Artificial Intelligence Methodologies Applied to Technologies for Screening, Diagnosis and Care of the Diabetic Foot: A Narrative Review
Abstract
1. Introduction
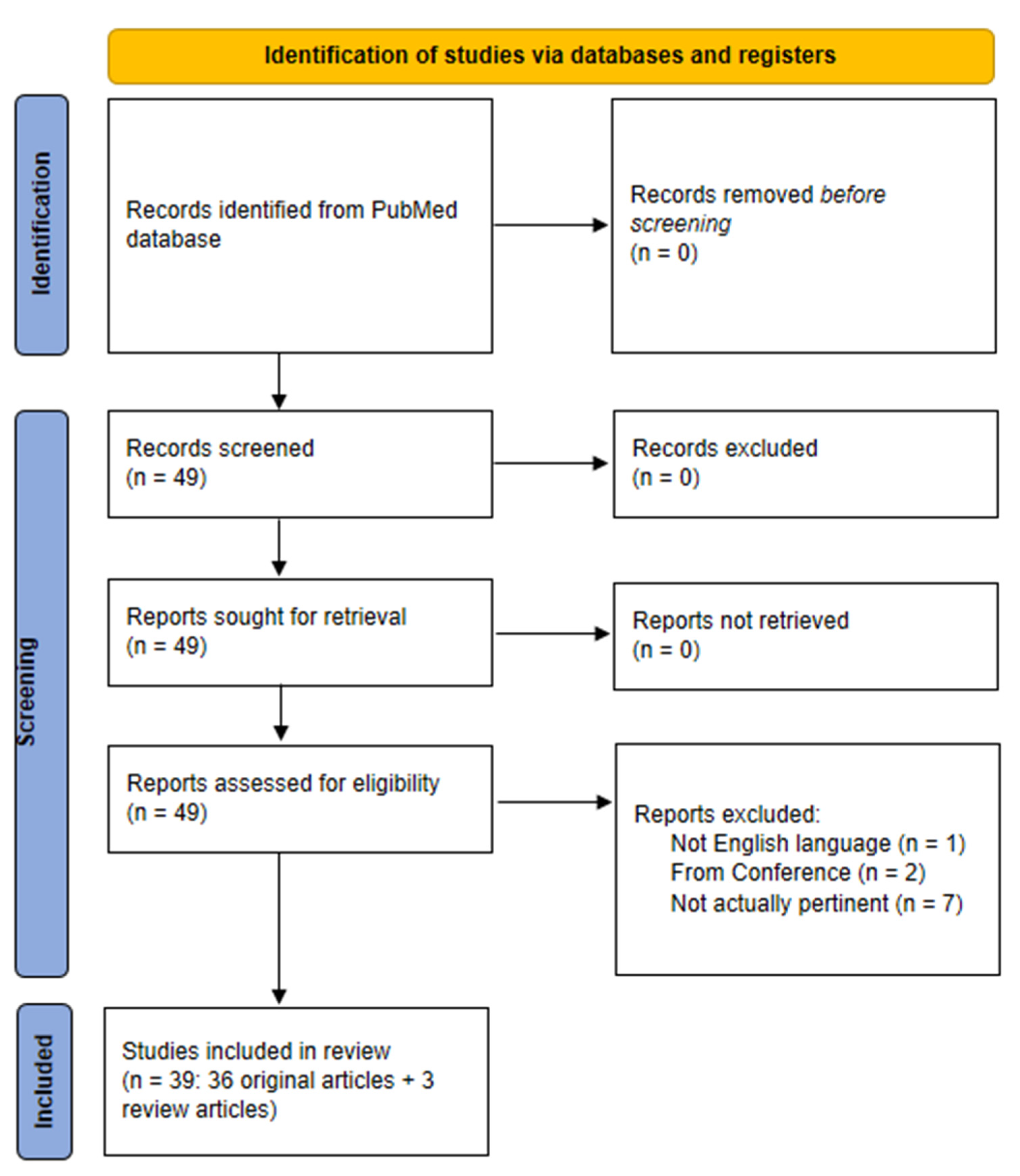
2. Scientific Literature Search Strategy
diabet*[ti] AND (foot*[ti] OR feet*[ti] OR ulcer*[ti] OR skin*[ti]) AND (((machine*[tw] OR deep*[tw]) AND learning*[tw]) OR (artificial*[tw] AND intelligen*[tw]) OR (data*[tw] AND mining*[tw] OR (neural*[tw] AND network*[tw])))
3. Artificial Intelligence in Diabetic Foot Syndrome: Methodological Approaches and the Main Physiological and Clinical Outcomes
3.1. Screening for Diabetic Foot Syndrome and Risk Prediction for Ulceration
3.1.1. Screening and Risk Prediction: From Clinical, Socioeconomic, Sociodemographic Data
3.1.2. Screening and Risk Prediction: From Imaging
3.2. Overt Diabetic Foot Ulcer Detection, Grading, Prognosis and Care
3.2.1. Overt Diabetic Foot Ulcer Focus: From Clinical, Socioeconomic, Sociodemographic Data
3.2.2. Overt Diabetic Foot Ulcer Focus: From Imaging
4. Discussion
4.1. Introductory Comments and Comparison with Previous Review Studies
4.2. Comments on the Specific Sections Summarizing the Studies Pertinent for The Review
4.3. Other Comments and Our Personal View for Future Studies in the Field of the Diabetic Foot
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Acc | Accuracy per class |
| AdaBoost | Adaptive Boosting |
| AHRF | Associative Hierarchical Random Field |
| AI | Artificial Intelligence |
| ANN | Artificial Neural Network |
| Apo A1 | Alipoprotein A1 |
| AUCROC | Area-Under-the-Curve of ROC curve |
| BDC | Binary Decision Classification |
| BMI | Body Mass Index |
| BN | Bayesian Network |
| BPNN | Back-Propagation Neural Network |
| CKBs | Class Knowledge Banks |
| CNL | Competitive Neural Layer |
| CNN | Convolutional Neural Network |
| CRF | Conditional Random Field |
| CTA | Computed Tomography Angiography |
| CTREE | Conditional Inference Tree |
| DA | Discriminant Analysis |
| DE-ResUnet | Double Encoder-ResUnet |
| DFINET | Diabetic Foot Infection Network |
| DFTNet | Diabetic Foot Thermograms Network |
| DFU | Diabetic Foot Ulcer |
| DFUC | Diabetic Foot Ulcer Consortium |
| DICE | Dice Similarity Coefficient |
| DT | Decision Tree |
| EC | Ensemble Classification |
| EHR | Electronic Health Record |
| EMG | Electromyography |
| GA | Genetic Algorithm |
| GaNDLF | Generally Nuanced Deep Learning Framework |
| GAP | Global Average Pooling |
| GBDT | Gradient Boosting Decision Tree |
| GBM | Gradient Boosting Machine |
| GRF | Ground Reaction Forces |
| HCE | Hyperglycemic Crisis Episode |
| IL-10 | Interleukin-10 |
| IoU | Intersection over Union |
| KC | Kernel Classification |
| KNN | K-Nearest Neighbor |
| LASSO | Least Absolute Shrinkage and Selection Operator |
| LBP | Local Binary Patterns |
| LC | Linear Classification |
| LDA | Linear Discriminant Analysis |
| LightGBM | Light Gradient Boosting Machine |
| LR | Logistic Regression |
| mAP | mean Average Precision |
| MCC | Matthews Correlation Coefficient |
| MLP | Multilayer Perceptron |
| MLR | Multivariate Linear Regression |
| MMP9 | Metalloproteinase-9 |
| MRI | Magnetic Resonance Imaging |
| NB | Naïve Bayes |
| NPV | Negative Predictive Value |
| PCA | Principal Component Analysis |
| PNN | Probability Neural Network |
| PPV | Positive Predictive Value |
| PU | Pressure Ulcer |
| RBF | Radial Basis Function |
| RBFNN | Radial Basis Function Neural Network |
| R-CNN | Region-based Convolutional Neural Network |
| RF | Random Forest |
| R-FCN | Region-based Fully Convolutional Networks |
| RGB | Red, Green, Blue |
| ROC | Receiver-Operating-Characteristic |
| ROI | Region of Interest |
| SHAP | Shapley Additive Explanations |
| SMOTE | Synthetic Minority Oversampling Technique |
| SNPs | Single Nucleotide Polymorphisms |
| SPCD | Superpixel Color Descriptors |
| SSD | Single Shot Detector |
| SVD | Singular-Value Decomposition |
| SVM | Support Vector Machine |
| T2DM | Type 2 Diabetes Mellitus |
| WIFI | Wound, Ischemia, Foot Infection |
| XGBoost | Extreme Gradient Boosting |
| YOLO | You-Only-Look-Once |
References
- Newrick, P. International consensus on the diabetic foot. BMJ 2000, 321, 642A. [Google Scholar] [PubMed]
- Reiber, G.E. Epidemiology of Foot Ulcers and Amputations in the Diabetic Foot. In The Diabetic Foot; Bowker, J.H., Pfeifer, M.A., Eds.; Mosby Inc.: St. Louis, MO, USA, 2001; pp. 13–32. [Google Scholar]
- Jeffcoate, W.J.; Harding, K.G. Diabetic foot ulcers. Lancet 2003, 361, 1545–1551. [Google Scholar] [CrossRef]
- Formosa, C.; Gatt, A.; Chockalingam, N. A Critical Evaluation of Existing Diabetic Foot Screening Guidelines. Rev. Diabet. Stud. 2016, 13, 158–186. [Google Scholar] [CrossRef]
- Bus, S.A.; Van Netten, J.J.; Hinchliffe, R.J.; Apelqvist, J.; Lipsky, B.A.; Schaper, N.C.; IWGDF Editorial Board. Standards for the development and methodology of the 2019 International Working Group on the Diabetic Foot guidelines. Diabetes Metab. Res. Rev. 2020, 36 (Suppl. S1), e3267. [Google Scholar] [CrossRef]
- Ibrahim, A. IDF Clinical Practice Recommendation on the Diabetic Foot: A guide for healthcare professionals. Diabetes Res. Clin. Pract. 2017, 127, 285–287. [Google Scholar] [CrossRef]
- Apelqvist, J.; Bakker, K.; van Houtum, W.H.; Schaper, N.C.; International Working Group on the Diabetic Foot (IWGDF) Editorial Board. Practical guidelines on the management and prevention of the diabetic foot: Based upon the International Consensus on the Diabetic Foot (2007) Prepared by the International Working Group on the Diabetic Foot. Diabetes Metab. Res. Rev. 2008, 24 (Suppl. S1), S181–S187. [Google Scholar] [CrossRef] [PubMed]
- Schaper, N.C.; van Netten, J.J.; Apelqvist, J.; Bus, S.A.; Hinchliffe, R.J.; Lipsky, B.A.; IWGDF Editorial Board. Practical Guidelines on the prevention and management of diabetic foot disease (IWGDF 2019 update). Diabetes Metab. Res. Rev. 2020, 36 (Suppl. S1), e3266. [Google Scholar] [CrossRef] [PubMed]
- Won, J.C.; Park, T.S. Recent Advances in Diagnostic Strategies for Diabetic Peripheral Neuropathy. Endocrinol. Metab. 2016, 31, 230–238. [Google Scholar] [CrossRef]
- Kalinchenko, S.; Zemlyanoy, A.; Gooren, L.J. Improvement of the diabetic foot upon testosterone administration to hypogonadal men with peripheral arterial disease. Report of three cases. Cardiovasc. Diabetol. 2009, 8, 19. [Google Scholar] [CrossRef] [PubMed]
- Santosa, Y.; Harca, A.D.; Sukmaja, E.; Yuwono, A. The Role of Endovascular Procedure for Peripheral Arterial Disease in Diabetic Patients with Chronic Limb-Threatening Ischemia. Cureus 2022, 14, e23857. [Google Scholar] [CrossRef] [PubMed]
- Rubitschung, K.; Sherwood, A.; Crisologo, A.P.; Bhavan, K.; Haley, R.W.; Wukich, D.K.; Castellino, L.; Hwang, H.; La Fontaine, J.; Chhabra, A.; et al. Pathophysiology and Molecular Imaging of Diabetic Foot Infections. Int. J. Mol. Sci. 2021, 22, 11552. [Google Scholar] [CrossRef] [PubMed]
- Paes Leme, R.C.; Chaves, J.R.E.; Gonçalves, L.C.S.; Alvim, L.C.; Almeida, J.R.C.D.; da Cruz Renó, L. Diabetic foot infection caused by bacteria of the Burkholderia cepacia complex: Report of an unusual case and a scoping literature review. Rev. Inst. Med. Trop. São Paulo 2022, 64, e36. [Google Scholar] [CrossRef] [PubMed]
- Bakker, K.; Apelqvist, J.; Lipsky, B.A.; Van Netten, J.J.; International Working Group on the Diabetic Foot. The 2015 IWGDF guidance documents on prevention and management of foot problems in diabetes: Development of an evidence-based global consensus. Diabetes Metab. Res. Rev. 2016, 32 (Suppl. S1), 2–6. [Google Scholar] [CrossRef]
- Monteiro-Soares, M.; Russell, D.; Boyko, E.J.; Jeffcoate, W.; Mills, J.L.; Morbach, S.; Game, F.; International Working Group on the Diabetic Foot (IWGDF). Guidelines on the classification of diabetic foot ulcers (IWGDF 2019). Diabetes Metab. Res. Rev. 2020, 36 (Suppl. S1), e3273. [Google Scholar] [CrossRef]
- Fregoso-Aparicio, L.; Noguez, J.; Montesinos, L.; García-García, J.A. Machine learning and deep learning predictive models for type 2 diabetes: A systematic review. Diabetol. Metab. Syndr. 2021, 13, 148. [Google Scholar] [CrossRef] [PubMed]
- Gautier, T.; Ziegler, L.B.; Gerber, M.S.; Campos-Náñez, E.; Patek, S.D. Artificial intelligence and diabetes technology: A review. Metabolism 2021, 124, 154872. [Google Scholar] [CrossRef]
- Nomura, A.; Noguchi, M.; Kometani, M.; Furukawa, K.; Yoneda, T. Artificial Intelligence in Current Diabetes Management and Prediction. Curr. Diabetes Rep. 2021, 21, 61. [Google Scholar] [CrossRef]
- Tan, K.R.; Seng, J.J.B.; Kwan, Y.H.; Chen, Y.J.; Zainudin, S.B.; Loh, D.H.F.; Liu, N.; Low, L.L. Evaluation of Machine Learning Methods Developed for Prediction of Diabetes Complications: A Systematic Review. J. Diabetes Sci. Technol. 2021, 19322968211056917. [Google Scholar] [CrossRef]
- Wang, Y.-L.; Yang, J.-Y.; Yang, J.-Y.; Zhao, X.-Y.; Chen, Y.-X.; Yu, W.-H. Progress of artificial intelligence in diabetic retinopathy screening. Diabetes Metab. Res. Rev. 2021, 37, e3414. [Google Scholar] [CrossRef]
- Singh, K.; Singh, V.K.; Agrawal, N.K.; Gupta, S.K.; Singh, K. Association of Toll-like receptor 4 polymorphisms with diabetic foot ulcers and application of artificial neural network in DFU risk assessment in type 2 diabetes patients. BioMed Res. Int. 2013, 2013, 318686. [Google Scholar] [CrossRef]
- Ferreira, A.C.B.H.; Ferreira, D.D.; Oliveira, H.C.; Resende, I.C.D.; Anjos, A.; Lopes, M.H.B. de M. Competitive neural layer-based method to identify people with high risk for diabetic foot. Comput. Biol. Med. 2020, 120, 103744. [Google Scholar] [CrossRef] [PubMed]
- Schäfer, Z.; Mathisen, A.; Svendsen, K.; Engberg, S.; Rolighed Thomsen, T.; Kirketerp-Møller, K. Toward Machine-Learning-Based Decision Support in Diabetes Care: A Risk Stratification Study on Diabetic Foot Ulcer and Amputation. Front. Med. 2020, 7, 601602. [Google Scholar] [CrossRef] [PubMed]
- Stefanopoulos, S.; Ayoub, S.; Qiu, Q.; Ren, G.; Osman, M.; Nazzal, M.; Ahmed, A. Machine learning prediction of diabetic foot ulcers in the inpatient population. Vascular 2021, 17085381211040984. [Google Scholar] [CrossRef]
- Haque, F.; Reaz, M.B.I.; Chowdhury, M.E.H.; Ezeddin, M.; Kiranyaz, S.; Alhatou, M.; Ali, S.H.M.; Bakar, A.A.A.; Srivastava, G. Machine Learning-Based Diabetic Neuropathy and Previous Foot Ulceration Patients Detection Using Electromyography and Ground Reaction Forces during Gait. Sensors 2022, 22, 3507. [Google Scholar] [CrossRef]
- Nanda, R.; Nath, A.; Patel, S.; Mohapatra, E. Machine learning algorithm to evaluate risk factors of diabetic foot ulcers and its severity. Med. Biol. Eng. Comput. 2022, 60, 2349–2357. [Google Scholar] [CrossRef] [PubMed]
- Troitskaya, N.I.; Shapovalov, K.G.; Mudrov, V.A. Possibilities of Multilayer Perceptron in Complexing Risk Factors of Diabetic Foot Syndrome. Bull. Exp. Biol. Med. 2022, 173, 415–418. [Google Scholar] [CrossRef] [PubMed]
- Toledo Peral, C.L.; Ramos Becerril, F.J.; Vega Martínez, G.; Vera Hernández, A.; Leija Salas, L.; Gutiérrez Martínez, J. An Application for Skin Macules Characterization Based on a 3-Stage Image-Processing Algorithm for Patients with Diabetes. J. Healthc. Eng. 2018, 2018, 9397105. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Vega, I.; Hernandez-Contreras, D.; Peregrina-Barreto, H.; Rangel-Magdaleno, J.D.J.; Ramirez-Cortes, J.M. Deep Learning Classification for Diabetic Foot Thermograms. Sensors 2020, 20, 1762. [Google Scholar] [CrossRef]
- Khandakar, A.; Chowdhury, M.E.H.; Ibne Reaz, M.B.; Md Ali, S.H.; Hasan, M.A.; Kiranyaz, S.; Rahman, T.; Alfkey, R.; Bakar, A.A.A.; Malik, R.A. A machine learning model for early detection of diabetic foot using thermogram images. Comput. Biol. Med. 2021, 137, 104838. [Google Scholar] [CrossRef]
- Arteaga-Marrero, N.; Hernández, A.; Villa, E.; González-Pérez, S.; Luque, C.; Ruiz-Alzola, J. Segmentation Approaches for Diabetic Foot Disorders. Sensors 2021, 21, 934. [Google Scholar] [CrossRef]
- Dremin, V.; Marcinkevics, Z.; Zherebtsov, E.; Popov, A.; Grabovskis, A.; Kronberga, H.; Geldnere, K.; Doronin, A.; Meglinski, I.; Bykov, A. Skin Complications of Diabetes Mellitus Revealed by Polarized Hyperspectral Imaging and Machine Learning. IEEE Trans. Med. Imaging 2021, 40, 1207–1216. [Google Scholar] [CrossRef] [PubMed]
- Khandakar, A.; Chowdhury, M.E.H.; Reaz, M.B.I.; Ali, S.H.M.; Abbas, T.O.; Alam, T.; Ayari, M.A.; Mahbub, Z.B.; Habib, R.; Rahman, T.; et al. Thermal Change Index-Based Diabetic Foot Thermogram Image Classification Using Machine Learning Techniques. Sensors 2022, 22, 1793. [Google Scholar] [CrossRef] [PubMed]
- Khandakar, A.; Chowdhury, M.E.H.; Reaz, M.B.I.; Ali, S.H.M.; Kiranyaz, S.; Rahman, T.; Chowdhury, M.H.; Ayari, M.A.; Alfkey, R.; Bakar, A.A.A.; et al. A Novel Machine Learning Approach for Severity Classification of Diabetic Foot Complications Using Thermogram Images. Sensors 2022, 22, 4249. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Dong, W.; Guan, H.; Yakupu, A.; Wang, H.; Chen, L.; Lu, S.; Tang, J. CT-Angiography-Based Outcome Prediction on Diabetic Foot Ulcer Patients: A Statistical Learning Approach. Diagnostics 2022, 12, 1076. [Google Scholar] [CrossRef]
- Bouallal, D.; Douzi, H.; Harba, R. Diabetic foot thermal image segmentation using Double Encoder-ResUnet (DE-ResUnet). J. Med. Eng. Technol. 2022, 46, 378–392. [Google Scholar] [CrossRef]
- Muralidhara, S.; Lucieri, A.; Dengel, A.; Ahmed, S. Holistic multi-class classification & grading of diabetic foot ulcerations from plantar thermal images using deep learning. Health Inf. Sci. Syst. 2022, 10, 21. [Google Scholar] [CrossRef]
- Yusuf, N.; Zakaria, A.; Omar, M.I.; Shakaff, A.Y.M.; Masnan, M.J.; Kamarudin, L.M.; Abdul Rahim, N.; Zakaria, N.Z.I.; Abdullah, A.A.; Othman, A.; et al. In-vitro diagnosis of single and poly microbial species targeted for diabetic foot infection using e-nose technology. BMC Bioinform. 2015, 16, 158. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-K.; Chang, C.-C.; Lin, P.-X.; Lin, B.-S.; Huang, Y.-K.; Chang, C.-C.; Lin, P.-X.; Lin, B.-S. Quantitative Evaluation of Rehabilitation Effect on Peripheral Circulation of Diabetic Foot. IEEE J. Biomed. Health Inform. 2018, 22, 1019–1025. [Google Scholar] [CrossRef]
- Lin, C.; Yuan, Y.; Ji, L.; Yang, X.; Yin, G.; Lin, S. The amputation and survival of patients with diabetic foot based on establishment of prediction model. Saudi J. Biol. Sci. 2020, 27, 853–858. [Google Scholar] [CrossRef]
- Du, C.; Li, Y.; Xie, P.; Zhang, X.; Deng, B.; Wang, G.; Hu, Y.; Wang, M.; Deng, W.; Armstrong, D.G.; et al. The amputation and mortality of inpatients with diabetic foot ulceration in the COVID-19 pandemic and postpandemic era: A machine learning study. Int. Wound J. 2021, 19, 1289–1297. [Google Scholar] [CrossRef] [PubMed]
- Xie, P.; Li, Y.; Deng, B.; Du, C.; Rui, S.; Deng, W.; Wang, M.; Boey, J.; Armstrong, D.G.; Ma, Y.; et al. An explainable machine learning model for predicting in-hospital amputation rate of patients with diabetic foot ulcer. Int. Wound J. 2022, 19, 910–918. [Google Scholar] [CrossRef] [PubMed]
- Margolis, D.J.; Mitra, N.; Malay, D.S.; Mirza, Z.K.; Lantis, J.C.; Lev-Tov, H.A.; Kirsner, R.S.; Thom, S.R. Further evidence that wound size and duration are strong prognostic markers of diabetic foot ulcer healing. Wound Repair Regen. 2022, 30, 487–490. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Xie, P.; Chen, Y.; Rui, S.; Yang, C.; Deng, B.; Wang, M.; Armstrong, D.G.; Ma, Y.; Deng, W. Impact of acute hyperglycemic crisis episode on survival in individuals with diabetic foot ulcer using a machine learning approach. Front. Endocrinol. 2022, 13, 974063. [Google Scholar] [CrossRef]
- Wang, L.; Pedersen, P.C.; Agu, E.; Strong, D.M.; Tulu, B. Area Determination of Diabetic Foot Ulcer Images Using a Cascaded Two-Stage SVM-Based Classification. IEEE Trans. Biomed. Eng. 2017, 64, 2098–2109. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Pedersen, P.C.; Agu, E.; Strong, D.; Tulu, B. Boundary determination of foot ulcer images by applying the associative hierarchical random field framework. J. Med. Imaging 2019, 6, 024002. [Google Scholar] [CrossRef]
- Ohura, N.; Mitsuno, R.; Sakisaka, M.; Terabe, Y.; Morishige, Y.; Uchiyama, A.; Okoshi, T.; Shinji, I.; Takushima, A. Convolutional neural networks for wound detection: The role of artificial intelligence in wound care. J. Wound Care 2019, 28, S13–S24. [Google Scholar] [CrossRef]
- Goyal, M.; Reeves, N.D.; Rajbhandari, S.; Yap, M.H. Robust Methods for Real-Time Diabetic Foot Ulcer Detection and Localization on Mobile Devices. IEEE J. Biomed. Health Inform. 2019, 23, 1730–1741. [Google Scholar] [CrossRef]
- Goyal, M.; Reeves, N.D.; Davison, A.K.; Rajbhandari, S.; Spragg, J.; Yap, M.H. DFUNet: Convolutional Neural Networks for Diabetic Foot Ulcer Classification. IEEE Trans. Emerg. Top. Comput. Intell. 2020, 4, 728–739. [Google Scholar] [CrossRef]
- Goyal, M.; Reeves, N.D.; Rajbhandari, S.; Ahmad, N.; Wang, C.; Yap, M.H. Recognition of ischaemia and infection in diabetic foot ulcers: Dataset and techniques. Comput. Biol. Med. 2020, 117, 103616. [Google Scholar] [CrossRef]
- Kim, R.B.; Gryak, J.; Mishra, A.; Cui, C.; Soroushmehr, S.M.R.; Najarian, K.; Wrobel, J.S. Utilization of smartphone and tablet camera photographs to predict healing of diabetes-related foot ulcers. Comput. Biol. Med. 2020, 126, 104042. [Google Scholar] [CrossRef]
- Al-Garaawi, N.; Ebsim, R.; Alharan, A.F.H.; Yap, M.H. Diabetic foot ulcer classification using mapped binary patterns and convolutional neural networks. Comput. Biol. Med. 2021, 140, 105055. [Google Scholar] [CrossRef] [PubMed]
- Yap, M.H.; Hachiuma, R.; Alavi, A.; Brüngel, R.; Cassidy, B.; Goyal, M.; Zhu, H.; Rückert, J.; Olshansky, M.; Huang, X.; et al. Deep learning in diabetic foot ulcers detection: A comprehensive evaluation. Comput. Biol. Med. 2021, 135, 104596. [Google Scholar] [CrossRef] [PubMed]
- Cassidy, B.; Reeves, N.D.; Pappachan, J.M.; Gillespie, D.; O’Shea, C.; Rajbhandari, S.; Maiya, A.G.; Frank, E.; Boulton, A.J.; Armstrong, D.G.; et al. The DFUC 2020 Dataset: Analysis Towards Diabetic Foot Ulcer Detection. touchREV. Endocrinol. 2021, 17, 5–11. [Google Scholar] [CrossRef]
- Xu, Y.; Han, K.; Zhou, Y.; Wu, J.; Xie, X.; Xiang, W. Classification of Diabetic Foot Ulcers Using Class Knowledge Banks. Front. Bioeng. Biotechnol. 2021, 9, 811028. [Google Scholar] [CrossRef]
- Viswanathan, V.; Govindan, S.; Selvaraj, B.; Rupert, S.; Kumar, R. A Clinical Study to Evaluate Autofluorescence Imaging of Diabetic Foot Ulcers Using a Novel Artificial Intelligence Enabled Noninvasive Device. Int. J. Low. Extrem. Wounds 2021. [Google Scholar] [CrossRef]
- Güley, O.; Pati, S.; Bakas, S. Classification of Infection and Ischemia in Diabetic Foot Ulcers Using VGG Architectures. Diabet. Foot Ulcers Grand Chall 2022, 13183, 76–89. [Google Scholar] [CrossRef]
- Wang, C.; Yu, X.; Sui, Y.; Zhu, J.; Zhang, B.; Su, Y. Magnetic Resonance Imaging Data Features to Evaluate the Efficacy of Compound Skin Graft for Diabetic Foot. Contrast Media Mol. Imaging 2022, 2022, 5707231. [Google Scholar] [CrossRef]
- Yogapriya, J.; Chandran, V.; Sumithra, M.G.; Elakkiya, B.; Shamila Ebenezer, A.; Suresh Gnana Dhas, C. Automated Detection of Infection in Diabetic Foot Ulcer Images Using Convolutional Neural Network. J. Healthc. Eng. 2022, 2022, 2349849. [Google Scholar] [CrossRef]
- Chan, K.S.; Chan, Y.M.; Tan, A.H.M.; Liang, S.; Cho, Y.T.; Hong, Q.; Yong, E.; Chong, L.R.C.; Zhang, L.; Tan, G.W.L.; et al. Clinical validation of an artificial intelligence-enabled wound imaging mobile application in diabetic foot ulcers. Int. Wound J. 2022, 19, 114–124. [Google Scholar] [CrossRef] [PubMed]
- Chan, K.S.; Lo, Z.J. Wound assessment, imaging and monitoring systems in diabetic foot ulcers: A systematic review. Int. Wound J. 2020, 17, 1909–1923. [Google Scholar] [CrossRef]
- Howard, T.; Ahluwalia, R.; Papanas, N. The Advent of Artificial Intelligence in Diabetic Foot Medicine: A New Horizon, a New Order, or a False Dawn? Int. J. Low. Extrem. Wounds 2021, 15347346211041866. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Qiu, Y.; Peng, L.; Zhou, Q.; Wang, Z.; Qi, M. A comprehensive review of methods based on deep learning for diabetes-related foot ulcers. Front. Endocrinol. 2022, 13, 945020. [Google Scholar] [CrossRef] [PubMed]
- Salahuddin, Z.; Woodruff, H.C.; Chatterjee, A.; Lambin, P. Transparency of deep neural networks for medical image analysis: A review of interpretability methods. Comput. Biol. Med. 2021, 140, 105111. [Google Scholar] [CrossRef] [PubMed]
- Fan, F.-L.; Xiong, J.; Li, M.; Wang, G. On Interpretability of Artificial Neural Networks: A Survey. IEEE Trans. Radiat. Plasma Med. Sci. 2021, 5, 741–760. [Google Scholar] [CrossRef] [PubMed]
- Samek, W.; Montavon, G.; Lapuschkin, S.; Anders, C.J.; Müller, K.-R. Explaining Deep Neural Networks and Beyond: A Review of Methods and Applications. Proc. IEEE 2021, 109, 247–278. [Google Scholar] [CrossRef]
- Armstrong, D.G.; Holtz-Neiderer, K.; Wendel, C.; Mohler, M.J.; Kimbriel, H.R.; Lavery, L.A. Skin temperature monitoring reduces the risk for diabetic foot ulceration in high-risk patients. Am. J. Med. 2007, 120, 1042–1046. [Google Scholar] [CrossRef]
- Lavery, L.A.; Higgins, K.R.; Lanctot, D.R.; Constantinides, G.P.; Zamorano, R.G.; Athanasiou, K.A.; Armstrong, D.G.; Agrawal, C.M. Preventing diabetic foot ulcer recurrence in high-risk patients: Use of temperature monitoring as a self-assessment tool. Diabetes Care 2007, 30, 14–20. [Google Scholar] [CrossRef]
- Sibbald, R.G.; Mufti, A.; Armstrong, D.G. Infrared skin thermometry: An underutilized cost-effective tool for routine wound care practice and patient high-risk diabetic foot self-monitoring. Adv. Skin Wound Care 2015, 28, 37–44; quiz 45–46. [Google Scholar] [CrossRef] [PubMed]
- Maddah, E.; Beigzadeh, B. Use of a smartphone thermometer to monitor thermal conductivity changes in diabetic foot ulcers: A pilot study. J. Wound Care 2020, 29, 61–66. [Google Scholar] [CrossRef]
- van Doremalen, R.F.M.; van Netten, J.J.; van Baal, J.G.; Vollenbroek-Hutten, M.M.R.; van der Heijden, F. Infrared 3D Thermography for Inflammation Detection in Diabetic Foot Disease: A Proof of Concept. J. Diabetes Sci. Technol. 2020, 14, 46–54. [Google Scholar] [CrossRef]
- Ghosh, A.; Ray, S.; Garg, M.K.; Chowdhury, S.; Mukhopadhyay, S. The role of infrared dermal thermometry in the management of neuropathic diabetic foot ulcers. Diabet. Med. 2021, 38, e14368. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, D.G.; Lipsky, B.A.; Polis, A.B.; Abramson, M.A. Does dermal thermometry predict clinical outcome in diabetic foot infection? Analysis of data from the SIDESTEP* trial. Int. Wound J. 2006, 3, 302–307. [Google Scholar] [CrossRef] [PubMed]
- Wijlens, A.M.; Holloway, S.; Bus, S.A.; van Netten, J.J. An explorative study on the validity of various definitions of a 2·2°C temperature threshold as warning signal for impending diabetic foot ulceration. Int. Wound J. 2017, 14, 1346–1351. [Google Scholar] [CrossRef] [PubMed]
- Petrofsky, J.S.; McLellan, K. Galvanic skin resistance--a marker for endothelial damage in diabetes. Diabetes Technol. Ther. 2009, 11, 461–467. [Google Scholar] [CrossRef]
- Tentolouris, N.; Voulgari, C.; Liatis, S.; Kokkinos, A.; Eleftheriadou, I.; Makrilakis, K.; Marinou, K.; Katsilambros, N. Moisture Status of the Skin of the Feet Assessed by the Visual Test Neuropad Correlates With Foot Ulceration in Diabetes. Diabetes Care 2010, 33, 1112–1114. [Google Scholar] [CrossRef] [PubMed]
- Moulaei, K.; Malek, M.; Sheikhtaheri, A. A smart wearable device for monitoring and self-management of diabetic foot: A proof of concept study. Int. J. Med. Inform. 2021, 146, 104343. [Google Scholar] [CrossRef]
- Belvedere, C.; Giacomozzi, C.; Carrara, C.; Lullini, G.; Caravaggi, P.; Berti, L.; Marchesini, G.; Baccolini, L.; Durante, S.; Leardini, A. Correlations between weight-bearing 3D bone architecture and dynamic plantar pressure measurements in the diabetic foot. J. Foot Ankle Res. 2020, 13, 64. [Google Scholar] [CrossRef]
- Chatwin, K.E.; Abbott, C.A.; Boulton, A.J.M.; Bowling, F.L.; Reeves, N.D. The role of foot pressure measurement in the prediction and prevention of diabetic foot ulceration-A comprehensive review. Diabetes Metab. Res. Rev. 2020, 36, e3258. [Google Scholar] [CrossRef]
- Zwaferink, J.B.J.; Custers, W.; Paardekooper, I.; Berendsen, H.A.; Bus, S.A. Optimizing footwear for the diabetic foot: Data-driven custom-made footwear concepts and their effect on pressure relief to prevent diabetic foot ulceration. PLoS ONE 2020, 15, e0224010. [Google Scholar] [CrossRef]
- Zequera, M.; Stephan, S.; Paul, J. The position of the centres of the sensors of the Parotec pressure measurement insoles system related to the anatomy centres of the metatarsal heads by means of radiological records. In Proceedings of the 2007 29th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Lyon, France, 22–26 August 2007; pp. 4453–4456. [Google Scholar] [CrossRef]
- Potdevin, F.J.; Femery, V.G.; Decatoire, A.; Bosquet, L.; Coello, Y.; Moretto, P. Using effect size to quantify plantar pressure asymmetry of gait of nondisabled adults and patients with hemiparesis. J. Rehabil. Res. Dev. 2007, 44, 347–354. [Google Scholar] [CrossRef]
- Fang, F.; Wang, Y.-F.; Gu, M.-Y.; Chen, H.; Wang, D.-M.; Xiao, K.; Yan, S.; Yao, L.-L.; Li, N.; Zhen, Q.; et al. Pedobarography—A novel screening tool for diabetic peripheral neuropathy? Eur. Rev. Med. Pharmacol. Sci. 2013, 17, 3206–3212. [Google Scholar] [PubMed]
- Patrick, K.; Donovan, L. Test-retest reliability of the Tekscan® F-Scan® 7 in-shoe plantar pressure system during treadmill walking in healthy recreationally active individuals. Sports Biomech. 2018, 17, 83–97. [Google Scholar] [CrossRef]
- Price, C.; Parker, D.; Nester, C. Validity and repeatability of three in-shoe pressure measurement systems. Gait Posture 2016, 46, 69–74. [Google Scholar] [CrossRef]
- Suresh, R.; Bhalla, S.; Hao, J.; Singh, C. Development of a high resolution plantar pressure monitoring pad based on fiber Bragg grating (FBG) sensors. Technol. Health Care 2015, 23, 785–794. [Google Scholar] [CrossRef]
- Najafi, B.; Mohseni, H.; Grewal, G.S.; Talal, T.K.; Menzies, R.A.; Armstrong, D.G. An Optical-Fiber-Based Smart Textile (Smart Socks) to Manage Biomechanical Risk Factors Associated With Diabetic Foot Amputation. J. Diabetes Sci. Technol. 2017, 11, 668–677. [Google Scholar] [CrossRef]
- Tavares, C.; Domingues, M.F.; Frizera-Neto, A.; Leite, T.; Leitão, C.; Alberto, N.; Marques, C.; Radwan, A.; Rocon, E.; André, P.; et al. Gait Shear and Plantar Pressure Monitoring: A Non-Invasive OFS Based Solution for e-Health Architectures. Sensors 2018, 18, 1334. [Google Scholar] [CrossRef]
- Wang, D.; Ouyang, J.; Zhou, P.; Yan, J.; Shu, L.; Xu, X. A Novel Low-Cost Wireless Footwear System for Monitoring Diabetic Foot Patients. IEEE Trans. Biomed. Circuits Syst. 2021, 15, 43–54. [Google Scholar] [CrossRef]
- De Pascali, C.; Francioso, L.; Giampetruzzi, L.; Rescio, G.; Signore, M.A.; Leone, A.; Siciliano, P. Modeling, Fabrication and Integration of Wearable Smart Sensors in a Monitoring Platform for Diabetic Patients. Sensors 2021, 21, 1847. [Google Scholar] [CrossRef] [PubMed]
- Khandakar, A.; Mahmud, S.; Chowdhury, M.E.H.; Reaz, M.B.I.; Kiranyaz, S.; Mahbub, Z.B.; Ali, S.H.M.; Bakar, A.A.A.; Ayari, M.A.; Alhatou, M.; et al. Design and Implementation of a Smart Insole System to Measure Plantar Pressure and Temperature. Sensors 2022, 22, 7599. [Google Scholar] [CrossRef]
- Chung, W.K.; Erion, K.; Florez, J.C.; Hattersley, A.T.; Hivert, M.-F.; Lee, C.G.; McCarthy, M.I.; Nolan, J.J.; Norris, J.M.; Pearson, E.R.; et al. Precision Medicine in Diabetes: A Consensus Report From the American Diabetes Association (ADA) and the European Association for the Study of Diabetes (EASD). Diabetes Care 2020, 43, 1617–1635. [Google Scholar] [CrossRef] [PubMed]
- Nolan, J.J.; Kahkoska, A.R.; Semnani-Azad, Z.; Hivert, M.-F.; Ji, L.; Mohan, V.; Eckel, R.H.; Philipson, L.H.; Rich, S.S.; Gruber, C.; et al. ADA/EASD Precision Medicine in Diabetes Initiative: An International Perspective and Future Vision for Precision Medicine in Diabetes. Diabetes Care 2022, 45, 261–266. [Google Scholar] [CrossRef] [PubMed]
- McCarthy, M.I. Painting a new picture of personalised medicine for diabetes. Diabetologia 2017, 60, 793–799. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Sloan, G.; Ye, Y.; Wang, S.; Duan, B.; Tesfaye, S.; Gao, L. New Perspective in Diabetic Neuropathy: From the Periphery to the Brain, a Call for Early Detection, and Precision Medicine. Front. Endocrinol. 2019, 10, 929. [Google Scholar] [CrossRef]
- Van Netten, J.J.; Woodburn, J.; Bus, S.A. The future for diabetic foot ulcer prevention: A paradigm shift from stratified healthcare towards personalized medicine. Diabetes Metab. Res. Rev. 2020, 36 (Suppl. S1), e3234. [Google Scholar] [CrossRef] [PubMed]
| Reference | Aim | Population | Measured/Collected Data | AI Methods | Metrics |
|---|---|---|---|---|---|
| Singh et al., 2013 [21] | Finding DFU risk associated with 5 SNPs in the TLR4 gene | 255 T2DM patients (125 with DFU, 130 without DFU) | Genomic DNA, clinical and laboratory evaluation, family history, habits, duration of disease | ANN | Accuracy |
| Ferreira et al., 2020 [22] | Early identification of T2DM patients at high risk of developing DFU | 239 T2DM patients | Health conditions, changes perceived in feet, information on foot care, type of footwear, socioeconomic and sociodemographic conditions | CNL | Accuracy, sensitivity, specificity |
| Schäfer et al., 2020 [23] | Risk of DFU development/amputation in diabetic people | 246,705 diabetic patients | Patient’s health and socioeconomic data | LR, RF | Accuracy, AUCROC |
| Stefanopoulos et al., 2021 [24] | Prediction of DFU | Over 10 million diabetic patients, 326,853 of which with DFU | Nationwide Inpatient Sample dataset (2008–2014, USA) | CTREE | Accuracy, sensitivity, specificity, AUCROC |
| Haque et al., 2022 [25] | Prediction of diabetic neuropathy or overt DFU | 21 subjects (6 with diabetic neuropathy, 9 with DFU, 6 controls) | Electromyography and ground reaction forces | DA, EC, KC, KNN, LC, NB, SVM, BDC | Accuracy, sensitivity, precision, AUCROC, F1-score |
| Nanda et al., 2022 [26] | Detection of DFU risk and of its severity (according to Wagner Score) | 160 T2DM patients (80 with DFU, 80 without DFU) | Clinical and biochemical risk factors for DFU | SVM, NB, KNN, RF, ensemble learners; Relieff, Info Gain, Gain Ratio and Chi-squared (for feature ranking) | AUCROC, F1-score, MCC |
| Troitskaya et al., 2022 [27] | Prediction of onset of diabetic foot syndrome | 198 diabetic patients without complications, and 199 diabetic patients with signs of diabetic foot | Polymorphisms of genes, markers of endothelial dysfunction | MLP | Accuracy, sensitivity, specificity, AUCROC |
| Reference | Aim | Population | Measured/Collected Data | AI Methods | Metrics |
|---|---|---|---|---|---|
| Toledo Peral et al., 2018 [28] | Identification and classification of skin macules | 19 diabetic patients (without DFU) | 82 photographs of skin macules | ANN | Accuracy, confusion matrix |
| Cruz-Vega et al., 2020 [29] | Classification of diabetic foot thermograms (five classes) | Diabetic patients (number not specified) | 110 thermograms | MLP, SVM, CNN (GoogLeNet and AlexNet, and new CNN: DFTNet) | Accuracy, sensitivity, specificity, precision, AUCROC, F1-score |
| Khandakar et al., 2021 [30] | Classification in diabetic or control subject for early detection of DFU risk | 122 diabetic and 45 control subjects | Gender, age, weight, height, pairs of thermograms | Machine learning algorithms on features extracted from images; deep CNN algorithms on images | Accuracy, sensitivity, specificity, precision, AUCROC, F1-score |
| Arteaga-Marrero et al., 2021 [31] | Proof-of-concept of foot sole segmentation of multimodal images | 37 healthy subjects | 74 visual-light, infrared and depth images | CNN (U-Net), deep CNN (SegNet) | Accuracy, sensitivity, specificity, precision, DICE, spatial overlap |
| Dremin et al., 2021 [32] | Identification of skin differences between diabetic and healthy subjects | 32 healthy subjects (1st study phase), 20 diabetic and 20 healthy subjects (2nd study phase) | Photonic data (hyperspectral imaging and parameters) | ANN (MLP) | Accuracy, sensitivity, specificity, AUCROC |
| Khandakar et al., 2022 (two articles) [33,34] | Early detection of DFU risk, clustering of severity in foot temperature anomalies | 122 diabetic and 45 control subjects | Gender, age, weight, height, pairs of thermograms | Machine learning algorithms; deep CNN algorithms; K-mean clustering | Accuracy, sensitivity, specificity, precision, AUCROC, F1-score |
| Zhang et al., 2022 [35] | Detection of DFU risk and of its severity (according to Wagner Score) | 203 diabetic patients | Sociodemographic and clinical data, and CTA images | ANN, with MLP algorithm | Accuracy, PPV, NPV, sensitivity, specificity, AUCROC |
| Bouallal et al., 2022 [36] | Segmentation of diabetic foot | 145 diabetic and 54 healthy subjects | 398 pairs of thermal and RGB images | DE-ResUnet | IoU, Acc |
| Muralidhara et al., 2022 [37] | Detection of DFU risk and of its severity (6 classes) | 122 diabetic and 45 control subjects | Thermograms | CNN algorithm coupled with class balancing (weighted classification and data augmentation) | Accuracy, sensitivity, specificity, precision, F1-score |
| Reference | Aim | Population | Measured/Collected Data | AI Methods | Metrics |
|---|---|---|---|---|---|
| Yusuf et al., 2015 [38] | Validation of e-nose in detection of bacteria responsible for DFU infection | Patients with DFU (number not specified) | In vitro bacteria samples | SVM, KNN, LDA, PNN | Accuracy, sensitivity, specificity, precision |
| Huang et al., 2018 [39] | Quantification of rehabilitative efficiency of Buerger’s exercise; discrimination between healthy and diabetic subjects | 30 diabetic and 15 healthy subjects | Tissue oxygen saturation in lower limbs and relative total hemoglobin concentration | RBFNN | F1-score |
| Lin et al., 2020 [40] | Prediction of amputation/mortality in patients with DFU | 200 patients with DFU | Biochemical markers, clinical data and presence of complications | Cox regression, BPNN (also with GA) | Sensitivity, specificity, AUCROC |
| Du et al., 2021 [41] | Prediction of amputation/mortality in inpatient with DFU before/after pandemic | 23 inpatients with DFU | Clinical and laboratory data, WIFI classification | LR, SVM, RF, GBDT, ANN, XGBoost | Accuracy, NPV, PPV, sensitivity, specificity, AUCROC |
| Xie et al., 2022 [42] | Prediction of in-hospital amputation | 618 patients with DFU | Demographic features, medical and medication history, clinical and laboratory data, Wagner and WIFI classifications | LightGBM | Accuracy, NPV, PPV, sensitivity, specificity, AUCROC |
| Margolis et al., 2022 [43] | Prediction of wound healing | 204 patients with DFU | Wound area, duration, depth, site, arterial flow, BMI, history of dialysis | LR, LASSO | AUCROC |
| Deng et al., 2022 [44] | Prediction of mortality in DFU+HCE patients | 27 inpatients with DFU+HCE, 93 inpatients with isolated DFU | HCE presence, mortality occurrence, clinical data | XGBoost | AUCROC, accuracy, sensitivity, specificity |
| Reference | Aim | Population | Measured/Collected Data | AI Methods | Metrics |
|---|---|---|---|---|---|
| Wang et al., 2017 [45] | Detection of DFU area | 15 patients with DFU | 100 DFU images | Two-stage SVM | Sensitivity, specificity |
| Wang et al., 2019 [46] | Automatic DFU localization under different conditions | 15 patients with DFU | 162 moulage wound images + 100 actual DFU images | AHRF | Sensitivity, specificity |
| Ohura et al., 2019 [47] | Automatic DFU localization | Patients with DFU (number not specified) | 400 pressure ulcer images and 20 DFU images | SegNet, LinkNet, U-Net and U-Net with VGG16 | Accuracy, sensitivity, specificity, AUCROC, MCC, DICE |
| Goyal et al., 2019 (and 2020) [48,49] | Real time automatic DFU localization | Patients with DFU (number not specified) | (Up to) 1775 DFU images | From machine learning: SVM; from deep learning: R-CNN, R-FCN, SSD; DFUNet | mAP, overlap percentage, size of model, speed; accuracy, sensitivity, specificity, precision, AUCROC, F1-score |
| Goyal et al., 2020 [50] | Detection of ischemia/infection in DFU | Patients with DFU (number not specified) | 1459 DFU images | From machine learning: RF, BN, MLP; from deep learning: three CNN (InceptionV3, ResNet50, and InceptionResNetV2), ensemble CNN based on the three CNN | Accuracy, sensitivity, specificity, precision, AUCROC, F1-score, MCC |
| Kim et al., 2020 [51] | Prediction of DFU prognosis | 155 patients with 2291 visits for 381 DFUs | Clinical variables, smartphone-based photographs | ResNet50, RF, SVM | Accuracy, precision, recall, AUCROC F1-score |
| Al-Garaawi et al., 2021 [52] | DFU classification, detection of ischemia, detection of infection | Patients with DFU (number not specified) | RGB images and derived information about texture of the ROI | CNN | Accuracy, sensitivity, specificity, precision, AUCROC, F1-score |
| Yap et al., 2021 (and Cassidy et al., 2021) [53,54] | DFU detection | Patients with DFU (number not specified) | 4000 DFU images with expert annotations | R–CNN, three variants of R–CNN, an ensemble method; YOLOv3, YOLOv5; efficientDet; Cascade Attention Network | Precision, recall, true and false positives, F1-score, mAP |
| Xu et al., 2021 [55] | Detection of ischemia/infection in DFU | Patients with DFU (number not specified) | 1459 DFU images | CKBs | Accuracy, sensitivity, specificity, precision, AUCROC, F1-score |
| Viswanathan et al., 2021 [56] | Identification of wound Gram type infections | 178 patients with DFU, for 203 wound tissue samples | Autofluorescence images | Not specified | Not specified |
| Güley et al., 2022 [57] | Identification of wound infection and/or ischemia | Patients with DFU (number not specified) | 15,863 DFU images (possibly with wound infection and/or ischemia) | VGG11, VGG16, VGG19 | Recall, AUCROC, F1-score |
| Wang et al., 2022 [58] | Ability of MRI images to describe therapeutic effect of skin grafting | 78 patients with DFU (39 +39, for composite and autologous graft, respectively) | MRI images of DFU | Deep learning model (SSD) | Accuracy, sensitivity, specificity, AUCROC |
| Yogapriya et al., 2022 [59] | Prediction of DFU non-infection or infection (risk for amputation) | Patients with DFU (number not specified) | 5890 DFU images (2945 with foot infection, 2945 without infection) | CNN with normalization and dropout layers (DFINET) | Accuracy, NPV, PPV, sensitivity, specificity, precision, F1-score, MCC |
| Chan et al., 2022 [60] | DFU detection and measurement of its length, width, and area | Patients with DFU (number not specified) | 547 DFU images | Not specified | Intra- and inter-rater reliability |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chemello, G.; Salvatori, B.; Morettini, M.; Tura, A. Artificial Intelligence Methodologies Applied to Technologies for Screening, Diagnosis and Care of the Diabetic Foot: A Narrative Review. Biosensors 2022, 12, 985. https://doi.org/10.3390/bios12110985
Chemello G, Salvatori B, Morettini M, Tura A. Artificial Intelligence Methodologies Applied to Technologies for Screening, Diagnosis and Care of the Diabetic Foot: A Narrative Review. Biosensors. 2022; 12(11):985. https://doi.org/10.3390/bios12110985
Chicago/Turabian StyleChemello, Gaetano, Benedetta Salvatori, Micaela Morettini, and Andrea Tura. 2022. "Artificial Intelligence Methodologies Applied to Technologies for Screening, Diagnosis and Care of the Diabetic Foot: A Narrative Review" Biosensors 12, no. 11: 985. https://doi.org/10.3390/bios12110985
APA StyleChemello, G., Salvatori, B., Morettini, M., & Tura, A. (2022). Artificial Intelligence Methodologies Applied to Technologies for Screening, Diagnosis and Care of the Diabetic Foot: A Narrative Review. Biosensors, 12(11), 985. https://doi.org/10.3390/bios12110985







