In Silico Analysis of Extended-Spectrum β-Lactamases in Bacteria
Abstract
1. Introduction
2. Results
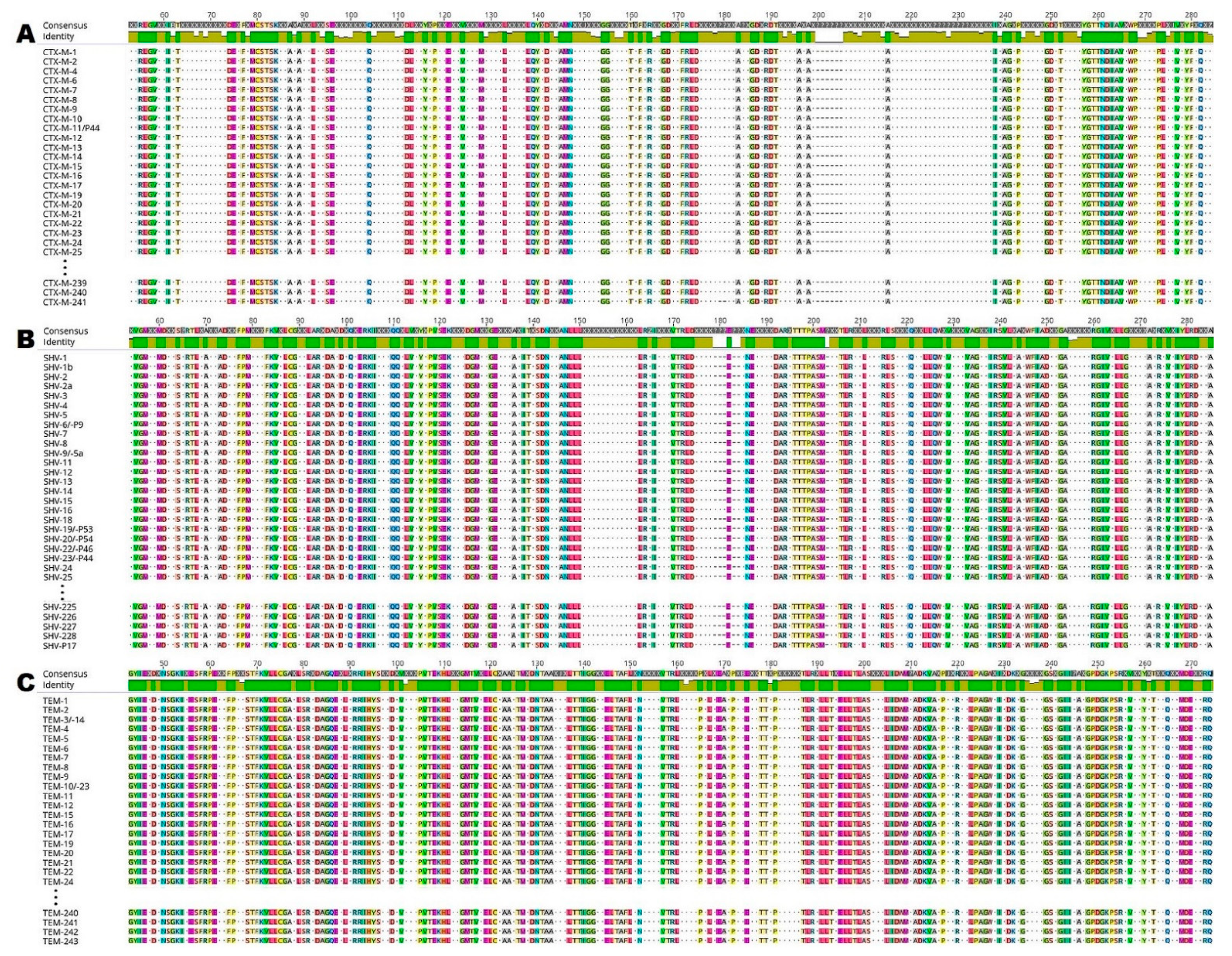
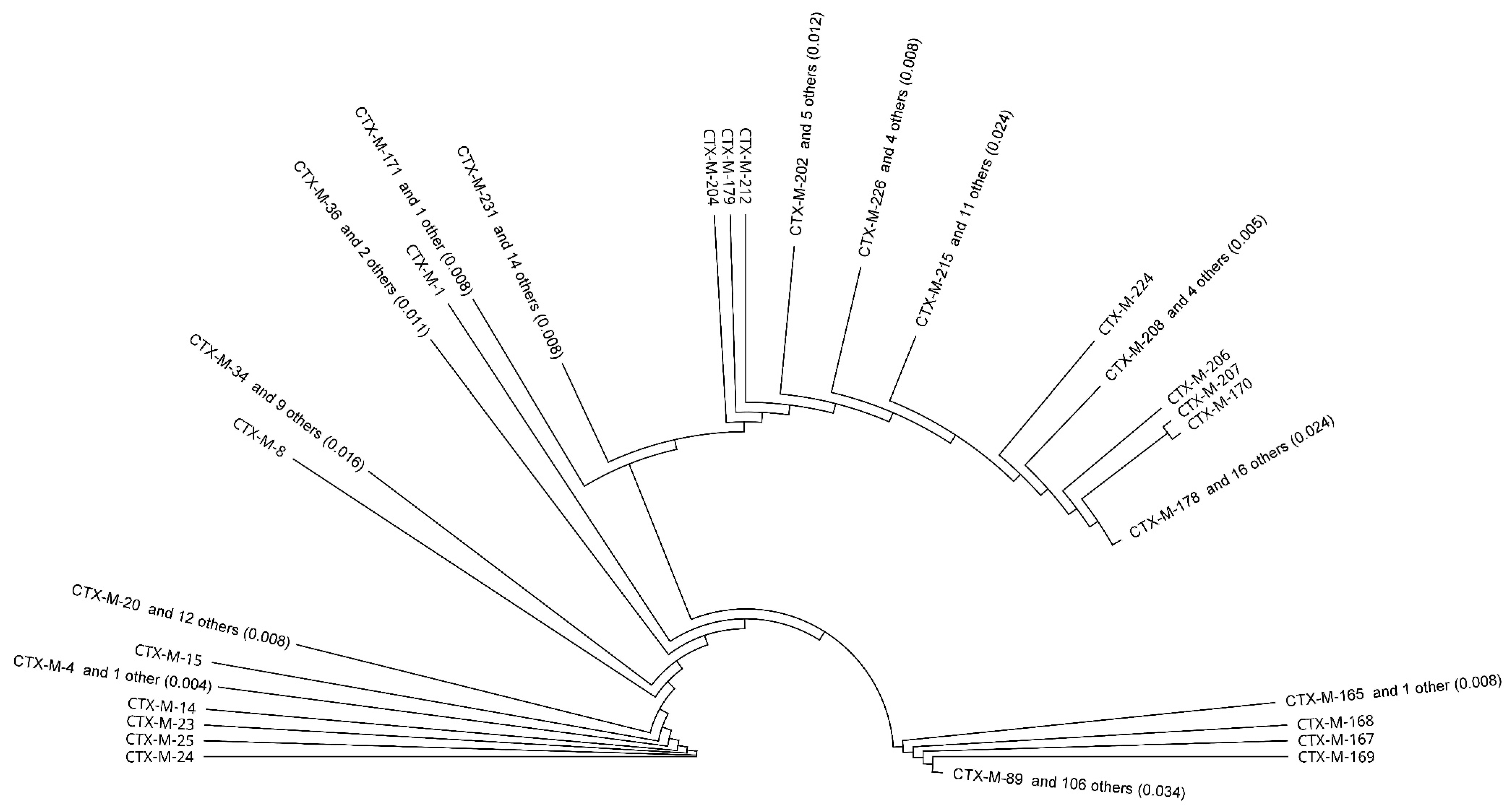
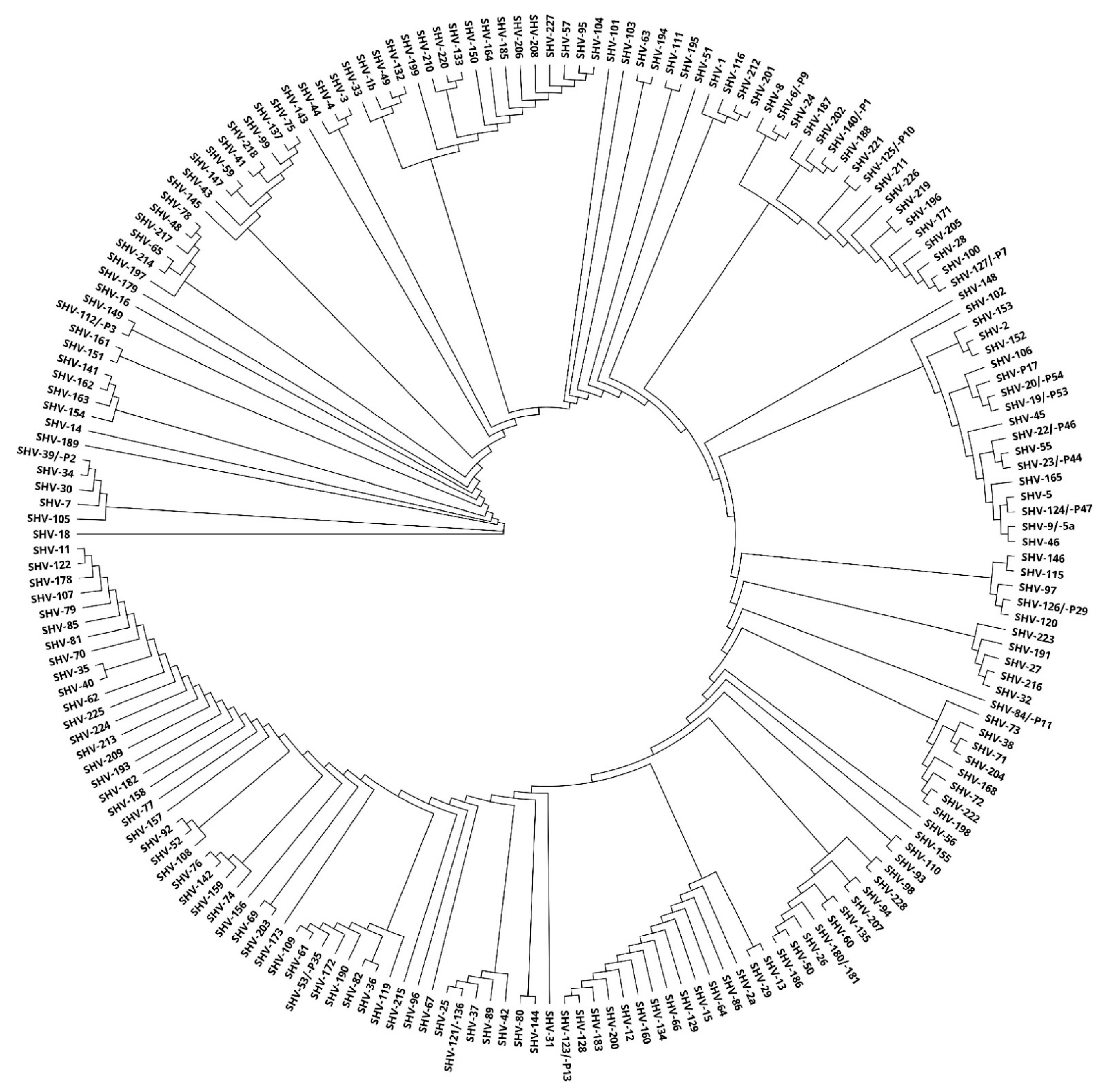
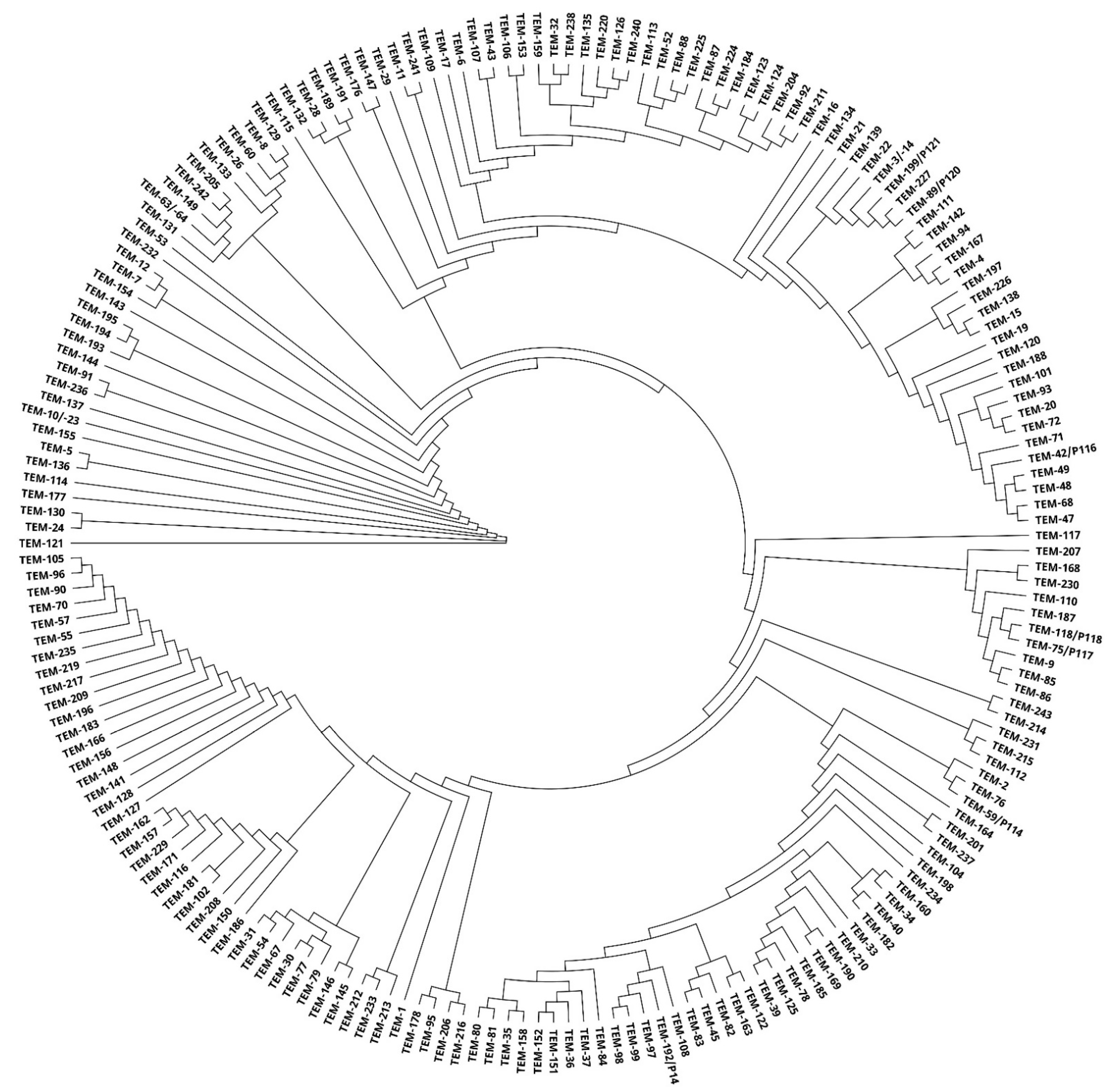
2.1. In Silico Analysis of ESBL Enzymes
2.2. Detection of ESBL-Positive Bacteria with PCR
3. Discussion
4. Materials and Methods
4.1. Sequence Analysis
4.2. Phylogenetic Tree Construction
4.3. Designing Primers for PCR
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| A | alanine |
| AmpC | ampicillin chromosomal cephalosporinase |
| BSBL | broad-spectrum β-lactamase |
| C | cysteine |
| CARBA | carbapenemase |
| CTX-M | cefotaxime-hydrolyzing β-lactamase–Munich |
| E | glutamic acid |
| ESBLs | extended-spectrum β-lactamases |
| F | phenylalanine |
| G | glycine |
| GES | Guiana extended-spectrum β-lactamase |
| H | histidine |
| I | isoleucine |
| K | lysine |
| KPC | Klebsiella pneumoniae carbapenemase |
| L | leucine |
| M | methionine |
| MBLs | metallo- β-lactamases |
| NDM | New Delhi metallo- β-lactamase |
| NSBL | narrow-spectrum β-lactamase |
| OXA | oxacillinases |
| P | proline |
| Q | glutamine |
| R | arginine |
| S | serine |
| SHV | sulfhydryl variant of the TEM enzyme |
| T | threonine |
| TEM | Temoneira class A extended-spectrum β-lactamase |
| V | valine |
| W | tryptophan |
References
- Naas, T.; Oueslati, S.; Bonnin, R.A.; Dabos, M.L.; Zavala, A.; Dortet, L.; Retailleau, P.; Iorga, B.I. Beta-lactamase database (BLDB)—structure and function. J. Enzym. Inhib. Med. Chem. 2017, 32, 917–919. [Google Scholar] [CrossRef]
- Jeon, J.H.; Hong, M.K.; Lee, J.H.; Lee, J.J.; Park, K.S.; Karim, A.M.; Jo, J.Y.; Kim, J.H.; Ko, K.S.; Kang, L.W.; et al. Structure of ADC-68, a novel carbapenem-hydrolyzing class C extended-spectrum beta-lactamase isolated from Acinetobacter baumannii. Acta Crystallogr. D Biol. Crystallogr. 2014, 70, 2924–2936. [Google Scholar] [CrossRef] [PubMed]
- Mammeri, H.; Guillon, H.; Eb, F.; Nordmann, P. Phenotypic and biochemical comparison of the carbapenem-hydrolyzing activities of five plasmid-borne AmpC beta-lactamases. Antimicrob. Agents Chemother. 2010, 54, 4556–4560. [Google Scholar] [CrossRef] [PubMed]
- Jousset, A.B.; Oueslati, S.; Bernabeu, S.; Takissian, J.; Creton, E.; Vogel, A.; Sauvadet, A.; Cotellon, G.; Gauthier, L.; Bonnin, R.A.; et al. False-Positive Carbapenem-Hydrolyzing Confirmatory Tests Due to ACT-28, a Chromosomally Encoded AmpC with Weak Carbapenemase Activity from Enterobacter kobei. Antimicrob. Agents Chemother. 2019, 63, e02388-18. [Google Scholar] [CrossRef]
- Kaitany, K.C.; Klinger, N.V.; June, C.M.; Ramey, M.E.; Bonomo, R.A.; Powers, R.A.; Leonard, D.A. Structures of the class D Carbapenemases OXA-23 and OXA-146: Mechanistic basis of activity against carbapenems, extended-spectrum cephalosporins, and aztreonam. Antimicrob. Agents Chemother. 2013, 57, 4848–4855. [Google Scholar] [CrossRef] [PubMed]
- Marathe, N.P.; Janzon, A.; Kotsakis, S.D.; Flach, C.F.; Razavi, M.; Berglund, F.; Kristiansson, E.; Larsson, D.G.J. Functional metagenomics reveals a novel carbapenem-hydrolyzing mobile beta-lactamase from Indian river sediments contaminated with antibiotic production waste. Environ. Int. 2018, 112, 279–286. [Google Scholar] [CrossRef] [PubMed]
- Suh, B.; Bae, I.K.; Kim, J.; Jeong, S.H.; Yong, D.; Lee, K. Outbreak of meropenem-resistant Serratia marcescens comediated by chromosomal AmpC beta-lactamase overproduction and outer membrane protein loss. Antimicrob. Agents Chemother. 2010, 54, 5057–5061. [Google Scholar] [CrossRef]
- Bonnet, R. Growing group of extended-spectrum beta-lactamases: The CTX-M enzymes. Antimicrob. Agents Chemother. 2004, 48, 1–14. [Google Scholar] [CrossRef]
- Tanimoto, K.; Nomura, T.; Hashimoto, Y.; Hirakawa, H.; Watanabe, H.; Tomita, H. Isolation of Serratia fonticola Producing FONA, a Minor Extended-Spectrum beta-Lactamase (ESBL), from Imported Chicken Meat in Japan. Jpn. J. Infect. Dis. 2021, 74, 79–81. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Sun, Z.; Lu, J.; Liu, H.; Lu, W.; Lin, H.; Zhang, X.; Li, Q.; Zhou, W.; Zhu, X.; et al. Characterization of a Novel Chromosomal Class C beta-Lactamase, YOC-1, and Comparative Genomics Analysis of a Multidrug Resistance Plasmid in Yokenella regensburgei W13. Front. Microbiol. 2020, 11, 2021. [Google Scholar] [CrossRef]
- Toth, M.; Antunes, N.T.; Stewart, N.K.; Frase, H.; Bhattacharya, M.; Smith, C.A.; Vakulenko, S.B. Class D beta-lactamases do exist in Gram-positive bacteria. Nat. Chem. Biol. 2016, 12, 9. [Google Scholar] [CrossRef] [PubMed]
- Bradford, P.A. Extended-spectrum beta-lactamases in the 21st century: Characterization, epidemiology, and detection of this important resistance threat. Clin. Microbiol. Rev. 2001, 14, 933–951. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.L.; Bonomo, R.A. Extended-spectrum beta-lactamases: A clinical update. Clin. Microbiol. Rev. 2005, 18, 657–686. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Poirel, L.; Nordmann, P. Minor extended-spectrum beta-lactamases. Clin. Microbiol. Infect. 2008, 14 (Suppl. 1), 42–52. [Google Scholar] [CrossRef]
- Guillon, H.; Eb, F.; Mammeri, H. Characterization of CSP-1, a novel extended-spectrum beta-lactamase produced by a clinical isolate of Capnocytophaga sputigena. Antimicrob. Agents Chemother. 2010, 54, 2231–2234. [Google Scholar] [CrossRef] [PubMed]
- Lamoureaux, T.L.; Vakulenko, V.; Toth, M.; Frase, H.; Vakulenko, S.B. A novel extended-spectrum beta-lactamase, SGM-1, from an environmental isolate of Sphingobium sp. Antimicrob. Agents Chemother. 2013, 57, 3783–3788. [Google Scholar] [CrossRef]
- Pfennigwerth, N.; Lange, F.; Campos, C.B.; Hentschke, M.; Gatermann, S.G.; Kaase, M. Genetic and biochemical characterization of HMB-1, a novel subclass B1 metallo-beta-lactamase found in a Pseudomonas aeruginosa clinical isolate. J. Antimicrob. Chemother. 2017, 72, 1068–1073. [Google Scholar] [CrossRef][Green Version]
- Queenan, A.M.; Bush, K. Carbapenemases: The versatile beta-lactamases. Clin. Microbiol. Rev. 2007, 20, 440–458. [Google Scholar] [CrossRef]
- Poirel, L.; Heritier, C.; Podglajen, I.; Sougakoff, W.; Gutmann, L.; Nordmann, P. Emergence in Klebsiella pneumoniae of a chromosome-encoded SHV beta-lactamase that compromises the efficacy of imipenem. Antimicrob. Agents Chemother. 2003, 47, 755–758. [Google Scholar] [CrossRef]
- Poirel, L.; Ortiz de la Rosa, J.M.; Richard, A.; Aires-de-Sousa, M.; Nordmann, P. CTX-M-33, a CTX-M-15 derivative conferring reduced susceptibility to carbapenems. Antimicrob. Agents Chemother. 2019, 63, e01515-19. [Google Scholar] [CrossRef]
- Alegria, A.; Arias-Temprano, M.; Fernandez-Natal, I.; Rodriguez-Calleja, J.M.; Garcia-Lopez, M.L.; Santos, J.A. Molecular Diversity of ESBL-Producing Escherichia coli from Foods of Animal Origin and Human Patients. Int. J. Environ. Res. Public Health 2020, 17, 1312. [Google Scholar] [CrossRef]
- Bardon, J.; Mlynarcik, P.; Prochazkova, P.; Roderova, M.; Mezerova, K.; Kolar, M. Occurrence of bacteria with a dangerous extent of antibiotic resistance in poultry in the Central Region of Moravia. Acta Vet. Brno 2018, 87, 165–172. [Google Scholar] [CrossRef]
- Zhang, H.; Zhou, Y.; Guo, S.; Chang, W. Prevalence and characteristics of extended-spectrum beta-lactamase (ESBL)-producing Enterobacteriaceae isolated from rural well water in Taian, China, 2014. Environ. Sci. Pollut. Res. Int. 2015, 22, 11488–11492. [Google Scholar] [CrossRef] [PubMed]
- Abbott, I.; Cerqueira, G.M.; Bhuiyan, S.; Peleg, A.Y. Carbapenem resistance in Acinetobacter baumannii: Laboratory challenges, mechanistic insights and therapeutic strategies. Expert Rev. Anti Infect. Ther. 2013, 11, 395–409. [Google Scholar] [CrossRef] [PubMed]
- Kolar, M.; Htoutou Sedlakova, M.; Urbanek, K.; Mlynarcik, P.; Roderova, M.; Hricova, K.; Mezerova, K.; Kucova, P.; Zapletalova, J.; Fiserova, K.; et al. Implementation of Antibiotic Stewardship in a University Hospital Setting. Antibiotics 2021, 10, 93. [Google Scholar] [CrossRef]
- Mlynarcik, P.; Kolar, M. Molecular mechanisms of polymyxin resistance and detection of mcr genes. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc. Czech. Repub. 2019, 163, 28–38. [Google Scholar] [CrossRef]
- Mlynarcik, P.; Chalachanova, A.; Vagnerova, I.; Holy, O.; Zatloukalova, S.; Kolar, M. PCR Detection of Oxacillinases in Bacteria. Microb Drug Resist. 2020. [Google Scholar] [CrossRef]
- Bendjama, E.; Loucif, L.; Chelaghma, W.; Attal, C.; Bellakh, F.Z.; Benaldjia, R.; Kahlat, I.; Meddour, A.; Rolain, J.M. First detection of an OXA-48-producing Enterobacter cloacae isolate from currency coins in Algeria. J. Glob. Antimicrob. Resist. 2020, 23, 162–166. [Google Scholar] [CrossRef]
- Gniadkowski, M. Evolution and epidemiology of extended-spectrum beta-lactamases (ESBLs) and ESBL-producing microorganisms. Clin. Microbiol. Infect. 2001, 7, 597–608. [Google Scholar] [CrossRef]
- Canton, R.; Novais, A.; Valverde, A.; Machado, E.; Peixe, L.; Baquero, F.; Coque, T.M. Prevalence and spread of extended-spectrum beta-lactamase-producing Enterobacteriaceae in Europe. Clin. Microbiol. Infect. 2008, 14 (Suppl. 1), 144–153. [Google Scholar] [CrossRef]
- Bush, K. The ABCD’s of beta-lactamase nomenclature. J. Infect. Chemother. 2013, 19, 549–559. [Google Scholar] [CrossRef]
- Mlynarcik, P.; Bardon, J.; Htoutou Sedlakova, M.; Prochazkova, P.; Kolar, M. Identification of novel OXA-134-like beta-lactamases in Acinetobacter lwoffii and Acinetobacter schindleri isolated from chicken litter. Biomed. Pap. Med. Fac. Univ. Palacky Olomouc. Czech. Repub. 2019, 163, 141–146. [Google Scholar] [CrossRef]
- Mlynarcik, P.; Roderova, M.; Kolar, M. Primer Evaluation for PCR and its Application for Detection of Carbapenemases in Enterobacteriaceae. Jundishapur. J. Microbiol. 2016, 9, e29314. [Google Scholar] [CrossRef]
- Mlynarcik, P.; Dolejska, M.; Vagnerova, I.; Kutilová, I.; Kolar, M. Detection of clinically important β-lactamases by using PCR. FEMS Microbiol. Lett. 2021, 368, fnab068. [Google Scholar] [CrossRef] [PubMed]
- Peleg, A.Y.; Seifert, H.; Paterson, D.L. Acinetobacter baumannii: Emergence of a successful pathogen. Clin. Microbiol. Rev. 2008, 21, 538–582. [Google Scholar] [CrossRef] [PubMed]
- Peymani, A.; Naserpour-Farivar, T.; Zare, E.; Azarhoosh, K.H. Distribution of blaTEM, blaSHV, and blaCTX-M genes among ESBL-producing P. aeruginosa isolated from Qazvin and Tehran hospitals, Iran. J. Prev. Med. Hyg. 2017, 58, E155–E160. [Google Scholar] [PubMed]
- Pagani, L.; Dell’Amico, E.; Migliavacca, R.; D’Andrea, M.M.; Giacobone, E.; Amicosante, G.; Romero, E.; Rossolini, G.M. Multiple CTX-M-Type extended-spectrum b-lactamases in nosocomial isolates of Enterobacteriaceae from a hospital in northern Italy. J. Clin. Microbiol. 2003, 41, 4264–4269. [Google Scholar] [CrossRef] [PubMed]
| Bacterial Genera | ESBL Enzymes | Bacterial Genera | ESBL Enzymes |
|---|---|---|---|
| Achromobacter | TEM | Legionella | TEM |
| Acinetobacter | CTX-M, SHV, TEM | Lysobacter | TEM |
| Aeromonas | CTX-M, SHV, TEM | Morganella | CTX-M, SHV, TEM |
| Alcaligenes | TEM | Mycobacterium | TEM |
| Atlantibacter | CTX-M, TEM | Neisseia | TEM |
| Bacillus | CTX-M, SHV, TEM | Nocardia | TEM |
| Bordetella | TEM | Ochrobactrum | CTX-M, SHV |
| Brevibacterium | CTX-M, TEM | Pantoea | CTX-M, TEM |
| Brucella | TEM | Pasteurella | TEM |
| Burkholderia | CTX-M, SHV | Proteus | CTX-M, SHV, TEM |
| Campylobacter | TEM | Providencia | CTX-M, TEM |
| Cedecea | SHV | Pseudomonas | CTX-M, SHV, TEM |
| Citrobacter | CTX-M, SHV, TEM | Pseudochrobactrum | CTX-M |
| Cronobacter | SHV, TEM | Ralstonia | TEM |
| Elizabethkingia (formerly Chryseobacterium) | TEM | Raoultella | CTX-M, SHV, TEM |
| Enterobacter | CTX-M, SHV, TEM | Salmonella | CTX-M, SHV, TEM |
| Enterococcus | SHV, TEM | Serratia | CTX-M, SHV, TEM |
| Erwinia | TEM | Shewanella | CTX-M |
| Escherichia | CTX-M, SHV, TEM | Shigella | CTX-M, SHV, TEM |
| Haemophilus | TEM | Staphylococcus | TEM |
| Hafnia | TEM | Stenotrophomonas | SHV |
| Kerstersia | SHV | Streptococcus | TEM |
| Klebsiella | CTX-M, SHV, TEM | Streptomyces | TEM |
| Kluyvera | CTX-M, SHV, TEM | Superficieibacter | SHV, TEM |
| Lactobacillus | TEM | Vibrio | CTX-M, TEM |
| Leclercia | CTX-M, TEM | Yersinia | TEM |
| Amino Acid Change | Number of Mutations (Amino Acid Position) | ||||||
|---|---|---|---|---|---|---|---|
| from | to | CTX-M * | OXY * | PER * | SHV * | TEM * | VEB * |
| A | C | 7 (22) | 2 (7) | ||||
| A | D | 36 (32/131/157) | 3 (114) | 1 (264) | 1 (230) | ||
| A | E | 28 (35/46/157/222/236) | 21 (230) | 2 (114/166) | 1 (38) | ||
| A | G | 158 (16/64/73/131/136/157/297) | 10 (240) | 1 (7) | 3 (205/259) | 2 (235/280) | |
| A | I | 1 (16) | |||||
| A | K | 22 (64/244) | |||||
| A | M | 1 (73) | |||||
| A | N | 36 (35) | |||||
| A | P | 78 (59/78/272) | 1 (15) | 1 (183) | |||
| A | Q | 1 (157) | 2 (151) | ||||
| A | R | 1 (222) | 1 (151) | 1 (185) | |||
| A | S | 102 (35/46/151/235/244) | 2 (240) | 4 (109) | 1 (23) | 3 (9/40/266) | |
| A | T | 110 (16/18/35/90/120/131/137/161/222/244) | 108 (16/30/32/38/56/129/171/208/240/284) | 1 (11) | 14 (20/23/63/90/125/135/157/161/204) | 12 (9/183/235/280) | 4 (248) |
| A | V | 74 (16/35/88/131/135/137/163) | 32 (16/233) | 20 (20/63/125/135/137/145/157/188/205/218/264/288) | 18 (23/40/182/222/266/276) | ||
| A | W | 1 (299) | |||||
| C | N | 1 (9) | |||||
| C | R | 1 (134) | |||||
| C | S | 1 (9) | |||||
| D | A | 69 (36) | 1 (39) | 4 (155) | 3 (195/231) | ||
| D | E | 160 (218/303) | 1 (242) | 2 (168/230) | 2 (155/177) | ||
| D | G | 179 (259/270) | 4 (115/195/230/231) | 2 (113/161) | |||
| D | H | 23 (36/292/303) | 2 (112) | 1 (161) | |||
| D | K | 2 (303) | |||||
| D | N | 40 (36/218/270/303) | 28 (39/58) | 1 (195) | 3 (36/113/174) | ||
| D | P | 1 (292) | 1 (33) | ||||
| D | Q | 1 (218) | |||||
| D | S | 25 (36/65/256/303) | |||||
| E | A | 106 (47/177) | |||||
| E | D | 35 (98/169) | 22 (92/99/149/278) | 2 (29/273) | 1 (26) | ||
| E | G | 1 (47) | 1 (33) | 3 (59/75/177) | 2 (164/237) | ||
| E | K | 3 (47/132/169) | 1 (278) | 37 (29/100/256/289) | 82 (26/62/102/166/237) | ||
| E | N | 1 (169) | |||||
| E | P | 1 (107) | |||||
| E | Q | 85 (98/132) | 4 (92) | 8 (119/193/224) | 1 (75) | ||
| E | R | 1 (256) | 1 (237) | ||||
| E | T | 1 (98) | |||||
| E | V | 3 (23) | 1 (237) | ||||
| F | A | 10 (12) | |||||
| F | L | 1 (12) | 3 (19) | 3 (14/228) | |||
| F | M | 72 (12) | |||||
| F | S | 21 (12) | 1 (162) | ||||
| F | W | 1 (22) | |||||
| F | Y | 2 (64) | |||||
| G | A | 31 (25/217) | 4 (149/190) | 6 (158/191/255) | |||
| G | C | 3 (47/132/169) | 1 (258) | ||||
| G | D | 1 (159) | 11 (98/154/167/245) | 4 (39/90/194/236) | |||
| G | E | 1 (25) | 1 (41) | 2 (158/296) | 1 (216) | ||
| G | H | 1 (304) | |||||
| G | K | 1 (126) | |||||
| G | N | 1 (53) | 1 (236) | ||||
| G | R | 4 (54/234/304) | 1 (296) | 2 (154/236) | |||
| G | S | 27 (25/213/234/253/304) | 30 (24/75) | 42 (65/155/167/255) | 40 (194/236) | ||
| G | V | 2 (194/263) | |||||
| H | D | 2 (134/154) | |||||
| H | F | 1 (123) | |||||
| H | K | 1 (152) | |||||
| H | L | 2 (134) | 4 (285) | ||||
| H | N | 1 (156) | |||||
| H | Q | 74 (152) | 1 (22) | ||||
| H | R | 5 (24/151) | |||||
| H | T | 1 (107) | |||||
| H | Y | 3 (123) | |||||
| I | A | 1 (69) | 2 (5/97) | ||||
| I | F | 1 (119) | 11 (6) | 1 (11) | |||
| I | L | 1 (150) | 5 (33) | 3 (61/194) | 2 (297/302) | ||
| I | M | 4 (111) | 1 (275) | 1 (259) | |||
| I | P | 4 (3) | |||||
| I | R | 1 (11) | |||||
| I | S | 1 (300) | |||||
| I | T | 13 (69) | 7 (5/40/97) | 2 (275/297) | 1 (275) | 1 (18) | |
| I | V | 170 (69/108/266/275) | 7 (111) | 15 (61/162/194/245/258) | 3 (6/58) | 5 (125/171/256) | 14 (18) |
| K | A | 5 (109/110/299) | 23 (102) | 3 (102) | |||
| K | D | 1 (214) | |||||
| K | E | 36 (93/122/269) | 27 (180) | 1 (113) | 1 (32) | ||
| K | G | 1 (214) | |||||
| K | H | 73 (4/214) | |||||
| K | L | 1 (148) | 20 (140) | ||||
| K | N | 80 (3/214/286) | 20 (101) | 2 (209/271) | |||
| K | P | 81 (110) | |||||
| K | Q | 217 (4/44/94/109/269/286) | 2 (196) | 1 (144) | |||
| K | R | 103 (3/44/94/109/122/229/251/299) | 7 (196/246) | 5 (251/271) | 2 (237) | ||
| K | S | 7 (110/214) | |||||
| K | T | 91 (3/269) | 1 (190) | ||||
| L | A | 4 (9/14/23/210) | 2 (256) | ||||
| L | F | 121 (14/24/30/130/305) | 1 (84) | 6 (124/133/189) | 23 (19/55/100/136) | 1 (56) | |
| L | G | 2 (30/70) | |||||
| L | I | 111 (9/149/166/276) | 24 (2) | 1 (256) | 4 (19/135/218) | ||
| L | M | 5 (9/22/102/207/242) | 23 (2/11) | 4 (75/260) | 2 (17/237) | 3 (47/219) | |
| L | N | 1 (305) | |||||
| L | P | 4 (130/149/212) | 1 (262) | 4 (38/62/102/301) | 1 (49) | ||
| L | Q | 2 (60/180) | 73 (33/62) | ||||
| L | R | 5 (8/102/133/185) | 1 (10) | ||||
| L | S | 3 (14/23/305) | |||||
| L | V | 193 (9/60/101/153/216) | 3 (305) | 2 (159/210) | 4 (28/38/100/247) | ||
| L | Y | 15 (224/305) | |||||
| M | A | 69 (14) | |||||
| M | G | 1 (197) | |||||
| M | I | 1 (86) | 35 (12/141) | 1 (160) | 3 (80) | 10 (66/67/153/180) | |
| M | K | 4 (3) | |||||
| M | L | 105 (228) | 20 (12) | 3 (160) | 1 (80) | 17 (67) | |
| M | T | 21 (14) | 34 (180) | ||||
| M | V | 7 (80/140/228) | 11 (67/184) | ||||
| N | D | 89 (62/125/143) | 36 (54/90/200/256) | 3 (308) | 2 (269/291) | 12 (272) | 6 (294) |
| N | E | 1 (86) | |||||
| N | G | 12 (66/100/125) | |||||
| N | H | 25 (36/65/256/303) | 10 (90) | 2 (269) | 4 (134/173) | ||
| N | I | 1 (270) | 1 (173) | ||||
| N | K | 2 (100/209) | 1 (169) | ||||
| N | L | 1 (100) | |||||
| N | Q | 141 (100/209) | |||||
| N | S | 27 (100/103/117/181) | 3 (98/272) | ||||
| N | T | 1 (66) | 1 (168) | ||||
| N | W | 1 (209) | |||||
| N | Y | 1 (115) | 1 (200) | ||||
| P | A | 3 (29/178/194) | 3 (123) | 1 (143) | |||
| P | G | 1 (268) | |||||
| P | H | 1 (178) | |||||
| P | K | 105 (99) | 23 (91) | ||||
| P | L | 3 (178/188) | 1 (269) | 3 (156/268/284) | |||
| P | N | 4 (91) | |||||
| P | Q | 95 (156/178/185/285) | |||||
| P | S | 18 (178/283) | 1 (170) | 7 (18/27/190/236/243/284) | 1 (143) | ||
| P | T | 29 (29/178/283) | 1 (123) | ||||
| Q | A | 77 (11/165/220) | |||||
| Q | D | 1 (199) | |||||
| Q | E | 35 (165/284) | 1 (165) | ||||
| Q | H | 1 (68) | 23 (34/156) | 2 (223/293) | 1 (88) | ||
| Q | I | 1 (68) | |||||
| Q | K | 2 (11/223) | 2 (94/183) | 38 (4/37) | |||
| Q | L | 3 (11/165/199) | 4 (156/270) | 1 (129) | |||
| Q | P | 1 (116) | 2 (175/214) | 1 (203) | |||
| Q | R | 198 (11/68/239) | 27 (35/156) | 1 (158) | 1 (37) | 3 (97/204) | |
| Q | S | 128 (33/51/165) | |||||
| Q | T | 10 (165) | |||||
| Q | V | ||||||
| R | A | 4 (176/271) | |||||
| R | C | 1 (72) | 9 (162/241) | ||||
| R | G | 2 (10/175) | 2 (232) | 2 (118/241) | |||
| R | H | 25 (10/271/291) | 9 (43/68/194) | 5 (54/72/219) | 22 (162/238/241) | ||
| R | K | 131 (10/50/105/208) | |||||
| R | L | 37 (10/72/195) | 7 (76/175/222/232/290) | 4 (241/271) | |||
| R | M | 1 (97) | |||||
| R | N | 1 (43) | |||||
| R | P | 73 (76/105) | 1 (175) | 1 (41) | |||
| R | Q | 74 (10/50/208/221/290) | 4 (257/307) | 6 (202/271) | |||
| R | S | 4 (10/105/175) | 10 (43/225) | 17 (54/109/219) | 37 (63/162/241) | ||
| R | T | 1 (41) | |||||
| R | V | 2 (72) | |||||
| S | A | 110 (52/67/111/296) | 45 (25/89/103) | 1 (280) | |||
| S | C | 1 (245) | 1 (4) | 1 (221) | |||
| S | D | 4 (2) | |||||
| S | E | 2 (280) | |||||
| S | F | 1 (12) | |||||
| S | G | 101 (111/134/141/158/237/245) | 39 (143/231) | 2 (18) | 2 (36/144) | 7 (128/264) | |
| S | H | 2 (289) | |||||
| S | I | 3 (134/254) | 1 (192) | 2 (286) | |||
| S | K | 36 (158/296) | |||||
| S | L | 1 (220) | |||||
| S | N | 6 (26/237/254/289) | 29 (4/29/157/275) | 3 (81/213) | 1 (122) | ||
| S | P | 1 (104) | |||||
| S | Q | 1 (97) | |||||
| S | R | 83 (5/237/289) | |||||
| S | T | 275 (67/97/129/141) | 37 (4/44/174) | 10 (12/37/121/135) | 1 (117) | 3 (128/233) | 2 (131) |
| S | V | 2 (219) | 2 (89) | ||||
| T | A | 192 (13/17/19/21/34/144/206/211/226/281/302) | 11 (13/165/293) | 9 (16/69/82/252) | 9 (25/176/219) | ||
| T | C | 81 (19) | |||||
| T | D | 1 (233) | |||||
| T | G | 3 (17/19) | 1 (178) | ||||
| T | H | 2 (226/233) | |||||
| T | I | 6 (176/182/226/278) | 48 (9/61) | 4 (152/212) | 1 (267) | ||
| T | K | 3 (34/170/211) | 3 (202) | 1 (186) | |||
| T | M | 134 (13/17/226) | 8 (168) | 1 (283) | 19 (261) | 10 (104) | |
| T | N | 1 (176) | 2 (16/212) | 1 (186) | |||
| T | P | 90 (19/21) | 1 (16) | 1 (112) | |||
| T | S | 130 (21/127/170/179/182/192/193/206) | 34 (9/184) | 3 (95/221) | 4 (129/160/283) | ||
| T | V | 108 (144/206) | 1 (61) | 3 (24) | |||
| V | A | 98 (27/85/106/248) | 20 (161) | 1 (290) | 2 (86/91) | 2 (19) | |
| V | E | 1 (78) | 7 (19) | ||||
| V | F | 1 (293) | 3 (9) | 2 (153) | 1 (258) | ||
| V | G | 3 (37/247) | |||||
| V | I | 175 (2/20/37/114/159/293/301) | 2 (261) | 2 (235) | 1 (130) | 6 (82) | |
| V | L | 22 (20/91) | 11 (17/94) | 3 (17) | 1 (86) | ||
| V | M | 35 (2/37) | 5 (86/91/95/277) | ||||
| V | P | 1 (27) | |||||
| V | Q | 2 (27/106) | |||||
| V | R | ||||||
| V | S | 2 (225/247) | 1 (42) | ||||
| V | T | 68 (247) | |||||
| W | C | 1 (163) | |||||
| W | G | 2 (163) | |||||
| W | L | 1 (163) | |||||
| W | P | 1 (176 | |||||
| W | R | 1 (246) | 1 (71) | 5 (163) | |||
| Y | C | 10 (31) | |||||
| Y | F | 1 (31) | 8 (5) | 1 (103) | |||
| Y | G | ||||||
| Y | H | 22 (31) | 1 (144) | 4 (115) | 2 (5/108) | ||
| Y | N | 1 (103) | |||||
| Y | S | 1 (31) | 1 (103) | ||||
| Y | W | 1 (71) | |||||
| Target (Subtype) | Primer Name | Sequence (5′ to 3′ Direction) a | Natural (N) or Acquired (A)/Phenotype | Amplicon Size (bp) | Tm (°C) | Reference |
|---|---|---|---|---|---|---|
| ACI (1) | ACI-F/R | CCGTTGACATGGAGAATGG, GCGTGTCGGTTATGGAATT | N/ESBL | 507 | 54 | This study |
| ADC (178) | ADC-F1/R1 | MAACCTAAAAACYCAATCGGTG, YGGATAAGMAAACTCTTCCCA | N/AmpC, ESBL or CARBA | 417–420 | 58 | [34] |
| ADC (29) | ADC-F2/R2 | RGGTTTCTAYCAAGTCGGYA, GCGTTCTTCATTBGGAATACGT | 268 | 59 | ||
| ADC (5) | ADC-F3/R3 | TGGTCTACAATCCGTTCAAGA, GCCGGGGTTAACTCGAAT | 517 | 54 | ||
| ADC (7) | ADC-F4/R4 | TATRATGTGCCGGGTATGG, RTCTGTTTGTACTTCAYCTGG | 318 | 54 | This study | |
| BEL (4) | BEL-F/R | CGTTCCTTGAAGAGTACGC, ACCCGTTACCCATGAATCA | A/ESBL | 401 | 53 | This study |
| BES (1) | BES-F/R | ATAAGCGGGTGCATTATGC, CTTTAAGCCAGCTCACCAG | A/ESBL | 363 | 53 | This study |
| BPS (11) | BPS-F/R | GCTYCAGTACAGCGACAAC, GTCKTGTTGCCGAGCATCCA | N/cephalosporinase or ESBL | 270 | 57 | This study |
| BPU (1) | BPU-F/R | AAGAAAAGTCCCCATGGGT, CGAACTTGTTCGATGGGAG | N/ESBL | 364 | 55 | This study |
| CARB (8) | CARB-F2/R2 | GGGAAAACGTTGGGAACAT, TAATAGCACGCGACCCATA | N or A/BSBL or ESBL | 578 | 54 | |
| CDD (2) | CDD-F/R | AACAAGTGCAAACAATGGC, TTCCTTTACCTTTGGCCCT | N/ESBL | 266 | 52 | This study |
| CdiA (2) | CdiA-F/R | CGTGCTCGCTTTCTTTACT, CACCTGCTCCGGTTTTATC | N/penicillinase or ESBL | 692 | 53 | This study |
| CepA (6) | CepA-F/R | AGTGACAATAATGCCTGCG, TGCTTCGGAATCTTTCACG | N/ESBL | 438 | 52 | This study |
| CfxA (13) | CfxA-F/R | GAAATTGGTGTGGCGGTTA, CAGCACCAAGAGGAGATGT | N or A/BSBL or ESBL | 442 | 53 | This study |
| CGA (1) | CGA-F/R | AGCTACAGTCGGTGTTTCT, TTCATTTTCTGCGCCTGTT | N/ESBL | 640 | 53 | This study |
| CIA (4) | CIA-F/R | GATGGTTTCTGCCTTTGCT, CTTCCGGAAATTTTTCGCG | N/ESBL | 299 | 53 | This study |
| CME (3) | CME-F/R | CCAAAGTGACAACAACGGA, TCCTGAATCGTTCTCAGCA | N/ESBL | 376 | 53 | This study |
| CSP (1) | CSP-F/R | TCTGCTGAGGTTGATTGGA, TCCCACATCATTGGTAGCA | N/ESBL | 346 | 53 | This study |
| CTX-M (208), KLUB (2), KLUG (5), KLUY (5) | CTX-M-F1/R1 | ATGTGCAGYACCAGTAARGT, TGGGTRAARTARGTSACCAGA | N or A/ESBL or CARBA | 593 | 55 | [37] |
| CTX-M (7) | CTX-M-F2/R2 | ATGTGCAGYACCAGYAAAG, GGCCARATCACCGCRATAT | 551 | 56 | [34] | |
| CumA (3) | CumA-F/R | ATCTCCAATGCTATGGGCT, TCACGAGGATCACCATGAA | N/BSBL or ESBL | 483 | 53 | This study |
| DES (1) | DES-F/R | GTTCCAGTTATTCCAGGCG, TGCCAGCACTTTAAAGGTG | N/ESBL | 269 | 53 | This study |
| ERP (1) | ERP-F/R | GTATCGGGCTGTCTCTGAT, GCTGTGCTGTCTGTAATCC | N/ESBL | 477 | 54 | This study |
| FAR (1) | FAR-F/R | CTGAAGAAATCTGGTCGCC, AGCAGTTTCAGGATCTGGT | N/ESBL | 473 | 53 | This study |
| FONA (8) | FONA-F/R | CCGATCTGGTCAACTACAAC, CCCTTCATCCATTCAACCAG | N/ESBL | 340 | 55 | This study |
| GES (45) | GES-F1/R1 | ACGTTCAAGTTTCCGCTAG, GGCAACTAATTCGTCACGT | N or A/ESBL or CARBA | 624 | 53 | [25] |
| GES (1) | GES-F2/R2 | ATGATCGTGGAGTGGAGCCC, AAGAAGCCGATGTCGTTGCG | 448 | 58 | This study | |
| HMB (1), KHM (1) | HMB, KHM-F/R | AAATCGAAGCYTTTTATCCGGG, TTTCCAGCAGCGATGCRTCG | N or A/ESBL or CARBA | 237 | 60 | This study |
| KLUA (12) | KLUA-F/R | CGCTCAATGTTAACGGTGA, TTCATGGCAGTATTGTCGC | N/ESBL | 395 | 52 | This study |
| KLUC (6) | KLUC-F/R | CGATTGCGGAAAAACATGT, CGCCGAGGCTAAWACATC | N or A/ESBL | 521 | 53 | This study |
| KPC (52) | KPC-F/R | CGCTAAACTCGAACAGGAC, CGGTCGTGTTTCCCTTTAG | N or A/ESBL or CARBA | 548 | 54 | [34] |
| LUT (6) | LUT-F/R | TGCTCATGAAAAAGCTGGG, ACCTGTCTTATCGCCTACC | N/BSBL or ESBL | 299 | 54 | This study |
| L2 (18) | L2-F1/R1 | TTCCCGATGTGCAGCAC, TTGCTGCCGGTCTTGTC | N/ESBL | 518 | 53 | [34] |
| OHIO (1) | OHIO-F/R | CTTTCCCATGATGAGCACC, CCCGCAGATAAATCACCAC | A/ESBL | 599 | 54 | This study |
| OXA-1-like (11) | OXA(1)-F/R | TCTGTTGTTTGGGTTTCGC, TCTATGGTGTTTTCTATGGCTG | N or A/NSBL, BSBL or ESBL | 245 | 53 | [27] |
| OXA-2-like (22) | OXA(2)-F/R | GATAGTTGTGGCAGACGAAC, TCCATYCTGTTTGGCGTATC | N or A/ESBL | 603 | 55 | This study |
| OXA-10-like (38) | OXA(3)-F/R | ACAAAGAGTTCTCTGCCGAA, TCCACTTGATTAACTGCGGA | N or A/ESBL | 418 | 53 | This study |
| OXA-18 | OXA(4)-F/R | ACCATCTGGCTGAAGGATT, CAGAAGTTTTCCGACAGGG | A/ESBL | 506 | 54 | This study |
| OXA-23-like (41) | OXA(5)-F/R | ACTAGGAGAAGCCATGAAGC, ATTTTTCCATCTGGCTGCTC | N or A/ESBL or CARBA | 369 | 55 | [34] |
| OXA-45 | OXA(6)-F/R | GCGGTAAACACACTGTCAT, GGGTCAATTGCTGCGAATA | A/ESBL | 333 | 52 | This study |
| OXA-48-like (38) | OXA(7)-F/R | ACCARGCATTTTTACCCGCA, GGCATATCCATATTCATCGC | N or A/ESBL or CARBA | 538 | 55 | This study |
| OXY (28) | OXY-F1/R1 | TAAAGTGATGGCYGCCGC, RTTGGTGGTGCCGTAATC | N/ESBL | 517 | 54 | This study |
| OXY (20) | OXY-F2/R2 | CCCTGCCTTTATTGCTCTG, TTTATCTCCCACGACCCAG | 665 | 54 | ||
| PER (12) | PER-F1/R1 | CTCGACGCTACTGATGGTA, TTCATTGGTTCGGCTTGAC | N or A/ESBL | 820 | 54 | This study |
| PER (3) | PER-F2/R2 | CTGTTAATCGTGCTGCAGT, GACAAATACCGCCACCAAT | 530 | 53 | ||
| PME (1) | PME-F/R | GATCCACTTCAGCGATGAC, GACATCGTGGGTCTTGTTC | A/ESBL | 478 | 54 | This study |
| RAHN (2) | RAHN-F/R | ATGACGTCAGTTCAGCAAC, CATCCATTCCACCAGTTGC | N/ESBL | 555 | 54 | This study |
| RSA1 (1) | RSA-F1/R1 | TCGACGATCCTCACTGTTT, GTTGGTGTTCAAATCGGGT | N or A/ESBL or CARBA | 483 | 53 | This study |
| RSA2 (1) | RSA-F2/R2 | TGTGGACCTTTCCGAAGAA, CGCGATCAGATTACGAGTG | 475 | 55 | ||
| SGM (7) | SGM-F/R | CATGTGCTCGACCTTCAAG, ATCGGCAGCARCAGRTTGG | N/ESBL | 225 | 58 | This study |
| SHV (199) | SHV-F/R | TGGATGCCGNTGACNAACAGC, NTATCGGCGATAAACCAGNCC | N or A/BSBL, ESBL or CARBA | 451 | 59 | [34] |
| SMO (1) | SMO-F/R | CTCACAGACCGTATACCGT, GAATGTCTCATCGCCGATC | N/ESBL | 316 | 54 | This study |
| TEM (199) | TEM-F/R | CACCAGTCACAGAAAAGCA, AGGGCTTACCATCTGGC | N or A/BSBL or ESBL | 450 | 54 | [34] |
| SFO (1) | SFO-F/R | CTCGAGAAAAACTCCGGTG, GTTAGGGTTTGCAGGCTTT | A/ESBL | 473 | 54 | This study |
| TLA (2) | TLA-F/R | GCTAAAGGTACGGATTCGC, CTTAACGCCAAGCTTGCTA | A/ESBL | 417 | 54 | This study |
| TLA2 (1) | TLA2-F/R | ATCGTGCTTGCTGTTTTGA, TCATTTGCCGCATTGTTCT | A/ESBL | 623 | 52 | This study |
| VEB (27) | VEB-F/R | TTTCCGATTGCTTTAGCCG, CCCCAACATCATTAGTGGC | N or A/ESBL | 553 | 54 | [34] |
| YOC (1) | YOC-F/R | CCGGCATCAGAAGAGAAAAA, GGATTCGGGTAGCTTTTGTT | N/ESBL | 467 | 54 | This study |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mlynarcik, P.; Chudobova, H.; Zdarska, V.; Kolar, M. In Silico Analysis of Extended-Spectrum β-Lactamases in Bacteria. Antibiotics 2021, 10, 812. https://doi.org/10.3390/antibiotics10070812
Mlynarcik P, Chudobova H, Zdarska V, Kolar M. In Silico Analysis of Extended-Spectrum β-Lactamases in Bacteria. Antibiotics. 2021; 10(7):812. https://doi.org/10.3390/antibiotics10070812
Chicago/Turabian StyleMlynarcik, Patrik, Hana Chudobova, Veronika Zdarska, and Milan Kolar. 2021. "In Silico Analysis of Extended-Spectrum β-Lactamases in Bacteria" Antibiotics 10, no. 7: 812. https://doi.org/10.3390/antibiotics10070812
APA StyleMlynarcik, P., Chudobova, H., Zdarska, V., & Kolar, M. (2021). In Silico Analysis of Extended-Spectrum β-Lactamases in Bacteria. Antibiotics, 10(7), 812. https://doi.org/10.3390/antibiotics10070812









