Evidence of Protein Adsorption in Pegylated Liposomes: Influence of Liposomal Decoration
Abstract
:1. Introduction
2. Results and Discussion
2.1. Physicochemical Characterization of Liposomes and Magnetoliposomes
2.2. Physicochemical Characterization of BSA Corona-Coated Liposomes and Magnetoliposomes
2.2.1. Changes in Size, Polydispersity Index, and Zeta Potential Corroborate the Interaction of BSA with LUVs and MLs
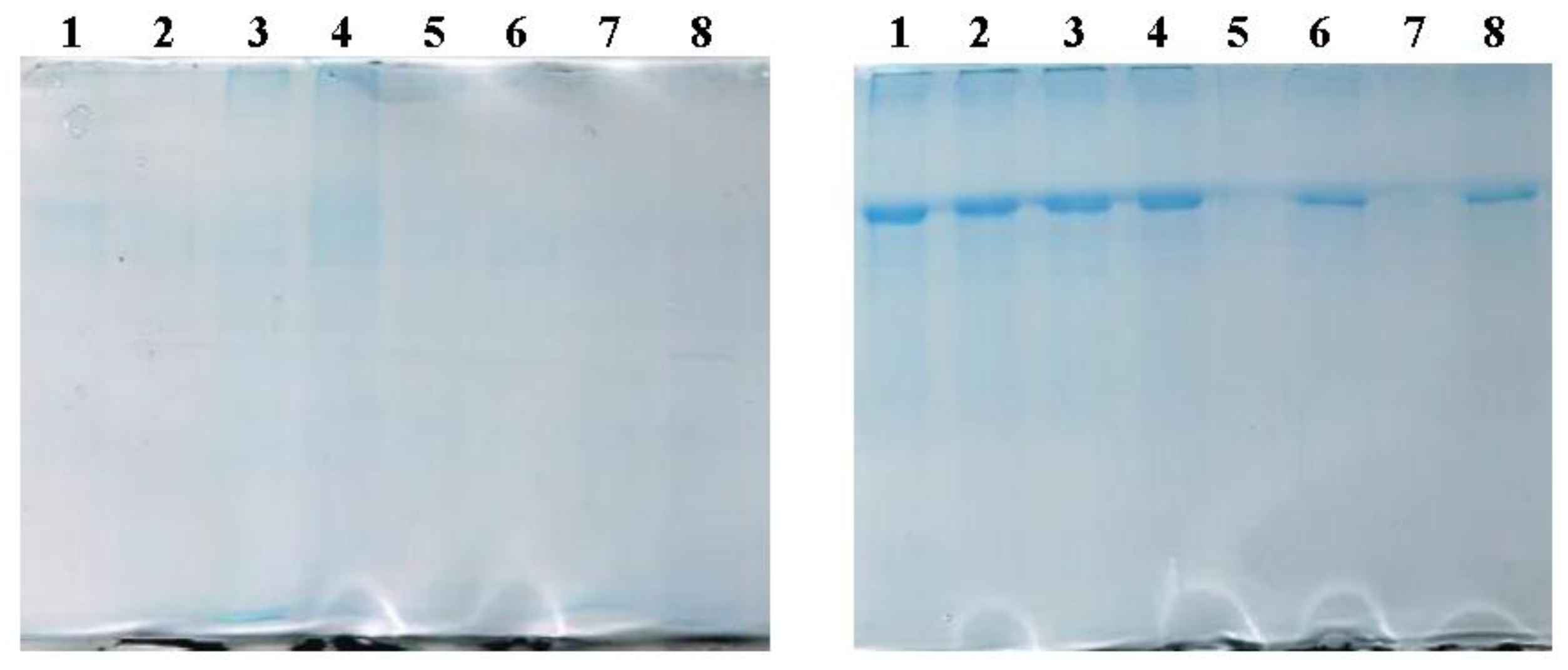
2.2.2. SDS-PAGE Evidences the Presence of BSA in LUVs and MLs
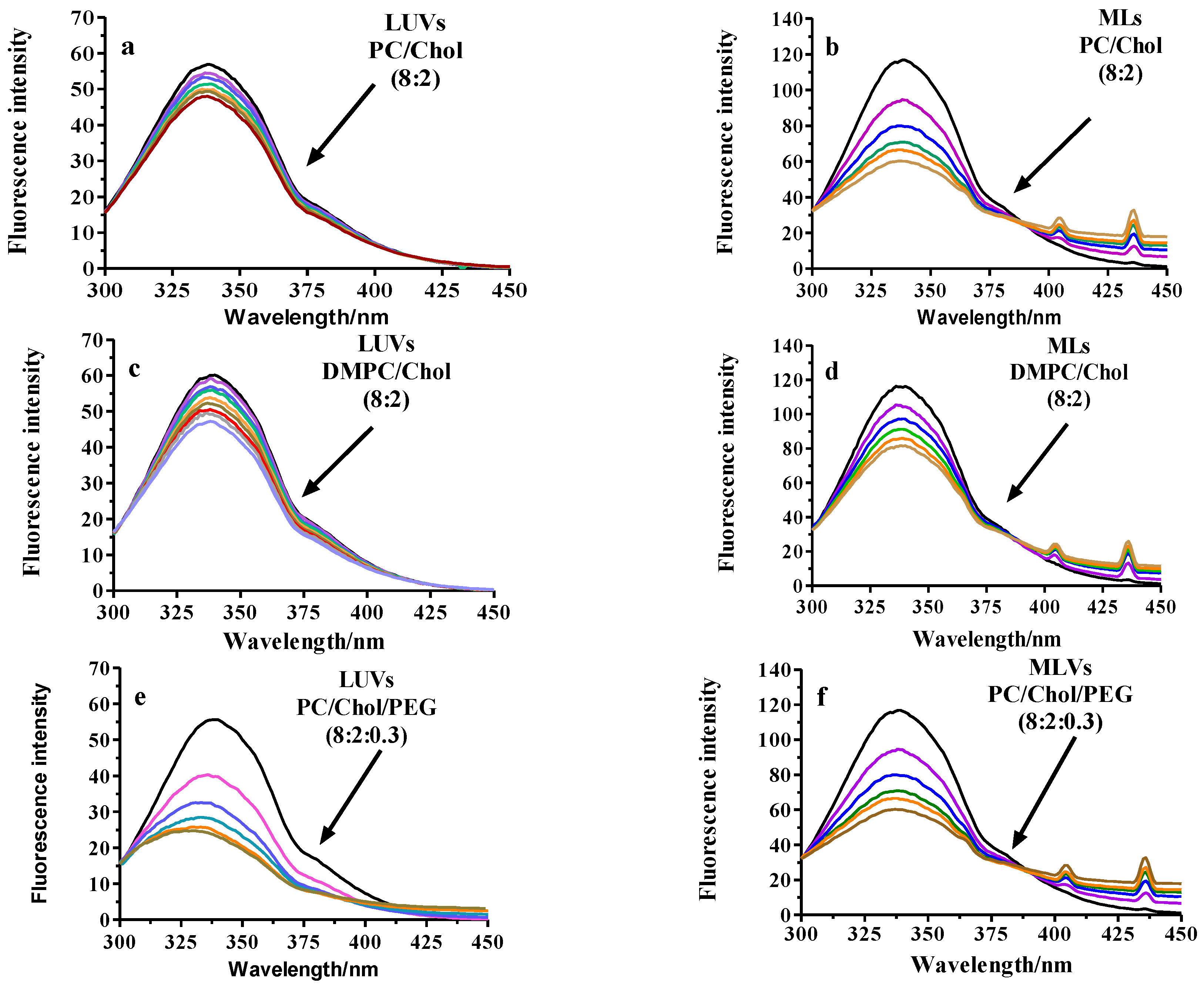
2.2.3. Fluorescence Analysis of BSA Binding to LUVs or MLs
Fluorescence Quenching
Fluorescence Anisotropy
2.2.4. Thermotropic Behavior of LUVs and MLs in Presence of BSA
Differential Scanning Calorimetry (DSC)
Isothermal Titration Calorimetry (ITC)
3. Methods
3.1. Materials
3.2. Methods
3.2.1. Preparation and Characterization of Liposomes and Magnetoliposomes
3.2.2. Isothermal Titration Calorimetry (ITC)
3.2.3. Differential Scanning Calorimetry (DSC)
3.2.4. SDS Polyacrylamide Gel Electrophoresis (SDS-PAGE)
3.2.5. Fluorescence Spectroscopy
3.2.6. Fluorescence Anisotropy
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| BSA | Bovine Serum Albumin |
| DMPC | 1,2-dimyristoyl-sn-glycero-3-phosphocholine |
| CHOL | Cholesterol |
| FF | Ferrofluid |
| hd | Hydrodynamic diameter |
| LUVs | Large unilamellar vesicles |
| MLs | Magnetoliposomes |
| NPs | Nanoparticles |
| PC | l-α-phosphatydylcholine |
| PDI | Polydispersity Index |
| PEG | 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[methoxy(polyethylene glycol)-2000] (ammonium salt) |
| PEG* | 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-N-[maleimide (polyethylene glycol)-2000] (ammonium salt) (DSPE-Mal-PEG-2000) |
| PL | Phospholipid |
| RGDc | Cyclic arginine-glycine-aspartate peptide |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| Cryo-TEM | Cryogenic transmission electron microscopy |
References
- Marty, J.J.; Oppenheim, R.C.; Speiser, P. Nanoparticles—A new colloidal drug delivery system. Pharm. Acta Helv. 1978, 53, 17–23. [Google Scholar] [PubMed]
- Antonietti, M.; Foerster, S. Vesicles and liposomes: A self-assembly principle beyond lipids. Adv. Mater. 2003, 15, 1323–1333. [Google Scholar] [CrossRef]
- Torchilin, V.P. Recent advances with liposomes as pharmaceutical carriers. Nat. Rev. Drug Discov. 2005, 4, 145–160. [Google Scholar] [CrossRef] [PubMed]
- Clausell, A.; Garcia-Subirats, M.; Pujol, M.; Busquets, M.A.; Rabanal, F.; Cajal, Y. Gram negative bacteria outer and inner membrane models: Insertion of cyclic cationic peptide. J. Phys. Chem. B 2007, 111, 551–563. [Google Scholar] [CrossRef] [PubMed]
- Allen, T.M.; Cullis, P.R. Liposomal drug delivery systems: From concept to clinical applications. Adv. Drug Deliv. Rev. 2013, 65, 36–48. [Google Scholar] [CrossRef] [PubMed]
- Sercombe, L.; Veerati, T.; Moheimani, F.; Wu, S.Y.; Sood, A.K.; Hua, S. Advances and Challenges of Liposome Assisted Drug Delivery. Front. Pharmacol. 2015, 6, 286. [Google Scholar] [CrossRef] [PubMed]
- Oussoren, C.; Zuidema, J.; Crommelin, D.J.A.; Storm, G. Lymphatic uptake and biodistribution of liposomes after subcutaneous injection. II. Influence of liposomal size, lipid composition and lipid dose. Biochim. Biophys. Acta BBA Biomembr. 1997, 1328, 261–272. [Google Scholar] [CrossRef]
- Szoka, F., Jr.; Papahadjopoulos, D. Comparative properties and methods of preparation of lipid vesicles (liposomes). Ann. Rev. Biophys. Bioeng. 1980, 9, 467–508. [Google Scholar] [CrossRef] [PubMed]
- Akbarzadeh, A.; Rezaei-Sadabady, R.; Davaran, S.; Woo Joo, S.; Zarghami, N.; Hanifehpour, Y.; Samiei, M.; Kouhi, M.; Nejati-Koshki, K. Liposome: Classification, preparation, and applications. Nanoscale Res. Lett. 2013, 8, 102. [Google Scholar] [CrossRef] [PubMed]
- Woodle, M.C.; Lasic, D.D. Sterically stabilized liposomes. Biochim. Biophys. Acta BBA Rev. Biomembr. 1992, 1113, 171–199. [Google Scholar] [CrossRef]
- Harris, J.M.; Martin, N.E.; Modi, M. Pegylation. A novel process for modifying pharmacokinetics. Clin. Pharmacokinet. 2001, 40, 539–551. [Google Scholar] [CrossRef] [PubMed]
- Moghimi, S.M.; Szebeni, J. Stealth liposomes and long circulating nanoparticles: Critical issues in pharmacokinetics, opsonization and protein-binding properties. Prog. Lipid Res. 2003, 42, 463–478. [Google Scholar] [CrossRef]
- Dos Santos, N.; Allen, C.G.; Doppen, A.M.; Anantha, M.; Cox, K.A.K.; Gallagher, R.C.; Karlsson, G.; Edwards, K.; Kenner, G.; Samuels, L.; et al. Influence of ply(ethylene glycol) grafting density and polymer length on liposomes: Relating plasma circulation lifetimes to protein binding. Biochim. Biophys. Acta BBA Biomembr. 2007, 1768, 1367–1377. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, D.; Colapicchioni, V.; Caracciolo, G.; Piovesana, S.; Capriotti, A.L.; Palchetti, S.; De Grossi, S.; Riccioli, A.; Amenitsch, H.; Laganà, A. Effect of polyethyleneglycol (PEG) chain length on the bio–nano-interactions between PEGylated lipid nanoparticles and biological fluids: From nanostructure to uptake in cancer cells. Nanoscale 2014, 6, 2782–2792. [Google Scholar] [CrossRef] [PubMed]
- Balducci, C.; Mancini, S.; Minniti, S.; La Vitola, P.; Zotti, M.; Sancini, G.; Mauri, M.; Cagnotto, A.; Colombo, L.; Fiordaliso, F.; et al. Multifunctional liposomes reduce brain β-amyloid burden and ameliorate memory impairment in Alzheimer’s disease mouse models. J. Neurosci. 2014, 34, 14022–14031. [Google Scholar] [CrossRef] [PubMed]
- Amin, M.; Mansourian, M.; Koning, G.A.; Badiee, A.; Jaafari, M.R.; ten Hagen, T.L.M. Development of a novel cyclic RGD peptide for multiple targeting approaches of liposomes to tumor region. J. Control. Release 2015, 220, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Gómara, M.J.; Pérez-Pomeda, I.; Gatell, J.M.; Sánchez-Merino, V.; Yuste, E.; Haro, I. Lipid raft-like liposomes used for targeted delivery of a chimeric entry-inhibitor peptide with anti-HIV-1 activity. Nanomed. Nanotechnol. Biol. Med. 2017, 13, 601–609. [Google Scholar] [CrossRef] [PubMed]
- Al-Ahmady, Z.; Kostarelos, K. Chemical Components for the Design of Temperature-Responsive Vesicles as Cancer Therapeutics. Chem. Rev. 2016, 116, 3883–3918. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Rao, X.; Cai, L.; Liu, X.; Wang, H.; Wu, W.; Zhu, C.; Chen, M.; Wang, P.G.; Yi, W. Targeting tumor cells by natural anti-carbohydrate antibodies using rhamnose-functionalized liposomes. ACS Chem. Biol. 2016, 11, 1205–1209. [Google Scholar] [CrossRef] [PubMed]
- Moles, E.; Urbán, P.; Jiménez-Díaz, M.B.; Viera-Morilla, S.; Angulo-Barturen, I.; Busquets, M.A.; Fernández-Busquets, X. Immunoliposome-mediated drug delivery to Plasmodium-infected and non-infected red blood cells as a dual therapeutic/prophylactic antimalarial stragegy. J. Control. Release 2015, 210, 217–229. [Google Scholar] [CrossRef] [PubMed]
- Ruoslahti, E. RGD and other recognition sequences for integrins. Annu. Rev. Cell Dev. Biol. 1996, 12, 697–715. [Google Scholar] [CrossRef] [PubMed]
- Danhier, F.; Le Breton, A.; Préat, V. RGD-based strategies to target alpha(v) beta(3) integrin in cancer therapy and diagnosis. Mol. Pharm. 2012, 9, 2961–2973. [Google Scholar] [CrossRef] [PubMed]
- Del Pino, P.; Pelaez, B.; Zhang, Q.; Maffre, P.; Nienhaus, G.U.; Parak, W.J. Protein corona formation around nanoparticles-from the past to the future. Mater. Horiz. 2014, 1, 301–313. [Google Scholar] [CrossRef]
- Salvati, A.; Pitek, A.S.; Monopoli, M.P.; Prapainop, K.; Bombelli, F.B.; Hristov, D.R.; Kelly, P.M.; Åberg, C.; Mahon, E.; Dawson, K.D. Transferrin-functionalized nanoparticles lose their targeting capabilities when a biomolecule corona adsorbs on the surface. Nat. Nanotechnol. 2012, 8, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Barran-Berdon, A.L.; Pozzi, D.; Caracciolo, G.; Capriotti, A.L.; Caruso, G.; Cavaliere, C.; Riccioli, A.; Palchetti, S.; Lagana, A. Time evolution of nanoparticle–protein corona in human plasma: Relevance for targeted drug delivery. Langmuir 2013, 9, 2030–2044. [Google Scholar] [CrossRef] [PubMed]
- Mirshafiee, V.; Kim, R.; Mahmoudi, M.; Kraft, M. The importance of a proper biological milieu for protein corona analysis in vitro: Human plasma versus human serum. Int. J. Biochem. Cell Biol. 2016, 75, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Chattopadhyay, N.; Fonge, H.; Cai, Z.; Scollard, D.; Lechtman, E.; Done, S.J.; Pignoland, J.P.; Reilly, R.M. Role of antibody-mediated tumor targeting and route of administration in nanoparticle tumor accumulation in vivo. Mol. Pharm. 2012, 9, 2168–2179. [Google Scholar] [CrossRef] [PubMed]
- Schöttler, S.; Becker, G.; Winzen, S.; Steinbach, T.; Mohr, K.; Landfester, K.; Mailänder, V.; Wurm, F.R. Protein adsorption is required for stealth effect of poly(ethylene glycol)- and poly(phosphoester)-coated nanocarriers. Nat. Nanotechnol. 2016, 11, 372–377. [Google Scholar] [CrossRef] [PubMed]
- Moyano, D.F.; Saha, K.; Prakash, G.; Yan, B.; Kong, H.; Yazdani, M.; Rotello, V.M. Fabrication of Corona-Free Nanoparticles with Tunable Hydrophobicity. ACS Nano 2014, 8, 6748–6755. [Google Scholar] [CrossRef] [PubMed]
- Gref, R.; Luck, M.; Quellec, P.; Marchand, M.; Dellacherie, E.; Harnisch, S.; Blunk, T.; Muller, R.H. Stealth corona-core nanoparticles surface modified by poyethylene glycol (PEG) influences of the corona (PEG chain length and surface density) and of the core composition on phagocytic uptake and plasma protein adsorption. Colloids Suf. B 2000, 18, 301–313. [Google Scholar] [CrossRef]
- Lundqvist, M.; Stigler, J.; Elia, G.; Lynch, I.; Cedervall, T.; Dawson, K. Nanoparticle size an surface properties determine the protein corona with possible implications for biological impacts. Proc. Natl. Acad. Sci. USA 2008, 105, 14265–14270. [Google Scholar] [CrossRef] [PubMed]
- Nappini, S.; Fogli, S.; Castroflorio, B.; Bonini, M.; Baldelli Bombelli, F.; Baglioni, P. Magnetic field responsive drug release from magnetoliposomes in biological fluids. J. Mater. Chem. B 2016, 4, 716–725. [Google Scholar] [CrossRef]
- Ritz, S.; Schöttler, S.; Kotman, N.; Baier, G.; Musyanovych, A.; Kuharev, J.; Landfester, K.; Schild, H.; Jahn, O.; Tenzer, S.; et al. Protein corona of nanoparticles: Distinct proteins regulate the cellular uptake. Biomacromolecules 2015, 16, 1311–1321. [Google Scholar] [CrossRef] [PubMed]
- Schöttler, S.; Klein, K.; Landfester, K.; Mailänder, V. Protein source and choice of anticoagulant decisively affect nanoparticle protein corona and cellular uptake. Nanoscale 2016, 8, 5526–5536. [Google Scholar] [CrossRef] [PubMed]
- Hajipour, M.J.; Laurent, S.; Aghaie, A.; Rezaee, F.; Mahmoudi, M. Personalized protein corones: A “key” factor at the nanobiointerface. Biomater. Sci. 2014, 2, 1210–1221. [Google Scholar] [CrossRef]
- Cifuentes-Rius, A.; de Puig, H.; Kah, J.C.; Borros, S.; Hamad-Schifferli, K. Optimizing the properties of the protein corona surrounding nanoparticles for tuning payload release. ACS Nano 2013, 7, 10066–10074. [Google Scholar] [CrossRef] [PubMed]
- Palchetti, S.; Pozzi, D.; Mahmoudi, M.; Caracciolo, G. Exploitation of nanoparticle–protein corona for emerging therapeutic and diagnostic applications. J. Mater. Chem. B 2016, 4, 4376–4381. [Google Scholar] [CrossRef]
- Di Silvio, D.; Rigby, N.; Bajika, B.; Mackie, A.; Baldelli Bombelli, F. Effect of protein corona magnetite nanoparticles derived from bread in vitro digestion on Caco-2 cells morphology and uptake. Int. J. Biochem. Cell Biol. 2016, 75, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Walkey, C.D.; Chan, W.C.W. Understanding and controlling the interaction of nanomaterials with proteins in a physiological environment. Chem. Soc. Rev. 2012, 41, 2780–2799. [Google Scholar] [CrossRef] [PubMed]
- Colapicchioni, V.; Tilio, M.; Digiacomo, L.; Gambini, V.; Palchetti, S.; Marchini, C.; Pozzi, D.; Occhipinti, S.; Amici, A.; Caracciolo, G. Personalized liposome-protein corona in the blood of breast, gastric and pancreatic cancer patients. Int. J. Biochem. Cell Biol. 2016, 75, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Zanganeh, S.; Spitler, R.; Erfanzadeh, M.; Alkilany, A.M.; Mahmoudi, M. Protein corona: Opportunities and challenges. Int. J. Biochem. Cell Biol. 2016, 75, 143–147. [Google Scholar] [CrossRef] [PubMed]
- Hadjidemetriou, M.; Al-Ahmady, Z.; Mazza, M.; Collins, R.F.; Dawson, K.; Kostarelos, K. In vivo biomolecule corona around blood-circulating, clinically used and antibody-targeted lipid bilayer nanoscale vesicles. ACS Nano 2015, 9, 8142–8156. [Google Scholar] [CrossRef] [PubMed]
- Caracciolo, G. Liposome-protein corona in a physiological environment: Challenges and opportunities for targeted delivery nanomedicines. Nanomed. Nanotechnol. Biol. Med. 2015, 11, 543–557. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, D.; Caracciolo, G.; Capriotti, A.L.; Cavaliere, C.; La Barbera, G.; Anchordoquy, T.J.; Laganà, A. Surface chemistry and serum type both determine the nanoparticle–protein corona. J. Proteom. 2015, 119, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Bigdeli, A.; Palchetti, S.; Pozzi, D.; Hormozi-Nezhad, M.R.; Baldelli Bombelli, F.; Caracciolo, G.; Morteza Mahmoudi, M. Exploring cellular interactions of liposomes using protein corona fingerprints and physicochemical properties. ACS Nano 2016, 10, 3723–3737. [Google Scholar] [CrossRef] [PubMed]
- M’Hallah, R.; Alkandari, A.; Mlandenovic, N. Packing unit spheres into the smallest sphere using VNS and NLP. Comput. Oper. Res. 2013, 40, 603–6015. [Google Scholar] [CrossRef]
- Palcheti, S.; Colapicchioni, V.; Digiacomo, L.; Caracciolo, G.; Pozzi, D.; Capriotti, A.L.; La Barcera, G.; Laganà, A. The protein corona of circulating PEGylated liposomes. Biochim. Biophys. Acta BBA Biomembr. 2016, 1858, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Tayeh, N.; Rungassamy, T.; Albani, J.R. Fluorescence spectral resolution of tryptophan residues in bovine and human serum albumins. J. Pharm. Biomed. Anal. 2009, 50, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Toprak, M. Fluorescence study on the interaction of human serum albumin with butein in liposomes. Spectrochim. Acta Part A 2016, 154, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Tian, J.; Hu, Z.; Chen, X. Binding of isofraxidin to bovine serum albumin. Biopolymers 2004, 73, 443–450. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.H.; Qi, S.D.; Chen, X.G. Spectroscopic studies of the interaction of anticoagulant rodenticide diphacinone with human serum albumin. J. Mol. Struct. 2005, 779, 87–95. [Google Scholar] [CrossRef]
- Hill, A.V.; Brown, T.G.; Roal, H.E. The possible effects of the aggregation of the molecules of hemoglobin on its dissociation curves. J. Physiol. 1910, 40, 4–7. [Google Scholar]
- Hühn, J.; Fedeli, C.; Zhang, Q.; Masood, A.; del Pino, P.; Khashab, N.M.; Papini, E.; Parak, W.J. Dissociation coefficients of protein adsorption to nanoparticles as quantitative metrics for description of the protein corona: A comparison of experimental techniques and methodological relevance. Int. J. Biochem. Cell Biol. 2016, 75, 148–161. [Google Scholar] [CrossRef] [PubMed]
- Fleischer, C.C.; Payne, C.K. Nanoparticle-cell interactions: Molecular structure of the protein corona and cellular outcomes. Acc. Chem. Res. 2014, 47, 2651–2659. [Google Scholar] [CrossRef] [PubMed]
- Stewart, J.C. Colorimetric determination of phospholipids with ammonium ferrothio-cyanate. Anal. Biochem. 1980, 104, 10–14. [Google Scholar] [CrossRef]
- García-Jimeno, S.; Estelrich, J. Ferrofluid based on polyethylene glycolcoated iron oxide nanoparticles: Characterization and properties. Colloids Surf. A 2013, 420, 74–81. [Google Scholar] [CrossRef]
- Kiwada, H.; Sato, J.; Yamada, S.; Kato, Y. Feasibility of magnetic liposomes as a targeting device for drugs. Chem. Pharm. Bull. 1986, 34, 4253–4258. [Google Scholar] [CrossRef] [PubMed]
- Charbonneau, D.M.; Tajmir-Riahi, H.A. Study of the interaction of cationic lipids with bovine serum albumin. J. Phys. Chem. B 2010, 114, 1148–1155. [Google Scholar] [CrossRef] [PubMed]
- Alay, M.; Haro, I.; Alsina, M.A.; Girona, V.; Prat, J.; Busquets, M.A. Interaction of two overlapped synthetic peptides from GB virus C with charged mono and bilayers. Colloids Surf. B 2013, 105, 7–13. [Google Scholar] [CrossRef] [PubMed]

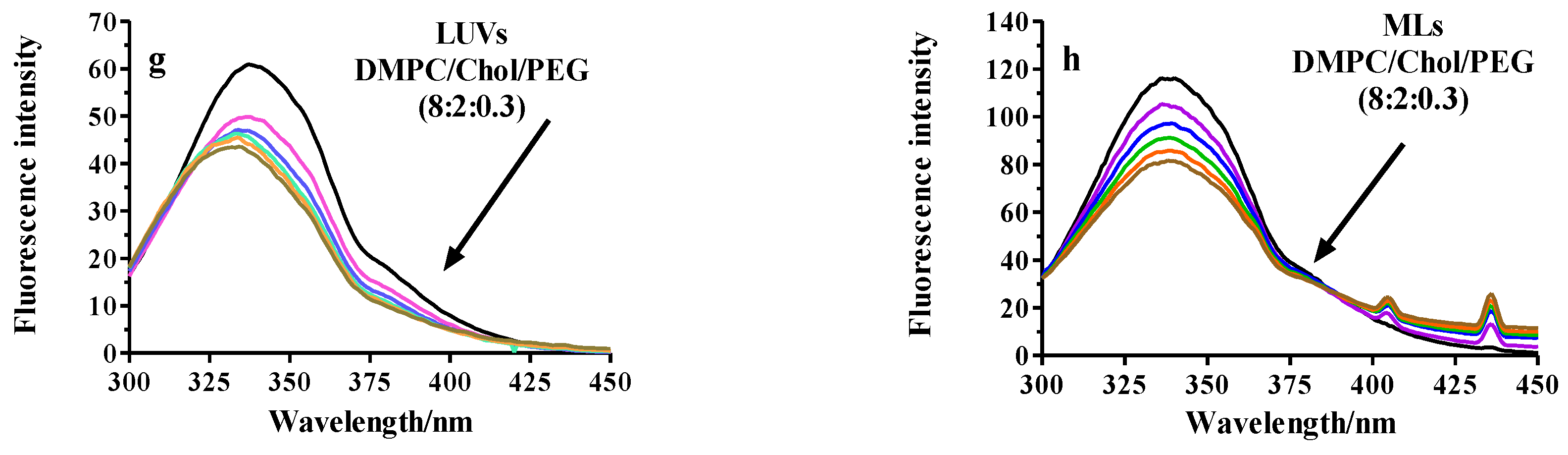
| Lipid Composition | hd/nm | PDI | ζ/mV | |||
|---|---|---|---|---|---|---|
| LUVs | MLs | LUVs | MLs | LUVs | MLs | |
| PC/CHOL (8:2) | 170.4 ± 1.0 | 178.5 ± 6.4 | 0.090 ± 0.041 | 0.136 ± 0.048 | −1.85 ± 0.85 | −7.5 ± 0.43 |
| PC/CHOL/PEG (8:2:0.3) | 161.8 ± 3.7 | 114.0 ± 7.6 | 0.119 ± 0.032 | 0.129 ± 0.05 | −16.6 ± 0.20 | −18.9 ± 4.23 |
| PC/CHOL/PEG*/RGD (8:2:0.3:0.03) | 136.2 ± 1.5 | 175.1 ± 1.4 | 0.146 ± 0.011 | 0.200 ± 0.01 | −13.0 ± 0.90 | −27.0 ± 0.35 |
| DMPC/CHOL (8:2) | 180.0 ± 3.6 | 191.5 ± 6.9 | 0.122 ± 0.018 | 0.200 ± 0.02 | −4.66 ± 0.92 | −1.23 ± 0.96 |
| DMPC/CHOL/PEG (8:2:0.3) | 154.9 ± 4.5 | 168.7 ± 6.6 | 0.163 ± 0.024 | 0.139 ± 0.06 | −14.8 ± 0.14 | −15.7 ± 0.40 |
| DMPC/CHOL/PEG*/RGD (8:2:0.3:0.03) | 183.1 ± 4.2 | 198.0 ± 5.2 | 0.149 ± 0.023 | 0.135 ± 0.02 | −26.3 ± 0.95 | −22.5 ± 0.83 |
| Lipid Composition | LUVs | MLs | ||||
|---|---|---|---|---|---|---|
| Δ Size/nm | Δ PDI | Δ ζ/mV | Δ Size/nm | Δ PDI | Δ ζ/mV | |
| Pristine-PC | Aggregated | >1 | Aggregated | >1 | ||
| PEGylated-PC | Aggregated | >1 | Aggregated | >1 | ||
| RGD-PC | Aggregated | >1 | Aggregated | >1 | ||
| Pristine-DMPC | ~0 | 0.017 | −25.25 ± 16.53 | 4.50 ± 2.6 | 0.070 | −4.24 ± 4.35 |
| PEGylated-DMPC | ~0 | 0.014 | −0.73 ± 1.00 | 24.0 ± 5.2 | 0.085 | −16.93 ± 3.59 |
| RGD-DMPC | ~0 | 0.014 | −25.25 ± 2.05 | Aggregated | >1 | |
| Lipid Composition | LUVs | MLs | ||
|---|---|---|---|---|
| Ksv/L·mol−1 | r | Ksv/L·mol−1 | r | |
| Pristine-PC | 345 ± 6 | 0.994 | 2465 ± 30 | 0.996 |
| PEGylated-PC | 4460 ± 22 | 0.997 | 1908 ± 27 | 0.997 |
| RGD-PC | 963 ± 15 | 0.995 | 2151 ± 13 | 0.996 |
| Pristine-DMPC | 500 ± 12 | 0.992 | 1237 ± 20 | 0.999 |
| PEGylated-DMPC | 840 ± 10 | 0.996 | 1685 ± 35 | 0.997 |
| RGD-DMPC | 1300 ± 21 | 0.994 | 1739 ± 24 | 0.999 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sangrà, M.; Estelrich, J.; Sabaté, R.; Espargaró, A.; Busquets, M.A. Evidence of Protein Adsorption in Pegylated Liposomes: Influence of Liposomal Decoration. Nanomaterials 2017, 7, 37. https://doi.org/10.3390/nano7020037
Sangrà M, Estelrich J, Sabaté R, Espargaró A, Busquets MA. Evidence of Protein Adsorption in Pegylated Liposomes: Influence of Liposomal Decoration. Nanomaterials. 2017; 7(2):37. https://doi.org/10.3390/nano7020037
Chicago/Turabian StyleSangrà, Marc, Joan Estelrich, Raimon Sabaté, Alba Espargaró, and Maria Antònia Busquets. 2017. "Evidence of Protein Adsorption in Pegylated Liposomes: Influence of Liposomal Decoration" Nanomaterials 7, no. 2: 37. https://doi.org/10.3390/nano7020037
APA StyleSangrà, M., Estelrich, J., Sabaté, R., Espargaró, A., & Busquets, M. A. (2017). Evidence of Protein Adsorption in Pegylated Liposomes: Influence of Liposomal Decoration. Nanomaterials, 7(2), 37. https://doi.org/10.3390/nano7020037







