Third Generation Cephalosporin Resistant Enterobacterales Infections in Hospitalized Horses and Donkeys: A Case–Case–Control Analysis
Abstract
1. Introduction
2. Results
2.1. Population Characteristics
2.2. Predictors of 3GCRE Infections
2.3. Clinical Outcomes of 3GCRE Infections
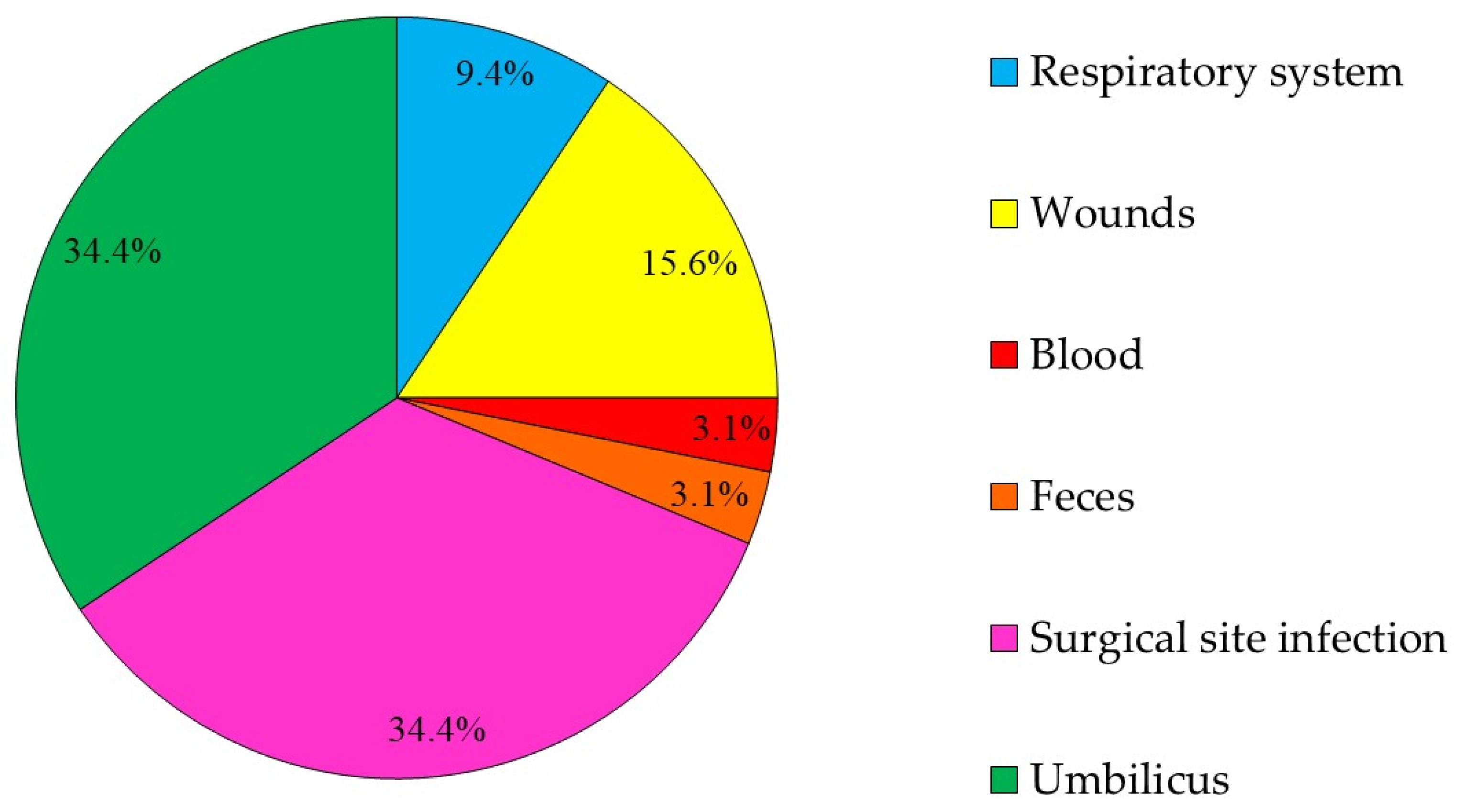
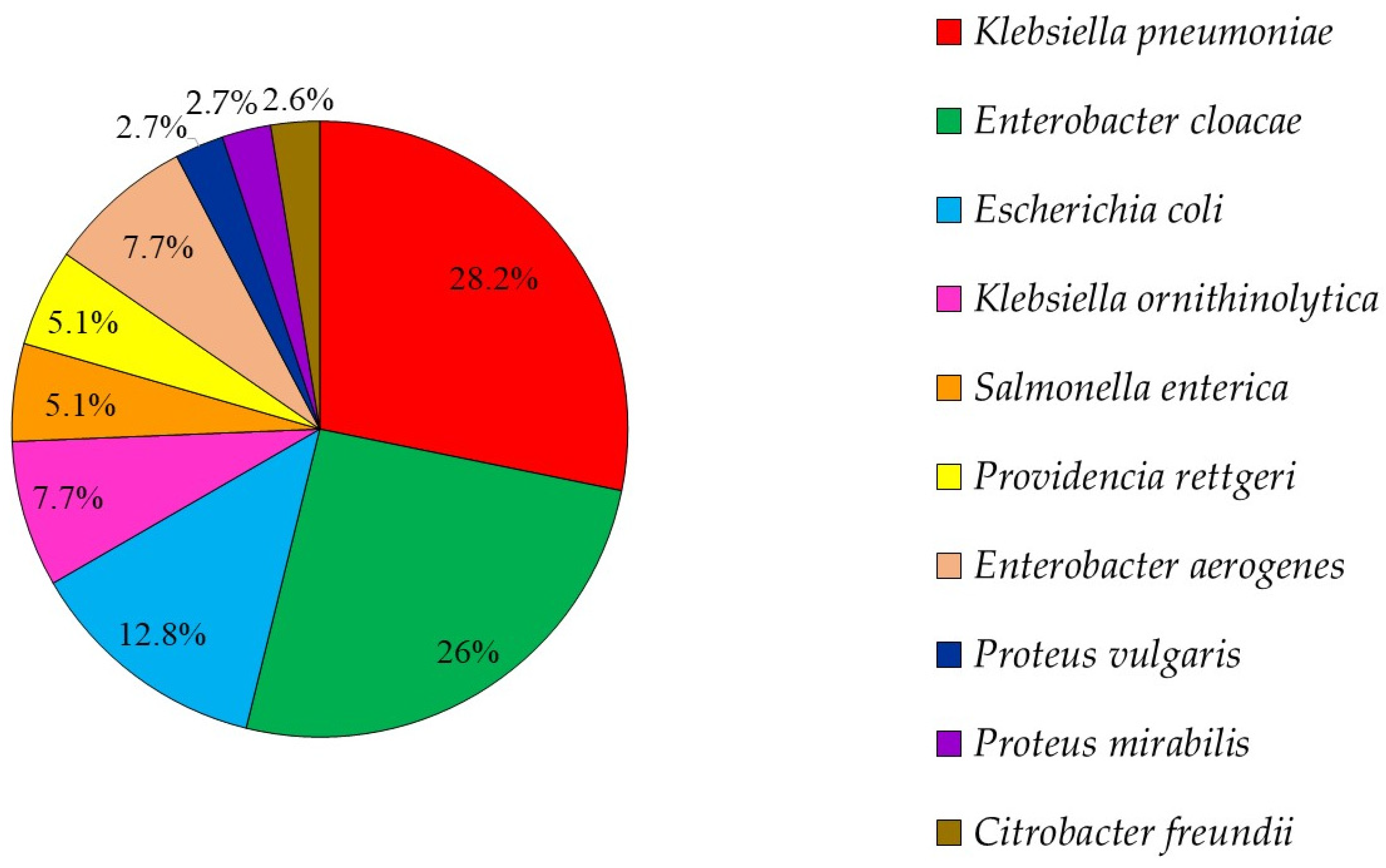
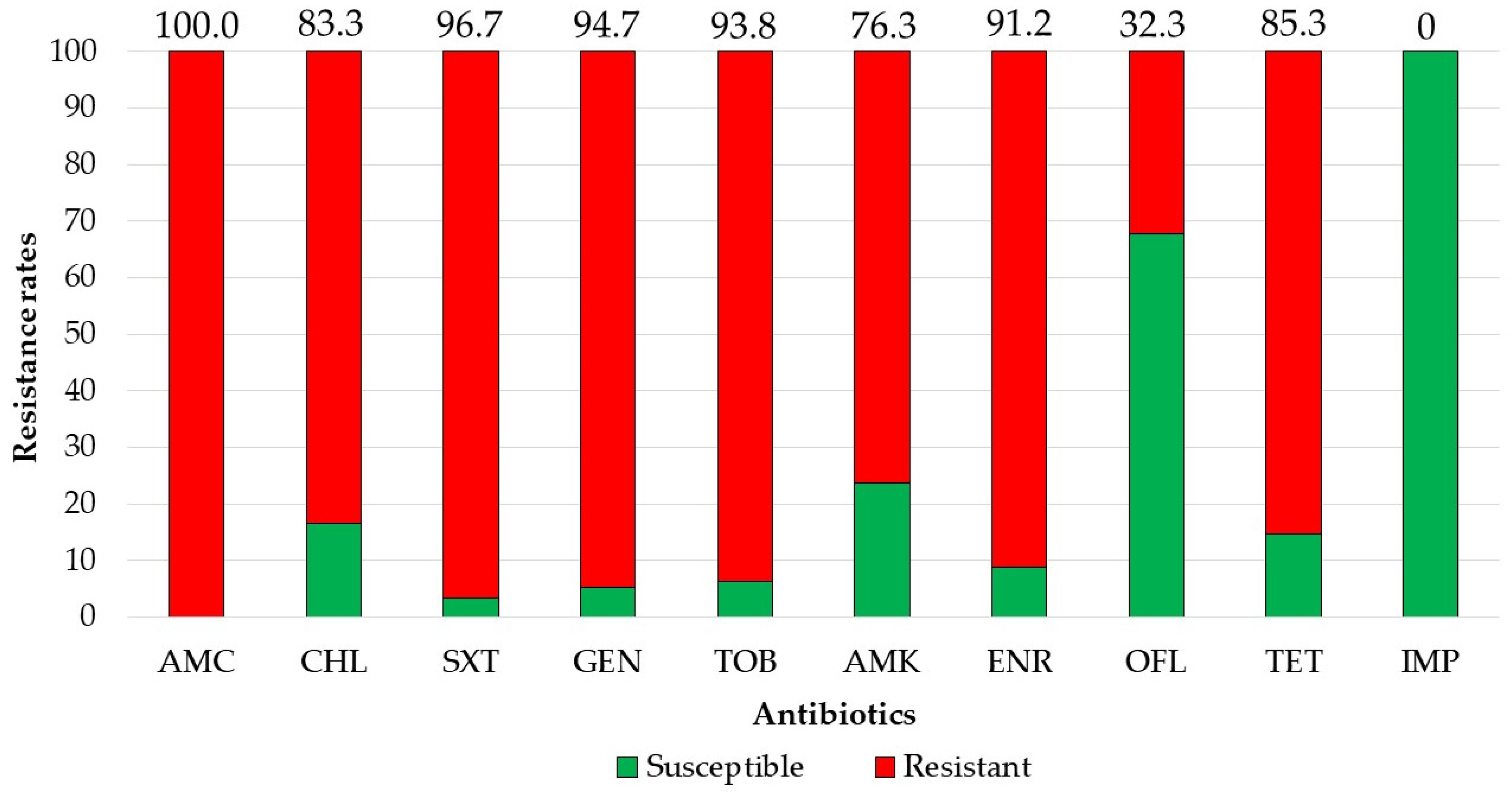
2.4. GCRE Samples Description, Species Distribution and Resistance Rates
2.5. Molecular Characteristics of 3GCRE Isolates
3. Discussion
4. Materials and Methods
4.1. Study Design
4.2. Bacterial Isolates Collection, Identification and Susceptibility Testing
4.3. Molecular Characterization of ESBL-PE
4.4. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Singh, R.; Singh, A.P.; Kumar, S.; Giri, B.S.; Kim, K.-H. Antibiotic Resistance in Major Rivers in the World: A Systematic Review on Occurrence, Emergence, and Management Strategies. J. Clean. Prod. 2019, 234, 1484–1505. [Google Scholar] [CrossRef]
- Lee, J.H.; Bae, I.K.; Lee, S.H. New Definitions of Extended-Spectrum β-Lactamase Conferring Worldwide Emerging Antibiotic Resistance. Med. Res. Rev. 2012, 32, 216–232. [Google Scholar] [CrossRef] [PubMed]
- Schwaber, M.J.; Carmeli, Y. Mortality and Delay in Effective Therapy Associated with Extended-Spectrum Beta-Lactamase Production in Enterobacteriaceae Bacteraemia: A Systematic Review and Meta-Analysis. J. Antimicrob. Chemother. 2007, 60, 913–920. [Google Scholar] [CrossRef]
- Paul, M.; Shani, V.; Muchtar, E.; Kariv, G.; Robenshtok, E.; Leibovici, L. Systematic Review and Meta-Analysis of the Efficacy of Appropriate Empiric Antibiotic Therapy for Sepsis. Antimicrob. Agents Chemother. 2010, 54, 4851–4863. [Google Scholar] [CrossRef]
- Biehl, L.M.; Schmidt-Hieber, M.; Liss, B.; Cornely, O.A.; Vehreschild, M.J.G.T. Colonization and Infection with Extended Spectrum Beta-Lactamase Producing Enterobacteriaceae in High-Risk Patients - Review of the Literature from a Clinical Perspective. Crit. Rev. Microbiol. 2016, 42, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Coque, T.M.; Baquero, F.; Cantón, R. Increasing Prevalence of ESBL-Producing Enterobacteriaceae in Europe. Eurosurveillance 2008, 13, 19044. [Google Scholar]
- Dupouy, V.; Abdelli, M.; Moyano, G.; Arpaillange, N.; Bibbal, D.; Cadiergues, M.-C.; Lopez-Pulin, D.; Sayah-Jeanne, S.; de Gunzburg, J.; Saint-Lu, N.; et al. Prevalence of Beta-Lactam and Quinolone/Fluoroquinolone Resistance in Enterobacteriaceae From Dogs in France and Spain—Characterization of ESBL/PAmpC Isolates, Genes, and Conjugative Plasmids. Front. Vet. Sci. 2019, 6, 279. [Google Scholar] [CrossRef]
- World Organization for Animal Health. Available online: https://www.oie.int/scientific-expertise/veterinary-products/antimicrobials/ (accessed on 17 January 2021).
- Walther, B.; Tedin, K.; Lübke-Becker, A. Multidrug-Resistant Opportunistic Pathogens Challenging Veterinary Infection Control. Vet. Microbiol. 2017, 200, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Steinman, A.; Navon-Venezia, S. Antimicrobial Resistance in Horses. Animals 2020, 10, 1161. [Google Scholar] [CrossRef]
- Shnaiderman-Torban, A.; Navon-Venezia, S.; Dor, Z.; Paitan, Y.; Arielly, H.; Abu Ahmad, W.; Kelmer, G.; Fulde, M.; Steinman, A. Extended-Spectrum β-Lactamase-Producing Enterobacteriaceae Shedding in Farm Horses Versus Hospitalized Horses: Prevalence and Risk Factors. Animals 2020, 10, 282. [Google Scholar] [CrossRef] [PubMed]
- de Lagarde, M.; Fairbrother, J.M.; Arsenault, J. Prevalence, Risk Factors, and Characterization of Multidrug Resistant and ESBL/AmpC Producing Escherichia Coli in Healthy Horses in Quebec, Canada, in 2015–2016. Animals 2020, 10, 523. [Google Scholar] [CrossRef] [PubMed]
- Hordijk, J.; Farmakioti, E.; Smit, L.A.M.; Duim, B.; Graveland, H.; Theelen, M.J.P.; Wagenaar, J.A. Fecal Carriage of Extended-Spectrum-β-Lactamase/AmpC-Producing Escherichia Coli in Horses. Appl. Environ. Microbiol. 2020, 86. [Google Scholar] [CrossRef]
- Sukamawinata, E.; Sato, W.; Mitoma, S.; Kanda, T.; Kusano, K.; Kambayashi, Y.; Sato, T.; Ishikawa, Y.; Goto, Y.; Uemura, R.; et al. Extended-Spectrum β-Lactamase-Producing Escherichia Coli Isolated from Healthy Thoroughbred Racehorses in Japan. J. Equine Sci. 2019, 30, 47–53. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Anyanwu, M.U.; Ugwu, I.C.; Onah, C.U. Occurrence and Antibiogram of Generic Extended-Spectrum Cephalosporin-Resistant and Extended-Spectrum β-Lactamase-Producing Enterobacteria In Horses. Maced. Vet. Rev. 2018, 41, 123–132. [Google Scholar] [CrossRef]
- Shnaiderman-Torban, A.; Paitan, Y.; Arielly, H.; Kondratyeva, K.; Tirosh-Levy, S.; Abells-Sutton, G.; Navon-Venezia, S.; Steinman, A. Extended-Spectrum β-Lactamase-Producing Enterobacteriaceae in Hospitalized Neonatal Foals: Prevalence, Risk Factors for Shedding and Association with Infection. Animals 2019, 9, 600. [Google Scholar] [CrossRef]
- Schoster, A.; van Spijk, J.; Damborg, P.; Moodley, A.; Kirchgaessner, C.; Hartnack, S.; Schmitt, S. The Effect of Different Antimicrobial Treatment Regimens on the Faecal Shedding of ESBL-Producing Escherichia Coli in Horses. Vet. Microbiol. 2020, 243, 108617. [Google Scholar] [CrossRef]
- Shnaiderman-Torban, A.; Navon-Venezia, S.; Dahan, R.; Dor, Z.; Taulescu, M.; Paitan, Y.; Edery, N.; Steinman, A. CTX-M-15 Producing Escherichia Coli Sequence Type 361 and Sequence Type 38 Causing Bacteremia and Umbilical Infection in a Neonate Foal. J. Equine Vet. Sci. 2020, 85, 102881. [Google Scholar] [CrossRef] [PubMed]
- Loncaric, I.; Cabal Rosel, A.; Szostak, M.P.; Licka, T.; Allerberger, F.; Ruppitsch, W.; Spergser, J. Broad-Spectrum Cephalosporin-Resistant Klebsiella Spp. Isolated from Diseased Horses in Austria. Animals 2020, 10, 332. [Google Scholar] [CrossRef] [PubMed]
- Hann, M.; Timofte, D.; Isgren, C.M.; Archer, D.C. Bacterial Translocation in Horses with Colic and the Potential Association with Surgical Site Infection: A Pilot Study. Vet. Rec. 2020, 187, 68. [Google Scholar] [CrossRef] [PubMed]
- Walther, B.; Klein, K.-S.; Barton, A.-K.; Semmler, T.; Huber, C.; Wolf, S.A.; Tedin, K.; Merle, R.; Mitrach, F.; Guenther, S.; et al. Extended-Spectrum Beta-Lactamase (ESBL)-Producing Escherichia Coli and Acinetobacter Baumannii among Horses Entering a Veterinary Teaching Hospital: The Contemporary “Trojan Horse”. PLOS ONE 2018, 13, e0191873. [Google Scholar] [CrossRef]
- Gilbertie, J.M.; Schnabel, L.V.; Stefanovski, D.; Kelly, D.J.; Jacob, M.E.; Schaer, T.P. Gram-Negative Multi-Drug Resistant Bacteria Influence Survival to Discharge for Horses with Septic Synovial Structures: 206 Cases (2010–2015). Vet. Microbiol. 2018, 226, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Kaye, K.S.; Harris, A.D.; Samore, M.; Carmeli, Y. The Case–case–control Study Design: Addressing the Limitations of Risk Factor Studies for Antimicrobial Resistance. Infect. Control Hosp. Epidemiol. 2005, 26, 346–351. [Google Scholar] [CrossRef]
- Magiorakos, A.-P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-Resistant, Extensively Drug-Resistant and Pandrug-Resistant Bacteria: An International Expert Proposal for Interim Standard Definitions for Acquired Resistance. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Sheats, M.K. A Comparative Review of Equine SIRS, Sepsis, and Neutrophils. Front. Vet. Sci. 2019, 6, 69. [Google Scholar] [CrossRef] [PubMed]
- Roy, M.-F.; Kwong, G.P.S.; Lambert, J.; Massie, S.; Lockhart, S. Prognostic Value and Development of a Scoring System in Horses With Systemic Inflammatory Response Syndrome. J. Vet. Intern. Med. 2017, 31, 582–592. [Google Scholar] [CrossRef] [PubMed]
- Savage, V.L.; Marr, C.M.; Bailey, M.; Smith, S. Prevalence of Acute Kidney Injury in a Population of Hospitalized Horses. J. Vet. Intern. Med. 2019, 33, 2294–2301. [Google Scholar] [CrossRef]
- Brewer, B.D.; Koterba, A.M.; Carter, R.L.; Rowe, E.D. Comparison of Empirically Developed Sepsis Score with a Computer Generated and Weighted Scoring System for the Identification of Sepsis in the Equine Neonate. Equine Vet. J. 1988, 20, 23–24. [Google Scholar] [CrossRef]
- Edwardson, S.; Cairns, C. Nosocomial Infections in the ICU. Anaesth. Intensive Care Med. 2019, 20, 14–18. [Google Scholar] [CrossRef]
- Oreff, G.L.; Tatz, A.J.; Dahan, R.; Segev, G.; Berlin, D.; Kelmer, G. Surgical Management and Long-Term Outcome of Umbilical Infection in 65 Foals (2010–2015). Vet. Surg. 2017, 46, 962–970. [Google Scholar] [CrossRef]
- Willyard, C. The Drug-Resistant Bacteria That Pose the Greatest Health Threats. Nature 2017, 543, 15. [Google Scholar] [CrossRef]
- Weese, J.S. Antimicrobial Use and Antimicrobial Resistance in Horses. Equine Vet. J. 2015, 47, 747–749. [Google Scholar] [CrossRef]
- Prescott, J.F. Outpacing the Resistance Tsunami: Antimicrobial Stewardship in Equine Medicine, an Overview. Equine Vet. Educ. 2020. [Google Scholar] [CrossRef]
- Adler, A.; Friedman, N.D.; Marchaim, D. Multidrug-Resistant Gram-Negative Bacilli: Infection Control Implications. Infect. Dis. Clin. N. Am. 2016, 30, 967–997. [Google Scholar] [CrossRef]
- Singh, A.; Walker, M.; Rousseau, J.; Monteith, G.J.; Weese, J.S. Methicillin-Resistant Staphylococcal Contamination of Clothing Worn by Personnel in a Veterinary Teaching Hospital. Vet. Surg. VS 2013, 42, 643–648. [Google Scholar] [CrossRef]
- Schwaber, M.J.; Navon-Venezia, S.; Masarwa, S.; Tirosh-Levy, S.; Adler, A.; Chmelnitsky, I.; Carmeli, Y.; Klement, E.; Steinman, A. Clonal Transmission of a Rare Methicillin-Resistant Staphylococcus Aureus Genotype between Horses and Staff at a Veterinary Teaching Hospital. Vet. Microbiol. 2013, 162, 907–911. [Google Scholar] [CrossRef]
- Gallagher, J.C.; Kuriakose, S.; Haynes, K.; Axelrod, P. Case–case–control Study of Patients with Carbapenem-Resistant and Third-Generation-Cephalosporin-Resistant Klebsiella Pneumoniae Bloodstream Infections. Antimicrob. Agents Chemother. 2014, 58, 5732–5735. [Google Scholar] [CrossRef] [PubMed]
- Bonine, N.G.; Berger, A.; Altincatal, A.; Wang, R.; Bhagnani, T.; Gillard, P.; Lodise, T. Impact of Delayed Appropriate Antibiotic Therapy on Patient Outcomes by Antibiotic Resistance Status From Serious Gram-Negative Bacterial Infections. Am. J. Med. Sci. 2019, 357, 103–110. [Google Scholar] [CrossRef]
- Tacconelli, E.; Cataldo, M.A.; Mutters, N.T.; Carrara, E.; Bartoloni, A.; Raglio, A.; Cauda, R.; Mantengoli, E.; Luzzaro, F.; Pan, A.; et al. Role of Place of Acquisition and Inappropriate Empirical Antibiotic Therapy on the Outcome of Extended-Spectrum β-Lactamase-Producing Enterobacteriaceae Infections. Int. J. Antimicrob. Agents 2019, 54, 49–54. [Google Scholar] [CrossRef] [PubMed]
- Leigue, L.; Warth, J.F.G.; Melo, L.C.; Silva, K.C.; Moura, R.A.; Barbato, L.; Silva, L.C.; Santos, A.C.M.; Silva, R.M.; Lincopan, N. MDR ST2179-CTX-M-15 Escherichia Coli Co-Producing RmtD and AAC(6′)-Ib-Cr in a Horse with Extraintestinal Infection, Brazil. J. Antimicrob. Chemother. 2015, 70, 1263–1265. [Google Scholar] [CrossRef]
- Haenni, M.; Saras, E.; Ponsin, C.; Dahmen, S.; Petitjean, M.; Hocquet, D.; Madec, J.-Y. High Prevalence of International ESBL CTX-M-15-Producing Enterobacter Cloacae ST114 Clone in Animals. J. Antimicrob. Chemother. 2016, 71, 1497–1500. [Google Scholar] [CrossRef]
- Börjesson, S.; Greko, C.; Myrenås, M.; Landén, A.; Nilsson, O.; Pedersen, K. A Link between the Newly Described Colistin Resistance Gene Mcr-9 and Clinical Enterobacteriaceae Isolates Carrying BlaSHV-12 from Horses in Sweden. J. Glob. Antimicrob. Resist. 2020, 20, 285–289. [Google Scholar] [CrossRef] [PubMed]
- Izdebski, R.; Baraniak, A.; Herda, M.; Fiett, J.; Bonten, M.J.M.; Carmeli, Y.; Goossens, H.; Hryniewicz, W.; Brun-Buisson, C.; Gniadkowski, M. MLST Reveals Potentially High-Risk International Clones of Enterobacter Cloacae. J. Antimicrob. Chemother. 2015, 70, 48–56. [Google Scholar] [CrossRef]
- Torres-González, P.; Valle, M.B.; Tovar-Calderón, E.; Leal-Vega, F.; Hernández-Cruz, A.; Martínez-Gamboa, A.; Niembro-Ortega, M.D.; Sifuentes-Osornio, J.; Ponce-de-León, A. Outbreak Caused by Enterobacteriaceae Harboring NDM-1 Metallo-β-Lactamase Carried in an IncFII Plasmid in a Tertiary Care Hospital in Mexico City. Antimicrob. Agents Chemother. 2015, 59, 7080–7083. [Google Scholar] [CrossRef]
- Marcade, G.; Brisse, S.; Bialek, S.; Marcon, E.; Leflon-Guibout, V.; Passet, V.; Moreau, R.; Nicolas-Chanoine, M.-H. The Emergence of Multidrug-Resistant Klebsiella Pneumoniae of International Clones ST13, ST16, ST35, ST48 and ST101 in a Teaching Hospital in the Paris Region. Epidemiol. Infect. 2013, 141, 1705–1712. [Google Scholar] [CrossRef]
- Nagasaka, Y.; Kimura, K.; Yamada, K.; Wachino, J.-I.; Jin, W.; Notake, S.; Yanagisawa, H.; Arakawa, Y. Genetic Profiles of Fluoroquinolone-Nonsusceptible Klebsiella Pneumoniae Among Cephalosporin-Resistant K. Pneumoniae. Microb. Drug Resist. 2014, 21, 224–233. [Google Scholar] [CrossRef]
- Loucif, L.; Kassah-Laouar, A.; Saidi, M.; Messala, A.; Chelaghma, W.; Rolain, J.-M. Outbreak of OXA-48-Producing Klebsiella Pneumoniae Involving a Sequence Type 101 Clone in Batna University Hospital, Algeria. Antimicrob. Agents Chemother. 2016, 60, 7494–7497. [Google Scholar]
- Kremer, K.; Kramer, R.; Neumann, B.; Haller, S.; Pfennigwerth, N.; Werner, G.; Gatermann, S.; Schroten, H.; Eckmanns, T.; Hans, J.B. Rapid Spread of OXA-244-Producing Escherichia Coli ST38 in Germany: Insights from an Integrated Molecular Surveillance Approach; 2017 to January 2020. Eurosurveillance 2020, 25, 2000923. [Google Scholar] [CrossRef]
- Alsharapy, S.A.; Gharout-Sait, A.; Muggeo, A.; Guillard, T.; Cholley, P.; Brasme, L.; Bertrand, X.; Moghram, G.S.; Touati, A.; De Champs, C. Characterization of Carbapenem-Resistant Enterobacteriaceae Clinical Isolates in Al Thawra University Hospital, Sana’a, Yemen. Microb. Drug Resist. 2019, 26, 211–217. [Google Scholar] [CrossRef]
- Shen, Z.; Gao, Q.; Qin, J.; Liu, Y.; Li, M. Emergence of an NDM-5-Producing Hypervirulent Klebsiella Pneumoniae Sequence Type 35 Strain with Chromosomal Integration of an Integrative and Conjugative Element, ICEKp1. Antimicrob. Agents Chemother. 2020, 64, e01675-19. [Google Scholar] [CrossRef]
- Royden, A.; Ormandy, E.; Pinchbeck, G.; Pascoe, B.; Hitchings, M.D.; Sheppard, S.K.; Williams, N.J. Prevalence of Faecal Carriage of Extended-Spectrum β-Lactamase (ESBL)-Producing Escherichia Coli in Veterinary Hospital Staff and Students. Vet. Rec. Open 2019, 6, e000307. [Google Scholar] [CrossRef]
- Uzunović, S.; Ibrahimagić, A.; Hodžić, D.; Bedenić, B. Molecular Epidemiology and Antimicrobial Susceptibility of AmpC- and/or Extended-Spectrum (ESBL) ß-Lactamaseproducing Proteus Spp. Clinical Isolates in Zenica-Doboj Canton, Bosnia and Herzegovina. Med. Glas. Off. Publ. Med. Assoc. Zenica-Doboj Cant. Bosnia Herzeg. 2016, 13, 103–112. [Google Scholar]
- Uzunović, S.; Bedenić, B.; Budimir, A.; Kamberović, F.; Ibrahimagić, A.; Delić-Bikić, S.; Sivec, S.; Meštrović, T.; Varda Brkić, D.; Rijnders, M.I.A.; et al. Emergency (Clonal Spread) of Methicillin-Resistant Staphylococcus Aureus (MRSA), Extended Spectrum (ESBL)—And AmpC Beta-Lactamase-Producing Gram-Negative Bacteria Infections at Pediatric Department, Bosnia and Herzegovina. Wien. Klin. Wochenschr. 2014, 126, 747–756. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kaftandzhieva, A.; Kotevska, V.; Jankoska, G.; Kjurcik-Trajkovska, B.; Cekovska, Z.; Petrovska, M. Extended-Spectrum Beta-Lactamase-Producing E. Coli and Klebsiella Pneumoniae in Children at University Pediatric Clinic in Skopje. Maced. J. Med. Sci. 2009, 2, 36–41. [Google Scholar] [CrossRef]
- Rampacci, E.; Passamonti, F.; Bottinelli, M.; Stefanetti, V.; Cercone, M.; Nannarone, S.; Gialletti, R.; Beccati, F.; Coletti, M.; Pepe, M. Umbilical Infections in Foals: Microbiological Investigation and Management. Vet. Rec. 2017, 180, 543-543. [Google Scholar] [CrossRef]
- Willis, A.T.; Magdesian, K.G.; Byrne, B.A.; Edman, J.M. Enterococcus Infections in Foals. Vet. J. Lond. Engl. 1997 2019, 248, 42–47. [Google Scholar] [CrossRef]
- Elce, Y.A. Infections in the equine abdomen and pelvis: Perirectal abscesses, umbilical infections, and peritonitis. Vet. Clin. North Am. Equine Pract. 2006, 22, 419–436. [Google Scholar] [CrossRef]
- Wohlfender, F.D.; Barrelet, F.E.; Doherr, M.G.; Straub, R.; Meier, H.P. Diseases in Neonatal Foals. Part 1: The 30 Day Incidence of Disease and the Effect of Prophylactic Antimicrobial Drug Treatment during the First Three Days Post Partum. Equine Vet. J. 2009, 41, 179–185. [Google Scholar] [CrossRef]
- McGowan, C. Welfare of Aged Horses. Animals 2011, 1, 366–376. [Google Scholar] [CrossRef]
- Pogue, J.M.; Kaye, K.S.; Cohen, D.A.; Marchaim, D. Appropriate Antimicrobial Therapy in the Era of Multidrug-Resistant Human Pathogens. Clin. Microbiol. Infect. 2015, 21, 302–312. [Google Scholar] [CrossRef]
- CLSI. Autoverification of Medical Laboratory Results for Specific Disciplines, 1st ed.; CLSI Guideline AUTO15; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2019. [Google Scholar]
- Woodford, N.; Fagan, E.J.; Ellington, M.J. Multiplex PCR for Rapid Detection of Genes Encoding CTX-M Extended-Spectrum β-Lactamases. J. Antimicrob. Chemother. 2006, 57, 154–155. [Google Scholar] [CrossRef]
- Versalovic, J.; Koeuth, T.; Lupski, J.R. Distribution of Repetitive DNA Sequences in Eubacteria and Application to Fingerprinting of Bacterial Genomes. Nucleic Acids Res. 1991, 19, 6823–6831. [Google Scholar] [CrossRef]
- Miyoshi-Akiyama, T.; Hayakawa, K.; Ohmagari, N.; Shimojima, M.; Kirikae, T. Multilocus Sequence Typing (MLST) for Characterization of Enterobacter Cloacae. PLoS ONE 2013, 8, e66358. [Google Scholar] [CrossRef]
- Diancourt, L.; Passet, V.; Verhoef, J.; Grimont, P.A.D.; Brisse, S. Multilocus Sequence Typing of Klebsiella Pneumoniae Nosocomial Isolates. J. Clin. Microbiol. 2005, 43, 4178–4182. [Google Scholar] [CrossRef]
- Wirth, T.; Falush, D.; Lan, R.; Colles, F.; Mensa, P.; Wieler, L.H.; Karch, H.; Reeves, P.R.; Maiden, M.C.J.; Ochman, H.; et al. Sex and Virulence in Escherichia Coli: An Evolutionary Perspective. Mol. Microbiol. 2006, 60, 1136–1151. [Google Scholar] [CrossRef] [PubMed]
| Parameter | 3GCRE 1 No. (Valid % 3) | 3GCSE 2 No. (Valid % 3) | Uninfected No. (Valid % 1) | 3GCRE vs. Uninfected | 3GCSE vs. Uninfected | 3GCRE vs. 3GCSE | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| OR (95% CI) | p-Value | OR (95% CI) | p-Value | OR (95% CI) | p-Value | |||||
| Demographics | ||||||||||
| Age (Years), Median (Range) | 2.25 (0–24) | 3 (0–20) | 3 (0–17) | 0.93 | 0.756 | 0.786 | ||||
| Age Group | Neonates (<30 days) | 12 (41.4) | 12 (41.4) | 12 (41.4) | 1 (0.35–2.844) | >0.99 | 1.0 (0.352–2.844) | >0.99 | 1 (0.352–2.844) | >0.99 |
| Elderly (>20 years) | 1 (3.4) | 1 (3.4) | 0 | 1.036 (0.97–1.11) | >0.99 | 1.036 (0.97–1.11) | >0.99 | 1 (0.06–16.791) | >0.99 | |
| Weight (Kg), Median (Range) /mean ±SD | 125 (22–520) | 118 (40–118) | 200 (30–614) | 0.859 | 0.94 | 0.92 | ||||
| Female Gender | 19 (65.5) | 21 (72.4) | 12 (41.4) | 0.372 (0.128–1.077) | 0.065 | 0.269 (0.09–0.808) | 0.017 | 1.382 (0.452–4.225) | 0.57 | |
| Castrated Adult Male 4 | 2 (66.7) | 3 (60) | 6 (66.7) | 1 (0.063–15.988) | >0.99 | 0.75 (0.078–7.21) | >0.99 | 1.333 (0.067–26.618) | >0.99 | |
| Pregnant Mare | 4 (28.6) | 5 (41.7) | 2 (25) | 1.2 (0.166–8.659) | >0.99 | 2.143 (0.299–15.355) | 0.642 | 0.56 (0.11–2.8620 | 0.683 | |
| Shelter Resident | 4 (13.8) | 2 (6.9) | 3 (10.3) | 1.387 (0.282–6.83) | >0.99 | 0.642 (0.099–4.159) | >0.99 | 2.16 (0.363–12.84) | 0.670 | |
| Recent exposure to healthcare environments and/or settings | ||||||||||
| Recent Hospitalization (<3 months) | 6 (20.7) | 1 (3.4) | 0 (0) | 1.3 (1.053–1.605) | 0.023 | 1.036 (0.967–1.109) | >0.99 | 7.304 (0.819–65.114) | 0.102 | |
| Surgery Prior (<3 months) to the Date of Event 5 | 16 (55.2) | 12 (42.9) | 0 (0) | 2.231 (1.49–3.34) | <0.001 | 1.75 (1.27–2.412) | <0.001 | 1.641 (0.576–4.675) | 0.352 | |
| Urologic Procedure During Hospitalization, Prior to the Date of Event 5 | 14 (48.3) | 11 (39.3) | 2 (6.9) | 12.6 (2.517–63.063) | 0.001 | 8.735 (1.721–44.328) | 0.004 | 1.442 (0.5.4–4.128) | 0.494 | |
| Upper Airways Procedure During Hospitalization, Prior to the Date of Event 5 | 9 (31) | 3 (10.7) | 1 (3.4) | 12.6 (1.476–107.543) | 0.005 | 3.36 (0.328–34.415) | 0.352 | 3.75 (0.895–15.715) | 0.06 | |
| Plasma Therapy During Hospitalization, Prior to the Date of Event 5 | 10 (34.5) | 4 (14.3) | 0 (0) | 1.526 (1.172–1.988) | 0.001 | 1.167 (1.003–1.357) | 0.052 | 3.281 (0.868–12.4) | 0.116 | |
| Feeding/Nasogastric Tube During Hospitalization, Prior to the Date of Event 5 | 16 (57.1) | 14 (46.4) | 9 (32.1) | 2.815 (0.946–8.376) | 0.06 | 1.83 (0617–5.423) | 0.247 | 1.538 (0.536–4.416) | 0.422 | |
| Prior MDRO 6 Isolation (<1 year) | 2 (6.9) | 0 (0) | 0 (0) | 1.074 (0.973–1.186) | 0.491 | a | a | 1.074 (0.973–1.186) | 0.492 | |
| Prior ESBL Isolation (<1 year) | 0 (0) | 0 (0) | 0 (0) | a | a | a | a | a | a | |
| Background conditions and co-morbidities prior to the date of event3 | ||||||||||
| Chronic Lung Disease | 2 (6.9) | 3 (10.3) | 0 (0) | 1.074 (0.973–1.186) | 0.491 | 1.115 (0.986–1.262) | 0.237 | 0.642 (0.099–4.159) | >0.99 | |
| Neurologic Disease 7 | 6 (20.7) | 1 (3.4) | 4 (13.8) | 1.63 (0.408–6.521) | 0.487 | 0.223 (0.023–2.132) | 0.352 | 7.304 (0.819–65.114) | 0.102 | |
| Immunosuppression 8 | 7 (24.1) | 1 (3.6) | 1 (3.4) | 8.909 (1.019–77.905) | 0.052 | 1.037 (0.062–17.429) | >0.99 | 8.591 (0.981–75.221) | 0.052 | |
| Hyperlactatemia 9 | 4 (66.7) | 4 (40) | 4 (36.4) | 3.5 (0.431–28.447) | 0.335 | 1.167 (0.2–6.805) | >0.99 | 3 (0.361–24.919) | 0.608 | |
| Azotemia 10 | 7 (25.9) | 4 (40) | 10 (35.7) | 0.63 (0.198–2.003) | 0.432 | 0.3 (0.0815–1.113) | 0.064 | 2.1 (0.537–8.217) | 0.281 | |
| Antimicrobial therapy prior (< 3 months) to the date of event 5 | ||||||||||
| Any Antibiotic Treatment | 28 (96.6) | 20 (71.4) | 3 (10.3) | 242.667 (23.722–2482.349) | <0.001 | 21.667 (5.086–92.303) | <0.001 | 11.2 (1.296–96.787) | 0.012 | |
| Penicillins | 20 (71.4) | 15 (53.6) | 1 (3.4) | 70 (8.1–604.917) | <0.001 | 32.308 (3.845–271.441) | <0.001 | 2.167 (0.717–6.55) | 0.168 | |
| Fluoroquinolone | 6 (21.4) | 3 (11.1) | 0 (0) | 1.273 (1.049–1.544) | 0.01 | 1.125 (0.985–1.285) | 0.106 | 2.182 (0.486–9.796) | 0.469 | |
| Aminoglycoside | 23 (82.1) | 9 (32.1) | 1 (3.4) | 128 (14.034–1182.052) | <0.001 | 13.263 (1.55–113.47) | 0.005 | 9.711 (2.78–33.92) | <0.001 | |
| Polymyxin | 6 (37.5) | 9 (56.3) | 0 (0) | 1.6 (1.095–2.339) | 0.006 | 2.286 (1.311–3.984) | <0.001 | 0.467 (0.113–1.92) | 0.288 | |
| Metronidazole | 4 (14.3) | 5 (17.9) | 0 (0) | 1.167 (1.003–1.357) | 0.052 | 1.217 (1.024–1.447) | 0.023 | 0.767 (0.183–3.216) | >0.99 | |
| Cephalosporins | 4 (14.3) | 2 (7.1) | 0 (0) | 1.167 (1.003–1.357) | 0.052 | 1.077 (0.972–1.193) | 0.237 | 2.167 (0.363–12.922) | 0.669 | |
| Acute illness indices at the date of event 5 | ||||||||||
| Sepsis 11 | 10 (34.5) | 6 (20.7) | 0 (0) | 1.526 (1.172–1.988) | 0.001 | 1.261 (1.047–1.518) | 0.023 | 2.018 (0.62–6.569) | 0.24 | |
| Parameter | 3GCRE 1 No. (Valid % 1) | 3GCSE 2 No. (Valid % 3) | Uninfected No. (Valid % 1) | 3GCRE vs. Uninfected | 3GCSE vs. Uninfected | 3GCRE vs. 3GCSE | |||
|---|---|---|---|---|---|---|---|---|---|
| OR (95% CI) | p-Value | OR (95% CI) | p-Value | OR (95% CI) | p-Value | ||||
| Total length of stay (LOS) after excluding the patients who died in hospital, days, median (range) | 17.5 (2–181) | 9 (2–59) | 4 (2–17) | <0.001 | <0.001 | 0.027 | |||
| LOS from date of event 4 to discharge after excluding the patients who died in-hospital, days, median (range) | 11.5 (1–181) | 8 (0–59) | 4 (2–17) | 0.002 | 0.068 | 0.11 | |||
| Additional hospitalization in the following 3 months | 2 (10.5) | 4 (17.4) | 2 (8) | 1.353 (0.173–10.592) | >0.99 | 2.421 (0.399–14.688) | 0.407 | 0.559 (0.091–3.446) | 0.673 |
| Clinical failure 5 | 14 (51.9) | 6 (21.4) | 1 (3.4) | 30.3 (3.57–250) | <0.001 | 7.634 (0.855–66.667) | 0.052 | 3.77 (1.157–12.195) | 0.024 |
| Bacteriological cure 10 | 3 (60) | 1 (50) | a | a | a | a | 1.5 (0.055–40.633) | >0.99 | |
| Surgery following the date of event 4 | 18 (64.3) | 15 (57.7) | 7 (26.9) | 4.725 (1.537–14.552) | 0.005 | 3.231 (1.081–9.656) | 0.033 | 1.463 (0.504–4.24) | 0.483 |
| Urologic procedure following the date of event 4 | 18 (62.1) | 15 (57.7) | 7 (26.9) | 4.295 (1.42–12.997) | 0.008 | 3.231 (1.081–9.656) | 0.033 | 1.33 (0.466–3.792) | 0.594 |
| Upper airways procedures following the date of event 4 | 5 (17.2) | 12 (41.4) | 13 (44.8) | 0.256 (0.076–0.86) | 0.045 | 0.869 (0.307–2.458) | 0.791 | 0.295 (0.088–0.994) | 0.043 |
| Feeding tube/nasogastric tube following the date of event 4 | 16 (59.3) | 15 (55.6) | 25 (86.2) | 0.233 (0.063–0.858) | 0.023 | 0.2 (0.055–0.734) | 0.011 | 1.164 (0.395–3.425) | 0.783 |
| In hospital mortality | 9 (31) | 0 (10.3) | 0 (0) | 1.45 (1.136–1.851) | 0.002 | 1.083 (0.97–1.21) | 0.49 | 3.9 (0.933–16.31) | 0.052 |
| 14-days mortality 6 | 8 (30.8) | 4 (16) | 0 (0) | 1.444 (1.118–1.866) | 0.004 | 1.19 (1.003–1.413) | 0.11 | 2.333 (0.602–9.049) | 0.214 |
| 90-days mortality 6 | 12 (48) | 5 (20.8) | 2 (8) | 8.462 (1.61–44.53) | 0.002 | 3.026 (0.527–17.394) | 0.247 | 3.257 (0.932–11.38) | 0.059 |
| 1-year mortality 6,7 | 11 (47.8) | 5 (20.8) | 5 (18.5) | 4.062 (1.166–14.154) | 0.024 | 1.158 (0.29–4.617) | >0.99 | 3.508 (0.996–12.359) | 0.046 |
| Appropriate therapy 8 (given 2 days before to 5 days after culture date) | 3 (11.5) | 19 (76) | a | a | a | a | 0.041 (0.009–0.187) | <0.001 | |
| Days of appropriate therapy 8, median (range)/mean ± SD | 0 (0–16) | 7.1±5.9 | a | a | a | a | <0.001 | ||
| Days to appropriate therapy 8, median (range) | 0 (0–1) | 0 (0–5) | a | a | a | a | 0.929 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shnaiderman-Torban, A.; Marchaim, D.; Navon-Venezia, S.; Lubrani, O.; Paitan, Y.; Arielly, H.; Steinman, A. Third Generation Cephalosporin Resistant Enterobacterales Infections in Hospitalized Horses and Donkeys: A Case–Case–Control Analysis. Antibiotics 2021, 10, 155. https://doi.org/10.3390/antibiotics10020155
Shnaiderman-Torban A, Marchaim D, Navon-Venezia S, Lubrani O, Paitan Y, Arielly H, Steinman A. Third Generation Cephalosporin Resistant Enterobacterales Infections in Hospitalized Horses and Donkeys: A Case–Case–Control Analysis. Antibiotics. 2021; 10(2):155. https://doi.org/10.3390/antibiotics10020155
Chicago/Turabian StyleShnaiderman-Torban, Anat, Dror Marchaim, Shiri Navon-Venezia, Ori Lubrani, Yossi Paitan, Haya Arielly, and Amir Steinman. 2021. "Third Generation Cephalosporin Resistant Enterobacterales Infections in Hospitalized Horses and Donkeys: A Case–Case–Control Analysis" Antibiotics 10, no. 2: 155. https://doi.org/10.3390/antibiotics10020155
APA StyleShnaiderman-Torban, A., Marchaim, D., Navon-Venezia, S., Lubrani, O., Paitan, Y., Arielly, H., & Steinman, A. (2021). Third Generation Cephalosporin Resistant Enterobacterales Infections in Hospitalized Horses and Donkeys: A Case–Case–Control Analysis. Antibiotics, 10(2), 155. https://doi.org/10.3390/antibiotics10020155








