japan, unpublished) | AB587676.1 | N/A | ||
| IMP−19 | Acinetobacter baumannii | Iran | JQ766528.1 | N/A |
| Japan | AB184977.1 | Hospital | ||
| Achromobacter xylosoxidans | Japan | AB201263.1 | N/A | |
| Enterobacter cloacae | Japan | AB201264.1 | N/A | |
| Aeromonas caviae | France | NG_049182.1 | Hospital | |
| Klebsiella pneumoniae | (Direct submission from Japan, unpublished) | LC062960.1 | Hospital | |
| Pseudomonas aeruginosa | Japan | AB184976.1 | Hospital | |
| Pseudomonas putida | Japan | AB201265.1 | N/A | |
| Serratia marcescens | Poland | MH071810.1 | N/A | |
| MF678587.1 | Hospital | |||
| IMP−20 | Pseudomonas aeruginosa | Japan | AB196988, NG_049184.1 | N/A |
| IMP−21 | Pseudomonas aeruginosa | Japan | AB204557, NG_049185.1 | N/A |
| IMP−22 | Providencia rettgeri | Japan | AB754495.1 | N/A |
| Pseudomonas aeruginosa | Austria | FM876313.1 | Hospital | |
| Pseudomonas fluorescens | Italy | DQ361087.2, NG_049186.1 | Non-hospital | |
| IMP−23 | Citrobacter freundii | China | NG_049187.1 | N/A |
| IMP−24 | Serratia marcescens | Taiwan | EF192154.1, NG_049188.1 | Hospital |
| IMP−25 | Pseudomonas aeruginosa | China | EU352796 | Hospital |
| Korea | EU541448.1, NG_049189.1 | Hospital | ||
| (Direct submission from China, unpublished) | KY081418.1, KY081417.1, HM175876.1 | N/A | ||
| Stenotrophomonas maltophilia | (Direct submission fom China) | GU944726.1 | N/A | |
| IMP−26 | Enterobacter cloacae | China | HQ685900.1 | Hospital |
| Pseudomonas aeruginosa | Malaysia | JQ629930.1 | Hospital | |
| Pseudomonas aeruginosa | Nepal | LC636067.1 | Hospital | |
| Pseudomonas aeruginosa | Singapore | GU045307.1, NG_049190.1 | Hospital | |
| Pseudomonas aeruginosa | Vietnam | LC075717.1 | N/A | |
| IMP−27 | Morganella morganii | Mexico | KY847875.1, KY847873.1 | N/A |
| Proteus mirabilis | USA | JF894248.1 | Hospital | |
| (Direct submission from USA) | NG_049191.1 | N/A | ||
| Providencia rettgeri | USA | KY847874.1 | N/A | |
| IMP−28 | Klebsiella oxytoca | Spain | HQ263342.1, NG_049192.1 | Hospital |
| IMP−29 | Pseudomonas aeruginosa | France | HQ438058.1, JQ041634, NG_049193.1 | Hospital |
| IMP−30 | Escherichia coli | China | KM589497.1 | Hospital |
| Pseudomonas aeruginosa | Russia | NG_049195.1 | N/A | |
| IMP−31 | Pseudomonas aeruginosa | Germany | KF148593.1, NG_049196.1 | Hospital |
| IMP−32 | Klebsiella pneumoniae | Thailand | NG_049197.1, JQ002629.1 | Hospital |
| IMP−33 | Pseudomonas aeruginosa | Italy | JN848782, NG_049198.1 | Hospital |
| IMP−34 | Klebsiella oxytoca | Japan | AB700341.1, NG_049199.1 | Hospital |
| Acinetobacter colistiniresistens | Japan | LC276939.1 | Hospital | |
| IMP−35 | Pseudomonas aeruginosa | Germany | JF816544.1, NG_049200.1 | Hospital |
| IMP−36 | Not found in NCBI database and pubmed | |||
| IMP−37 | Pseudomonas aeruginosa | France | JX131372.1, NG_049201.1 | Hospital |
| IMP−38 | Klebsiella pneumoniae | China | HQ875573.1, NG_049202.1 | N/A |
| IMP−39 | Pseudomonas aeruginosa | France | MK507818.1, NG_064724.1 | Hospital |
| IMP−40 | Pseudomonas aeruginosa | Japan | AB753457, NG_049204.1 | N/A |
| IMP−41 | Pseudomonas aeruginosa | Japan | AB753458, NG_049205.1 | N/A |
| IMP−42 | Acinetobacter soli | Japan | AB753456.1, NG_049206.1 | N/A |
| IMP−43 | Pseudomonas aeruginosa | Japan | NG_049207.1 | Hospital |
| IMP−44 | Pseudomonas aeruginosa | Japan | NG_049208.1 | Hospital |
| IMP−45 | Pseudomonas aeruginosa | China | KJ510410.1, NG_049209.1 | Animal |
| France | KU984333.1 | Hospital | ||
| IMP−46 | Pseudomonas putida | France | MK543944.1, MK507819.1, NG_064725.1 | Hospital |
| IMP−47 | Serratia marcescens | (Direct submit USA) | KP050486.1 | N/A |
| IMP−48 | Pseudomonas aeruginosa | (Direct submit USA, unpublished) | NG_049210.1, KM087857.1 | N/A |
| IMP−49 | Pseudomonas aeruginosa | Brazil | NG_049211, KP681694.1 | N/A |
| IMP−50 | Not found in NCBI database and pubmed | |||
| IMP−51 | Pseudomonas aeruginosa | Vietnam | NG_049213.1, LC031883.1 | Hospital |
| IMP−52 | Escherichia coli | Japan | NG_049214.1, LC055762.1 | N/A |
| IMP−53 | Pseudomonas aeruginosa | (Direct submit USA) | NG_049215.1 | N/A |
| IMP−54 | Pseudomonas aeruginosa | Thailand | KU052795.1, NG_049216.1 | N/A |
| IMP−55 | Acinetobacter baumannii | Iran | KU299753.1, NG_049217.1 | Hospital |
| IMP−56 | Pseudomonas aeruginosa | Mexico | KU351745.1 | Hospital |
| Guatemala | KU315553.1, NG_049218.1 | N/A | ||
| IMP−57 | Not found in NCBI database and pubmed | |||
| IMP−58 | Pseudomonas putida | Denmark | KU647281.1, NG_049219.1 | N/A |
| IMP−59 | Escherichia coli | Australia | KX196782.1, NG_055477.1 | N/A |
| IMP−60 | Enterobacter cloacae | Japan | LC159227.1, NG_050945.1 | Hospital |
| IMP−61 | Acinetobacter baumannii | (Direct submission from Germany, unpublished) | KX462700.1, NG_051166.1 | Hospital |
| IMP−62 | Pseudomonas aeruginosa | Mexico | KX753224.1, NG_051513.1 | Hospital |
| IMP−63 | Pseudomonas aeruginosa | France | KX821663.1, NG_052049.1 | Hospital |
| IMP−64 | Proteus mirabilis | USA | NG_054710.1, KX949735.2 | N/A |
| IMP−65 | Pseudomonas aeruginosa | Thailand | KY315991.1, NG_066508.1 | Hospital |
| IMP−66 | Escherichia coli | Japan | LC190726.1, NG_054676.1 | N/A |
| IMP−67 | Providencia rettgeri | (Direct submission from USA, unpublished) | MF281100.1, NG_055271.1 | N/A |
| IMP−68 | Klebsiella pneumoniae | Japan | MF669572.1, NG_055584.1 | N/A |
| IMP−69 | Providencia spp. | China | MF678349.1, NG_055665.1 | N/A |
| IMP−70 | Pseudomonas aeruginosa | Germany | MG748725.1, NG_056176.1 | Hospital |
| Providencia rettgeri | Japan | LC348383.1 | N/A | |
| IMP−71 | Pseudomonas aeruginosa | France | MG818167.1 | Hospital |
| IMP−72 | Pseudomonas aeruginosa | Mexico | MH021847.1 | N/A |
| IMP−73 | Pseudomonas aeruginosa | Japan | MH021848.1, NG_057463.1 | N/A |
| IMP−74 | Pseudomonas aeruginosa | Brazil | MH243349.1, NG_057606.1 | N/A |
| IMP−75 | Pseudomonas aeruginosa | Mexico | MH243350.1, MW692112.1, NG_057607.1 | N/A |
| IMP−76 | Pseudomonas aeruginosa | Japan | NG_061409.1 | Hospital |
| IMP−77 | Pseudomonas aeruginosa | Japan | NG_061410.1 | Hospital |
| IMP−78 | Pseudomonas aeruginosa | Japan | NG_061411.1 | Hospital |
| IMP−79 | Pseudomonas aeruginosa | France | MG873561.1, NG_061626.1 | Hospital |
| IMP−80 | Pseudomonas aeruginosa | Japan | NG_062274.1 | Hospital |
| IMP−81 | Pseudomonas aeruginosa | Columbia | MN267699.1 | N/A |
| IMP−82 | Pseudomonas aeruginosa | (Direct submission from Germany, unpublished) | MN057782.1 | Hospital |
| (Direct submission from USA, unpublished) | NG_065873.1 | Hospital | ||
| IMP−83 | Pseudomonas aeruginosa | Mexico | MN104595.1, NG_065874.1 | N/A |
| IMP−84 | Pseudomonas aeruginosa | (Direct submission from Switzerland, unpublished) | MN219692.1 | N/A |
| Pseudomonas aeruginosa | (Direct submission from USA, unpublished) | NG_065875.1 | N/A | |
| IMP−85 | Pseudomonas aeruginosa | France | MN510335.1, NG_066696.1 | Hospital |
| IMP−86 | Pseudomonas aeruginosa | China | MT241520.1, NG_076650.1 | N/A |
| IMP−87 | Pseudomonas aeruginosa | China | MT241521.1, NG_076651.1 | N/A |
| IMP−88 | Pseudomonas aeruginosa | Japan | LC558310.1, NG_070737.1 | Hospital |
| IMP−89 | Pseudomonas putida | China | NG_070738.1 | N/A |
| IMP−90 | Pseudomonas aeruginosa | (Direct submission from Germany, unpublished) | MW811441.1 | Hospital |
| (Direct submission from USA, unpublished) | NG_074713.1 | Hospital | ||
| IMP−91 | Pseudomonas aeruginosa | China | MZ702721.1, NG_076634.1 | N/A |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pongchaikul, P.; Mongkolsuk, P. Comprehensive Analysis of Imipenemase (IMP)-Type Metallo-β-Lactamase: A Global Distribution Threatening Asia. Antibiotics 2022, 11, 236. https://doi.org/10.3390/antibiotics11020236
Pongchaikul P, Mongkolsuk P. Comprehensive Analysis of Imipenemase (IMP)-Type Metallo-β-Lactamase: A Global Distribution Threatening Asia. Antibiotics. 2022; 11(2):236. https://doi.org/10.3390/antibiotics11020236
Chicago/Turabian StylePongchaikul, Pisut, and Paninee Mongkolsuk. 2022. "Comprehensive Analysis of Imipenemase (IMP)-Type Metallo-β-Lactamase: A Global Distribution Threatening Asia" Antibiotics 11, no. 2: 236. https://doi.org/10.3390/antibiotics11020236
APA StylePongchaikul, P., & Mongkolsuk, P. (2022). Comprehensive Analysis of Imipenemase (IMP)-Type Metallo-β-Lactamase: A Global Distribution Threatening Asia. Antibiotics, 11(2), 236. https://doi.org/10.3390/antibiotics11020236







